Abstract
Purposes
Mutations in the feline McDonough sarcoma–like tyrosine kinase 3 (FLT3) gene represent one of the most common genetic alterations in acute myeloid leukemia and are associated with an increased risk of relapse and poor prognosis. FLT3 inhibitors have significantly improved clinical outcomes in acute myeloid leukemia patients with FLT3 internal tandem duplication mutations. However, currently available FLT3 inhibitors often exhibit off-target effects, leading to numerous adverse reactions. Therefore, identifying more specific FLT3 inhibitors has important clinical significance.
Methods
We conducted high-throughput virtual screening for identifying small molecules that bind to the autoinhibited conformation of the FLT3 protein using the National Cancer Institute Developmental Therapeutics Program library. Then, we performed molecular screening of 159 small-molecule compounds targeting the autoinhibited conformation of FLT3. Following in vitro screening in 32D and 32D cells with FLT3 internal tandem duplication overexpression, TIP-20 was identified as the lead compound. TIP-20 was further evaluated in FLT3-internal tandem duplication mutation and FLT3 wild-type acute myeloid leukemia cell lines to assess its effects on apoptosis, cell cycle progression, downstream FLT3 signaling pathways, and kinase activity inhibition. The in vivo survival benefit of TIP-20 was evaluated using MV4-11 xenograft mouse models. RNA sequencing and Gene Set Enrichment Analysis were performed to further elucidate the antileukemic mechanisms of TIP-20.
Results
Cellular thermal shift assays confirmed the direct binding of TIP-20 to the FLT3 protein. TIP-20 inhibited FLT3 kinase activity, as demonstrated by kinase activity assays. In FLT3 internal tandem duplication mutation acute myeloid leukemia cell lines, TIP-20 suppressed cell proliferation, induced apoptosis, and caused G0/G1 cell cycle arrest. In addition, TIP-20 inhibited the phosphorylation of FLT3 and its downstream signaling components, including signal transducer and activator of transcription 5. Furthermore, TIP-20 reduced leukemia burden in the MV4-11 xenograft mouse model, with no significant differences observed in liver or kidney function between the treatment and control groups. Gene Set Enrichment Analysis indicated that TIP-20 enhanced or restored antigen presentation in MV4-11 cells.
Conclusions
TIP-20 is a novel FLT3 inhibitor with potent antileukemic activity. These findings suggest that TIP-20 may provide a new therapeutic option for leukemia patients.
Keywords
Introduction
Acute myeloid leukemia (AML) is one of the most common types of leukemia in adults. 1 It is a biologically heterogeneous malignancy, and its mutational landscape can evolve dynamically during disease progression. 2 Among recurrent genomic lesions, feline McDonough sarcoma–like tyrosine kinase 3 (FLT3) alterations are particularly common, occurring in approximately 30% of newly diagnosed patients. 3 FLT3 is a class III receptor tyrosine kinase characterized by five extracellular immunoglobulin-like domains, a juxtamembrane segment, and two intracellular kinase domains separated by a kinase insert region. 4 Mutations in the tyrosine kinase domain (TKD) occur in approximately 7% of cases and have demonstrated variable prognostic impact. 5 In contrast, internal tandem duplication (ITD) mutations are present in approximately 25% of patients and are widely regarded as key oncogenic drivers. 5 FLT3-ITD mutations result in ligand-independent receptor signaling, with downstream hyperactivation of the mitogen-activated protein kinase/extracellular signal-regulated kinase, phosphatidylinositol 3-kinase/protein kinase B, and signal transducer and activator of transcription 5 (STAT5) pathways. This signaling promotes uncontrolled leukemic proliferation, impaired hematopoietic differentiation, and resistance to apoptosis.6,7 Clinically, patients harboring FLT3-ITD mutations typically exhibit more aggressive disease, shorter overall survival, and an increased risk of relapse compared with FLT3-ITD–negative counterparts.8,9 FLT3-TKD mutations, particularly at residues such as D835 and Y842, occur within the kinase domain of FLT3. These mutations alter kinase conformation, increase ATP affinity, and enhance FLT3 kinase activity, thereby promoting downstream signaling. 10 FLT3-TKD mutations are associated with resistance to FLT3 inhibitors and poorer clinical responses, as structural alterations in the kinase binding site reduce the effectiveness of existing inhibitors. 11 A key distinction between FLT3-ITD and FLT3-TKD mutations lies in their differential impact on clinical outcomes. Patients with FLT3-TKD–positive AML demonstrate a better 10-year overall survival rate and cumulative incidence rate compared with patients with FLT3-ITD 12 –positive disease. Unlike FLT3-ITD mutations, which lead to sustained activation of the Janus kinase (JAK)/STAT signaling pathway, FLT3-TKD mutations enhance the activity of Src homology region 2 domain-containing phosphatase 1 (SHP1) and SHP2, which are negative regulators of JAK signaling. 13 This difference in downstream signaling may contribute to the more favorable prognosis observed in patients with FLT3-TKD AML.
Therapeutic strategies targeting FLT3 have advanced into the second generation, including gilteritinib, quizartinib (AC220), and crenolanib. Although these agents have improved patient outcomes, their efficacy and selectivity remain suboptimal.14–16 Despite these advances, current FLT3 inhibitors are limited by significant off-target and mechanism-associated toxicities that adversely affect their clinical utility. 17 Gastrointestinal disturbances, such as nausea, vomiting, and diarrhea, were among the most common nonhematologic adverse events observed with midostaurin in clinical trials. 18 Second-generation agents such as quizartinib and gilteritinib, although more FLT3-selective, exhibit notable toxicities. Gilteritinib can induce a differentiation syndrome resembling that observed in acute promyelocytic leukemia, an inflammatory complication caused by rapid leukemic cell maturation. 19 Quizartinib, in contrast, has a propensity to cause dose-dependent QT-interval prolongation, likely due to off-target inhibition of cardiac ion channels. 20 Both gilteritinib and quizartinib are also associated with substantial myelosuppression. In the ADMIRAL and QuANTUM-R studies, grade 3 or higher hematologic toxicities such as febrile neutropenia, anemia, and thrombocytopenia were frequently reported.16,21 Although generally manageable with supportive care, these toxicities often necessitate dose interruptions, intensive monitoring, or concomitant medications and may preclude some patients from prolonged therapy. Moreover, resistance to FLT3 inhibitors, particularly type II agents, remains a major clinical challenge and arises from FLT3-TKD mutations such as D835 and Y842, F691 mutation, or off-target genetic alterations, with TKD mutations representing a predominant mechanism. 22
Therefore, there is a compelling need to develop novel FLT3 inhibitors. Overall, the clinical success of first- and second-generation FLT3 inhibitors validates FLT3 as an actionable therapeutic target in AML; however, persistent safety concerns and resistance mechanisms highlight the need for improved therapies. This study aims to identify a novel FLT3 inhibitor with enhanced efficacy and safety, thereby providing an additional therapeutic option for patients with FLT3-mutated leukemia.
Methods
Compounds
Virtual screening protocol
The FLT3 protein complex (PDB ID: 4xuf) was used to generate the docking grid. Protein preparation was performed using the Protein Preparation Wizard in Maestro (Schrödinger). Ligand preparation was conducted using the LigPrep tool with default settings, and energy minimization was performed using the OPLS3 force field. For molecular docking, the Ligand Docking module was used. The docking grid was defined as a 20-Å box centered on the ligand. Compounds were docked using the Glide HTVS mode, with all parameters left at default settings. Compounds with docking scores lower than −7.5 were selected for further analysis. These compounds were then classified based on similarity, and 159 molecules were purchased for subsequent experimental validation.
Cell lines and culture conditions
The human leukemia cell lines MOLM13, HL60, THP-1, and K562 were purchased from BNCC (China), while the MV4-11, 32D, and HEK-293T cell lines were obtained from Cobioer Company (Nanjing, China). The 32D cell lines stably expressing different FLT3 mutations were generated via lentiviral infection with the appropriate mutated plasmid as previously described. 23 32D cells were cultured in RPMI 1640 (Gibco, USA) supplemented with 10% fetal bovine serum (FBS; PAN, Germany) and 5 ng/mL mouse interleukin-3 (IL-3; PeproTech, USA). THP-1 cells were cultured in RPMI 1640 medium supplemented with 10% FBS and 0.05 mM β-mercaptoethanol. HEK-293T cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, USA) supplemented with 10% FBS. Other leukemia cell lines were cultured in RPMI 1640 medium containing 10% FBS. These cell lines have been used in multiple studies of novel FLT3 inhibitors.24–26
Primary human peripheral blood mononuclear cells (PBMCs)
Primary blasts from AML patients and PBMCs from healthy donors were obtained from The First Affiliated Hospital of Soochow University. PBMCs were isolated using Ficoll–Paque PLUS (Cytiva, USA) according to the manufacturer’s instructions. The isolated PBMCs were cultured in Iscove’s modified Dulbecco’s medium (IMDM; Gibco, USA) supplemented with 10% FBS, 50 ng/mL recombinant human stem cell factor (rhSCF), 100 ng/mL recombinant human FMS-like tyrosine kinase 3 (rhFLT3)-ligand, 100 ng/mL recombinant human thrombopoietin (rhTPO), 25 ng/mL recombinant human interleukin (rhIL)-3, and 10 ng/mL rhIL-6. Ethical approval for this study, in accordance with the Declaration of Helsinki, was obtained from the Ethics Committee of the First Affiliated Hospital of Soochow University (Suzhou, 2024-556; 8 October 2024).
Cell viability assay
Cells (5–10 × 103 per well for cell lines; 3–5 × 104 per well for PBMCs) were seeded into 96-well plates and exposed to serial dilutions of the indicated compounds in triplicate. After incubation for 72 h at 37°C, cell viability was assessed using the Cell Counting Kit-8 (CCK-8; Dojindo, Japan) according to the manufacturer’s instructions. Dose–response curves and half-maximal inhibitory concentration (IC50) values were calculated using GraphPad Prism 8.0 (GraphPad, USA).
Apoptosis analysis
For apoptosis analysis, cells were seeded at a density of 2.0 × 105 cells per well in 6-well plates and treated with the indicated concentrations of compounds for 24 h at 37°C. Flow cytometry was performed using an APC-Annexin V Apoptosis Detection Kit (BioLegend, USA). The results were analyzed using FlowJo Version 10 (BD, USA).
Western blot analysis
Cultured cells were harvested, washed with PBS, and lysed in RIPA buffer. The protein lysates were clarified via centrifugation at 12,000 ×g for 30 min at 4°C, and the supernatant was collected. Protein concentration was determined using a BCA Protein Assay Kit (Beyotime Institute of Biotechnology, China). Equal amounts of protein (30 µg per lane) were separated via 10% sodium dodecyl sulfate (SDS)/polyacrylamide gel electrophoresis (PAGE) and transferred onto polyvinylidene fluoride (PVDF) membranes (MilliporeSigma, USA). Membranes were blocked with 5% bovine serum albumin (BSA; Beyotime Institute of Biotechnology, China) and incubated with primary antibodies at 4°C overnight. The membranes were then incubated with secondary antibodies for 1 h at room temperature and visualized using an enhanced chemiluminescent western blot detection reagent with a Gel Doc™ EZ imaging system. For the protein phosphorylation assay, cells were collected after 6 h of compound treatment, and a phosphatase inhibitor cocktail was added during lysis. The following antibodies were used: anti-phospho-FLT3 (#60413, CST), anti-FLT3 (#3462, CST), anti-phospho-STAT5 (#9351, CST), anti-STAT5 (#94205, CST), and anti-GAPDH (ab8245, Abcam).
Cellular thermal shift assay (CETSA)
CETSA was performed as previously described, with modifications. 27 HEK-293T cells stably expressing FLT3-ITD (1 × 107 cells) were treated overnight with TIP-20 (10, 20, or 50 µM) or vehicle (dimethyl sulfoxide (DMSO)). Cells were lysed in RIPA buffer, aliquoted, and heated for 3 min across a temperature gradient of 40°C–58.4°C. Proteins were analyzed via immunoblotting. For isothermal dose–response analysis, lysates were prepared after exposure to TIP-20 (0–100 µM) at 51°C. Band intensities were quantified using ImageJ, and statistical analyses were performed using GraphPad Prism 8.0.
In vitro enzymatic assays
FLT3 kinase inhibition was quantified using a homogeneous time-resolved fluorescence (HTRF) assay that monitors phosphorylation of a biotin-tagged tyrosine kinase substrate peptide. Assay conditions, including substrate, ATP, and enzyme levels as well as reaction time, were optimized beforehand. Test compounds were prepared as DMSO stock solutions and diluted in reaction buffer (50 mM HEPES, pH 7.0; 0.1 mM sodium orthovanadate; 0.01% BSA; 0.02% NaN3; 5 mM MgCl2; 5 mM MnCl2; 1 mM DTT). In white 96-well plates, 4 µL of compound at the indicated concentrations were dispensed per well, followed by 2 µL of FLT3 kinase in kinase buffer (1 ng/µL). After a 10-min preincubation, the reaction was initiated by adding 2 µL of substrate (final concentration, 1 µM) and 2 µL of ATP (final concentration, 100 µM). Mixtures were incubated for 60 min at 37°C and then quenched by adding 10 µL of detection cocktail, composed of 5 µL of Eu3+–cryptate–labeled anti-TK antibody and 5 µL of streptavidin-XL665 (final concentration, 125 nM), to enable detection of the phosphorylated peptide. Time-resolved fluorescence resonance energy transfer (FRET) signals were recorded after an additional 1-h incubation at 37°C using a BioTek Cytation 3 multimode reader (excitation wavelength, 337 nm; dual emission wavelengths, 620 and 665 nm).
Animal studies
Six- to eight-week-old NCG mice (GemPharmatech Co., Ltd, China) were transplanted with MV4-11 cells via intravenous injection. Two weeks after transplantation, engraftment rates (>1%) in peripheral blood were assessed via flow cytometry. Mice were randomly assigned to three groups, with no significant differences in engraftment rates. TIP-20 (30 mg/kg/day or 100 mg/kg/day) or vehicle control (2-hydroxypropyl-β-cyclodextrin) was administered daily via oral gavage for 14 consecutive days. After treatment, engraftment rates in total mononuclear cells were evaluated via flow cytometry using human CD45 markers (eBioscience, USA). Bone marrow smears were prepared and stained using a Wright–Giemsa Stain Kit according to the manufacturer’s instructions (Solabio, China). Mice were euthanized via CO2 inhalation upon observation of typical leukemic symptoms, including paralysis, hunched posture, labored breathing, and decreased activity. All animal experiments were conducted in accordance with established guidelines and approved by the Institutional Animal Care and Welfare Committee of Suzhou Health Vocational Technical College (Suzhou, Ethics Approval No. SWAE202547; 14 August 2025).
RNA sequencing (RNA-seq) and data analysis
Total RNA from 5 × 106 cells was extracted using TRIzol (Cwbiotech, China). After quality assessment using an Agilent 2100 bioanalyzer, the qualified RNA was used to construct an RNA library. Transcriptome sequencing was performed on the Illumina NovaSeq platform. Reads were mapped to the human genome (UCSC hg38) using HISAT2 software. Gene expression levels were estimated using the featureCounts tool of Subread software and StringTie. Differential expression analysis was conducted using the edgeR and DESeq2 R packages, with p < 0.05 or log2 (fold change) >1 considered statistically significant. Gene Set Enrichment Analysis (GSEA) was performed using clusterProfiler. Transcriptomic data were normalized from fragments per kilobase of exon per million reads mapped (FPKM) to transcripts per million (TPM) for each gene, followed by log2 transformation of the TPM values (log (TPM + 1)).
Statistical analysis
All experiments were performed in triplicate wells, with three independent replicates. Data were presented as mean ± SD. One-way analysis of variance was used to compare multiple independent groups using GraphPad Prism 8.0. Further statistical comparisons were performed using Bonferroni’s and Dunnett’s tests, and the corresponding bar charts or line graphs were generated using GraphPad Prism 8.0. p < 0.05 was considered statistically significant. IC50 values were calculated using GraphPad Prism, and synergy scores were determined using SynergyFinder 2.0. 28
The study was conducted in accordance with the Declaration of Helsinki (2024 revision).
Results
Identification of TIP-20 as a potential FLT3 inhibitor
As shown in Figure 1(a), a molecular docking–based virtual screening was initially performed to evaluate the ChemDivCN library, which contains 7 million commercially available compounds. The results of the virtual screening were analyzed via cluster analysis and visual inspection, and 250 representative compounds were selected (Figure 1(b)). Of these, 159 available compounds were tested via cell viability assays using 32D cells and 32D cells overexpressing FLT3-ITD (32D-ITD) (Figure 1(c)). We found that TIP-20 had good specificity for 32D-ITD cells (Figure 1(d)). To validate the specificity of TIP-20 for FLT3-mutated cells, we assessed the survival of 32D-ITD cells. The results showed that TIP-20 effectively suppressed the growth of 32D-ITD cells. Notably, TIP-20 had minimal effects on parental 32D cells, indicating that its selective and potent activity is attributable to specific inhibition of FLT3-ITD (Figure 1(e)).

Screening identifies TIP-20 as the most promising small-molecule compound. (a) The virtual screening workflow for FLT3 inhibitors. (b) The binding pattern of 250 potential small-molecule compounds with FLT3 protein. (c) A total of 159 compounds were screened at a single concentration of 100 μM for 72 h using 32D and 32D-ITD stable cell lines. (d) Chemical structure of TIP-20 and (e) The inhibitory effect of the identified small-molecule compound TIP-20 on the viability of 32D and 32D-ITD cells via WST-based CCK-8 proliferation assays. FLT3: feline McDonough sarcoma–like tyrosine kinase 3; WST: water-soluble tetrazolium; CCK-8, Cell Counting Kit-8; ITD: internal tandem duplication.
TIP-20 directly binds to FLT3 protein and inhibits its activity
To assess the interaction between TIP-20 and FLT3, we performed virtual molecular docking studies using the FLT3 protein. 29 The predicted minimum binding energy of TIP-20 for FLT3 was −9.1 kcal/mol. Docking results indicated that TIP-20 fits well into the ATP-binding pocket of FLT3. The terminal pyridine ring penetrates the DFG-out hydrophobic pocket, while the triazole ring forms a key hydrogen bond with the amino acid residue Glu-661 in the hinge region, with an interaction distance of 2.0 Å. After the formation of aniline benzene ring bridge, the terminal imidazole ring forms a hydrogen bond with Cys-694 (Figure 2(a)). Compared with the DMSO-treated group, the CETSA curve for the TIP-20-treated group displayed a significant thermal shift. FLT3 protein levels decreased significantly at 45.9°C in the DMSO-treated group, whereas the TIP-20-treated group showed no noticeable reduction until 51.9°C (Figure 2(b)), confirming a direct interaction between TIP-20 and FLT3. A kinase activity inhibition assay using HTRF further demonstrated that TIP-20 inhibits FLT3 activity (Figure 2(c)).

TIP-20 significantly inhibits the viability of FLT3-ITD–mutated AML cells. (a) F Diagram mode of TIP-20 binding to FLT3 (PDB: 4xuf). Yellow dashed lines represent hydrogen bonding interactions. (b) 293 T-FLT3-ITD cells were treated with TIP-20 at concentrations of 10, 20, and 50 µM or DMSO for 16 h, and the temperature range tested was 40°C–58.4°C. FLT3 protein levels were quantified using ImageJ, and the corresponding melting curve was plotted and (c) TIP-20 inhibition of FLT3 kinase activity was measured using HTRF. FLT3: feline McDonough sarcoma–like tyrosine kinase 3; ITD: internal tandem duplication; AML: acute myeloid leukemia; DMSO: dimethyl sulfoxide.
TIP-20 suppresses survival/growth and promotes apoptosis and cell cycle arrest of human FLT3-ITD–mutated AML cells in vitro
To investigate the therapeutic potential and specificity of TIP-20 for AML, we conducted proliferation assays using multiple human leukemia cell lines. TIP-20 significantly inhibited the proliferation of FLT3-ITD–mutated MV4-11 and Molm13 cells, with IC50 values of 5.783 and 1.677 µM, respectively. In contrast, TIP-20 showed minimal activity against FLT3 wild-type (FLT3-WT) AML cell lines, including K562, HL60, Kasumi-1, and THP-1, compared with FLT3-ITD–mutated cells (Figure 3(a)). To further confirm its inhibitory effect on FLT3-mutated AML, proliferation assays were conducted with primary human leukemia cells. TIP-20 effectively reduced the viability of primary AML blasts harboring FLT3-ITD mutations (Supplementary Table 1) but had little effect on FLT3-WT AML blasts or healthy donor cells, highlighting the target selectivity of TIP-20 (Figure 3(b)). After 24 h of TIP-20 treatment, the proportion of Annexin V–positive cells increased in 32D-ITD, MV4-11, and Molm13 cells in a dose-dependent manner (Figure 3(c)). Cell cycle analysis revealed that 24 h of TIP-20 treatment resulted in a dose-dependent G1-phase arrest (Figure 3(d)). To evaluate the ability of TIP-20 to overcome resistance, MV4-11 cells were gradually exposed to gilteritinib or quizartinib (AC220) using a concentration escalation method, resulting in the establishment of MV4-11 AC-R (AC220-resistant) and MV4-11 Gil-R (gilteritinib-resistant) cell lines. We then assessed inhibitory effects of TIP-20 on these resistant cell lines; as shown in Figure S1 (A) to (C), TIP-20 exhibited robust inhibitory activity in parental MV4-11 cells but was ineffective against MV4-11 AC-R and MV4-11 Gil-R cells.

Effects of TIP-20 on cell viability, cell cycle, and apoptosis in AML cell lines and primary cells with FLT3-ITD. (a) Dose–response curves of FLT3-WT and FLT3-ITD–mutated AML cell lines (MV4-11 and Continued.Molm13) and (b) AML blasts with FLT3-ITD, FLT3-WT, and healthy donor PBMCs following 72-h treatment with TIP-20. At a concentration of 0, the mean viability from three replicates was normalized to 100% as the control. (c) Proportion of apoptotic cells in 32D-ITD, MV4-11, and Molm13 cell lines after treatment with different TIP-20 concentrations for 24 h. (d) Percentage of cells in different cell cycle phases, as determined by flow cytometry, following treatment of 32D-ITD, MV4-11, and Molm13 cells with DMSO or varying concentrations of TIP-20 (1 or 2 µM) for 24 h. Comparisons were performed using a two-tailed Student’s t-test. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. AML: acute myeloid leukemia; FLT3: feline McDonough sarcoma–like tyrosine kinase 3; ITD: internal tandem duplication; WT: wild-type; PBMCs: peripheral blood mononuclear cells; DMSO: dimethyl sulfoxide.
TIP-20 inhibits FLT3 phosphorylation and downstream signaling in FLT3-ITD–mutated AML cells
Previous studies have shown that activating FLT3 mutations promote leukemic cell proliferation and survival through constitutive autophosphorylation and activation of downstream signaling proteins. 30 We treated 32D-ITD cells with increasing concentrations of TIP-20 for 6 h and assessed the phosphorylation of FLT3 and its downstream target STAT5 via western blotting. TIP-20 suppressed the phosphorylation of FLT3 and STAT5 in a dose-dependent manner, which likely contributes to its pronounced antileukemic activity in FLT3-ITD–mutated cells (Figure 4(a)). Similar effects were observed in MV4-11 and Molm13 cells (Figure 4(b) and (c)).

TIP-20 inhibits FLT3 phosphorylation and downstream pathway activity. (a) Levels of phosphorylated FLT3 (p-FLT3), total FLT3, phosphorylated STAT5 (p-STAT5), and total STAT5 were measured in 32D-ITD, MV4-11, and Molm13 cells following treatment with TIP-20 at the specified doses for 6 h.
TIP-20 exhibits effective anti‑FLT3‑ITD–mutated AML cell activity in a mouse model
To evaluate the in vivo efficacy of TIP-20, we established an FLT3-ITD–mutated AML mouse model by intravenously injecting MV4-11 cells. Two weeks after transplantation, recipient mice were randomized into three groups (n = 5 per group) to receive vehicle or TIP-20 (30 or 100 mg/kg) via oral gavage (Figure 5(a)). After 14 days of treatment, both TIP-20–treated groups exhibited a marked reduction in leukemic burden in the spleen and bone marrow, accompanied by decreased spleen size, compared with the vehicle group (Figure 5(b) to (d)). Renal function (blood urea nitrogen (UREA) and creatinine (CREA)) and hepatic function (aspartate transaminase (AST) and alanine transaminase (ALT)) were then assessed (Figure 5(e)). Renal parameters did not differ among groups, whereas liver enzyme levels were significantly lower in the TIP-20–treated groups than in controls.
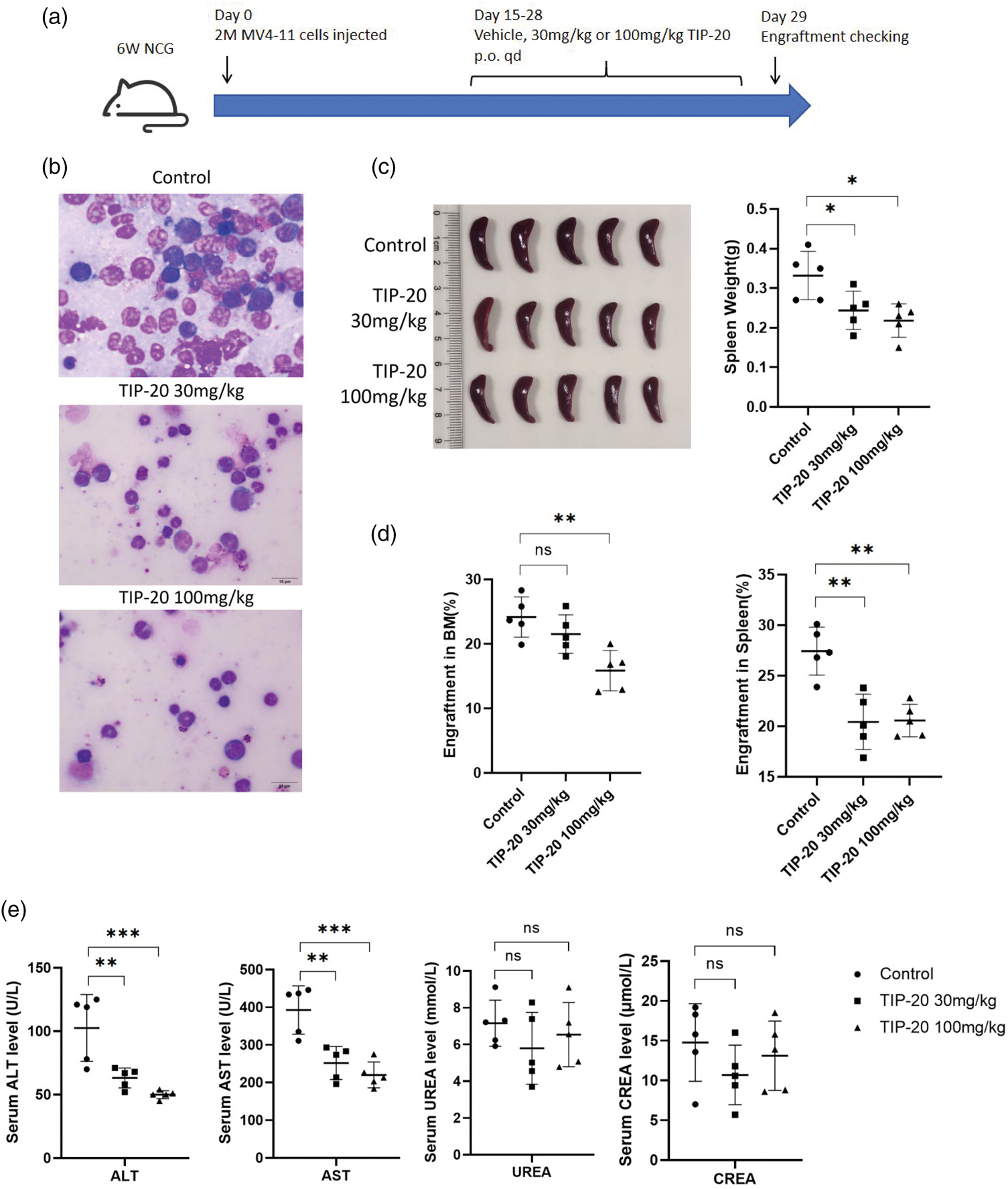
TIP-20 shows potent antileukemia effects in a mouse model. (a) In the in vivo study, 2 × 106 MV4-11 cells were injected intravenously into 6-week-old NCG mice (Day 0). From days 14 to 27, mice received vehicle or TIP-20 (30 mg/kg or 100 mg/kg, p.o., q.d.). Engraftment was assessed on Day 29. (b) Representative Wright–Giemsa–stained bone marrow smears from each group, with fewer leukemic blasts observed in the treatment groups. (c) Spleen images and spleen weights for all three groups. (d) The percentage of human CD45+ cells in the bone marrow and spleen for all three groups. (e) Serum kidney function markers (UREA and CREA) and liver enzymes (AST and ALT) showing no overt elevations with TIP-20 treatment. Data were analyzed using two-tailed Student’s t-test. **p < 0.01, ***p < 0.001. p.o.: per os (oral); q.d.: once daily; UREA: blood urea nitrogen; CREA: creatinine; AST: aspartate transaminase; ALT: alanine transaminase.
TIP-20 induces immune activation of FLT3-ITD–mutated AML cells and enhances antigen presentation
FLT3-ITD–mutated AML commonly exhibits a pronounced immune-evasive phenotype, including augmented “don’t-eat-me” signaling, a suppressive costimulatory network, and impaired antigen presentation—mechanisms that disrupt antigen display and enable escape from immune surveillance, thereby exacerbating disease aggressiveness and risk of relapse.31–34 MV4-11 and Molm13 cells were treated with TIP-20 (1 and 2 µM, respectively) for 24 h and subjected to RNA-seq. Hallmark GSEA indicated convergent immune activation in both cell lines: interferon-α/γ responses and inflammatory pathways were significantly upregulated, whereas proliferation-related pathways such as MYC targets were attenuated (Figure 6(a)). Concordant increases were observed in molecules associated with antigen presentation, chemokines/inflammation, natural killer (NK)/T-cell cytotoxicity, interferon (IFN) responses, immune checkpoints/inhibition, macrophage polarization, and the TLR/IRAK axis (Figure 6(b) and (c)). Scores for antigen processing and presentation (APM) Class I and Class II modules (Supplementary Table 2) increased in parallel, indicating enhanced antigen-presenting capacity (Figure 6(d)). Consistent with the transcriptomic data, flow cytometry in MV4-11 cells confirmed that TIP-20 (1 µM, 24 h) significantly increased surface human leukocyte antigen–DR isotype (HLA-DR) expression (elevated mean fluorescence intensity (MFI), p < 0.001) (Figure 6(e)). Taken together, short-term FLT3 blockade rapidly induces an IFN-driven inflammatory program and augments APM, suggesting that FLT3 inhibition has the potential to enhance the immunogenicity of leukemic cells.

TIP-20 treatment activates interferon signaling and antigen presentation pathways in AML cells. (a) GSEA plots showing enrichment of Hallmark gene sets in Molm13 and MV4-11 cells after treatment with TIP-20 (2 and 1 μM, respectively) for 24 h, with notable upregulation of INTERFERON_ALPHA_RESPONSE and INTERFERON_GAMMA_RESPONSE. Bubble size represents the proportion of matched genes, and color indicates statistical significance (–log10 p-value). (b) Heatmaps depicting immune-related transcriptional module activity across treatment (TRT) and control (CON) conditions in Molm13 and MV4-11 cells, with each row representing a feature module and color denoting scaled expression (Z-score). (c) Paired module score analysis demonstrating increased activity of immune-related pathways, including antigen presentation, IFN_response, and immune checkpoint suppression modules, following TIP-20 treatment compared with control. (d) APM scores (Z-scores) increased in both Molm13 and MV4-11 cells after TIP-20 treatment. (e) Flow cytometry analysis of HLA-DR surface expression in MV4-11 cells treated with DMSO or TIP-20 (1 µM), shown as a representative histogram (left) and quantification of mean fluorescence intensity (MFI) (right), indicating significant upregulation of HLA-DR following TIP-20 treatment (***p < 0.001). AML: acute myeloid leukemia; GSEA: Gene Set Enrichment Analysis; APM: antigen processing and presentation; HLA-DR: human leukocyte antigen–DR isotype; DMSO: dimethyl sulfoxide; MFI: mean fluorescence intensity.
Discussion
AML is a highly heterogeneous disease, and its clinical prognosis is influenced by diverse genomic alterations and molecular mutations. Among these, ITD and TKD mutations are common pathogenic drivers, accounting for a substantial proportion of cases. 1 These mutations not only serve as prognostic indicators but are also recognized as molecular biomarkers for assessing minimal residual disease (MRD) during monitoring of FLT3-ITD–mutated AML progression. 24 Targeted therapies against FLT3-ITD have markedly improved patient survival, fueling intensive efforts to develop novel FLT3 inhibitors.
Building on virtual screening–based discovery and multi-tier functional validation, we proposed TIP-20 as a candidate FLT3 inhibitor that targets the autoinhibited conformation and exerts both selective antileukemic activity and immune reprogramming effects against FLT3-ITD–mutated AML in vitro and in vivo. Compared with prior multitarget or narrowly selective FLT3 inhibitors, TIP-20 demonstrated a more favorable therapeutic window in cell-based and xenograft models, along with a more manageable safety profile, providing a rationale for further preclinical development and exploration of combination regimens.
TIP-20 and AC220 contain multiple nitrogen-containing aromatic rings, suggesting that they may interact with FLT3 kinase through hydrogen bonding with nitrogen atoms, enhancing their specific binding to FLT3 kinase. The polycyclic nitrogen rings in TIP-20 and the nitrogen atoms in AC220 also provide potential coordination interactions that help stabilize binding at the active site of FLT3 kinase. Both compounds have relatively large planar structures, allowing them to fit effectively into the ATP-binding pocket of the kinase and thereby inhibit its activation. 29
At the pharmacodynamic level, TIP-20 suppressed proliferation, induced G0/G1 arrest, and triggered apoptosis in FLT3-ITD–mutated cells (MV4-11, MOLM-13, and 32D-ITD) and dose-dependently downregulated the p-FLT3/STAT5 axis, consistent with the role of FLT3-ITD as an oncogenic driver. Its effects on FLT3-WT cells and healthy PBMCs were comparatively modest, indicating a therapeutic index shaped by “oncogene dependence.” Relative to clinically validated inhibitors such as gilteritinib, TIP-20’s distinctive value lies in its preferential activity against FLT3 and its narrower signaling footprint. Although pivotal trials such as ADMIRAL and QuANTUM-First have established the “druggability” of FLT3 and significantly prolonged overall survival (OS), they also revealed mechanism-related or off-target toxicities—including differentiation syndrome (gilteritinib) and QT-interval prolongation (quizartinib)—that constrain dosing, duration, and combination latitude.21,35 In our xenograft models, TIP-20 reduced leukemic burden without evidence of hepatic or renal function deterioration; this safety trend warrants confirmation in formal absorption, distribution, metabolism, and excretion (ADME); safety pharmacology; and chronic toxicology studies.
Expression of the macrophage “don’t-eat-me” signal CD47 is elevated on leukemic blasts and early progenitors, suppressing macrophage-mediated phagocytic clearance. Prior studies have shown that FLT3-ITD mutations can directly or indirectly upregulate CD47 and that dual inhibition of CD47 and the FLT3 pathway enhances phagocytosis and antileukemic activity in vitro and in animal models. 31 Through sustained activation of downstream STAT5 signaling, FLT3-ITD mutations also drive upregulation of immunosuppressive costimulatory or coinhibitory molecules such as PD-L1. Recent mechanistic studies have indicated that STAT5 activation can induce PD-L1 expression via a metabolism–epigenetics “lactylation–transcription” axis, thereby attenuating effector T-cell–mediated immune surveillance. 32 In addition, emerging basic studies have suggested that FLT3-ITD mutations may influence molecules such as CD80 through oxidative stress pathways, reshaping the balance of costimulatory and coinhibitory signaling and fostering a more immunosuppressive leukemic microenvironment. 33 Consistent with this biology, RNA-seq/GSEA revealed that TIP-20 activates interferon pathways and upregulates inflammatory responses, with a marked increase in APM scores; flow cytometry corroborated increased HLA-DR expression. These findings provide a rationale for multiple combination strategies. First, co-administration of “epigenetic reprogramming” agents such as hypomethylating agents or histone deacetylase inhibitors can reinforce transcriptional accessibility of MHC-II and antigen-processing genes. 36 Second, sequential or concurrent pairing with CD47 or LAG-3/PD-1/AXL immune checkpoint and myeloid-axis antagonists can overcome the immunosuppressive microenvironment. Third, “cross-calibration” dosing with agents targeting the same class or orthogonal targets can reduce the ecological niche for resistant clones by leveraging complementary conformational selectivity.
Safety and selectivity remain critical determinants of whether small-molecule FLT3 inhibitors can be feasibly deployed as long-term maintenance therapy. Midostaurin’s multikinase profile and quizartinib’s cardiac risk exemplify trade-offs at opposite ends of the spectrum: the former is commonly associated with gastrointestinal adverse events and myelosuppression, whereas the latter requires close electrocardiographic and electrolyte monitoring due to QT-interval prolongation and is further constrained by a risk evaluation and mitigation strategies (REMS) program. In this study, TIP-20 did not induce overt hepatic or renal toxicity or clinical deterioration. Nevertheless, a more comprehensive assessment of treatment-related adverse events will be required in subsequent studies.
From a translational perspective, TIP-20 is optimally positioned as a “highly selective FLT3-ITD inhibitor with immuno-sensitizing properties.” If phase I studies demonstrate a favorable exposure–inhibition relationship and evidence of early molecular responses—such as suppression of p-FLT3/STAT5 and reductions in circulating tumor burden—without dose-limiting electrocardiographic abnormalities or severe myelosuppression, TIP-20 could be integrated into front-line regimens, including standard “3 + 7” induction or low-intensity therapies (HMA+VEN), to deepen remission and accelerate MRD clearance. Alternatively, it could be combined with immunologic strategies—such as anti-CD47 agents, checkpoint inhibitors, or vaccines—to enhance mobilization of immune effectors and overcome the immunosuppressive microenvironment.
Our study has several limitations. Comprehensive kinome profiling and cardiac safety pharmacology have not yet been completed, precluding full exclusion of potential liabilities involving distal ion channels or hematopoietic progenitor targets. Evidence for immune reprogramming is currently derived primarily from short-term in vitro assays and transcriptomics; these findings should be substantiated in immunocompetent mouse models and patient samples using phenotypic functional readouts, such as antigen-specific T-cell responses, dendritic-cell cross-presentation, and phagocytic or cytotoxic activity. The resistance spectrum—including F691L, D835, and bypass pathways—also remains to be validated through in vitro evolution and patient-derived xenograft models. Future studies will pursue iterative optimization across the axes of selectivity, safety, immune effects, and resistance barriers while exploring rational sequencing and combinations with chemotherapy, epigenetic agents, and immunotherapies. Although this study did not investigate resistance mechanisms of TIP-20 in detail, we recognize that resistance is a critical issue in targeted therapies. Understanding TIP-20 resistance—including secondary mutations or activation of bypass signaling pathways—is essential for its clinical application. Resistance to targeted therapies is often associated with the evolution of FLT3 mutations, emergence of secondary mutations, or activation of compensatory signaling pathways. Therefore, future studies will focus on inducing resistance in FLT3-ITD and FLT3-TKD cell lines and further investigating these mechanisms. Overall, as a FLT3 inhibitor targeting the DFG-out conformation, TIP-20 combines pharmacologic selectivity for FLT3-ITD with the capacity to induce antigen presentation and interferon programs, providing compelling preclinical support for improving outcomes in FLT3 AML.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605251412963 - Supplemental material for TIP-20, a novel feline McDonough sarcoma–like tyrosine kinase 3 inhibitor with potent antileukemia activity
Supplemental material, sj-pdf-1-imr-10.1177_03000605251412963 for TIP-20, a novel feline McDonough sarcoma–like tyrosine kinase 3 inhibitor with potent antileukemia activity by Hongfeng Pang, Zhizhi Zhang, Xiaoyu Lv, Shuaishuai Ge, Yuanhong Huang, Qiaocheng Qiu, Depei Wu, Sheng-Li Xue and Song-Bai Liu in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605251412963 - Supplemental material for TIP-20, a novel feline McDonough sarcoma–like tyrosine kinase 3 inhibitor with potent antileukemia activity
Supplemental material, sj-pdf-2-imr-10.1177_03000605251412963 for TIP-20, a novel feline McDonough sarcoma–like tyrosine kinase 3 inhibitor with potent antileukemia activity by Hongfeng Pang, Zhizhi Zhang, Xiaoyu Lv, Shuaishuai Ge, Yuanhong Huang, Qiaocheng Qiu, Depei Wu, Sheng-Li Xue and Song-Bai Liu in Journal of International Medical Research
Supplemental Material
sj-pdf-3-imr-10.1177_03000605251412963 - Supplemental material for TIP-20, a novel feline McDonough sarcoma–like tyrosine kinase 3 inhibitor with potent antileukemia activity
Supplemental material, sj-pdf-3-imr-10.1177_03000605251412963 for TIP-20, a novel feline McDonough sarcoma–like tyrosine kinase 3 inhibitor with potent antileukemia activity by Hongfeng Pang, Zhizhi Zhang, Xiaoyu Lv, Shuaishuai Ge, Yuanhong Huang, Qiaocheng Qiu, Depei Wu, Sheng-Li Xue and Song-Bai Liu in Journal of International Medical Research
Footnotes
Acknowledgments
We thank the Suzhou College of Health Vocational Technology Biotechnology Center for their excellent technical support.
Author contributions
H.F.P., Z.Z.Z., and X.Y.L. performed experiments, analyzed the data, and wrote the manuscript. S.S.G. and Y.H.H. analyzed and interpreted the data. C.Q.Q., S.L.X., and S.B.L. designed the study and contributed to review and editing. S.B.L. and D.P.W. provided feedback on the report and showed academic leadership for the project.
Data availability statement
All data supporting the conclusions of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The authors declare that they have no conflicts of interest.
Funding
This study was supported by the Postgraduate Research & Practice Innovation Program of Jiangsu Province (KYCX25_3490), the National Natural Science Foundation of China (Nos. 82470224, 82270165, and 81970138), the Jiangsu Province Natural Science Foundation of China (No. BK20221235), the Jiangsu Province “333” Project, Boxi Clinical Research Project (Grant No. BXLC008) and Boxi Leading Talent Program (Grant No. 02622), the Project of Jiangsu Province Engineering Research Center of Molecular Target Therapy and Companion Diagnostics in Oncology, the Jiangsu Higher Education Institution Innovative Research Team for Science and Technology (2021), Program of Jiangsu Vocational College Engineering Technology Research Center (2023), The Natural Science Key Foundation of the Jiangsu Higher Education Institutions of China (Grant No. 24KJA310008), Key Programs of the Suzhou Vocational Health College (szwzy202406), and the Qing‐Lan Project of Jiangsu Province in China (2022).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
