Abstract
Rhinitis and sinusitis caused by fungal pathogens were studied in biopsy samples submitted from 52 horses distributed throughout subtropical and tropical regions of Florida. Methods included routine histopathology as well as polymerase chain reaction (PCR) with panfungal/panoomycete primers and DNA sequencing on extracted DNA (DNA barcoding). Granulomatous, pyogranulomatous, and fibrinopurulent lesions in nasal and sinus mucosa were associated with signs of upper airway obstruction and noise as well as nasal discharge. Morphologic and histochemical assessment of cases identified 31 cases of zygomycosis/pythiosis plus 1 mixed infection case, 16 cases of phaeohyphomycosis with 2 additional mixed infection cases, and 3 cases caused by other fungi. Morphologic evidence of Aspergillus sp. infection as a superficial copathogen was found in 2 of the mixed fungal infection cases. PCR and DNA sequencing facilitated identification of fungal pathogens in 11 of 52 cases (21%). No evidence of oomycete infection was found. Histomorphologic features of previously unrecognized forms of equine rhinitis/sinusitis were described, including those caused by Flavodon flavus, Curvularia lunata, Exserohilum rostrata, Alternaria alternata, Alternaria sp., Cladophialophora bantiana, Fusarium solani, and Toxicocladosporium irritans. PCR and DNA sequencing using panfungal and oomycete primers with DNA from formalin-fixed and paraffin-embedded specimens successfully identified the pathogen in phaeohyphomycosis (7/18 cases, 39%), zygomycosis/pythiosis (1/32 cases, 3%), and other nonpigmented fungal infections (3/3 cases, 100%). Zygomycosis and phaeohyphomycosis were the most common forms of fungal rhinitis found in Florida horses.
Rhinitis caused by fungal and oomycete pathogens is an infrequent, but serious, disease in horses because it causes obstructive and progressive lesions in the nasal passages of obligate nose breathers that can spread to other areas of the respiratory system (eg, sinusitis). Nasal fungal infection is thought to occur by contact with pathogens residing in decaying plant material where horses feed, by aerosol and by insect transmission. 2,30 One of the most common forms of mycotic rhinitis reported in horses is conidiobolomycosis caused by Conidiobolus (Entomophthora) coronata. Conidiobolomycosis was previously classified as a form phycomycosis and is currently classified as a form of zygomycosis. 1,18,24 Other fungal and oomycotic pathogens that have been associated with rhinitis and sinusitis in horses include Aspergillus spp., Pseudallescheria boydii, Cryptococcus neoformans, and Pythium insidiosum. 6,14,29,30
This study was stimulated by receipt of a surgical pathology accession from a horse in Florida with chronic obstructive rhinitis that had unique fungal histomorphology associated with granulomatous inflammation. Because additional material was unavailable for fungal culture and/or polymerase chain reaction (PCR) on fresh tissue, we extracted DNA from paraffin-embedded tissue blocks and used PCR with panfungal primers for short, standardized, and highly diverse DNA sequences and DNA sequencing (DNA barcoding) 13 to identify the fungal organism. 20,36 Since fungal diversity and abundance are reported to be highest in tropical and subtropical climate zones, 12 we postulated that a histopathology and PCR/DNA sequencing study of archived Florida cases of equine rhinitis would identify previously unrecognized fungal and oomycotic pathogens responsible for obstructive rhinitis in horses. The objectives of this study were to use histopathology and PCR/DNA sequencing to characterize cases of mycotic and oomycotic rhinitis and sinusitis in Florida horses and to evaluate whether PCR/DNA sequencing on surgical pathology specimens would be a useful method for etiologic diagnosis when nonfixed material for culture and PCR diagnostics was unavailable to the diagnostic laboratory.
Methods
Computerized surgical pathology accession files were searched from 1995 through 2016 as previously described 8 using search terms including rhinitis, sinus, fungal, fungus, pythium, hyphae, and granulomatous. Further report analysis as well as evaluation of archived slides and paraffin blocks resulted in identification of 51 cases of rhinitis and 1 case of sinusitis accessioned from horses throughout subtropical and tropical Florida locations that could be studied by histomorphology and PCR/DNA sequencing. Fungal culture results were available for 3 cases. Besides the 52 cases included in this study, additional cases were identified. However, insufficient paraffin-embedded tissue was available for DNA extraction in these additional cases.
Cases were categorized from the submission information based on whether they came from horses in subtropical or tropical areas. Florida has a humid subtropical climate in the northern and central regions of the state. South Florida has tropical climate zones (ie, Köppen-Geiger climate zones equatorial rainforest, equatorial monsoon, and equatorial savannah) 15 south of a line from near Fort Myers Beach on the west coast to Jupiter on the east coast.
Histomorphologic classification of the cases was initially made into 4 groups. Group 1 had cases with features of zygomycosis/pythiosis 1,18,29 with morphologic features in lesions of large ghost hyphae with nonparallel walls that generally failed to stain with hematoxylin, stained well with Gomori methenamine silver (GMS), and stained variably with periodic acid Schiff (PAS). Hyphae had infrequent septation and 90° branching and were surrounded by eosinophilic sheaths (Splendore-Hoeppli–like reactions). Tissue reactions also included heavy eosinophil infiltration and variably sized areas of necrosis (aka “kunkers”). Group 2 cases had features of phaeohyphomycosis, including pigmented septate hyphae, chlamydospores, and conidia with longitudinal and transverse septation (muriform conidia). 3,23 Fibrinopurulent to pyogranulomatous inflammatory reactions were associated with the pigmented fungi. Group 3 cases had features of Aspergillus sp., including septate hyphae with parallel walls and with acute, dichotomous branching. 3 Conidial heads and conidia were found in superficial areas of the inflammatory exudate that included fibrinonecrotic and pyogranulomatous inflammatory reactions. Group 4 cases were fungal infections with nonpigmented, septate hyphae and yeast-like or chlamydospore-like structures and granulomatous inflammatory reactions. 3
Characterization of Fungal Structures in Tissue Sections
Initial morphological assessment was performed on sections stained with hematoxylin and eosin (HE), GMS, and PAS stains using standard reference criteria for fungal pathogens. 3,28 Many cases also had Gram stains for bacteria. Enlarged or swollen round to ovoid fungal structures with or without thick walls and associated with and sometimes attached to hyphae were initially described as conidia or chlamydospores. Diameter and other dimensions of hyphae, conidia, and chlamydospores and lesion features were measured in photomicrographs calibrated with a stage micrometer with ImageJ software (National Institutes of Health, public domain). For organisms for which PCR and DNA sequencing permitted identification of the genus or genus and species, specific structure names from the literature were used, including chlamydospores, chlamydoconidia, conidia, and conidial chains. 16,17,21,23,26,33
PCR and DNA sequencing
Paraffin blocks were sectioned, and DNA was extracted from sections with a Qiagen formalin-fixed and paraffin-embedded (FFPE) QIAmp DNA extraction kit (Qiagen GmbH, Hilden, Germany) according to kit directions as described. 8 All DNA samples were processed by PCR with panfungal ITS3, ITS4 primers as described. 20,36 DNA samples not reacting with ITS3-ITS4 primer sets were processed for 28 S ribosomal DNA primers. 9 All group 1 cases negative for ITS3, ITS4 primers were also processed with oomycete primers 25 and also processed with P. insidiosum–specific primers. 10 Finally, 10 extracted DNA samples suspected of being oomycete positive were tested by PCR at the University of Florida Veterinary Diagnostic Laboratories by 2 methods optimized for Pythium and Lagenidium in fresh clinical samples. 27,34 The quality of the 10 DNA samples was also tested with β-actin PCR primers.
PCR primers used and the product size of PCR reactions are detailed in Table 1 following published methods. 9,10,25,36 Briefly, Taq Master mix (2×; Genescript, Piscataway, NJ) was diluted with nuclease-free water. Final concentrations of ITS300 and ITS4 primer sets were 0.4 μM, and all the remaining primer sets were used at 0.25 μM. DNA template concentrations were <50 ng/μL. The PCR conditions for the ITS3 and ITS4 were initial denaturation in the thermocycler at 95°C for 5=minutes. This was followed by 40 cycles of 94°C for 1 minute, 51°C for 1 minute, and 72°C for 1 minute. The final extension was at 72°C for 10 minutes. The PCR conditions for the 28 S were initial denaturation in the thermocycler at 94°C for 4 minutes. This was followed by 40 cycles of 94°C for 1 minute, 52°C for 1 minute, and 72°C for 1 minute. The final extension was at 72°C for 10 minutes. For PCR with ITS300 and ITS4, initial denaturation in the thermocycler was at 95°C for 15 minutes. This was followed by 35 cycles of 95°C for 30 seconds, 55°C for 30 seconds, and 72°C for 1 minute. The final extension was at 72°C for 10 minutes. The PCR conditions for the PI1 and PI2 were initial denaturation in the thermocycler at 95°C for 5 minutes. This was followed by 35 cycles of 95°C for 45 seconds, 64°C for 30 seconds, and 72°C for 30 seconds. The final extension was at 72°C for 10 minutes. All PCR cycles were held at 4°C after completion. PCR products were detected in 2% agarose electrophoretic gels. PCR products were purified using Qiagen QIAquick PCR Purification kits and USB ExoSAP-IT PCR Product Cleanup kits (Santa Clara, CA) according to kit instructions. DNA sequencing was completed using Sanger method by GeneWiz (South Plainfield, NJ). DNA sequences were compared by homology search using BLAST (Basic Local Alignment Search Tool; National Center for Biotechnology Information [NCBI], National Institutes of Health, Bethesda, MD) and by analysis in the DNA sequence database maintained by the International Society for Human and Animal Mycology (ISHAM-ITS database) to complete DNA barcoding. Organism identification was based on the highest percentage identity for the DNA sequence in either NCBI or ISHAM-ITS databases. DNA extracted from paraffin-embedded cases of Aspergillus terreus (canine renal aspergillosis) and P. insidiosum (equine cutaneous pythiosis and canine respiratory pythiosis) were used as positive controls. Template-free samples and samples of DNA extracted from blocks of cutaneous and subcutaneous equine sarcoids 8 were used as negative controls. For 10 samples from suspected zygomycosis/pythiosis case that had yielded negative PCR reactions using 4 primer sets for fungi and oomycetes, PCR assays for equine β-actin DNA were run to assess isolated DNA quality and the possibility of PCR inhibitors in the samples. Primers designed with Primer-BLAST (NCBI) for equine β-actin were run with initial denaturation in the thermocycler at 94°C for 5 minutes. This was followed by 40 cycles of 94°C for 1 minute, 55°C for 1 minute, and 72°C for 1 minute. The final extension was at 72°C for 10 minutes. PCR products were detected in 2% agarose electrophoretic gels.
Primers Used for Polymerase Chain Reaction in This Study.
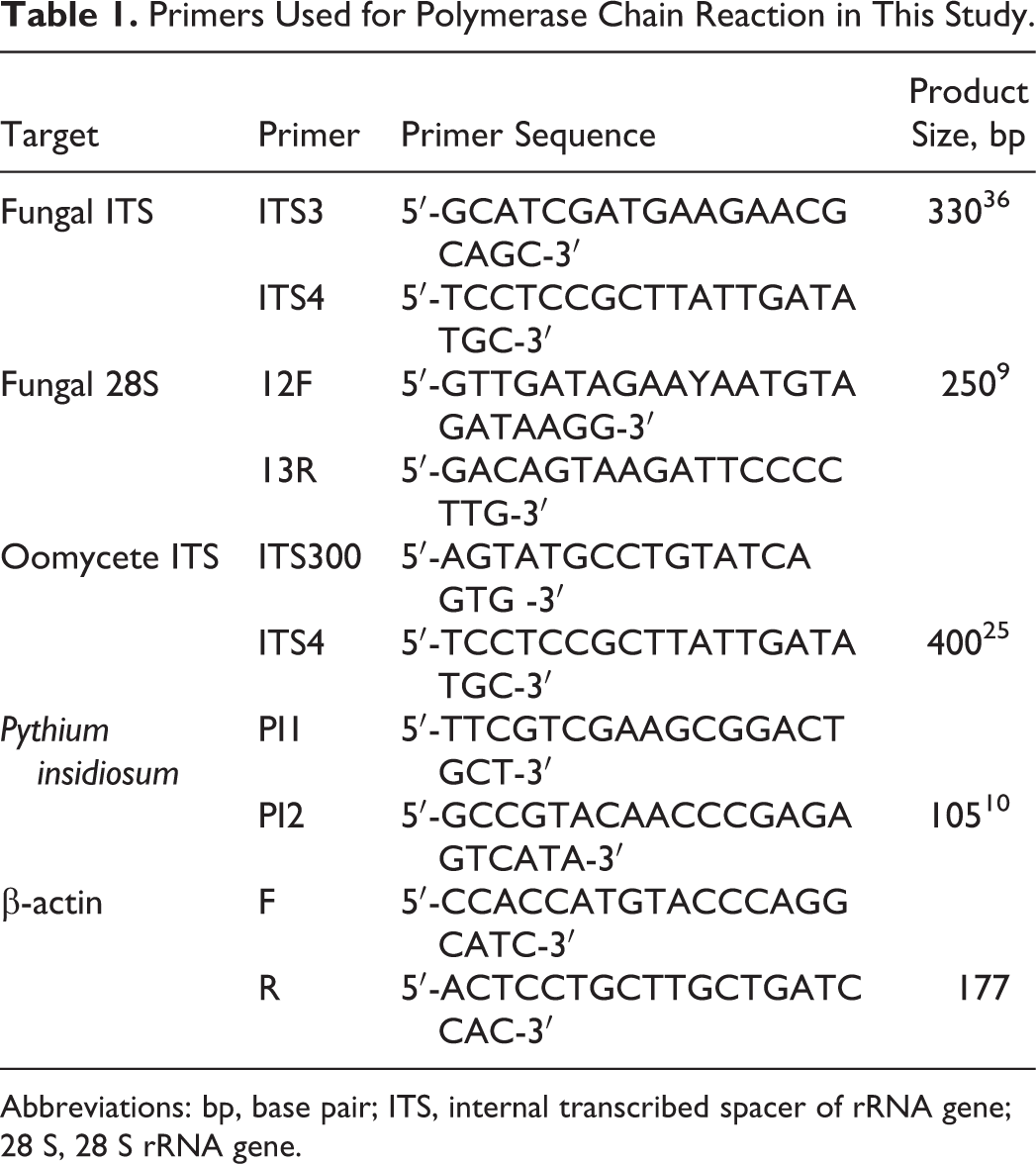

Abbreviations: bp, base pair; ITS, internal transcribed spacer of rRNA gene; 28 S, 28 S rRNA gene.
Results
A review of surgical pathology accessions from 1995 through 2016 identified 51 cases of rhinitis and 1 case of sinusitis with hyphae or other fungal structures in the lesions. Morphologic and histochemical assessment of cases yielded 31 cases of zygomycosis/pythiosis plus 1 mixed infection case, 16 cases of phaeohyphomycosis with 2 additional mixed infection cases, and 3 cases containing other fungal organisms. Morphologic evidence of Aspergillus sp. infection as a surface-oriented invader was found in 2 cases in which pigmented hyphae were found in deeper tissue lesions with more extensive inflammatory cell infiltrate. One case considered to be primarily involving zygomycosis/pythiosis had small numbers of pigmented hyphae in superficial lesions. Of the 52 cases of rhinitis and sinusitis, 11 had successful PCR amplification using fungal primers and DNA sequencing, in which the organism identified matched the histomorphology of the lesions (Table 2). Gram stains available for all 11 cases revealed no bacterial colonies, and these case were considered to be due to fungal infection. Three of the 52 cases had fungal isolates from cultures, and 2 of these cases had isolates consistent with the morphologic findings. Five of 52 cases of rhinitis/sinusitis came from horses in tropical regions of Florida (South Florida); of these, 3 of 5 cases were morphologically consistent with zygomycosis/pythiosis, and 2 were consistent with phaeohyphomycosis (cases 5 and 8 in Table 2). Forty-seven of 52 cases come from horses in subtropical regions of Florida (North, West, and Central Florida); the subtropical distribution of cases was zygomycosis/pythiosis, 28 of 47; phaeohyphomycosis, 16 of 47; and mixed, 3 of 47.
Case Data.
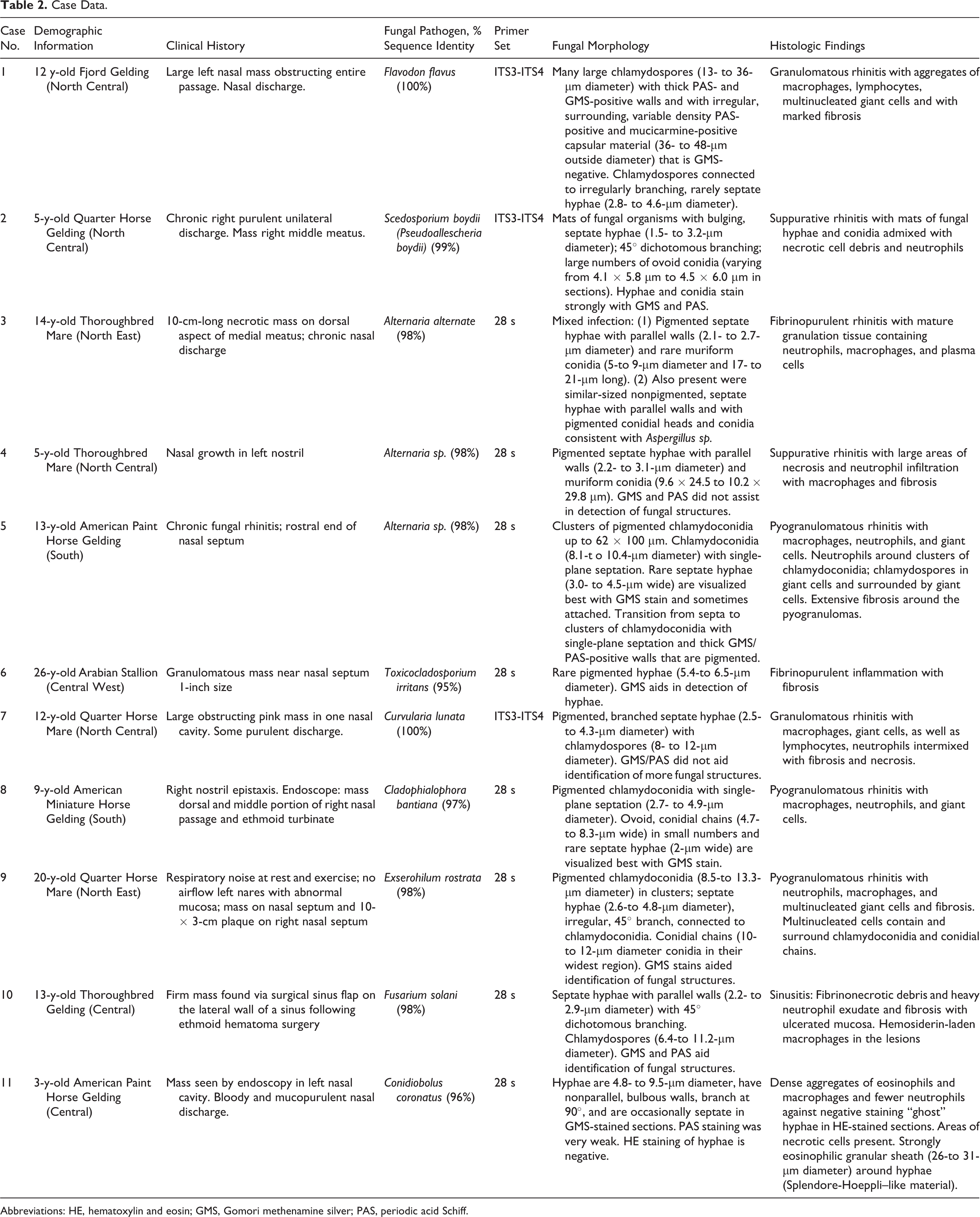
Abbreviations: HE, hematoxylin and eosin; GMS, Gomori methenamine silver; PAS, periodic acid Schiff.
The 11 cases with good agreement between histomorphology and DNA sequencing (Table 2) are summarized as follows.
Case 1: Flavodon flavus
Three biopsy samples, 3 × 3 × 1.5 mm to 9 × 5 × 3 mm, were submitted from a large mass obstructing the entire left nasal passage of a 12-year-old Fjord gelding. Within the biopsy sample, a granulomatous inflammatory infiltrate composed of lymphocytes, macrophages, and plasma cells as well as multinucleated giant cells with fibrosis was intermixed with aggregates of large chlamydospores, 13- to 36-μm diameter. The chlamydospores had thick, sometimes double-contoured, walls and were often surrounded by variable density eosinophilic material that was PAS and mucicarmine positive and consistent with capsular material (Figs. 1–5). Branching septate hyphae (2.8- to 4.6-μm diameter) were connected to the chlamydospores and distributed within capsular material and other areas of the inflammatory exudate (Figs. 2–4).
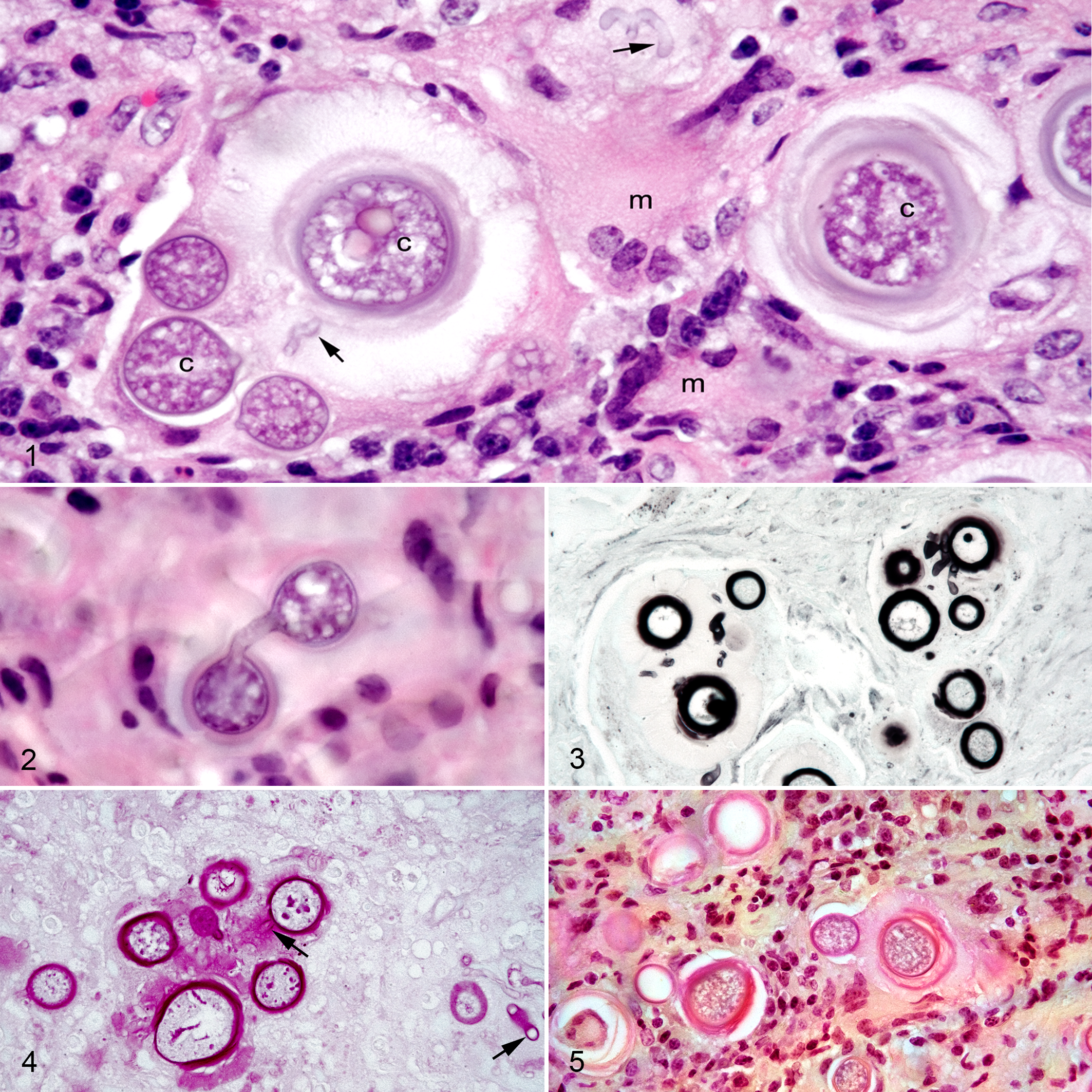
Flavodon flavus, fungal rhinitis, nasal mucosa, horse, Case 1.
Case 2: Pseudoallescheria boydii
The biopsy specimen from the right nasal mass revealed mats of septate hyphae (1.5- to 3.2-μm diameter) with parallel and sometimes bulging walls that were admixed with ovoid conidia varying in section from 4.1-μm diameter at the widest region and 5.8-μm long to 4.5-μm diameter and 6.0-μm long (Fig. 6). These fungal organisms were admixed with abundant necrotic cells and neutrophils. Hyphae and conidia were GMS and PAS positive.
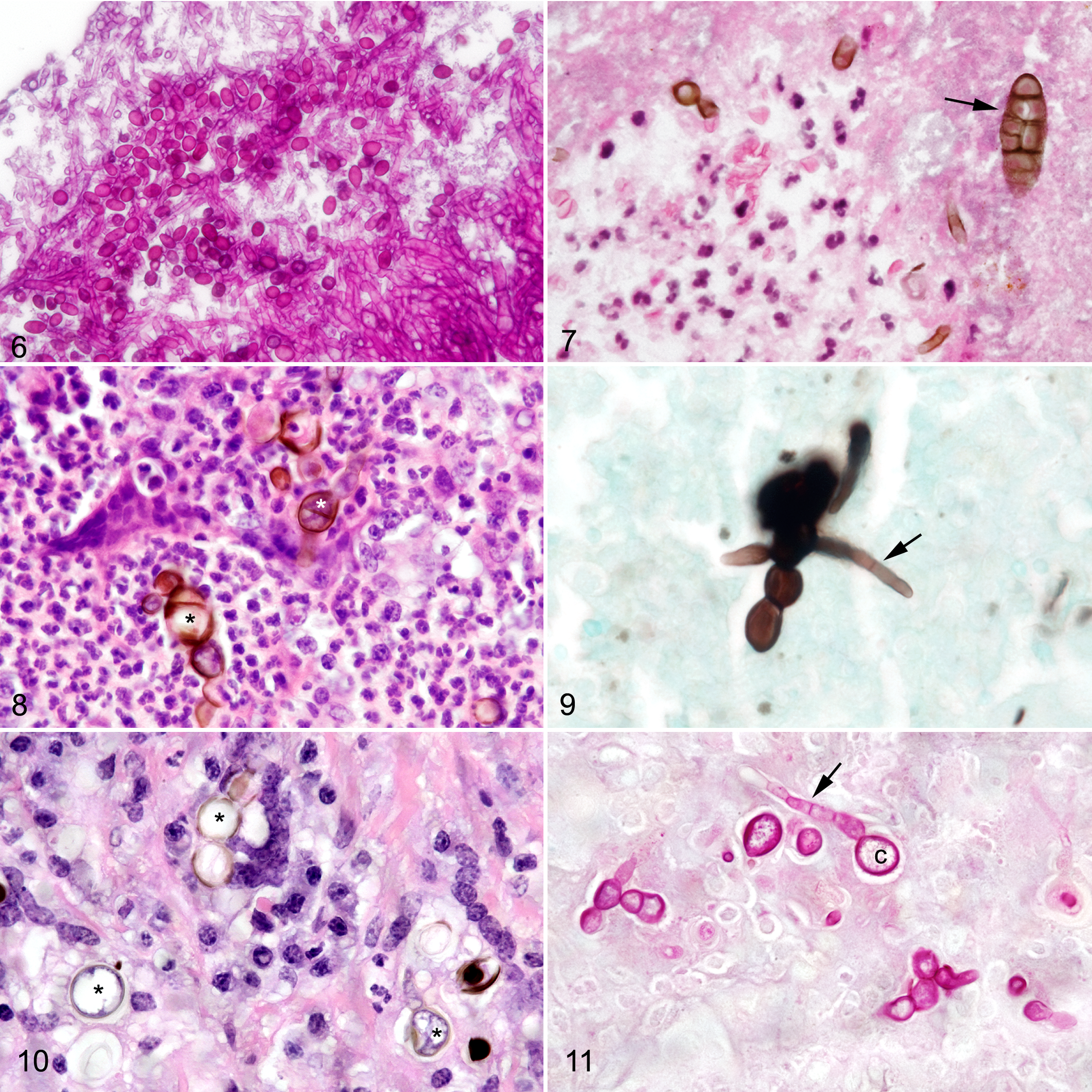
Fungal rhinitis, nasal mucosa, horse.
Cases 3 through 9 were all were associated with pigmented hyphae (phaeohyphomycosis).
Cases 3 Through 5: Alternaria sp. and Alternaria alternata
Biopsy specimens from cases 3 and 4 were similar with areas of neutrophil-rich exudate containing necrotic cells and/or fibrin associated with submucosal tissue composed of granulation tissue with increased collagen, fibroblasts, and macrophages. Small to moderate numbers of pigmented, septate hyphae with parallel walls of 2- to 3-μm diameter were present in the lesions. Elongate pigmented conidia of 5- to 10-μm diameter at their widest region and 17- to 30-μm long with both longitudinal and transverse septation (muriform conidia; Fig. 7) were present with hyphae in the exudate. Case 3 also had interspersed nonpigmented, septate hyphae with parallel walls and conidial head and conidia on the air interface of exudate that were suggestive of concurrent Aspergillus sp. infection. Case 5 from a horse in tropical South Florida differed from cases 3 and 4, since only rare pigmented septate hyphae were found in inflammatory exudate composed of neutrophils, macrophages, and multinucleated giant cells. Large clusters of pigmented thick-walled chlamydoconidia (8- to 10-μm diameter) with single-plane septation were present in the centers of the inflammatory exudate (Fig. 8). Chlamydoconidia stained strongly with GMS and PAS and similarly staining hyphae (3- to 4.5-μm diameter) connected with the chlamydoconidia (Fig. 9).
Case 6: Toxicocladosporium irritans
A 3- × 2- × 2-mm biopsy specimen from a 2.5-cm diameter mass near the nasal septum (Table 2) was composed of fibrous connective tissue and loose connective tissue with large amounts of fibrin, neutrophils, and small clusters of epithelioid macrophages. Necrotic cells were scattered in the exudate. Small numbers of pigmented hyphae (5.4- to 6.5-μm diameter) that stained with GMS were in the fibrinopurulent exudate.
Case 7: Curvularia lunata
In a large, obstructive nasal mass, pigmented chlamydospores (8- to 12-μm diameter) were surrounded by multinucleated giant cells, macrophages, and neutrophils along with scattered necrotic cells and increased density of collagen and fibroblasts (Fig. 10). Pigmented, branching, and septate hyphae (2.5- to 4.3-μm diameter) that stained well with both GMS and PAS were attached to the chlamydospores (Fig. 11).
Case 8: Cladophialophora bantiana
The biopsy specimen of the right nasal mass in a horse from tropical South Florida was composed of dense sheets of macrophages and neutrophils with dense aggregates of multinucleated cells around round pigmented chlamydoconidia (2.7- to 4.9-μm diameter) with thick walls and single-plane septation (Fig. 12). GMS stains revealed ovoid conidial chains (4.7- to 8.3-μm diameter in the widest areas) connected to small numbers of septate hyphae (2-μm diameter; Fig. 13).
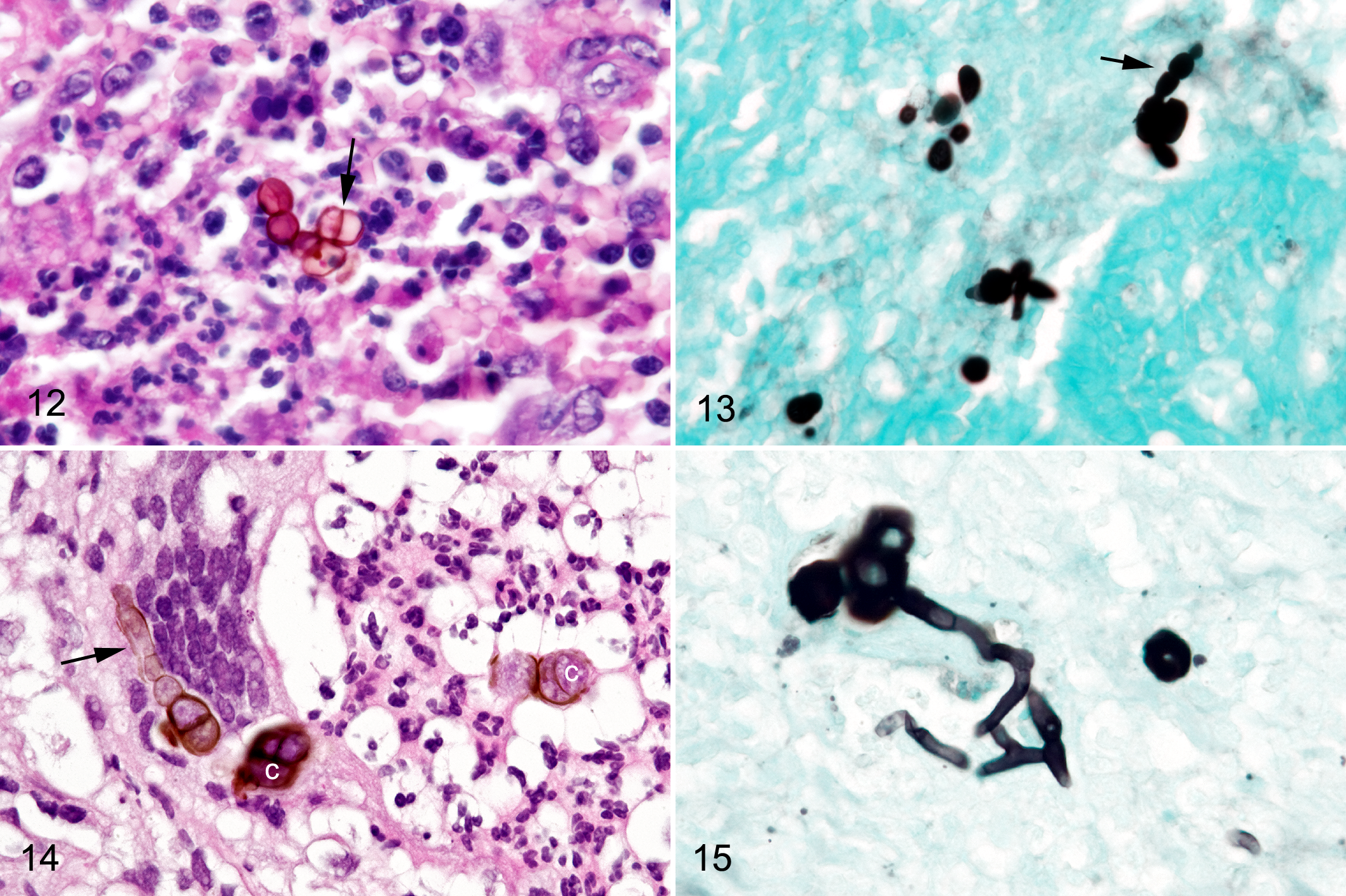
Fungal rhinitis, nasal mucosa, horse.
Case 9: Exserohilum rostrata
The biopsy sample from the nasal septal mass was composed of dense aggregates of neutrophils, macrophages, and multinucleated cells forming pyogranulomas around pigmented chlamydoconidia (8.5- to 13.3-μm diameter) arranged in clusters and singly, which were connected to pigmented, septate hyphae (2.6- to 4.8-μm diameter; Figs. 14 and 15). Conidial chains (10- to 12-μm diameter conidia in their widest region) were distributed throughout the inflammatory cell aggregates (Fig. 16). Macrophages and multinucleated giant cells surrounded fungal structures, and conidial chains, hyphae, and chlamydospores were present within multinucleated cell cytoplasm.
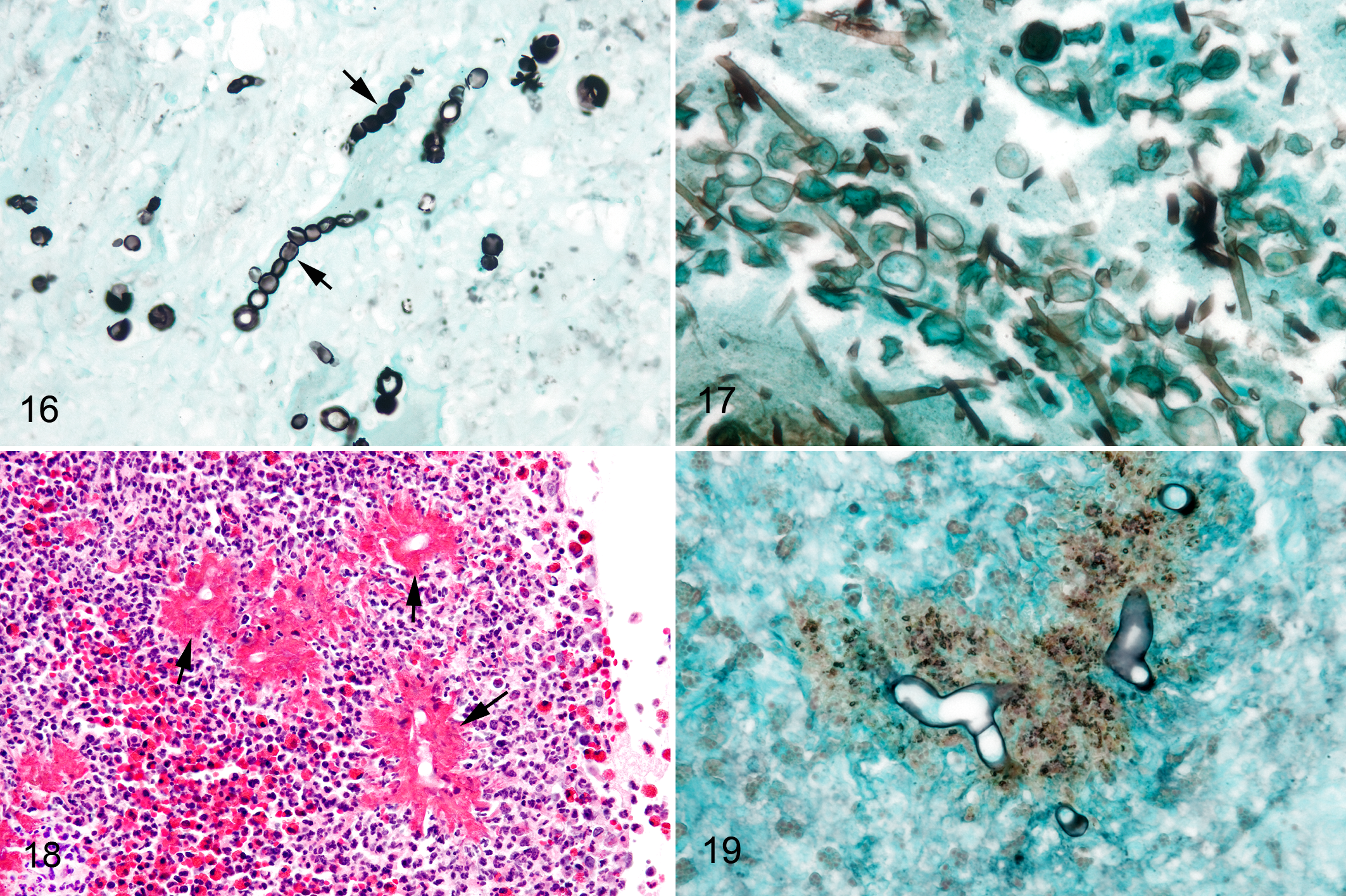
Fungal rhinitis, nasal mucosa, horse.
Case 10: Fusarium solani
A surface-oriented sinus lesion was biopsied in a horse that had undergone surgery for an ethmoid hematoma. The sample included an ulcerated surface of fibrous connective tissue covered by neutrophils and necrotic cell debris. Large numbers of septate hyphae (2.2- to 2.9-μm diameter) with parallel walls and dichotomous branching and interspersed chlamydospores (6.4- to 11.2-μm diameter) were present on the surface of the exudate (Fig. 17).
Case 11: Conidiobolus coronatus
The largest group of mycotic/oomycotic rhinitis cases in our surgical pathology archives (32/51) had histomorphologic features of conidiobolomycosis/pythiosis. Despite PCR testing by multiple fungal and oomycete primer sets (i.e., ITS3-ITS4; 28 S; ITS300-ITS4 and PI1-PI2), a PCR product was amplified in only 1 of 31 cases, and DNA sequencing matched C. coronatus. Lesions in this case were characterized by dense infiltrates of eosinophils and neutrophils surrounding ghost hyphae in HE-stained sections. Hyphae were surrounded by sheaths of eosinophilic granular material (Splendore-Hoeppli–like material; Fig. 18). Variable amounts of necrotic cells were in the lesions as well as smaller numbers of neutrophils and macrophages. Hyphae stained strongly with GMS (Figs. 19); had nonparallel, bulging walls; and were occasionally septate and branched infrequently at 90° angles. They stained variably with PAS. The hyphal diameter was 7.7 ± 1.3 μm (mean ± SD). There was a large, well-demarcated area of tissue necrosis (ie, kunker) in the biopsy sample. We had a second case of conidiobolomycosis that was culture positive for Conidiobolus sp. and was PCR negative. This biopsy sample was from a 17-year-old Quarter Horse gelding from West Central Florida with a large ulcerated and elevated lesion in the right nasal mucosa 15 cm from the external nares. The clinician suspected pythiosis. Histomorphology was similar to case 11, including a large area of well-demarcated necrosis. Hyphal size in this case was 6.1 ± 1.4 μm diameter.
All suspect zygomycosis/pythiosis cases were reviewed, and hyphal size was measured for each case. Average hyphal size varied from 3.0- to 8.9-μm diameter (6.4 ± 1.4 μm, mean ± SD). Of cases with large areas of tissue necrosis with hyphae within the areas of necrosis (kunkers), the hyphal size varied from 3.6- to 8.1-μm diameter (5.6 ± 1.7 μm, mean ± SD). No hyphae of 2.4-μm diameter or smaller (that usually characterize pythiosis cases) were found in any of the suspect cases.
Extracted DNA from positive control pythiosis cases were sequenced using both ITS300-ITS4 and PI1-PI2 primer sets (98%–99% sequence identity with P. insidiosum). For the control equine cutaneous pythiosis case, hyphal size varied from 1.9- to 4.0-μm diameter (2.8 ± 0.7 μm, mean ± SD). For the control canine respiratory pythiosis case, hyphal size varied from 1.9- to 5.4-μm diameter (3.1 ± 0.9 μm, mean ± SD). For both cases, many hyphae were found in lesions that were 2.6-μm diameter or smaller.
For 10 cases that were PCR negative and suspected of being pythiosis cases, DNA was tested in the University of Florida Veterinary Diagnostic Laboratories by 2 PCR methods optimized for Pythium and Lagenidium in fresh clinical samples. All test results were negative. The same 10 DNA samples were tested with β-actin PCR primers, and all resulted in 177-bp products indicating that good-quality DNA adequate to support PCR had been recovered from the samples.
Mismatching PCR/DNA Sequencing and Fungal Culture Results With Histomorphology
For 2 of 52 cases, PCR with ITS3-ITS4 primers and DNA sequencing identified plant DNA. In 1 mixed fungal infection case with histomorphologic evidence of phaeohyphomycosis and possible secondary Aspergillus sp. infection, Oenothera grandis (Showy Evening Primrose) DNA was amplified and sequenced. From a second case with histomorphologic features of zygomycosis/pythiosis, DNA from Quercus virginiana (Southern Live Oak) was sequenced. Primers for 28 S amplified a DNA sequence in a case of phaeohyphomycosis that was identified as Aspergillus sp.; this case had no histologic evidence of aspergillosis. Results for 3 fungal cultures were available in the archival studies. In addition to the conidiobolomycosis case previously discussed, a second culture recovered moderate growth of “dematiaceous mold” that was not otherwise specified from a case of phaeohyphomycosis that was PCR negative. The third case was another case of phaeohyphomycosis that had a fungal culture of heavy growth of Penicillium sp. and was PCR negative.
Discussion
The objectives of this study were to use histopathology and PCR/DNA sequencing (DNA barcoding) to characterize cases of mycotic and oomycotic rhinitis in Florida horses and to evaluate whether PCR/DNA sequencing on surgical pathology specimens would be a useful method for etiologic diagnosis when fresh material for culture and PCR diagnostics was not available. Use of PCR with panfungal primers and DNA sequencing allowed identification of a unique fungal pathogen (F. flavus) in an unusual case that was a stimulus for this study as well as identification of several other fungal pathogens not previously described as causing rhinitis in horses. However, PCR and DNA sequencing on FFPE resulted in overall low-yield results with only 11 of 52 cases (21%) being successfully sequenced for an organism compatible with the histomorphologic findings. Eight of 11 positively sequenced cases were with the 28 S primer pairs. No oomycotic pathogens (eg, Pythium sp. or Lagenidium sp.) were identified by molecular methods, and all oomycotic suspect cases had hyphal size most consistent with zygomycosis (conidiobolomycosis). Identification of mycotic and oomycotic infectious agents in equine rhinitis would most optimally be made by collecting fresh tissue for culture and PCR along with histopathology on fixed samples, followed by correlation of histopathologic findings with mycology and PCR/sequencing findings. In this retrospective study of 52 cases of rhinitis and sinusitis, only limited information was available on fungal culture results, since most samples were submitted from individual referring veterinarians throughout Florida over 21 years. Treatment and outcome information was also unavailable in the accessible records.
Application of DNA barcoding technology to FFPE biopsy specimens 9,36 facilitated identification of several previously unrecognized fungal pathogens from horses with rhinitis and sinusitis in subtropical and tropical regions of Florida.
The first previously unrecognized fungal pathogen was F. flavus, a lignicolous basidiomycete fungus present in decaying plant material. 21 F. flavus has not been described as a pathogen in vertebrate hosts. In the equine case in this study, it was associated with granulomatous rhinitis and large numbers of chlamydospores with thick walls and capsules was a striking histomorphologic finding. Chlamydospores/chlamydoconidia are enlarged, thick-walled structures formed with attachment to hyphae, either in line (intercalary) or at tips (terminal) in many fungal organisms. Chlamydospores are thought to serve as protecting survival forms of the fungus during dry or other unfavorable conditions. 16 It may be that the chlamydospores formed by F. flavus were in response to the granulomatous inflammatory response to infection and/or in response to other unfavorable fungal growth conditions in the horse. Chlamydospores and chlamydoconidia have previously been described in human respiratory and skin zygomycosis where hyphal organisms were close to an air surface. 4 The thick GMS- and PAS-positive walls of chlamydospores may be protective to the fungus. It is unclear whether the eosinophilic, PAS-positive, mucicarmine-positive, and GMS-negative material surrounding the chlamydospores represents fungal product or polysaccharide material produced by inflamed connective tissue in the host.
P. boydii has been previously described as a cause of fungal rhinitis in horses 14,30 and is associated with a wide array of infections in man including superficial, deep, and disseminated disease. 5 It is considered to be an opportunistic invader in previously damaged tissue as well as a primary pathogen.
The genus Alternaria includes saprophytes and pathogens of plants and animals, and individual species vary in morphology. 33,38 Alternaria has been associated with cutaneous infections in humans and in other animals as well as with keratitis, peritonitis, osteomyelitis, sinusitis, and onychomycosis in man. 7,23,31,33 Although muriform conidia characteristic of Alternaria were found in 2 cases, both pigmented hyphae as well as larger structures consistent with chlamydoconidia were found in the third case reported here.
T. irritans has not been described as a direct pathogen in animals, although it has been recovered commonly from skin microbiota of patients with atopic dermatitis. 39 It was the only pathogen identified in association with a nodular lesion in a case of rhinitis in our series. The sequenced DNA had 95% identity with T. irritans. This is a low sequence identity value, so another species of fungus may have been responsible. In addition, we cannot rule out that other factors or infectious agents contributed to the rhinitis.
Curvularia spp. are common in dead plant material and have been associated with keratitis, sinusitis, onychomycosis, endocarditis, skin infections, peritonitis, and systemic disease in man. 23 Curvularia infections in other animals have been described and include dermatitis with chlamydospores in the lesions of a dog. 11
C. bantiana is a common cause of cerebral phaeohyphomycosis in man and has been associated with infections in animals including endometritis in horses. 22 The pigmented chlamydoconidia in and surrounded by multinucleated macrophages resembled sclerotic bodies described in human chromoblastosis 3,17,23 except that only single-plane septation was found in the chlamydoconidia present in the equine rhinitis case.
E. rostrata is a common fungus isolated from cases of phaeohyphomycosis in man 23 and rarely in animals 37 and has been associated with contamination of drug preparations prepared by a compounding pharmacy in the United States with resultant widespread drug-related infections. 26
F. solani is an important plant pathogen and causes cutaneous, pulmonary, and disseminated infection in humans 19 as well as infections in animals including keratomycosis in horses. 35 Sinusitis in case 10 was a complication of previous surgery for ethmoid hematoma and may represent opportunistic infection as a sequela to previous sinus injury from the hematoma and surgery.
Both C. coronatus and P. insidiosum produce similar lesions of eosinophilic and granulomatous rhinitis in horses. Histomorphologic features that have been reported to distinguish conidiobolomycosis from pythiosis include smaller-diameter hyphae (2.6–6.4 μm) and larger areas of necrosis (ie, kunkers) in the latter. 18,29 No Pythium organisms were detected by PCR and DNA sequencing with 2 sets of oomycete primers that were used in the suspect cases. None of those cases had hyphae less than 2.6-μm diameter. It is most likely that all of our cases identified as zygomycosis/pythiosis suspects are zygomycosis cases caused by C. coronatus based on location and the relatively large hyphal size. 1,18 Although basidiobolomycosis cannot be completely excluded, it has been most widely associated with cutaneous lesions, rather than rhinitis in horses. 18
PCR on DNA extracted from FFPE and DNA sequencing yielded variable results depending on the organism group and tissue damage. For phaeohyphomycosis, sequencing was successful for 7 of 18 cases (39%). For zygomycosis/pythiosis suspect cases, 1 of 32 cases had successful PCR and sequencing (3%). For other nonpigmented fungal infections, PCR and sequencing were successful in 3 of 3 cases (100%). The lower quality of FFPE-derived DNA for PCR and sequencing compared with that from fresh tissue is well recognized. 8 However, the very low success in PCR/sequencing in the zygomycosis/pythiosis suspect cases compared with the other cases requires further consideration. The zygomycosis/pythiosis suspect cases are characterized by intense infiltrations of eosinophils and neutrophils around hyphae as well as the presence of ghost hyphae that have lost most of their hematoxylin-staining properties. It is reasonable to suggest that the release of hydrolytic enzymes (eg, nucleases, phospholipases, proteases) from eosinophils and neutrophils that account for the loss of hematoxylin-staining properties of hyphae in lesions also degrades a large percentage of fungal DNA that is available for PCR amplification.
The results of this study reinforce the principle that histopathology is important for correct etiologic diagnosis of mycotic rhinitis and sinusitis. Both fungal culture methods as well as PCR/DNA sequencing can lead to erroneous organism identification. One of 3 fungal culture results (33%) identified an irrelevant Penicillium sp. organism that was probably recovered from the surface of a case of granulomatous rhinitis due to a pigmented fungus (phaeohyphomycosis). Panfungal PCR primers can amplify DNA other than that from pathogenic fungi in a lesion. ITS3-ITS4 primers identified plant material in 2 of the cases. In another case of phaeohyphomycosis, 28 S primers identified a surface Aspergillus sp. sequence unrelated to the pathologic process.
One rationale for conducting this study was that we postulated that horses in the environment of subtropical and tropical Florida might have fungal and oomycotic forms of rhinitis and sinusitis that had not been previously described. Novel fungal pathogens were identified in this Florida-based study. Combined morphologic and molecular findings demonstrate that most of our mycotic rhinitis/sinusitis cases were due to zygomycosis (most likely conidiobolomycosis) in 62% of cases and phaeohyphomycosis in 35% of cases. Based on published reports, we anticipated finding many cases associated with Aspergillus spp. 14,30 However, we found morphologic evidence of superficial aspergillosis in only 2 cases, and both of these had deeper lesions of severe inflammation with the presence of pigmented fungi (phaeohyphomycosis). Five of 52 (9.6%) mycotic rhinitis/sinusitis cases in this study came from horses in tropical South Florida. The cases from tropical Florida had a similar predominance of zygomycosis and phaeohyphomycosis to that from subtropical areas of Florida. Although there are insufficient data from the current study to assess whether horses in tropical Florida have greater risks for nasal and sinus fungal infection than horses in subtropical Florida, it is worthwhile to observe that the 9.6% of the cases in this study came from tropical South Florida, where less than 5% of the horses in Florida are present. The overwhelming majority of horses in Florida are in subtropical North and Central Florida. 32
In conclusion, histomorphology coupled with PCR with panfungal primers and DNA sequencing (DNA barcoding) facilitated identification of several previously unrecognized fungal pathogens associated with rhinitis and sinusitis in horses. Zygomycosis and phaeohyphomycosis were the most common forms of fungal/oomycotic rhinitis in the biopsy samples submitted by veterinarians from Florida horses.
Footnotes
Acknowledgments
Pathology residents and faculty who also contributed to the initial pathologic workup of the 11 cases that are presented in detail in this report are K. Bischoff, J. Conway, M. Dark, C. Detrisac, E. Epperson, L. Farina, R. LaRock, R. Meisner, F. Origgi, B. Sheppard, S. Terrell, E. Uhl, and W. Zoll.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
