Abstract
Purpose
To investigate the association between Syndecan-2 gene methylation and protein expression dynamics in colorectal cancer and evaluate its clinicopathological significance.
Methods
We analyzed 197 tissues (96 colorectal cancer, 64 adenoma, and 37 normal tissues). Syndecan-2 methylation was quantified using quantitative methylation-specific polymerase chain reaction, and protein expression was evaluated via immunohistochemistry.
Results
Syndecan-2 methylation frequency was significantly higher in colorectal cancer (81.25%) and adenoma (64.06%) than in normal (5.41%) tissues. It showed high diagnostic accuracy for colorectal cancer (area under the curve = 0.88). Although methylation status was unrelated to most clinical parameters, higher methylation levels were correlated with poor tumor differentiation (p = 0.022). Syndecan-2 protein was aberrantly expressed in tumor stroma but lacked clinical correlations. A significant positive correlation between methylation and protein expression was observed across the sample (p < 0.05); however, this association was not maintained within the colorectal cancer subgroup specifically, showing no intrasample concordance (Kappa statistic = −0.046).
Conclusion
Syndecan-2 methylation serves as a highly sensitive epigenetic biomarker for the early detection of colorectal cancer. The findings reveal a complex and heterogeneous relationship between Syndecan-2 methylation and its protein expression during colorectal cancer progression, suggesting stage-specific regulatory mechanisms. Further multiomics studies are warranted to elucidate the underlying functional dynamics.
Introduction
Colorectal cancer (CRC) is a common and lethal malignancy; however, early diagnosis can increase 5-year survival rates from <10% to >90%, 1 underscoring the critical importance of effective screening. However, population heterogeneity must be considered in the pursuit of early detection. For instance, older individuals constitute a significant proportion of CRC patients. Although advanced age may increase treatment complexity, there is no consensus on its independent effect on survival, as their prognosis is largely confounded by factors such as disease stage at presentation, tumor location, pre-existing comorbidities, and treatment modalities. 2 Furthermore, evolving epidemiological landscapes, such as the recognition of a “CRC-like” subtype among cancers of unknown primary (CUP), may further nuance our understanding of incidence trends. 3 Despite the clear value of early detection, the majority of patients are still diagnosed at advanced stages due to the subtlety of early symptoms. International evidence has shown that population-based screening significantly reduces CRC mortality;4,5 however, current screening methods have notable limitations. Fecal occult blood tests lack sensitivity, while colonoscopy, although considered the gold standard, has low compliance rates due to its invasiveness, high cost, and potential associated complications.
Recent advances in epigenetic-based stool DNA methylation testing have introduced promising strategies for early CRC detection, with Syndecan-2 (SDC2) emerging as a widely studied biomarker. SDC2, located on chromosome 8q22.1, encodes a transmembrane proteoglycan that was initially identified in lung fibroblasts. 6 SDC2 is overexpressed in various tumors 7 and constitutively expressed in normal colonic mesenchymal cells; 8 however, its promoter hypermethylation is recognized as an early molecular event in CRC carcinogenesis. This epigenetic alteration can be reliably detected in noninvasive specimens such as stool and blood9–12 and promotes tumor progression by influencing epithelial–mesenchymal transition (EMT), angiogenesis, and key signaling pathways through interactions with vascular endothelial growth factor (VEGF), fibroblast growth factor (FGF-2), and other receptors.13–15 Notably, novel assays targeting two distinct methylated CpG islands within SDC2 have demonstrated enhanced sensitivity for CRC detection. 16 Despite multitarget approaches, such as combined analysis with SFRP2 methylation, which can further improve sensitivity, 17 their increased technical complexity and higher cost limit feasibility for large-scale implementation compared with single-gene assays.
The present study systematically evaluated SDC2 methylation levels and protein expression as well as their association across CRC tissues, colorectal adenomas (including low- and high-grade intraepithelial neoplasia), and paired normal tissues. Our findings highlight dynamic regulatory changes during malignant transformation and provide important insights to guide the development of molecular diagnostics and targeted therapies for CRC.
Materials and methods
Clinical data and specimens
In total, 197 samples, including CRC (specifically adenocarcinoma, with other subtypes excluded), colorectal adenoma, and paired adjacent normal colorectal tissues, were collected from 197 participants at the Quanzhou First Hospital affiliated to Fujian Medical University between January 2018 and December 2019. All cases were confirmed using either preoperative biopsy or postoperative pathological examination. Based on the pathological diagnosis, the samples were categorized into three groups: CRC (n = 96); colorectal adenoma (n = 64, including 33 low-grade and 31 high-grade adenomas); and normal colorectal tissue (n = 37) groups. General information (sex and age) and data on relevant clinicopathological parameters, including tumor differentiation, tumor–node–metastasis (TNM) stage, tumor location, maximum tumor diameter, presence or absence of lymph node metastasis, vascular invasion, and distant metastasis, were collected and recorded. Ethical approval was obtained from the Institutional Review Board of Quanzhou First Hospital, affiliated with Fujian Medical University (Approval No. [2023] K013; 16 March 2023).
Immunohistochemical (IHC) analysis
All 197 collected samples were paraffin-embedded. IHC staining was performed using a polyclonal anti-SDC2 antibody (1:400 dilution, #DF6630; Affinity Biosciences Co., Ltd., China). Tumor cells and SDC2-positive cells were counted under a microscope at 200× magnification. IHC staining was evaluated using a four-tier scoring system for staining intensity; a score of 0 indicated no staining; 1, weak staining; 2, moderate staining; and 3, strong staining. The percentage of positive cells was graded as follows: grade 0, no positive cells; grade 1, ≤25% positive cells; grade 2, 26%–50% positive cells; grade 3, 51%–75% positive cells; and grade 4, >75% positive cells. The final SDC2 expression level was calculated by summing the intensity and percentage scores. IHC results were categorized as negative/low expression if the total score was <4 and as or medium-to-strong expression if the total score was ≥4. 13
DNA extraction and bisulfite conversion
Genomic DNA was extracted from paraffin-embedded tissue sections using a commercial DNA extraction kit (Beijing Tiangen Biotechnology Co., Ltd., China) according to the manufacturer’s instructions. Bisulfite conversion and purification of the extracted DNA were then conducted using a DNA bisulfite conversion kit from the same manufacturer. For methylation analysis, both negative and positive controls from the Human SDC2 Gene Methylation Detection Kit (Creative Biosciences Co., Ltd., China) were included. The negative control consisted of unmethylated Caco-2 cell DNA, while the positive control contained methylated HCT116 cell DNA.
Real-time quantitative methylation-specific polymerase chain reaction (PCR) (qMSP)
PCR amplification for the test samples was performed in a total reaction volume of 22 μL, comprising 2 μL of bisulfite-converted DNA template and 20 μL of PCR master mix (Creative Biosciences Co., Ltd., China). For quality control reactions, the total volume was 30 μL, including 10 μL each of negative and positive control DNA and 20 μL of the same PCR master mix. ACTB was used as the reference gene. After thorough mixing and brief centrifugation, real-time qMSP was performed using a Bio-Rad PCR system (USA). Methylated SDC2 was detected using a 6-carboxyfluorescein (FAM)-labeled fluorescent probe, while ACTB was detected using a Texas Red-labeled probe. SDC2 methylation levels were quantified using the percentage of methylated reference (PMR) method, calculated as PMR = 2−ΔΔCt, where ΔCt = (Ct value of methylated SDC2) − (Ct value of ACTB). The criteria for interpreting methylation status are provided in Table S1 and detailed in Supplementary File S1.
Statistical analyses
Continuous variables were analyzed using the independent samples t-test, Wilcoxon rank-sum test, Kruskal–Wallis H test, or one-way analysis of variance (ANOVA), as appropriate. Results are presented as medians with interquartile ranges (M (P25–P75)). Categorical variables were compared using the χ2 test or Fisher’s exact test and were reported as frequencies (%). Diagnostic performance was assessed by constructing receiver operating characteristic (ROC) curves and calculating the area under the curve (AUC) with 95% confidence interval (CI). The association between SDC2 methylation levels and protein expression was evaluated using Spearman’s rank correlation analysis. All statistical analyses were conducted using Statistical Package for Social Sciences (SPSS) software (version 25.0, IBM Corporation, Armonk, NY) and GraphPad Prism (version 9, GraphPad Software Inc., USA). A two-tailed p value <0.05 was considered statistically significant.
Results
Clinical characteristics of study participants
In total, 197 samples were included in this study and categorized into 4 groups based on pathological diagnosis: 96 as CRC, 33 as low-grade adenomas (LGAs), 31 as high-grade adenomas (HGAs), and 37 as adjacent normal colorectal tissues.
As shown in Table 1, male predominance was noted in the CRC, HGA, and LGA groups, although sex distribution was comparable among the groups (p > 0.05). Significant age differences were observed (p < 0.05), with the LGA group comprising the youngest patients. Most lesions were located in the left colon (122 out of 160, 76.25%). Tumor size ranged from 3 mm to 10 cm, with significant variations among groups (p < 0.05); CRC specimens exhibited the largest volumes. In the CRC group, 57.29% of the patients were male. The CRC cohort had a nearly equal distribution of early-stage (I/II, 47.92%) and late-stage (III/IV, 52.08%) disease, with the majority of tumors being moderately to poorly differentiated (83.34%).
Baseline clinicopathological characteristics of study cohorts (n = 197).

encompassing the cecum, ascending colon, and transverse colon.
encompassing the region from the spleen flexure to sigmoid colon and rectum.
TNM: tumor–node–metastasis; HGA: high-grade adenoma; LGA: low-grade adenoma; CRC: colorectal cancer.
SDC2 gene methylation positivity rates and composition ratios across groups
As shown in Table 2 and Figure 1, the highest SDC2 methylation positivity rate was observed in the CRC group (81.25%, 78/96), followed by the colorectal adenoma group (64.06%, 41/64); the normal colorectal tissue group had the lowest rate (5.41%, 2/37). The differences in the positivity rates between the three groups were statistically significant (H test, p < 0.001). Pairwise χ2 test comparisons further confirmed significant differences between the CRC group and both colorectal adenoma (p < 0.05) and adjacent normal tissue (p < 0.001) groups.
Comparative analysis of SDC2 methylation positivity rates across study groups.

CRC: colorectal cancer; H: Kruskal–Wallis H test.
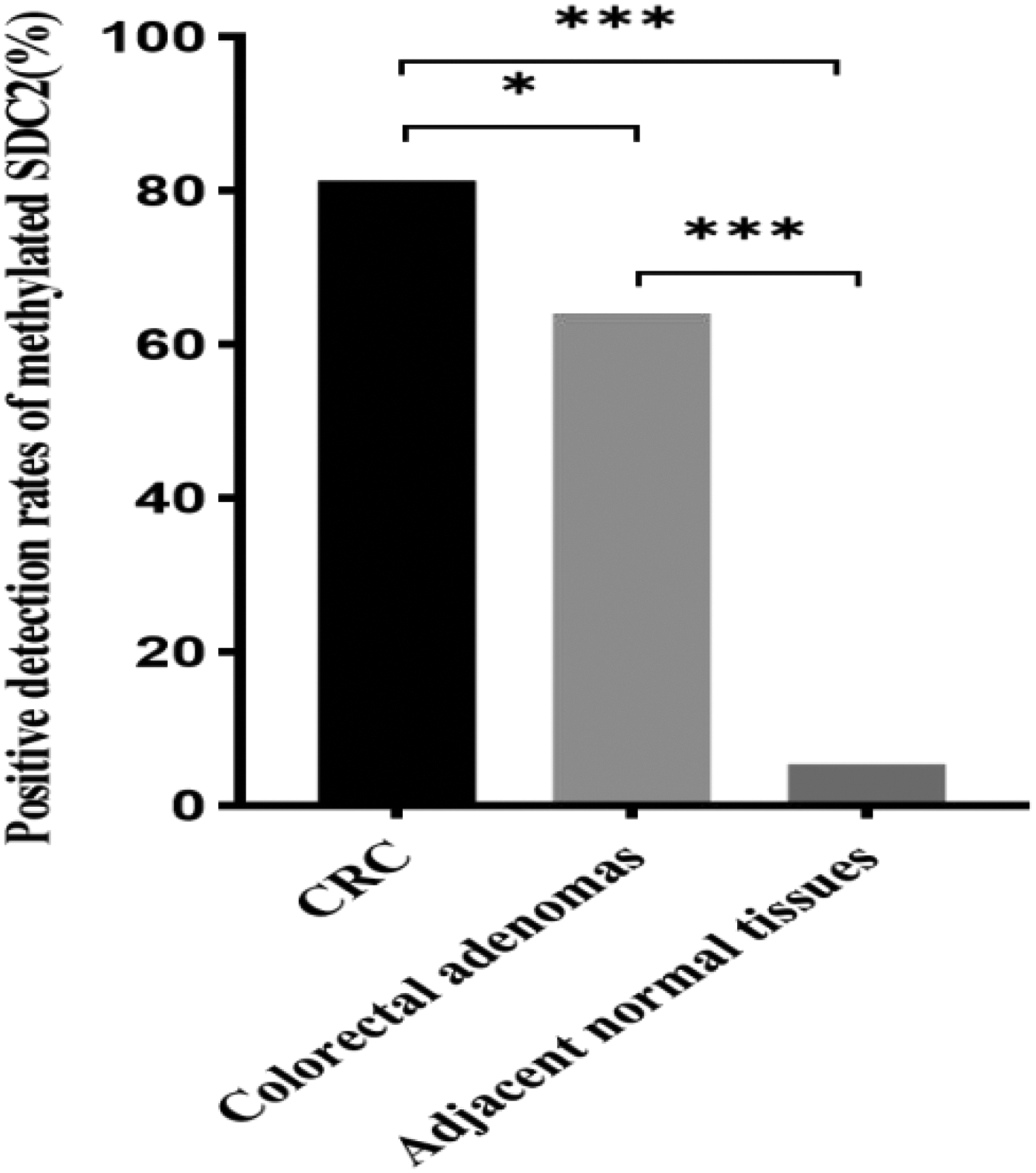
Detection rates of methylated SDC2 in CRC patients, colorectal adenoma subgroups (low-grade and high-grade), and adjacent normal tissues. Group comparisons were performed using χ2 tests (*p < 0.05, ***p < 0.001). CRC: colorectal cancer; SDC2: Syndecan-2.
Diagnostic performance of SDC2 methylation for CRC and colorectal adenomas
ROC curve analysis was performed to assess the diagnostic efficacy of SDC2 methylation in detecting CRC and colorectal adenomas. As shown in Figure 2, SDC2 methylation exhibited high diagnostic accuracy for both CRC (AUC = 0.88, 95% CI: 0.82–0.94) and colorectal adenomas (AUC = 0.80, 95% CI: 0.71–0.82), with significantly better performance for CRC detection (p < 0.001 for all comparisons).

Receiver operating characteristic (ROC) curve analysis of the diagnostic performance of methylated SDC2 detection. SDC2: Syndecan-2.
Association between SDC2 methylation and clinicopathological characteristics
Relationship between SDC2 methylation positivity and clinicopathological features
In the CRC cohort, the methylation status of SDC2 showed no significant correlation with age, sex, tumor location, tumor size, TNM stage, histological differentiation, lymph node metastasis, vascular invasion, or distant metastasis (all p > 0.05; Table 3).
Correlation of SDC2 methylation with clinicopathological features in CRC.

encompassing the cecum, ascending colon, and transverse colon.
encompassing the region from the spleen flexure to sigmoid colon and rectum.
TNM: tumor–node–metastasis; CRC: colorectal cancer.
Relationship between SDC2 methylation levels and clinicopathological features
Of the 197 samples, 177 yielded valid results (91 CRC, 57 colorectal adenoma, and 29 normal tissue samples). The median SDC2 methylation levels were 0.47, 0.24, and 0.00 in the CRC, colorectal adenoma, and normal colorectal tissue groups respectively, showing significant intergroup differences (p = 0.002). Post-hoc analysis revealed significantly higher methylation in CRC than in both colorectal adenoma and normal colorectal tissue groups (p < 0.05; Figure 3).

SDC2 methylation levels in colorectal cancer, adenomas, and normal control tissues.
In the CRC cohort (n = 78 methylation-positive cases, 81.25% positivity rate), samples were stratified by the median methylation level (0.47) into low- (n = 39) and high-methylation (n = 39) subgroups. Methylation levels were significantly associated with tumor differentiation (p = 0.022) but not with age, sex, tumor location, size, TNM stage, lymph node metastasis, vascular invasion, or distant metastasis (p > 0.05; Table 4).
Correlation between SDC2 methylation levels and clinicopathological characteristics in CRC.

encompassing the cecum, ascending colon, and transverse colon
encompassing the region from the spleen flexure to sigmoid colon and rectum
TNM: tumor–node–metastasis; CRC: colorectal cancer
Association between SDC2 protein expression and clinicopathological parameters
SDC2 protein localization was observed in both cytoplasmic and membranous patterns. In the CRC group, the expression intensity was heterogeneous, ranging from weak to strong staining. Two samples in the colorectal adenoma group showed negative staining, while normal tissues predominantly exhibited negative or weak expression. Notably, normal tissue sections displayed enhanced staining intensity in glandular structures adjacent to areas with prominent inflammatory infiltrates. Stromal cells (e.g. fibroblasts/myofibroblasts) showed variable expression levels, with notably stronger immunoreactivity in tumor budding foci and their surrounding stromal compartments compared with that in other regions (Figure 4).

Immunohistochemical profiles of SDC2 protein expression. (a) Colorectal carcinoma cases. (a1–a3) Immunohistochemical staining intensities in panels a1, a2, and a3 show strong positivity, moderate intensity, and weak positivity, respectively (400×). (b) Colorectal adenoma group. (b1) Low-grade tubular adenoma with partial weak staining (200×); (b2–b3) Low-grade (b2) and high-grade (b3) tubular adenomas showing strong positivity, contrasting with adjacent negative normal glands (100×). (c) Normal mucosa. (c1) Baseline weak expression (400×); (c2) Moderate intensity of staining with interstitial inflammatory reaction and focal lymphocyte aggregates (200×); (c3) Moderate intensity of partial staining with interstitial inflammatory reaction (200×). (d) Stromal expression patterns. (d1) Fibroblasts/myofibroblasts with moderate intensity (400×); (d2–d3) Panels (d2) and (d3) represent the same tissue section, showing stronger SDC2 immunoreactivity in tumor budding foci (d2) and adjacent peritumoral stromal cells than other stromal regions (d3) (400×); (d4–d5) Panels d4 and d5 similarly exhibit enhanced staining intensity in neoplastic budding clusters (d4) and their surrounding stromal compartments relative to background areas (d5) within the same section (400×). SDC2: Syndecan-2
IHC revealed no significant correlations between SDC2 protein expression levels and age, sex, tumor location, tumor size, TNM stage, differentiation status, lymph node metastasis, vascular invasion, or distant metastasis in CRC patients (p > 0.05 for all parameters; Table 5).
Correlation between SDC2 protein expression levels and clinicopathological characteristics in CRC.

encompassing the cecum, ascending colon, and transverse colon.
encompassing the region from the spleen flexure to sigmoid colon and rectum.
TNM: tumor–node–metastasis; CRC: colorectal cancer.
Relationship between SDC2 gene methylation level and SDC2 protein expression
Spearman’s correlation analysis was performed to examine the relationship between SDC2 methylation levels and SDC2 protein expression levels. The results showed a positive correlation between SDC2 methylation and SDC2 protein expression in the total sample (p < 0.05), while no significant association was observed in the CRC group (p > 0.05), as detailed in Table 6.
Correlation between SDC2 methylation and protein expression levels.
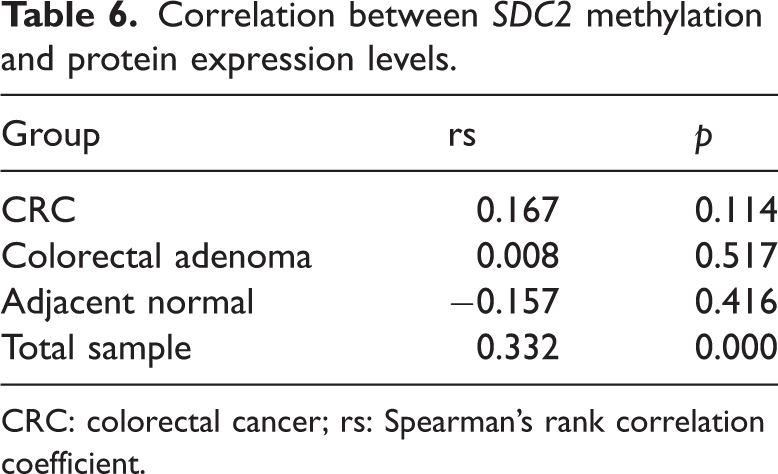
CRC: colorectal cancer; rs: Spearman’s rank correlation coefficient.
To further evaluate the concordance between SDC2 methylation levels and corresponding SDC2 protein expression in CRC, we stratified SDC2 methylation levels into high- and low-level groups based on the median methylation value, as previously described. Similarly, SDC2 protein expression was categorized into high-expression (≥4 points) and low-expression (<4 points) groups according to the total IHC score. Concordance was defined as either high SDC2 methylation with high protein expression or low methylation with low protein expression, while discordance was defined as high methylation with low protein expression or low methylation with high protein expression. The analysis revealed poor consistency between SDC2 methylation and protein expression levels in the CRC cohort, with a Kappa value of −0.046 (Table 7). The intrasample concordance between SDC2 methylation levels and SDC2 protein expression levels in the CRC group is described in Table S2 and further detailed in Supplementary File S1.
Intrasample concordance analysis of SDC2 methylation and protein expression levels in the CRC group.

CRC: colorectal cancer; SDC2: Syndecan-2.
Discussion
Active early clinical screening can effectively reduce the CRC incidence and improve survival rates in CRC patients. 18 However, maximizing this benefit for all patients requires consideration of population-specific barriers. Notably, older patients—a large subgroup—may face disparate outcomes; this is not because age dictates survival but because they have a higher risk of postoperative complications that can inappropriately limit their access to curative surgery, underscoring that treatment decisions should be based on comprehensive assessment rather than age alone. 2 This highlights the critical need for screening tools that facilitate early diagnosis across all demographics, thereby creating more opportunities for timely and appropriate intervention. Nonetheless, current screening methods have several significant limitations, making the identification of specific tumor biomarkers a promising approach for CRC screening. SDC2, one of the most extensively studied biomolecules, has emerged as a key focus in CRC pathogenesis. However, it is important to note that the adoption of any promising biomarker—including SDC2—into routine clinical practice involves considerable translational hurdles. These include challenges related to the standardization of assays, tumor heterogeneity, and equitable access, which are particularly pronounced for tumor-agnostic biomarkers. 19 The present study provides a comprehensive analysis of the role of SDC2 methylation and protein expression in CRC development, aiming to address some of these foundational questions, emphasize its potential value in clinical diagnosis, and understand molecular mechanisms.
Our results showed that the SDC2 methylation positivity rate was significantly higher in CRC tissues (81.25%) than in adenoma (64.06%) and normal tissues (5.41%) (p < 0.001), indicating tumor-associated methylation patterns that progressively increase with disease severity, consistent with previous studies.10–12 SDC2 methylation demonstrated excellent diagnostic performance for CRC (AUC = 0.88, 95% CI: 0.82–0.94), suggesting its potential as a valuable diagnostic biomarker. Notably, although adenomas exhibited lower methylation rates than CRC, their rates were significantly higher than those in normal tissues (p < 0.001), suggesting that SDC2 methylation is activated early in the adenoma-carcinoma sequence, thereby offering new insights for monitoring precancerous lesions.
The SDC2 methylation status showed no significant correlation with age, stage, or metastasis (p > 0.05), indicating consistent detection efficiency across different clinical characteristics. The noninvasive nature and high sensitivity and specificity of SDC2 methylation testing make it a promising complementary approach to colonoscopy for improving early CRC detection. Interestingly, tumors with high methylation levels exhibited worse differentiation (p = 0.022), suggesting that hypermethylation promotes dedifferentiation through epigenetic silencing of tumor suppressor pathways. 20 This finding supports SDC2’s potential oncogenic role, while its stage-independent nature suggests its utility in prognostic stratification.
SDC family members are predominantly expressed in stromal cells, with SDC2 being specifically detected in stromal-derived cells such as fibroblasts, smooth muscle cells, endothelial cells, and macrophages. 21 Through its extracellular domain, SDC2 binds to extracellular matrix components, activating the protein kinase C gamma (PKCγ)/ focal adhesion kinase (FAK)/ extracellular signal–regulated kinase (ERK) signaling pathway. This, in turn, induces matrix metalloproteinase-7 (MMP-7) expression, which facilitates matrix remodeling and enhances tumor invasiveness.22,23 All SDC family members contribute to cytoskeletal reorganization through specific receptor interactions, 24 playing a role in tumorigenesis. However, the significance of SDC2 protein expression in tumors remains debatable.
Here, we observed heterogeneous SDC2 expression in both cytoplasm and membranes of CRC cells, along with colocalization in the stroma, potentially reflecting its involvement in EMT or tumor–stroma crosstalk. 13 Although protein expression showed no significant correlation with clinicopathological characteristics (p > 0.05), its elevated expression in inflamed normal tissues suggests activation of inflammation-associated pathways, 25 offering insight into inflammation-mediated carcinogenesis.
Intriguingly, although the overall sample showed a positive correlation between methylation and protein expression, CRC specimens exhibited no significant association and demonstrated remarkably low intrasample concordance. Contrary to the conventional understanding of epigenetic silencing, promoter hypermethylation in CRC coincided with protein overexpression, in line with previous reports. 26 This paradoxical finding highlights the complexity of interactions between epigenetic modifications and functional phenotypes, challenging the traditional view of methylation as a mechanism for gene silencing.
The conventional paradigm posits that hypermethylation of CpG islands in tumor suppressor gene promoters suppresses gene expression, thereby facilitating tumorigenesis. Nevertheless, emerging evidence increasingly challenges this view. Yin et al. 27 previously used high-throughput sequencing to analyze the effects of methylation on 542 transcription factor binding events and found that approximately one-third of methylated genes exhibited enhanced transcription factor binding. Notably, CpG island methylation may paradoxically enhance gene expression depending on its genomic context and density, with methylation within gene bodies often associated with increased rather than decreased transcription.28,29
Notably, Smith et al. 30 systematically reviewed documented cases in which promoter hypermethylation was associated with transcriptional activation and proposed several possible mechanisms: (a) interference with the binding of repressive transcription factors;31,32 (b) recruitment of transcriptional activators; and (c) activation of alternative promoters. 33 Moreover, the aberrant overexpression of SDC2 protein observed in tumor stromal cells and inflamed regions suggests a role in remodeling of the tumor microenvironment. However, further investigation using single-cell sequencing or spatial transcriptomics is necessary to clarify these potential functions.
Conclusions
This study supports the clinical potential of SDC2 methylation as a diagnostic biomarker. The findings revealed the significant heterogeneity of its regulatory relationship with protein expression at the tumor stage, suggesting the complexity of epigenetic regulation and providing a theoretical basis for optimizing the molecular diagnosis of CRC and targeted interventions. Nonetheless, certain limitations should be acknowledged, including its single-center design and the lack of long-term prognostic data. Future research should aim to validate these findings in multicenter cohorts and, crucially, investigate the correlation between SDC2 status (methylation and/or protein expression) and patient outcomes such as disease recurrence and survival. Integrating multiomics approaches with functional assays will be essential to fully elucidate the dynamic regulatory mechanisms of SDC2 and assess its ultimate utility in liquid biopsy and personalized management paradigms for CRC.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605261420924 - Supplemental material for A study on the correlation between DNA methylation and protein expression levels of SDC2 in patients with colorectal cancer and its clinicopathological significance
Supplemental material, sj-pdf-1-imr-10.1177_03000605261420924 for A study on the correlation between DNA methylation and protein expression levels of SDC2 in patients with colorectal cancer and its clinicopathological significance by Yan Han, Jinling Yang, Hailong Wang, Yuru Xu, Weiqi Wu, Xin Chen and Yifeng Chen in Journal of International Medical Research
Footnotes
Authors’ contributions
Y.H. acquired, analyzed, and interpreted the data; performed statistical analyses; wrote the manuscript text; and prepared the figures and tables. J.Y. and H.W. conceptualized and designed the study as well as analyzed and interpreted the data. Y.C. participated in the review and revision of the paper and provided technical support. Y.X., W.W., and X.C. performed histologic examination and scoring. All authors read and approved the final version of the submitted manuscript.
Data availability
Data are available upon reasonable request.
Declaration of competing interests
The authors declare no conflict of interest.
Funding
This study was funded by the Quanzhou Science and Technology Program (Grant No. 2025QZNY071) and Startup Fund for scientific research, Fujian Medical University (Grant number: 2024QH1316).
Informed consent statement
Informed consent was obtained from all participants involved in the study.
Institutional review board statement
The study was conducted in accordance with the Declaration of Helsinki and was approved by the Institutional Review Board of Quanzhou First Hospital Affiliated to Fujian Medical University (Approval No. [2023] K013; 16 March 2023).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
