Abstract
Objective
Sepsis-induced myocardial dysfunction is a common complication of sepsis, characterized by high mortality and an unclear underlying mechanism. This study conducted an integrative multiomics analysis of mice with sepsis-induced myocardial dysfunction to provide new insights into potential mechanisms.
Method
We constructed an animal model of sepsis-induced myocardial dysfunction and analyzed the metabolomics, transcriptomics, and protein profiles of the heart tissues in the control and experimental groups.
Results
Untargeted metabolomics identified 74 significantly altered metabolites in the positive ion mode, of which 48 were upregulated and 26 downregulated; moreover, 70 significantly altered metabolites were detected in the negative ion mode, with 50 being upregulated and 20 downregulated. Transcriptomics revealed 4831 differentially expressed genes, with 3027 being downregulated and 1804 upregulated. Proteomics identified 107 significant proteins, 94 of which were significantly upregulated and 13 significantly downregulated. Integrated omics analysis revealed three significantly altered metabolites common to both groups: L-glutamate, L-aspartate, and nicotinamide. These metabolites were predominantly involved in nicotinate and nicotinamide metabolism, histidine metabolism, and nitrogen metabolism, potentially related to the pathogenesis of sepsis-induced myocardial dysfunction.
Conclusion
The pathogenesis of sepsis-induced myocardial dysfunction in mice may be associated with alterations in nicotinate and nicotinamide metabolism, histidine metabolism, and nitrogen metabolism.
Keywords
Background
Sepsis is a life-threatening organ dysfunction caused by a dysregulated host response to infection. 1 It represents a severe public health issue, affecting approximately 48.9 million people annually and leading to 11 million sepsis-related deaths, accounting for 19.7% of global mortality. 2 Sepsis-induced myocardial dysfunction (SIMD) is a common complication in patients with sepsis, primarily manifesting as myocardial systolic and diastolic dysfunction, which exacerbates multiple organ failure and is associated with a high mortality rate. 3 Current research has identified several pathogenic mechanisms of SIMD, including microcirculatory dysfunction, excessive inflammatory response, mitochondrial metabolic disorder, autonomic dysfunction, calcium dysregulation, excessive nitric oxide (NO) production, and apoptosis. 4 However, these abnormal changes do not fully explain the pathogenesis of SIMD, and therapeutic interventions for SIMD remain suboptimal. Therefore, elucidating the potential mechanisms of SIMD and identifying effective therapeutic targets for precise treatment are of crucial importance.
Metabolomics is the study of changes in the quantities of metabolites in an organism in response to external stimuli or disturbances or their temporal variations, which aims to establish correlations between metabolites and physiological or pathological changes. This approach helps in understanding disease progression and is considered a phenotype-proximal discipline. 5 Various studies have utilized metabolomics to discover novel biomarkers related to the diagnosis, mechanisms, and prognosis of sepsis, revealing diverse metabolic characteristics in sepsis patients.6,7 Transcriptomics, which examines gene expression at the RNA level, is an important method for studying cellular functions. The combination of metabolomics and transcriptomics enables a comprehensive assessment of gene expression at the levels of both metabolites and mRNA. 8 Proteomics is an analytical technique used to examine the expression, structural functions, and interactions of proteins within specific cells, tissues, or organisms. 9 As proteins are positioned downstream in the flow of genetic information and are closer to the physiological phenotype, proteomics can directly reflect physiological or pathological changes in an organism. The integrated analysis of metabolomics and proteomics can help elucidate the molecular regulatory mechanisms underlying phenotypic changes, facilitating the identification of key proteins, metabolites, and pathways that serve as research targets.
Currently, the pathogenic mechanisms of SIMD remain unclear. Multiomics analysis can reveal changes in biological phenotypes and mechanisms at different levels, compensating for potential false-positive results that may arise from single-omics approaches. This study aimed to profile cardiac tissue samples from mice with SIMD through integrated metabolomics, transcriptomics, and proteomics analyses. The objectives of this study were to uncover novel diagnostic biomarkers and therapeutic targets, thereby providing a mechanistic foundation for subsequent in vivo functional validation.
Methods
Establishment, grouping, and collection of blood samples and tissues for experimental models
In this study, 16 male C57BL/6 mice aged 9–10 weeks were uniformly procured and housed under identical laboratory conditions to minimize environmental variability. Following a 7-day acclimatization period, all mice were weighed, uniquely numbered, and randomly assigned to either the lipopolysaccharide (LPS) group or control group using a computer-generated randomization sequence. Sepsis was induced in the LPS group via intraperitoneal injection of LPS (10 mg/kg body weight), while the control group received an equivalent volume of sterile saline administered intraperitoneally.
Cardiac function was assessed 24 h postintervention using a high-resolution in vivo imaging system (VIVID E9; GE Healthcare, USA). Key echocardiographic parameters included left ventricular ejection fraction (LVEF), left ventricular fractional shortening (LVFS), left ventricular end-diastolic diameter (LVIDd), and left ventricular end-systolic diameter (LVIDs). Cardiac ultrasound examination was performed, blood was taken from the eye sockets to detect myocardial enzymes, and the heart was perfused and washed with physiological saline. The intact mouse heart tissue was taken out and stored in liquid nitrogen. After the experiment, it was transported to the sequencing unit with dry ice.
Enzyme-linked immunosorbent assay (ELISA)
Commercially available ELISA kits (Meimian Biotechnology, Jiangsu, China) were used to determine the levels of cardiac troponin I (c-TnI), according to the manufacturer’s instructions.
Processing and analysis of tissue samples for metabolomics
Tissues (100 mg) were individually ground with liquid nitrogen, and the homogenate was resuspended with prechilled 80% methanol via well vortex. The samples were incubated on ice for 5 min and then centrifuged for 20 min at 15,000 × g and 4°C. Some of the supernatant was diluted to final concentration containing 53% methanol via liquid chromatography–mass spectrometry (LC–MS) grade water. The samples were subsequently transferred to a fresh Eppendorf tube and then centrifuged for 20 min at 15,000 × g and 4°C. Finally, the supernatant was injected into the LC–tandem MS (LC–MS/MS) system.
Statistical analyses were performed using the statistical software R (R version R-3.4.3), Python (Python 2.7.6 version), and CentOS (CentOS release 6.6). When the data were not normally distributed, the values were standardized to obtain relative peak areas according to the following formula: raw quantitation value/(the sum of sample metabolite quantitation value/the sum of QC1 sample metabolite quantitation value). Compounds whose coefficients of variation (CVs) of relative peak areas in QC samples were greater than 30% were removed, and metabolite identification and relative quantification results were obtained.
These metabolites were annotated using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database, HMDB database, and LIPIDMaps database. Principal component analysis (PCA) was performed via metaX. We applied univariate analysis (t-test) to calculate statistical significance (P-value). Metabolites with variable importance in projection (VIP) >1 and P-value <0.05 and fold change (FC) ≥2 or FC ≤0.5 were considered differential metabolites. Volcano plots were used to filter metabolites of interest based on log2 (FC) and −log10(P) values of metabolites via ggplot2 in R.
Total RNA quantification
Total RNA was qualified and quantified as follows:
The RNA sample was first qualified using 1% agarose gel electrophoresis for possible contamination and degradation. RNA purity and concentration were then examined using NanoPhotometer spectrophotometer. RNA integrity and quantity were finally measured using RNA Nano 6000 Assay Kit of the Bioanalyzer 2100 system.
Total RNA was used as the input material for library preparation. After library preparation and pooling of different samples, the samples were subjected to Illumina sequencing. The libraries were sequenced on Illumina novaseq 6000 Platform for 6 G raw data, generating 150 nt pair-end reads. Differential expression analysis of two conditions/groups (two biological replicates per condition) was performed using the DESeq2 R package (v1.20.0). DESeq2 provides statistical routines for determining differential expression in digital gene expression data using a model based on the negative binomial distribution. The resulting P values were adjusted using the Benjamini and Hochberg (BH) approach for controlling the false discovery rate. Genes with an adjusted P-value of <0.05 determined via DESeq2 were considered differentially expressed. Before differential gene expression analysis, for each sequenced library, the read counts were adjusted via edgeR program package through one scaling normalized factor. Differential expression analysis of the two conditions was performed using the edgeR R package (v3.22.5). P values were adjusted using the BH method. BH correction is a statistical safeguard that ensures that the reported differential metabolites are likely true biological findings, not artifacts of mass testing. The corrected P-value of 0.05 and absolute FC of 2 were set as the threshold for significantly differential expression.
Proteomics analysis of tissue samples
Samples were ground individually in liquid nitrogen and lysed with sodium deoxycholate Tris (containing 100 mM NaCl) and 1/100 volume of dithiothreitol, followed by 5 min of ultrasonication on ice. After incubation for 8–15 min at 95°C and ice-bath cooling for 2 min, the lysate was centrifuged for 15 min at 12,000 × g and 4°C, and the supernatant was subsequently alkylated with sufficient iodoacetamide for 1 h at room temperature in the dark. Then, the samples were completely mixed with four times volume of precooled acetone by vortexing and incubated at −20°C for at least 2 h. The samples were then centrifuged for 15 min at 12,000 × g and 4°C, and the precipitate was collected. After washing with 1 mL of cold acetone, the pellet was dissolved completely using dissolution buffer (DB). Ultrahigh-performance LC (UHPLC)–MS/MS analyses were performed using the EASY-nLCTM 1200 UHPLC system coupled with a Q ExactiveTM HF-Xor Orbitrap Exploris 480 mass spectrometer.
Gene Ontology (GO) and InterPro (IPR) functional analyses were conducted using the interproscan program against the nonredundant protein database (including Pfam, PRINTS, ProDom, SMART, ProSite, and PANTHER), and the databases of Clusters of Orthologous Groups (COG) and KEGG were used to analyze the protein family and pathways. Differentially expressed proteins (DEPs) were used for volcanic map analysis, cluster heatmap analysis, and enrichment analysis of GO, IPR, and KEGG. The probable protein–protein interactions were predicted using the STRING-db server.
Integrated omics analysis methods
RNA sequencing (RNA-seq) data/protein enrichment analysis
Protein enrichment analysis was performed using the R package clusterProfiler for GO enrichment analysis.
Metabolite enrichment/joint analysis of metabolites and proteins
Metabolite enrichment was performed via the MetaboAnalyst website using KEGG enrichment analysis. We performed KEGG pathway enrichment analysis for metabolite–transcriptomics and metabolite–proteomics combinations in both positive and negative ion modes, with the significance threshold for pathways set at P < 0.1.
Statistical analysis
All data were presented as mean ± SD. Comparisons between two groups were performed using independent sample t-tests. A P-value of <0.05 was considered to indicate statistical significance. Statistical analyses were performed using GraphPad Prism 9.4.1 software.
Results
SIMD model
Echocardiography showed that there was no significant difference in the overall outcome values (LVEF, LVFS, LVIDd, and LVIDs) between the LPS and control groups, and the c-TnI level in the LPS group was significantly higher than that in the control group (P < 0.05) (Figure 1).
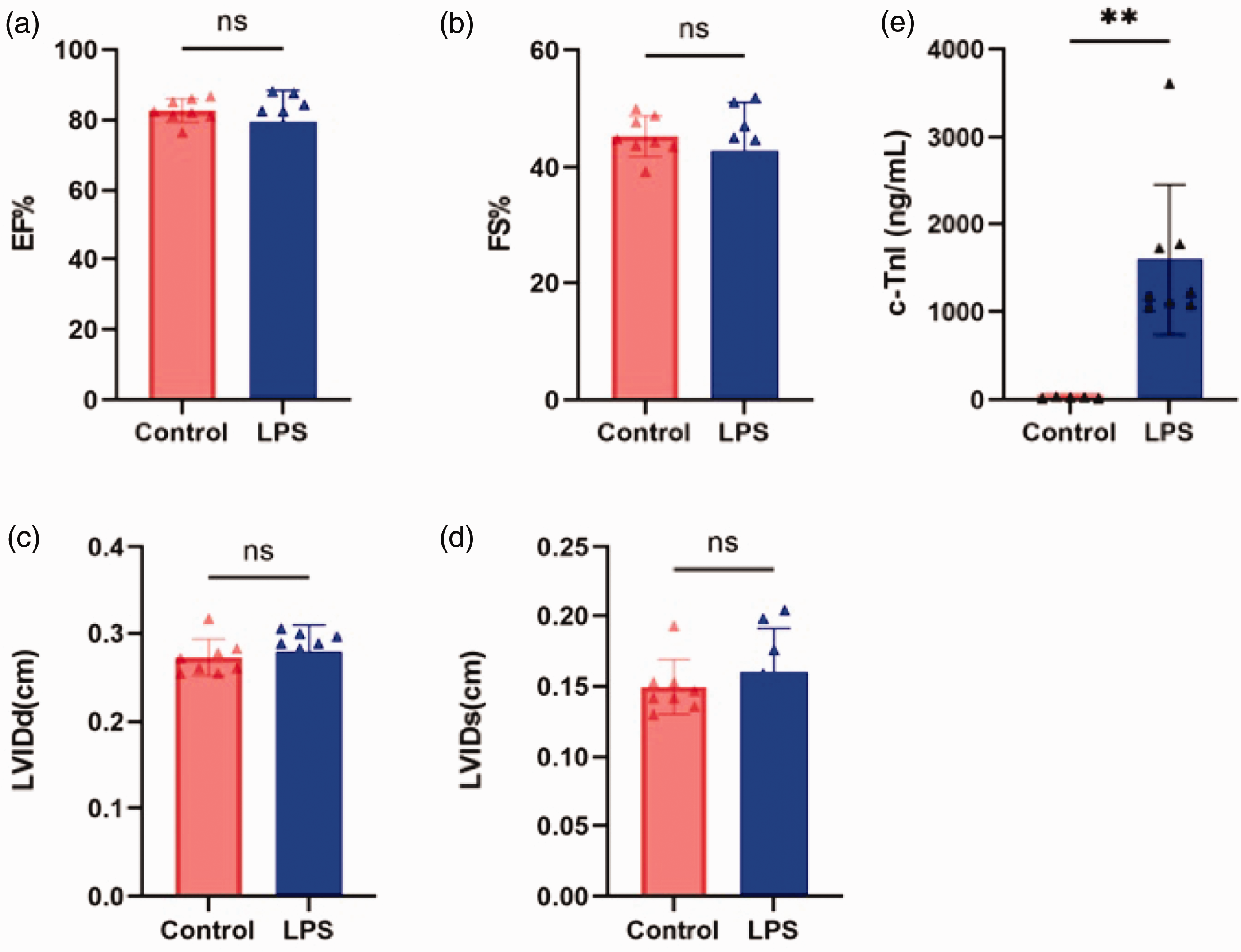
Echocardiographic data (a–d) and enzyme-linked immunosorbent assay (ELISA) results show the levels of c-TnI (e). c-TN: cardiac troponin; EF: ejection fraction; FS: fractional shortening; LPS: lipopolysaccharide; LVIDd: left ventricular end-diastolic diameter; LVIDs: left ventricular end-systolic diameter.
Differential metabolite screening
As shown in the component analysis plot (Figure 2(a)), both uniformity and uniqueness were observed between the two groups. The volcano plot (Figure 2(b)) shows the results of untargeted metabolomic analysis conducted on 16 samples, which identified a total of 1101 metabolites, including 591 in the positive ion mode and 510 in the negative ion mode. Differential metabolite screening was performed with P < 0.05 after BH correction and log2 (FC) < log2 (1/1.2) or log2 (FC) > log2 (1.2), because the number of metabolites is a necessary condition for enrichment. Differential metabolites without data on the number of metabolites were not included in the functional enrichment analysis. A total of 74 significantly altered metabolites were identified in the positive ion mode, with 48 metabolites being upregulated and 26 downregulated. In the negative ion mode, 70 significantly altered metabolites were identified, with 50 metabolites being upregulated and 20 downregulated.

(a) Principal component analysis diagram and (b) volcano map. FC: fold change; LPS: lipopolysaccharide.
Transcriptomic analysis
The boxplot of the sequenced mRNA (Figure 3(a)) and the PCA plot (Figure 3(b)) demonstrated that samples from both LPS and control groups exhibited good homogeneity and intergroup differentiation, with no outliers. The transcriptomic results, as shown in the volcano plot (Figure 3(c)), identified a total of 4831 differentially expressed genes (DEGs), of which 3027 were downregulated and 1804 upregulated. Figure 3(d) presents the heatmap of differential gene expression between the control and LPS groups, revealing distinct gene expression levels across the two groups.

(a) Box plot. (b) Principal component analysis diagram. (c) Volcano map and (d) heatmap. FC: fold change; LPS: lipopolysaccharide.
Proteomic analysis
Among the myocardial tissue samples collected from 16 mice, 4307 proteins were identified, with 4288 proteins being quantitatively analyzed. DEPs were screened based on an FC greater than 1.2 (upregulated > 1.2 or downregulated < 0.83) and a P-value <0.05. A comparison between the LPS and control groups revealed significant changes in the expression levels of 107 proteins, with 94 proteins being significantly upregulated and 13 significantly downregulated (Figure 4 and Figure 5(a)).
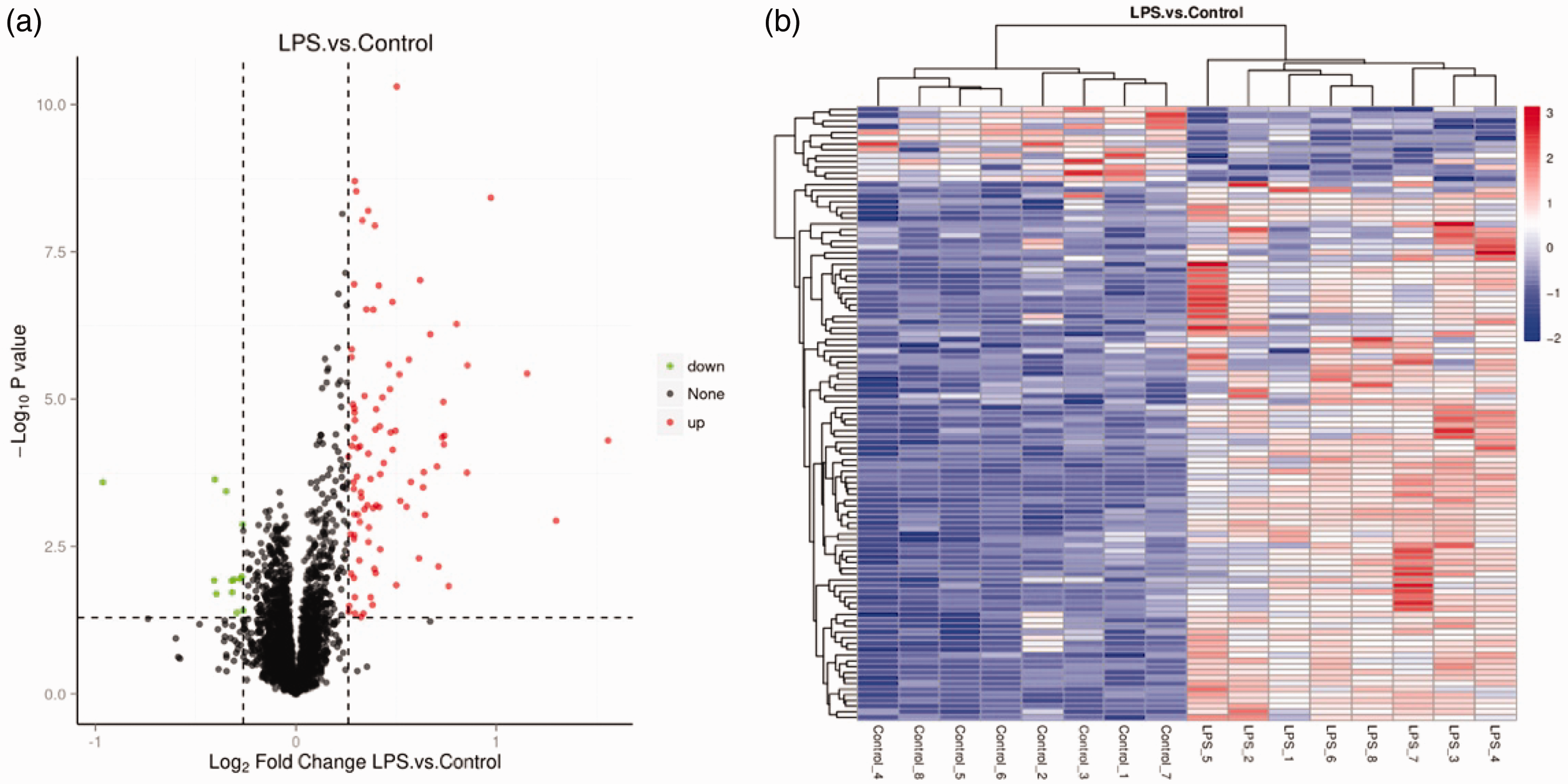
(a) Volcano map and (b) heatmap. LPS: lipopolysaccharide.
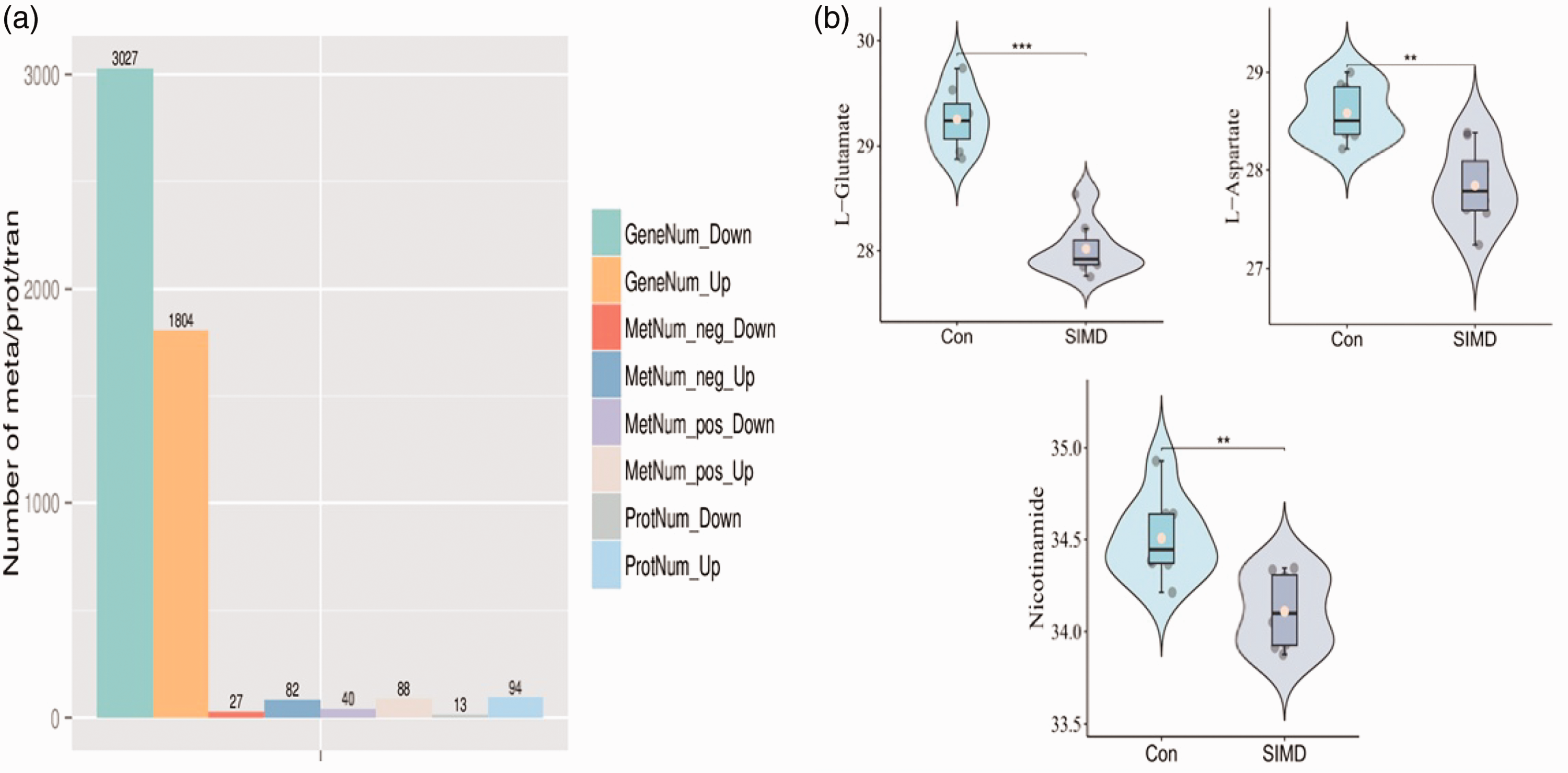
(a) Histograms of differential genes, metabolites, and proteins and (b) the differences in the three amino acids between the control group and experimental group were analyzed. SIMD: sepsis-induced myocardial dysfunction.
Integrated omics analysis
Through the combined analysis of metabolomics with transcriptomics and metabolomics with proteomics, three significantly altered metabolites were identified between the two groups of mice: L-glutamate (L-Glu), L-aspartate (L-Asp), and nicotinamide (NAM) (Figure 5(b)).
KEGG pathway intersection analysis
We performed KEGG pathway enrichment analysis for metabolite–transcriptomics and metabolite–proteomics combinations in both positive and negative ion modes, with the significance threshold for pathways set at P < 0.1. Eight results were obtained in total, and we determined the intersection of the pathways (Figure 6). We found that in the upregulated intersection of the negative metabolite–protein transcript combination, downregulated intersection of the negative metabolite–protein transcript combination, and upregulated intersection of the positive metabolite–protein transcript combination, there was only one intersecting pathway, namely, purine metabolism. In the downregulated intersection of the positive metabolite–protein transcript combination, we identified three intersecting pathways, namely, nicotinate and NAM metabolism, histidine metabolism, and nitrogen metabolism.
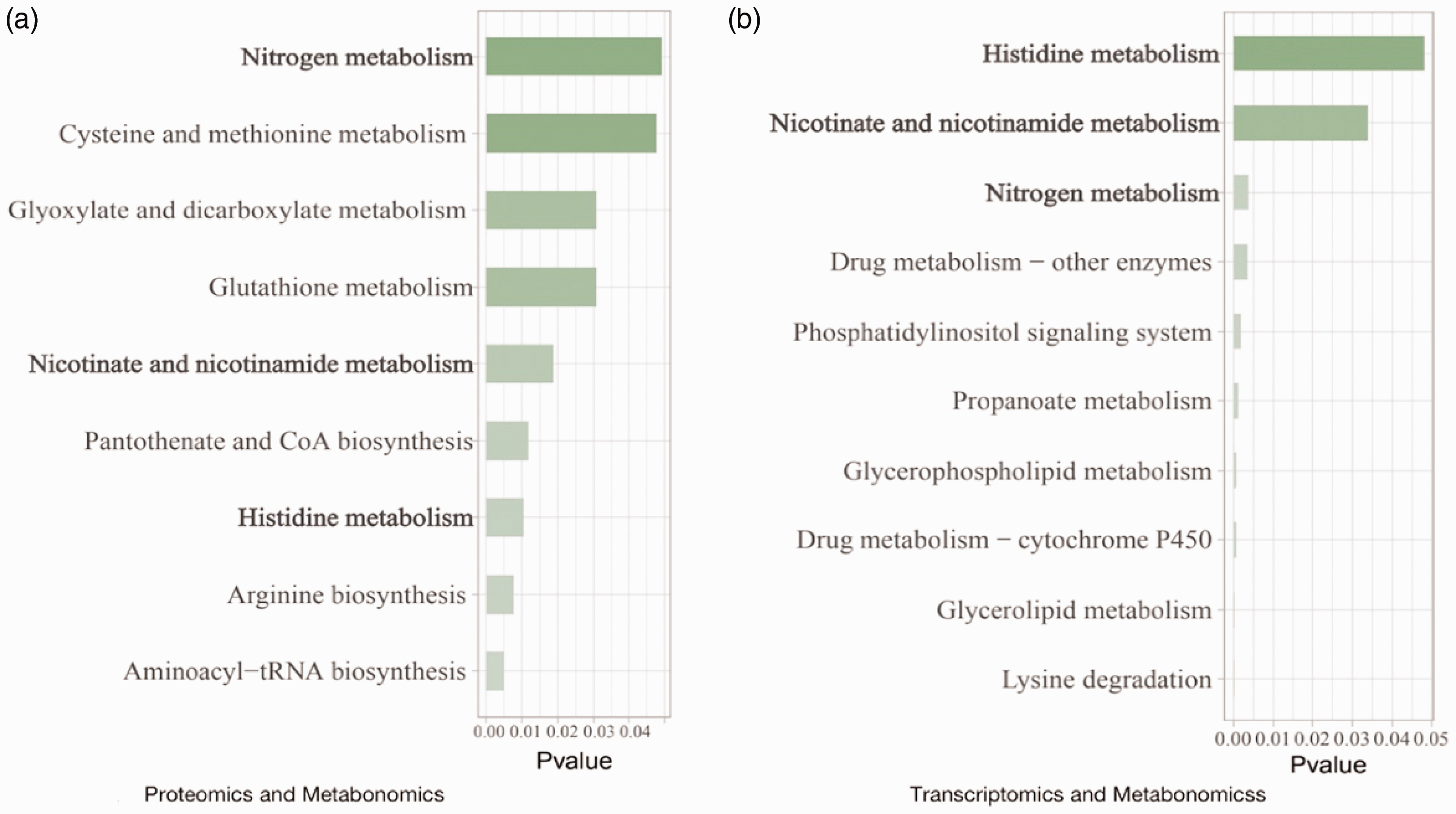
Omics-based combined analysis of enriched metabolic pathways.
Discussion
The pathogenic mechanisms of SIMD remain unclear, and omics studies can facilitate further exploration of these mechanisms. In our study, we identified significant changes in metabolites, genes, and proteins between control mice and those with SIMD. These findings suggest that three metabolic pathways, namely, nicotinate and NAM metabolism, histidine metabolism, and nitrogen metabolism, are closely related to the development and progression of SIMD, providing potential therapeutic targets for future research.
L-Glu is a primary excitatory neurotransmitter that plays a critical role in various brain functions by acting on glutamate receptors. It is the most abundant free amino acid in the brain. The majority of free L-Glu in the brain is synthesized locally from L-Glu and intermediates of the tricarboxylic acid (TCA) cycle. In the developing brain, L-Glu is essential for neuronal differentiation, migration, and survival, primarily by facilitating calcium transport.10,11 Animal studies have demonstrated that L-Glu mediates oxidative metabolism by replenishing TCA cycle intermediates lost during ischemia, thereby enhancing postischemic adenosine triphosphate (ATP) production and nearly reversing ischemic injury following cardiac arrest. 12
Although relevant studies have found the potential value of L-Glu, its wide application and specific role in clinical practice still need to be confirmed and elaborated by a large number of studies. As an isomer of glutamic acid, L-Glu is involved in both the synthesis and metabolism of glutamine. Glutamine is the most abundant free amino acid in the plasma and muscle tissue. During sepsis, plasma glutamine is transported into cells, where it enters the mitochondria via the TCA cycle, providing energy and intermediates for biosynthesis. A clinical study reported that glutamine levels were significantly elevated in the SIMD group compared with those in the non-SIMD group, with a strong correlation observed between glutamine levels and the key marker of myocardial injury creatine kinase myocardial band (CK-MB). 13 This may be attributed to SIMD-induced disruptions in the mitochondrial metabolism of glutamine to glutamate, resulting in glutamine accumulation. In our integrated omics analysis, we similarly observed a significant reduction in L-Glu levels in SIMD mice. This finding offers potential clinical evidence supporting the involvement of energy metabolism mechanisms in the pathogenesis of SIMD.
L-Asp is a nonessential amino acid that is widely involved in biosynthetic processes and serves as a central component of numerous metabolic pathways in most organisms. Similar to L-Glu, L-Asp functions as an excitatory neurotransmitter, inducing depolarization of the postsynaptic membrane and primarily regulating metabolic processes related to the nervous system and brain function. Additionally, L-Asp has been used in the treatment of certain cardiac diseases. Research has shown that the administration of L-Asp exerts a protective effect on mitochondrial function in rat hearts following myocardial infarction. 14 As mitochondrial dysfunction is one of the pathogenic mechanisms of SIMD, our study revealed a significant decrease in L-Asp levels in SIMD mice. This finding could serve as a reference for further investigation into the specific mechanisms of SIMD.
NAM is a metabolite of niacin, and both are forms of vitamin B3. Niacin and NAM play crucial roles in maintaining redox balance as well as in lipid metabolism, glycolysis, carbohydrate metabolism, and protein and amino acid metabolism within the body. NAM is also a component of coenzyme I (NAD+) and coenzyme II (NADP+), which are involved in energy and material metabolism. NAD+ can be reduced to NADH during cellular redox reactions or degraded into NAM or niacin by NAD+-consuming enzymes. Studies have shown that a deficiency in sirtuin (SIRT)-1 increases sepsis-related mortality. 15 Supplementation with NAD+ effectively upregulates SIRT1 expression, which in turn mediates T lymphocyte differentiation and immune responses. Immediate administration of NAM riboside (a precursor of NAD+) after sepsis not only suppresses tumor necrosis factor (TNF)-α expression but also alleviates immunosuppression, thereby enhancing bacterial clearance and improving sepsis outcomes. 16 Furthermore, a cohort study reported that high intake of NAM could improve diastolic dysfunction induced by aging, hypertension, or cardiometabolic syndrome. 17 Increasing NAD+ levels through NAM riboside supplementation has been proposed as a novel treatment for heart failure with reduced ejection fraction. 18 Although the mechanisms underlying SIMD are not fully understood, they likely involve energy metabolism and mitochondrial dysfunction, and the downregulation of NAM levels may contribute to the development and progression of SIMD.
This study identified three significantly different metabolites, primarily concentrated in three metabolic pathways, namely, nicotinate and NAM metabolism, histidine metabolism, and nitrogen metabolism. As mentioned previously, niacin and NAM form coenzymes with adenine, ribose, and phosphate in the body, serving as key components of NAD and NADP. NAM is involved in various metabolic processes related to the formation of high-energy phosphate bonds. 19 Importantly, in animal models, NAM mononucleotide has been shown to preserve mitochondrial function and improve survival in hemorrhagic shock. 20 The dysregulation of the niacin and NAM metabolic pathways plays a critical role in the pathogenesis and progression of SIMD. During sepsis, the massive release of inflammatory cytokines and enhanced oxidative stress may disrupt the activity of key enzymes in these pathways, thereby impairing the synthesis and regeneration of NAD+. The heart, being one of the most metabolically active organs in the human body, is highly dependent on mitochondrial function. Mitochondrial dysfunction is closely linked to myocardial injury mechanisms. NAD+ serves as an essential coenzyme for mitochondrial respiratory chain complex I, and its deficiency compromises mitochondrial function, leading to reduced ATP production and insufficient energy supply in cardiomyocytes. Furthermore, NAD+ acts as a substrate for the SIRT family. SIRTs regulate the expression of genes involved in metabolic and stress responses through their deacetylation activity. A decline in NAD+ levels suppresses the activity of SIRTs, thereby diminishing the stress adaptation and repair capacity of cardiomyocytes. Additionally, disturbances in the niacin and NAM metabolic pathways may indirectly exacerbate SIMD pathology by affecting the activity and function of immune cells. Reduced NAD+ levels inhibit the proliferation and activation of T cells and B cells, thereby weakening the host’s immune defense and rendering cardiomyocytes more vulnerable to pathogen invasion and damage. NAM administration also reduces inflammatory cell aggregates, thrombi, and cell death in mice infected with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). 21 Dysregulation of the niacin and NAM metabolic pathways drives inflammatory responses and induces cellular apoptosis. NAD+ deficiency may render cells more susceptible to inflammatory stimuli, exacerbating the inflammatory cascade, while simultaneously promoting cardiomyocyte apoptosis, collectively driving the progression of SIMD.
Histidine, an essential amino acid and one of the least abundant amino acids in the body’s total protein content, exhibits antioxidant and anti-inflammatory properties. In addition to its role in protein synthesis, histidine is involved in different metabolic pathways, where it can condense to form carnosine or decarboxylate to synthesize histamine. 22 Histamine plays a critical role in the cardiovascular system by regulating vascular tone and enhancing myocardial contractility. A decline in histidine levels may reduce histamine synthesis, thereby impairing cardiac contractile function. Histidine also participates in the synthesis of carnosine, which exerts antioxidant, anti-inflammatory, and cardioprotective effects. Dysregulation of the histidine metabolic pathway may lead to diminished carnosine production, exacerbating cardiomyocyte injury and dysfunction. Additionally, histidine enters the TCA cycle to provide energy for cardiomyocytes. In sepsis, impairment of histidine metabolism may disrupt myocardial energy supply, further aggravating cardiac dysfunction. Histidine may protect the heart by reducing free radical–induced toxicity. 23 The imbalance of the histidine metabolic pathway drives the progression of SIMD through three aspects: proinflammatory signals, oxidative damage, and energy crisis.
Nitrogen is one of the essential elements for life, and nitrogen metabolism is a fundamental biological process involving signal transduction, protein synthesis, and other functions. NO, a nitrogen oxide produced by nitrate reductase during nitrogen assimilation, is considered a key player in the pathogenesis of SIMD when NO synthesis is impaired. 24 Under septic conditions, inflammatory cytokines activate inducible nitric oxide synthase (iNOS), leading to dysregulation of NO homeostasis. 25 Pathological accumulation of NO inhibits cytochrome c oxidase, thereby disrupting oxidative phosphorylation and resulting in ATP depletion and contractile dysfunction in cardiomyocytes. Furthermore, the buildup of nitrogen metabolism intermediates (e.g. glutamine and citrulline) may perturb immunometabolic balance, promoting cardiomyocyte apoptosis. 26 Future studies may focus on these three pathways to further explore the mechanisms underlying SIMD.
This study has several limitations. First, the LPS-induced murine sepsis model used to simulate myocardial injury did not fully encompass the complex pathogenic factors observed in humans, such as species-specific immune responses and the absence of polymicrobial interactions. Future studies should establish more comprehensive animal models (e.g. cecal ligation and puncture for polymicrobial sepsis) to dissect the multifaceted mechanisms of SIMD. Second, while our findings are derived solely from animal experiments, future research should integrate clinical data (e.g. plasma metabolomics from septic patients) with experimental approaches to validate the translational relevance of SIMD-related pathways. At present, there is no unified guideline to determine the diagnostic criteria for SIMD, and most of them use common indicators (cardiac enzyme indicators and echocardiographic results) as diagnostic criteria. The echocardiogram results in the animal modeling part of this study did not show the desired results, but the cardiac enzyme indexes changed significantly. Moreover, the c-TnI results can confirm the successful establishment of mouse models of LPS-induced myocardial dysfunction based on the experience of successful modeling in previous studies.27–29 In this experiment, the early cardiac enzyme indexes of SIMD mice were significant, and the negative echocardiographic results may be caused by the following factors: 1. LPS-induced septic myocardial dysfunction exhibits a temporal window effect, wherein the release of myocardial injury biomarkers (c-TnI) may precede detectable functional alterations. 30 2. During the early phase of sepsis, the organism may activate compensatory mechanisms to preserve cardiac output, potentially obscuring the true extent of myocardial functional impairment. 31 3. Sepsis-associated myocardial injury often manifests as focal or regional pathological changes rather than uniform myocardial involvement. Conventional echocardiographic assessment of global cardiac systolic and diastolic function may fail to detect such localized myocardial damage. 31
Conclusion
Compared with control mice, significant changes in metabolites were observed in the cardiac tissue of SIMD mice. These differences are primarily associated with three pathways, namely, nicotinate and NAM metabolism, histidine metabolism, and nitrogen metabolism. Our results offer new insights and perspectives for investigating the specific mechanisms underlying SIMD.
Footnotes
Acknowledgements
The authors would like to express gratitude to the First Affiliated Hospital of Harbin Medical University for help with conducting the study.
Author contributions
All authors participated in the design of the study. KY and JZ searched for the articles, screened titles and abstracts, and extracted data. JZ, ZD, WW, and SS performed statistical analysis and interpretation of data. JZ, ZD, WW, and SS drafted the manuscript, and all authors revised it for important intellectual content. Final approval of the version submitted for publication was obtained from all authors.
Data availability statement
All data generated or analyzed during this study are included in this published article.
Declaration of conflicting interests
The authors declare that they have no competing interests.
Ethics approval and consent to participate
The animal study was reviewed and approved by the Ethics Committee of The First Affiliated Hospital of Harbin Medical University.
Funding
This project was funded by the National Natural Science Foundation of China (Nos. 81770276).
