Abstract
Objective
The mechanism of severe acute respiratory syndrome coronavirus 2 in infecting human cells requires binding of the virus to angiotensin-converting enzyme 2. Angiotensin-converting enzyme 2 is also involved in blood pressure regulation. There are several single-nucleotide polymorphisms in the angiotensin-converting enzyme 2 genome. As the occurrence of clinical symptoms and response to different treatments in patients with coronavirus disease 2019 are different, it seems that single-nucleotide polymorphisms can play a role in these differences. This receptor became important during the coronavirus disease 2019 pandemic because the coronavirus can enter host cells via this receptor. In this study, we aimed to discover the correlation between ACE2 gene polymorphism, hypertension, and anti–coronavirus disease 2019 drug usage in patients with coronavirus disease 2019.
Methods
We performed a cross-sectional study, and two polymorphisms, rs1978124 and rs2158083, were analyzed in 75 patients with essential hypertension and coronavirus disease 2019. All patients were genotyped using sequencing analysis methods, and the correlation between polymorphism, hypertension, severity of other diseases, and treatment response to anti–coronavirus disease 2019 drugs was determined using SPSS 20.0.
Results
There was no significant association between single-nucleotide polymorphisms and the risk of being hypertensive with other diseases and the risk of diabetes in unadjusted and adjusted models in patients with coronavirus disease 2019 (P > 0.05).
Conclusion
No significant association was found between single-nucleotide polymorphisms and hypertension in patients with coronavirus disease 2019.
Keywords
Introduction
Angiotensin-converting enzyme 2 (ACE2) is a key enzyme of the renin–angiotensin system and is an essential target for the prevention and treatment of hypertension. The genomic map of ACE2 includes 39.98 kilobases, 20 introns, and 18 exons. ACE2 is an enzyme with 802 amino acids and is located in the cell membranes of the lungs, arteries, heart, kidney, and intestine. 1
The ACE2 genome shows a high degree of polymorphism. ACE2 single-nucleotide polymorphism (SNP) is associated with blood pressure in Europeans (rs2074192, rs233575, and rs2158083), Australians (rs1978124 and rs2285666), Indians (rs2285666), and Koreans (rs1514282 and rs1514283). A meta-analysis of the association between blood pressure and ACE2 polymorphism confirmed that rs2285666 and rs2106809 are associated with the risk of hypertension in Han subjects.2–5 Therefore, ACE2 genetic variants that play critical roles in high blood pressure have been identified in some countries, but until now, no similar study has been conducted in Iran.
During the coronavirus disease 2019 (COVID-19) pandemic, several studies identified increased mortality in patients with hypertension who had COVID-19. For example, Wu et al. 6 and Zhou et al. 7 showed that hypertension had hazard ratios of 1.70 and 3.05 for death in 201 and 191 patients with COVID-19, respectively. Hence, in this study, we aimed to investigate the relationship between two SNPs of ACE2, rs2158083 and rs1978124, and hypertension in patients with COVID-19.
Materials and methods
Participants
This study was a cross-sectional investigation that included 75 patients. The inclusion criteria were essential hypertension and COVID-19, confirmed by a real-time polymerase chain reaction (PCR) test. All patients were Iranian and were living in Isfahan Province. Some participants also had diabetes mellitus or secondary hypertension. Patient characteristics (such as sex, SNP type, comorbidities, age, hydroxychloroquine (HCQ) usage, and illness severity) are summarized in Table S1; however, detailed patient information is omitted. After the patients had rested for 5 min, blood pressure was measured in a quiet setting using a mercury sphygmomanometer while the patients were seated. Then, 10 mL of blood was taken from the patients and collected in an ethylenediaminetetraacetic acid tube and then divided into sterile microtubes. Samples were stored in a freezer at −80°C until the next step. All patients completed the consent form with the scientific code 198286 and the ethical code IR.MUI.RESEARCH.REC.1398.828.
DNA extraction and PCR
The AddPrep Genomic DNA Extraction Kit (Addbio; Product Code: 10023) was used for isolating genomic DNA from blood samples. Primers for rs2158083 and rs1978124 were used to screen these SNPs. Primer sequences and PCR product sizes are provided in Table S2. PCR was performed with a final volume of 20 μL, which contained 10 μL of PCR master mix, 1 μM each of forward and reverse primers, and 3 μL of DNA. The reaction conditions were as follows: 95°C for 3 min, 30 cycles of 95°C for 30 s, 60°C for 30 s, 72°C for 20 s, followed by 72°C for 5 min. The 5-μL PCR products were analyzed using 1.0% agarose gel electrophoresis.
Determination of ACE2 genotypes
Sanger sequencing was used to determine the sequence of nucleotide bases in the PCR product. First, the amplified product from the previous step was electrophoresed on a 1% agarose gel, and the results were viewed using a ultraviolet transilluminator. DNA sequencing was then conducted by sending the PCR product to the core research facilities of Isfahan University of Medical Sciences. The outcomes were analyzed using Chromas software.
Statistical analysis
Continuous variables were expressed as means (SDs), and categorical variables were described as numbers (percentages). Comparisons of the means of continuous variables between two groups were performed using independent Student t-test for variables with normal distributions and the Mann–Whitney test for those with nonnormal distributions. Logistic regression models were utilized to assess the association between SNPs and the risk of high hypertension with other diseases and the risk of diabetes among high hypertensive individuals. All statistical analyses were conducted using SPSS 20.0 for Windows. P values of less than 0.05 were considered to indicate statistical significance.
Molecular docking of the ACE2–HCQ complex
AutoDock Vina 4.252 8 SwissDock servers (https://www.swissdock.ch/) were used for docking of HCQ and ACE2. AutoDockTools 1.5.6 was used to prepare lactate dehydrogenase (LDH) A and peptides. Polar hydrogens were added and Gasteiger charges were calculated. The torsion tree was identified, and molecules were saved in the PDBQT format and used for the docking process. The 52 × 52 × 50 Å grid map was set along the x, y, and z-axes for residues 300–305 of the LDHA C-terminal domain. The default parameters were used while implementing the Lamarckian genetic algorithm. Docking was performed with a fixed receptor, while the peptides were considered as being flexible. The docking experiments were repeated at least three times to achieve consistent results. Next, the complex binding modes were studied using Swiss-PdbViewer 9 and Pymol. 10
Results
Data from 75 patients were available for the present study; 52% of the patients were male. Table 1 shows the characteristics of the patients in the study. The mean (SD) age of the patients was 68.08 (12.16) years. There were no significant differences in the distribution of sex and mean age between the two groups of patients with hypertension and patients with hypertension and other diseases as well as between the two groups of patients with and without diabetes (P > 0.05). Tables 2 and 3 show the frequencies (percentages) of SNPs between the two groups of patients with hypertension and patients with hypertension and other diseases and between the two groups of patients with and without diabetes, respectively. Furthermore, they present the odds ratios and 95% confidence intervals. There was no significant association between SNPs and the risk of being hypertensive with other diseases and the risk of diabetes in unadjusted and adjusted models (P > 0.05) (Tables 2 and 3).
Characteristics of patients in the study.

Association of SNPs with the risk of hypertension and other diseases in the study patients.
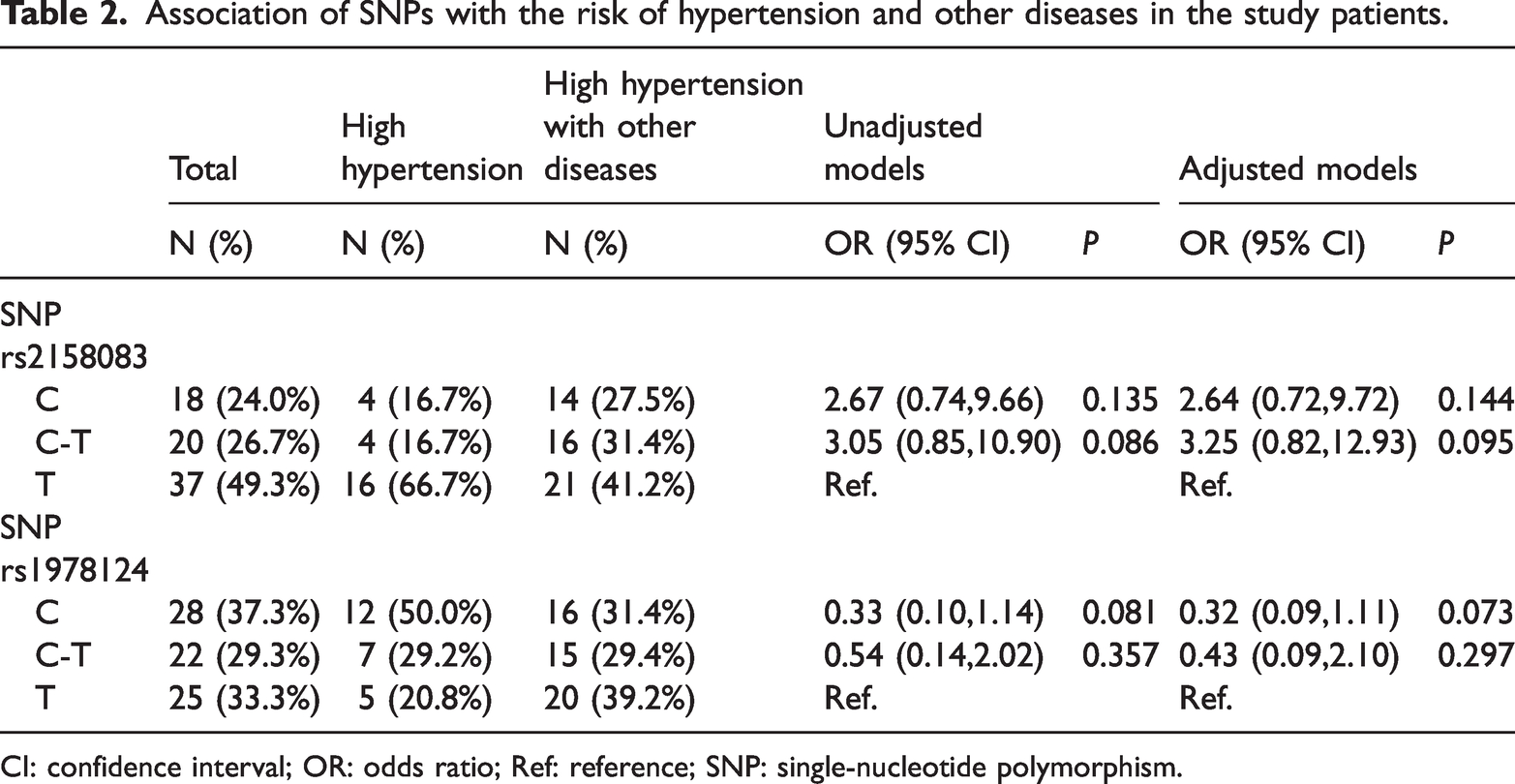
CI: confidence interval; OR: odds ratio; Ref: reference; SNP: single-nucleotide polymorphism.
Association of SNPs with the risk of diabetes and other diseases in the study patients.

CI: confidence interval; OR: odds ratio; SNP: single-nucleotide polymorphism.
In another analysis, the relationship between SNP types and disease severity was investigated. Patients were divided into three groups based on symptoms and medical history, as shown in Table 4. In SNP 21, most patients with T allele and C-T allele had severe disease severity (more than 40%). In SNP 19, disease severity was reported to be severe for most patients in all alleles. Of course, the difference in the severity of the disease in terms of alleles in both SNPs was not significant (P > 0.05). All patients in this study used HCQ as an anti–COVID-19 drug, and of the 75 patients in this study, only 4 died. The remaining patients showed a relatively good response to the treatment, and all of them were discharged from the hospital (Table S1). Therefore, ACE2 polymorphism does not play a role in the response to anti–COVID-19 drugs.
Association of SNPs with COVID-19 severity.

COVID-19: coronavirus disease 2019; SNP: single-nucleotide polymorphism.
Molecular docking of ACE2 with HCQ
Molecular docking was used to provide details on HCQ’s role in inhibiting viral infection. The interaction site of HCQ with ACE2 was investigated using AutoDock and SwissDock server. Based on the docking outcome, HCQ can attach to two areas, as shown in Figure 1(a) to (d). Interacting regions include Phe40, Ser44, Ser47, Thr347, Ala348, Asp350, and His401. Luc73, Ser77, Ala99, Luc100, Gln102, Phe390, Luc391, Arg393, and Asn394 are engaged with different areas. Because the regions of ACE2 responsible for binding HCQ are close to the binding site of the spike, HCQ may interfere with the spike’s binding to the receptor.
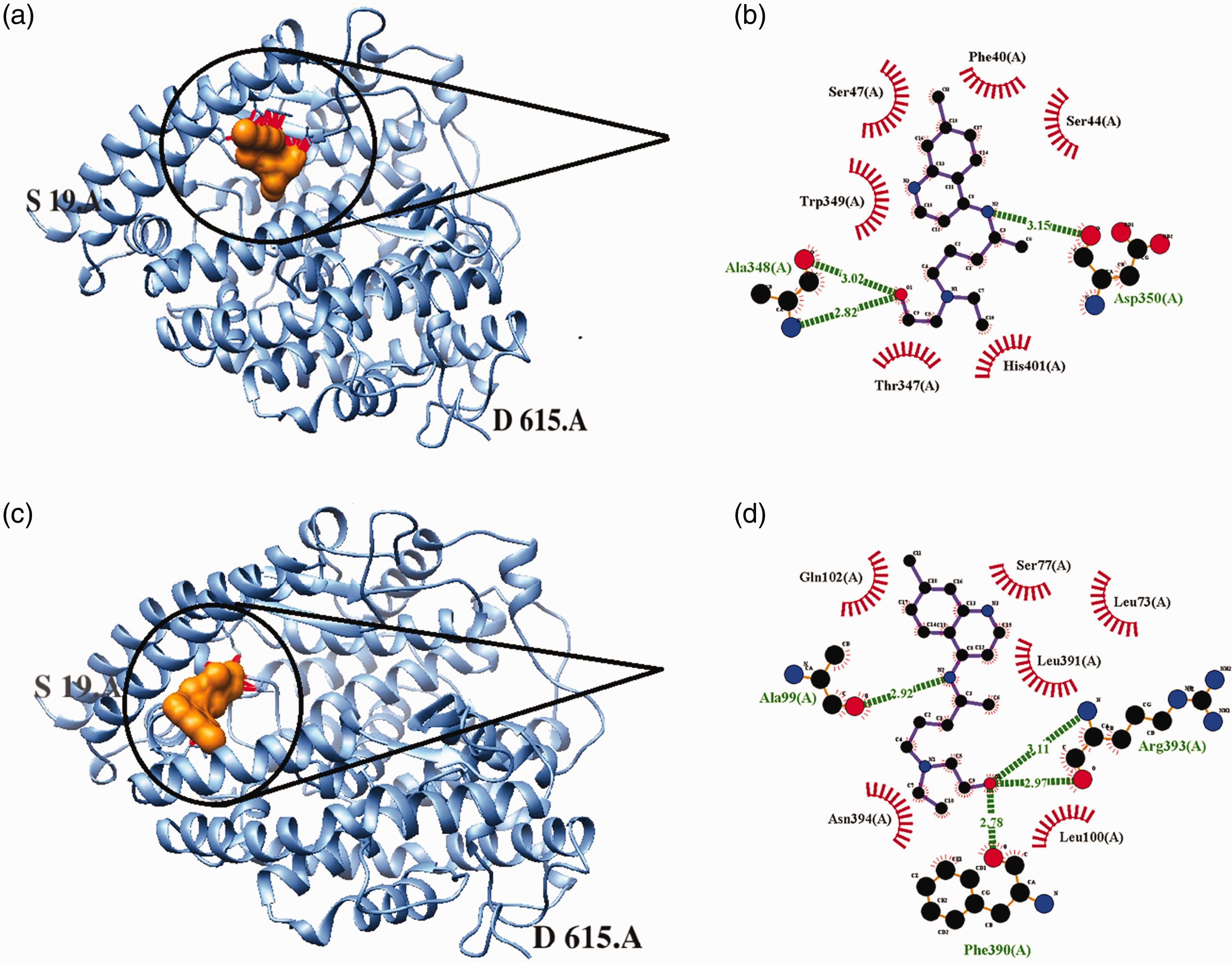
The three-dimensional structure of ACE2 complexes with hydroxychloroquine was obtained from AutoDock and SwissDock servers, as indicated in (a) and (b), respectively. The interaction pattern of ACE2 complexes with hydroxychloroquine was indicated by ligplot in (c) and (d). ACE: angiotensin-converting enzyme.
Discussion
ACE2 is present on different cell types such as monocytes, smooth muscle cells, T lymphocytes, and adipocytes as well as on the endothelium of lung vessels. ACE2 has a critical role in blood pressure regulation and the renin–angiotensin system and has been a possible therapeutic target in the treatment of hypertension. 11 The ACE2 genome exhibits a high level of polymorphism. 12 Rice et al. 13 were one of the first groups to show the association between ACE2 polymorphisms and the genetics of hypertension as well as circulating ACE2 concentrations. Several other studies have shown the link between ACE2 hereditary variations and the risk of developing hypertension in different ethnic groups. Moreover, the functions of ACE2 as a cellular receptor for severe acute respiratory syndrome coronavirus (SARS-CoV)-2 in many human tissues and organs lead to ACE2 being additionally included in the pathogenicity of COVID-19.14–16
The differences in the activity of specific proteins can be attributed to the presence of common polymorphic genetic variations within a population that lead to increased susceptibility to infection, increased efficiency of viral replication, or an overstated inflammatory reaction. 17 Because people’s responses to the coronavirus were different, there were hypotheses that people with high blood pressure have a specific polymorphism of this gene. To determine whether people with high blood pressure are more susceptible to COVID-19, we decided to analyze the relationship between two SNPs of ACE2, rs2158083 and rs1978124, and hypertension in patients with COVID-19. The frequencies of rs2158083 and rs1978124 were 0.3789 and 0.4502, respectively, and several studies have reported the relationship between rs1978124 and hypertension, dyslipidemia, and diabetes.3,14,18,19 A recent study showed the role of rs2158083 in obesity and hyperlipidemia. 20 With the spread of coronavirus, the study on ACE2 receptor polymorphism has intensified, and several studies have reported that ACE2 receptor polymorphism may play a role in SARS-CoV-2 infection.21–23 Other studies reported that the rate of mortality from COVID-19 was notably greater in patients carrying the T allele of ACE2 rs1978124. Based on the data, the frequency of the T allele was 0.68. In other regions, the frequencies were as follows: (a) Asian, 0.20; (b) South Asian, 0.78; (c) East Asian, 0.14; (d) other Asian, 0.30; (e) African, 0.261; (f) African–American, 0.261; (g) European, 0.625; and (h) Latin American, 0.744 (https://www.ncbi.nlm.nih.gov/snp/rs2074192). 24
Some studies reported that rs2158083 could increase the binding affinity of ACE2 toward SARS-CoV-2.25,26
According to the abovementioned data, the two SNPs rs2158083 and rs1978124 can affect and contribute to the increased risk of COVID-19 and other diseases such as hypertension, dyslipidemia, and diabetes. Various studies in Iran have evaluated the association of gene polymorphisms involved in SARS-CoV-2 infection, such as interferon lambda 3 and 4 (rs12979860, rs12980275, rs8099917, and rs368234815), tripartite motif containing-22 (rs7113258, rs7935564, and rs1063303), vitamin D receptor gene (ApaI, BsmI, BglI, TaqI, FokI, and Tru9I), interferon-induced transmembrane protein 3 (rs6598045 and rs12252), transmembrane serine protease 2 (rs12329760), interleukin-10 (rs1800871, rs1800872, and rs1800896), and ABO rs657152.2–8,11–13 However, the relationship between the two SNPs rs2158083 and rs1978124, hypertension, diabetes, and COVID-19 has not yet been detected, and we analyzed this relationship in the current study.
Based on our results, there was no significant association between the two SNPs (rs2158083 and rs1978124) and the risk of hypertension with other diseases and the risk of diabetes in unadjusted and adjusted models (P > 0.05). These results are opposite to those of other studies that indicated an association between ACE2 polymorphisms and hypertension, dyslipidemia, diabetes, and SARS-CoV-2 infection.3,14,17–21 One of the important reasons for this result is that these SNPs are not in the coding region on ACE2 and do not play a role in the structure and function of ACE2 receptor. Finally, we used molecular docking techniques to determine the binding site of HCQ on the ACE2 receptor. According to our results, the interaction of HCQ with ACE2 may disrupt the interaction between the ACE2 and spike receptor. Another study demonstrated that HCQ influences the movement of key amino acids within the interaction site between the viral spike protein and the ACE2 receptor. Navya et al. 27 suggested that HCQ-induced motion may reduce the spike protein’s affinity toward the ACE2 receptor.
Study limitations
One limitation of this study was the difficulty in collecting samples from patients with both COVID-19 and hypertension. Another limitation was limited access to laboratory tests and patient medical records.
Conclusion
To the best of our knowledge, we first identified the roles of two SNPs of ACE2—rs2158083 and rs1978124—in hypertension in patients with COVID-19. Based on our results, there is no significant correlation between SNPs and hypertension.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605251375221 - Supplemental material for Association of angiotensin-converting enzyme 2 gene polymorphisms with hypertension and treatment response to hydroxychloroquine in Iranian people with coronavirus disease 2019: A cross-sectional study
Supplemental material, sj-pdf-1-imr-10.1177_03000605251375221 for Association of angiotensin-converting enzyme 2 gene polymorphisms with hypertension and treatment response to hydroxychloroquine in Iranian people with coronavirus disease 2019: A cross-sectional study by Farzaneh Jafary, Mohammad Kazemi, Maryam Zarean, Mehri Khoshhali, Maryam Karimian, Fataneh Fani and Kiana Shahzamani in Journal of International Medical Research
Footnotes
Acknowledgments
The authors have no acknowledgments to disclose.
Author contributions
Study design: Farzaneh Jafary, Kiana Shahzamani, Maryam Zarean. Study conduct: Farzaneh Jafary, Maryam Karimian, Maryam Zarean. Data collection: Farzaneh Jafary, Kiana Shahzamani, Maryam Zarean, Mehri Khoshhali, Maryam Karimian, Fataneh Fani. Data analysis: Mehri Khoshhali. Data interpretation: Farzaneh Jafary, Maryam Zarean. Drafting of the manuscript: Farzaneh Jafary, Maryam Zarean, Mehri Khoshhali. Revision of the manuscript content: Farzaneh Jafary, Kiana Shahzamani. Approval of the final version of the manuscript: Farzaneh Jafary, Kiana Shahzamani, Maryam Zarean.
Data availability
The data used in this research are available from the biobank thesis entitled, “Establishing a biobank for collection, storing and analyzing bio specimens of patients with COVID-19,” with the ethical code IR.MUI.RESEARCH.REC.1398.828.
Declaration of conflicting interest
The authors declare no competing interests.
Funding
We appreciate the support provided by Isfahan University of Medical Sciences under grant No. 199408 for this study. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
