Abstract
An unusual case of infective endocarditis caused by a strain of extended-spectrum β-lactamase Escherichia coli O25b:H4-B2-ST131 originating from a urinary tract infection is described. This paper has thoroughly analyzed the distinctive genotypic, phenotypic, and phylogenetic elements of the strain, correlating them with the patient’s severe clinical presentation, diagnostic course, and successful outcome achieved through intensive antibiotic and surgical management, culminating in mitral valve replacement. Additionally, we proposed a hypothesis regarding the pathophysiology of infective endocarditis caused by extended-spectrum β-lactamase Escherichia coli, based on genotypic and phylogenetic analyses. A host-related explanatory approach was also assessed. Understanding the importance of early and real-time genotyping and molecular characterization of an Escherichia coli strain with a potential mechanism for developing infective endocarditis may help guide future clinical practice.
Introduction
Infective endocarditis (IE) is a rare disease with a high mortality rate. Gram-positive bacteria are frequently isolated from the majority of IE cases. However, its etiology differs according to local epidemiology.1,2 Escherichia coli is the most common bacterium isolated from human and animal gastrointestinal systems. 3 Extraintestinal pathogenic E. coli (ExPEC) is the leading cause of bacteremia worldwide, and the O serotype 25 has been reported as one of the most ubiquitous strains, which is also associated with the etiology of urinary tract infections (UTIs). 4 Moreover, the prevalence of extended-spectrum β-lactamase–producing E. coli (ESBL-EC) has increased in recent years. 5 However, IE caused by ESBL-EC remains rare,1,6,7 and available evidence is limited, making it important to understand the physiopathology and virulence factors involved in this infection. 8 This case report aimed to describe the genomic profile, resistome, and potential molecular mechanisms of ESBL-EC bacteremia with secondary IE of the native mitral valve. In addition, a host-related risk factor approach was used to understand the development of this condition. The report underscores the importance of early and real-time genotyping and molecular characterization of microorganisms for the diagnosis, treatment, and prognosis of IE.
Case presentation
A male in his late 60s with a history of insulin-dependent type 2 diabetes mellitus, hypertensive heart disease, and a 40-year smoking history was admitted to the intensive care unit (ICU) of the Fundación Clínica Shaio, Bogotá, Colombia, in January 2024, after experiencing chills, vomiting, and somnolence (Glasgow Coma Scale score, 13/15) for 36 h. Ten days earlier, he had been diagnosed with a UTI caused by ESBL-EC with associated bacteremia, which had been treated with carbapenem for 7 days. Upon arrival at the emergency department, the patient underwent immediate septic workup, including urine culture, paired blood cultures, and standard laboratory tests. Meropenem was used as an initial, sole, empirical, and final treatment, administered at a dose of 2 g intravenously every 8 h for 42 days. Vasopressor support was initiated with noradrenaline and vasopressin for septic shock. Urine and blood cultures confirmed the presence of ESBL-EC, consistent with UTI and bacteremia (growth after 15 h of incubation). Despite initial hypotension, the patient’s condition improved after 48 h without the need for further vasopressor support. Additional investigations, including abdominal computed tomography (CT), colonoscopy, ileoscopy, and lower limb doppler, revealed no other sources of infection. A transthoracic echocardiogram revealed a 2-cm vegetation on the posterior leaflet of the mitral valve, extending toward the atrial side, resulting in moderate regurgitation (class III b) without stenotic gradient, with preserved biventricular function (left ventricular ejection fraction of 62% and fractional shortening of 40%) (Figure 1).

Transthoracic echocardiogram demonstrating (a) vegetation (yellow arrow) on the posterior leaflet of the mitral valve, measuring 2 cm toward the atrial side; (b and d) color doppler images showing mitral regurgitation (yellow arrow); and (c) pulsed doppler image showing mitral regurgitation. Yellow arrow denote vegetation on the posterior leaflet of the mitral valve.
The endocarditis team evaluated the case and recommended the need for surgical intervention involving valve replacement. Intraoperatively, significant vegetation was observed on the posterior leaflet of the mitral valve. The anterior leaflet was resected with the entire posterior leaflet, revealing calcification of the papillary muscles and an abscess. A 31-size biological valve was implanted using separate sutures (Figure 2). Final valve culture confirmed ESBL-EC infection.

Intraoperative findings demonstrating mitral valve with large vegetation on the posterior leaflet (yellow arrow).
Subsequently, repeat blood cultures were obtained and found to be negative. The patient exhibited satisfactory postoperative progress and was discharged home. Figure 3 presents a summary of the key events during the clinical course. The reporting of this study conforms to the Case Report (CARE) guidelines. 9

Time line of clinical events. CT: computed tomography; ESBL: extended-spectrum β-lactamase-producing; ICU: intensive care unit; MALDI-TOF: Matrix-assisted laser desorption/ionization–time of flight; UTI: urinary tract infection.
Methods
Phenotypic analysis
Blood samples were collected under aseptic conditions and inoculated into aerobic and anaerobic blood culture bottles. The bottles were incubated in the BacT/ALERT® BXM automated system (automated blood culture system using continuous CO2-dependent colorimetric and optical growth detection; bioMérieux; France) until a positive growth signal was detected. Once positive, aliquots were subcultured on appropriate agar media for bacterial isolation and identification. Final species-level identification was confirmed using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS), which enables rapid protein-based spectral profiling for highly accurate bacterial identification. Antimicrobial susceptibility testing (AST) was subsequently performed using the VITEK® 2.0 automated system (automated microbial identification and antimicrobial susceptibility system using sealed biochemical/AST cards with optical monitoring and algorithm-based interpretation; bioMérieux; France), following the manufacturer’s instructions, and results were interpreted according to the Clinical and Laboratory Standards Institute (CLSI) guidelines.
Genotype and phylogenetic analyses
Genomic DNA was isolated from the cultivated E. coli using a DNeasy UltraClean Microbial Kit (silica-membrane–based kit for rapid microbial DNA extraction through chemical lysis, inhibitor removal, and column purification; QIAGEN; Netherlands) following the manufacturer’s instructions. Subsequently, it was sequenced using Oxford Nanopore technologies using the ligation sequencing kit SQK-LSK109 (ligation-based library preparation kit enabling high-fidelity adapter attachment for Oxford Nanopore Technologies long-read sequencing; United Kingdom) and a flow cell R9.4.1 (nanopore sequencing flow cell incorporating R9.4.1 pore chemistry for real-time single-molecule DNA sequencing with improved stability and accuracy).
The Guppy tool was used to perform base calls from Fast5 to Fastq files, quality preprocessing, and adapter removal. The FASTQ file from sequencing was assembled using the Fyle v2.9 tool (long-read genome assembler designed to construct high-quality contiguous assemblies from noisy Nanopore sequencing data) with default parameters. Assessment of assembly quality was performed using CheckM v1.1.3 (genome-quality assessment tool that estimates completeness and contamination based on lineage-specific marker genes). Taxonomic assignment was performed using GTDB-Tk (genome-based toolkit that classifies bacterial genomes using the standardized genome taxonomy database). Identification of molecular markers of interest, such as antimicrobial resistance genes (ARGs) and virulence factors, was performed using the web server of the Center of Genomic Epidemiology (https://www.genomicepidemiology.org/services/), as follows: SerotypeFinder v2.0 to identify serotypes from isolates of E. coli, 10 ResFinder 4.1 to identify acquired ARGs, 11 VirulenceFinder 2.0 to detect acquired virulence genes, 12 and PathogenFinder 1.1 to predict bacterium’s pathogenicity toward human hosts. 13
Results
The final antibiogram report detected an infection caused by ESBL-EC. The bacterium was resistant to ciprofloxacin and gentamicin. Notably, the organism remained susceptible to other antibiotics, including the carbapenems (ertapenem and meropenem), amikacin, and tigecycline.
The assembly and subsequent quality assessment showed an assembly consisting of three contigs totaling 5,480,739 bp (5.4 Mb), consistent with the reported size of the E. coli genome (4.5–5.5 Mb). An N50 value of 5,333,733 bp was determined, indicating that the assembled genome was reasonably complete. The taxonomic assignment corresponding to E. coli was confirmed with GTDB-Tk. 14 The fliC gene was found with 100% coverage and 100% identity, indicating an H_type (H4 serotype), and two copies of the wzx gene were identified with 100% coverage and 95.79% identity, indicating an O_type (O25 serotype). Finally, this isolate was typed as ST-131, a predominant E. coli lineage among ExPEC. 15 However, a loss of identity (99.77%) was identified in the mdh gene, indicating genetic diversity.
Analysis of resistome molecular markers of interest revealed elements predominantly associated with six antimicrobial classes: β-lactamase (n = 12), aminoglycoside (n = 6), quinolone (n = 3), tetracycline (n = 2), peroxide (n = 1), and amphenicol (n = 1) (Table 1).
Antibiotic resistome of E. coli.
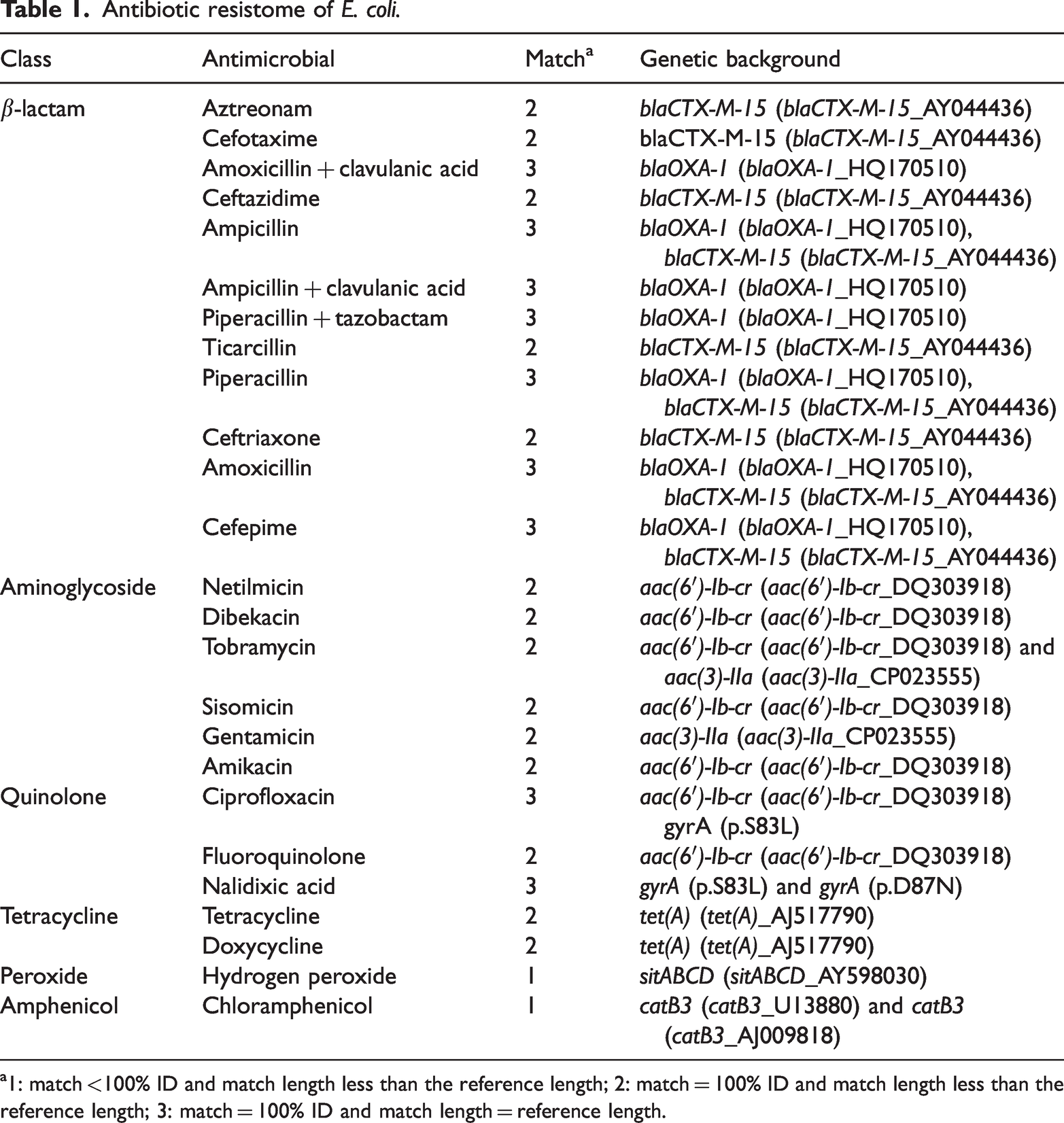
1: match <100% ID and match length less than the reference length; 2: match = 100% ID and match length less than the reference length; 3: match = 100% ID and match length = reference length.
Additional analysis using PathogenFinder demonstrated that this isolate has a 0.92 probability of being a human pathogen, carrying 479 pathogenic families comprising 7657 sequences (Table 2).
Virulence-associated (indirect/fitness-related) genes and their proposed pathophysiological roles in IE.

UPEC: Uropathogenic Escherichia coli.
Discussion
E. coli endocarditis is a rare condition with limited representation in the medical literature.7,16 IE caused by E. coli has not been extensively characterized, particularly with respect to its virulence mechanisms. Molecular and phylogenetic classification showed E. coli O25b:H4-B2-ST131 with a repertoire of virulence genes involved in diverse mechanisms for adherence to host cells and tissues as well as a resistome with ARGs.
This E. coli showed resistance for β-lactams (blaCTX-M-15_AY044436 and blaOXA-1_HQ170510 genes), aminoglycoside (aac(6')-Ib-cr_DQ303918 and aac(3)-IIa_CP023555 genes), quinolone (aac(6')-Ib-cr_DQ303918 gene), and tetracycline (tet(A)_AJ517790 gene), consistent with ESBL-EC. It has been reported that the increasing isolation of ESBL-EC worldwide is significantly affecting public health. 17 The challenge posed by ESBL-EC infections that are resistant to commonly used antibiotics complicates treatment, often necessitating the administration of more potent antibiotics. 18 The presence of ESBL-EC in endocarditis highlights the broader challenge of antibiotic resistance in clinical practice. It emphasizes the need for responsible antibiotic use through antimicrobial stewardship, considering factors such as infection type, patient characteristics, and local resistance patterns.
The B2 phylogenetic group of E. coli is strongly associated with ExPEC, which induces severe invasive infections, including bacteremia that may result in IE. B2 strains are genetically predisposed to virulence, often carrying a high number of virulence-associated genes (VAGs). 19
In contrast, virulome analysis facilitated the identification of genes coding for virulence factors that may explain the possible pathophysiological mechanism of IE associated with E. coli. The process begins with the onset of bacteremia. In this case, E. coli enters the bloodstream, typically originating from a source such as a UTI, as observed in our patient. Once in circulation, the bacteria focus on surviving in the bloodstream and evading the host immune response. This is often demonstrated by the expression of genes for capsule formation (e.g. kpsE and kpsMII_K5). These capsules act as protective layers around the bacteria, allowing them to evade the host immune system.20,21 This critical evasion step allows the bacteria to establish and maintain the infection within the blood and, subsequently, the heart tissue.22,23 Following survival in the bloodstream, a crucial step is adhesion to heart valves. E. coli generally binds poorly to intact endothelium; therefore, it usually adheres to damaged endothelium or preexisting sterile platelet-fibrin deposits (nonbacterial thrombi). Bacterial adhesion to host tissues is mediated by adhesins (e.g. FimH, Iha, AslA, PapC, FdeC, and Hra).24–29 The initial colonization and infection of damaged or prosthetic heart valves likely involve other adhesion factors encoded by papA_F43, acting as major pilin subunit, and anr, both of which facilitate the anchoring of the bacteria to the endocardial surface, initiating biofilm formation.25,30 All these factors allow the bacteria to attach to the compromised areas of the heart valves or endothelium, representing a critical step in IE initiation.28,31–35 Vegetation formation ensues as bacterial microcolonies grow on the heart valve. This process actively recruits platelets and fibrin, leading to the development of a clot-like biofilm known as a vegetation, which embeds the bacteria. This stage is associated with fimbriae-dependent cell adhesion proteins and secretion systems that promote colonization and further biofilm formation.36–38 Moreover, toxins (e.g. hha, sat, hlyA, and cnf) that may play a role in damaging heart tissues further encourage vegetation growth.36,39–41 The vegetation then undergoes biofilm maturation, developing an extracellular polymeric matrix. This matrix is vital for enabling bacterial persistence and conferring antibiotic tolerance.
This maturation process is often associated with the expression of genes for iron acquisition (e.g. iucC, iutA, and sitA)25,42,43 and siderophores (e.g. fyuA, iutA, usp, and yfcV).25,35,39,44 Bacteria require iron to thrive and replicate; therefore, the ability to scavenge iron from the host contributes significantly to their survival and persistence in the heart, where iron availability is typically limited.31,41 The gad gene, glutamate decarboxylase, is vital for maintaining bacterial physiological pH under acidic stress, which could include the local acidic environment produced during inflammation or within phagocytic cells. 45 The chronic infection process culminates in a period of complications. Importantly, the iss gene, increased serum survival, confers serum resistance, enabling the bacterium to evade the complement system and humoral immunity, a factor strongly associated with increased mortality. 46
Fragments of the friable vegetation can break off and embolize to distant sites, such as the brain or spleen, resulting in systemic sequelae. Furthermore, the persistent infection triggers ongoing inflammation, contributing to tissue damage and systemic illness. Other miscellaneous genes (ompT, traT, traJ, and usp) encoding proteins that may have diverse functions were identified, including those involved in interactions with the host immune system and modulation of the host response to infection.22,23,38,44,47,48
In recent years, E. coli has gained particular attention in cases of IE in patients with insidious clinical manifestations, such as intermittent fever related to persistent E. coli bloodstream infection despite targeted treatment, predominantly originating in the UTI. 49 In the clinical setting, risk factors for IE caused by ESBL-EC include older age, diabetes mellitus, prosthetic valves, and structural heart diseases with a history of UTI. 7 In addition to examining the genomic characteristics and virulence potential of this E. coli strain, it is crucial to investigate the host-related factors, including genetic predispositions, comorbidities, and immune status, that contribute to the development of E. coli–associated endocarditis. The patient in this case report was an older man with type 2 diabetes mellitus, hypertensive cardiac disease, and a history of UTI with previous bacteremia and use of antimicrobial agents. The combination of these risk factors and the unique characteristics of the ESBL-EC O25b:H4-B2-ST131 strain contributed to the development and severity of the clinical presentation in this patient.
It is important to highlight that genotyping is essential for understanding IE caused by E. coli, given the limited information in the medical literature on the virulence of strains producing β-lactamases. Studying these strains could provide therapeutic insights. 50 This study has certain inherent limitations. Particularly, it presents a description of the virulome and resistome of a single E. coli strain. To fully elucidate the factors contributing to E. coli–associated endocarditis, future analyses in larger cohorts are warranted. This would enable the construction of a more comprehensive profile, especially considering that E. coli O25b:H4-B2-ST131 is among the most frequently isolated E. coli strains worldwide. Moreover, there is paucity of literature regarding the increasing incidence of this specific type of E. coli infection. The driving factors may be more closely associated with the host characteristics rather than the bacterium itself.
Conclusion
IE caused by E. coli has not been well described in the literature, particularly with respect to the characterization of its virulence mechanisms. In ESBL-EC, several genes may be involved in the pathological progression of IE from UTI. Understanding the importance of early and real-time molecular virulome and resistome characterization of E. coli strains in bloodstream infections could improve risk evaluation for the development of IE in a specific high-risk population and shape future clinical practice.
Footnotes
Acknowledgments
We express our gratitude to the CRitical carE investigATors IN Group (CREATING) of the Fundación Clínica Shaio. We acknowledge the use of ChatGPT (version 5.0) for language corrections.
Author contributions
All authors have read and approved of the final version of manuscript. PCFN: Conceptualization, investigation, resources, data curation, writing–review and editing, supervision, and project administration. DJ and ESM: Conceptualization, methodology, software, validation, formal analysis, writing–review and editing, and visualization. NB, SC, and MM: Methodology, investigation, resources, data curation, and writing–review and editing. HRA: Conceptualization, writing–review and editing, and supervision. RBB: Conceptualization, methodology, and investigation. JOCB: Conceptualization, methodology, investigation, resources, data curation, writing–original draft, visualization, supervision, and project administration.
Check list
The study was performed in accordance with the Case Report (CARE) guidelines.
Data availability statement
The authors confirm that clinical and paraclinical data are available from the corresponding author, upon reasonable request.
Declaration of conflicting of interests
None.
Ethical approval
This study adhered to ethical requirements. It was approved by the Ethics committee in research of the Fundación Abood Shaio (ID approval DIB-22-23). We have deidentified all patient details.
Funding
None.
Informed consent statement
Written informed consent has been obtained from the involved patient for treatment and publication.
Manuscript publication
This manuscript has not been published elsewhere and is not under consideration by any other journal.
