Abstract
Objective
Matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) is currently used in clinical microbiology laboratories. This study aimed to determine whether dual-polarity time-of-flight mass spectrometry (DP-TOF MS) could be applied to clinical nucleotide detection.
Methods
This prospective study included 40 healthy individuals and 110 patients diagnosed with cardiovascular diseases. We used DP-TOF MS and Sanger sequencing to evaluate 17 loci across 11 genes associated with cardiovascular drug responses. In addition, we used DP-TOF MS to test 998 retrospectively collected clinical DNA samples with known results.
Results
A, T, and G nucleotide detection by DP-TOF MS and Sanger sequencing revealed 100% concordance, whereas the C nucleotide concordance was 99.86%. Genotyping based on the results of the two methods showed 99.96% concordance. Regarding clinical applications, DP-TOF MS yielded a 99.91% concordance rate for known loci. The minimum detection limit for DNA was 0.4 ng; the inter-assay and intra-assay precision rates were both 100%. Anti-interference analysis showed that aerosol contamination greater than 1013 copies/µL in the laboratory environment could influence the results of DP-TOF MS.
Conclusions
The DP-TOF MS platform displayed good detection performance, as demonstrated by its 99.96% concordance rate with Sanger sequencing. Thus, it may be applied to clinical nucleotide detection.
Keywords
Introduction
Mass spectrometry (MS) is an analytical technique used to determine the mass-to-charge ratio (m/z) of chemical compounds. 1 Multiple variations of this technique have been developed with different ionization methods and detectors.2–4 Matrix-assisted laser desorption ionization time-of-flight MS (MALDI-TOF MS) is currently used for the rapid and reproducible identification of bacteria, viruses, parasites, and fungi in clinical microbiology laboratories. 5 Future applications of MALDI-TOF MS in clinical diagnosis will likely be driven by technical advances in equipment, software, and sample preparation.6–9 Moreover, new MALDI-TOF MS applications may be developed that could revolutionize molecular diagnostics, such as nucleotide detection. 10 MALDI-TOF MS techniques offer new approaches in pharmacogenomics, genetic disease testing, methylation, laboratory medicine, toxicology, microbiology, health management, and molecular pathology; they have also enabled reliable, cost-effective, and rapid detection of amplified polymerase chain reaction (PCR) products.11–13 Thus, this method might complement the existing diagnostic arsenal and could be especially valuable when reagent shortages limit the use of other techniques, such as quantitative reverse transcription PCR (RT-qPCR), the current gold standard for detecting severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).14,15 In addition to serving as a cost-effective genotyping method, MALDI-TOF MS provides a platform for validating variants observed through high-throughput next-generation sequencing (NGS). 16
Numerous gene detection technologies developed in the 21st century, 17 including RT-qPCR, digital PCR, gene chip, Sanger sequencing, and NGS, have been applied to clinical testing in various contexts.18–20 Currently, clinical diagnosis is moving towards “omics” (multigene and multisite) methodologies.21,22 Thus, detection methods with high sensitivity, low cost, simplicity, and feasibility are needed to meet a wide range of clinical demands. In recent years, MS technologies exhibiting high sensitivity, specificity, and throughput, combined with inexpensive chip technology, have promoted the rapid development and use of MALDI-TOF MS. Accordingly, MALDI-TOF MS platforms can now accurately distinguish among A, T, C, and G bases. This technology has broad application potential in areas such as pharmacogenomics, tumor gene mutation detection, drug resistance studies, genetic screening, and liquid biopsy.23–27
This study aimed to evaluate the performance of dual-polarity time-of-flight MS (DP-TOF MS)—based on MALDI-TOF MS technology—combined with the chip sampling and MS module by comparing it with Sanger sequencing for the detection of genes associated with cardiovascular drug responses. In addition, we aimed to validate DP-TOF MS for clinical applications using a large number of retrospective samples.
Materials and methods
Sample collection
In total, 150 consecutive samples were prospectively collected from 92 men and 58 women (aged 18 to 65 years) at Fuwai Hospital, Chinese Academy of Medical Sciences, Beijing, China. To ensure diversity in sample types and results, 40 samples were obtained from healthy individuals and 110 samples were obtained from patients with cardiovascular disease diagnosed using the 2014 American College of Cardiology/American Heart Association criteria for coronary syndrome. 28 Low-frequency gene detection was performed on whole blood (138) and DNA (12) samples. The study protocol complied with all relevant national regulations and institutional policies for research involving human participants, was conducted in accordance with the tenets of the Helsinki Declaration of 1975 (revised in 2013), and was approved by the Ethics Committee of Fuwai Hospital (approval no. 2019-1153). Written informed consent was obtained from all individuals included in this study, and all participants’ details were de-identified. Reporting in this study complies with Standards for Reporting Qualitative Research (SRQR) guidelines (Supplementary Table 1). 29
In addition, 998 consecutive DNA samples, collected from September 2017 to October 2018 at Fuwai Hospital, were retrospectively included in the study. All 998 samples had already been tested by Sanger sequencing or RT-qPCR; in this study, they were evaluated by DP-TOF MS. The samples were collected from 502 men and 496 women. Among the samples, 280 were collected from patients aged 18 to 30 years, 422 were collected from patients aged 31 to 64 years, and 296 were collected from patients aged 65 years or older.
For all samples, the inclusion criteria were as follows: 1) whole blood or cryopreserved DNA samples with clear traceability; 2) samples considered usable by the research team, without age or sex restrictions; and 3) samples from healthy individuals or patients with cardiovascular disease. Exclusion criteria for all samples were as follows: 1) unclear sample collection time or missing information; 2) insufficient sample volume due to testing errors; 3) samples not meeting standard requirements for collection, processing, and/or storage: 4) duplicate samples; 5) samples with incomplete or untraceable information; and 6) samples considered otherwise unsuitable by the investigators.
Instruments and analytical methods
Genomic DNA was extracted from whole blood samples using an NP968-C nucleic acid extractor and the corresponding nucleic acid extraction kit (Tianlong Technology, Xi’an, China), in accordance with the manufacturer’s instructions. DNA concentrations were measured using a NanoDrop 2000 ultraviolet spectrophotometer (Thermo Fisher Scientific, Wilmington, NC, USA). DNA solutions were diluted to obtain the optimal concentration (15 ng/µL) for all samples.
Samples in the study group were tested using a cardiovascular drug response-associated gene detection kit (Digena, Hangzhou, China) and DP-TOF MS (Digena), whereas samples in the control group were subjected to Sanger sequencing using an ABI 3500xL Genetic Analyzer (Thermo Fisher Scientific), in accordance with the manufacturer’s instructions. The cardiovascular drug response-associated gene detection kit evaluated 17 loci across 11 genes, with uniform distributions for all 17 loci and extended primers between 4000 and 10,000 Da (Table 1).
Genes associated with cardiovascular drug responses.
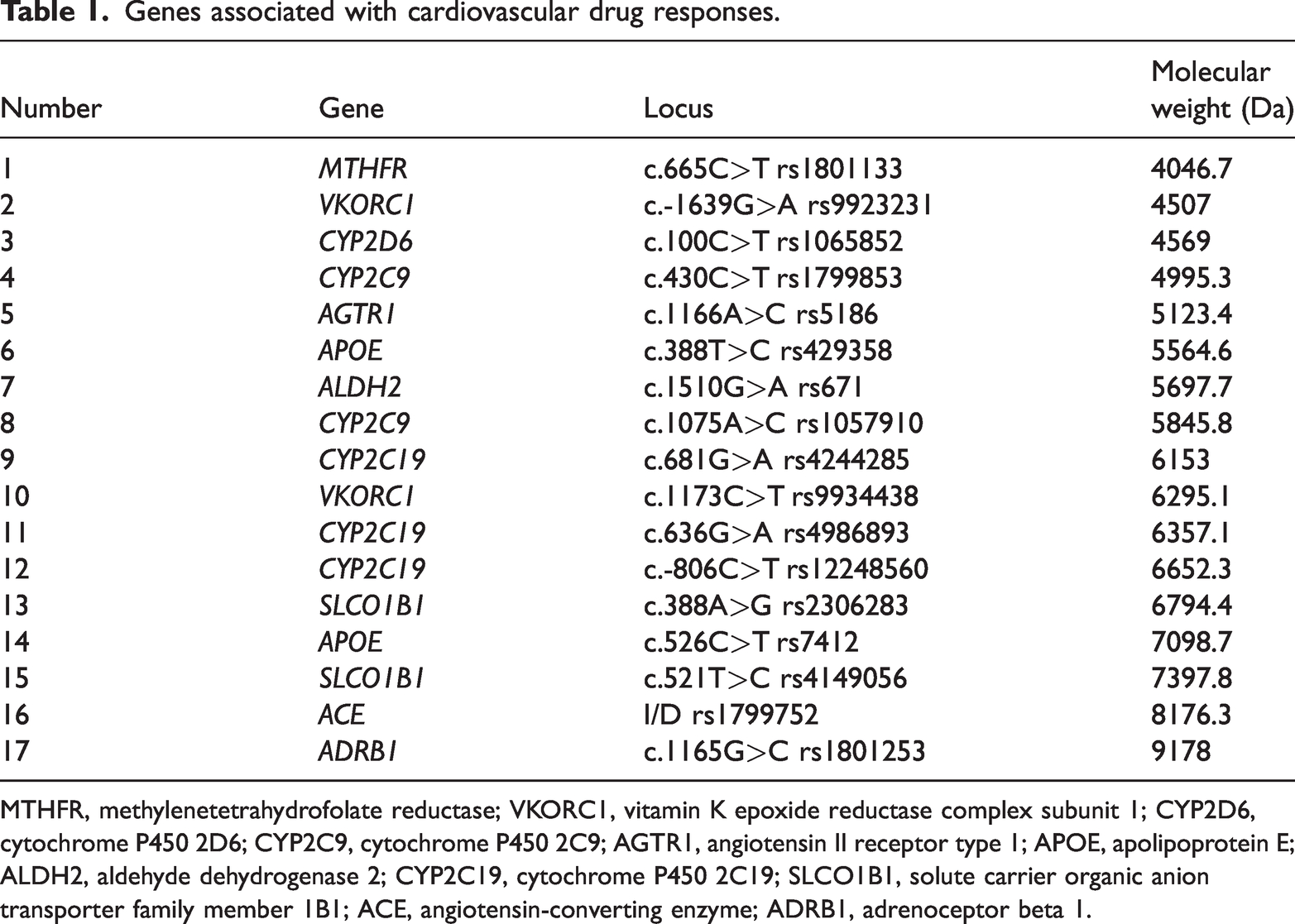
MTHFR, methylenetetrahydrofolate reductase; VKORC1, vitamin K epoxide reductase complex subunit 1; CYP2D6, cytochrome P450 2D6; CYP2C9, cytochrome P450 2C9; AGTR1, angiotensin II receptor type 1; APOE, apolipoprotein E; ALDH2, aldehyde dehydrogenase 2; CYP2C19, cytochrome P450 2C19; SLCO1B1, solute carrier organic anion transporter family member 1B1; ACE, angiotensin-converting enzyme; ADRB1, adrenoceptor beta 1.
In the study group, all target genes were amplified in a single reaction system. The PCR mixture contained nuclease-free water (0.8 µL), 10× PCR buffer (0.5 µL), MgCl2 (2 mM, 0.4 µL), deoxynucleotide triphosphates (dNTPs: dATP, dGTP, dCTP, and dTTP; 0.1 µL), multiple PCR primer mix (1 µL), DNA polymerase enzyme (1 U, 0.2 µL), and template DNA (2 µL) (all from Digena). Multiple allele-specific amplification was performed using a Veriti Thermal Cycler (Thermo Fisher Scientific) under the following conditions: denaturation for 2 minutes at 95°C; 45 cycles of 95°C for 30 s, 60°C for 30 s, and 72°C for 60 s; and extension at 72°C for 5 minutes. After the multiplex PCR reaction, shrimp alkaline phosphatase (SAP) was used to degrade dNTPs from the previous step. The SAP mixture contained nuclease-free water (1.53 µL), SAP buffer (0.17 µL), and SAP enzyme (0.3 µL). The SAP mixture was incubated at 37°C for 40 minutes, and the reaction was terminated by heating at 85°C for 5 minutes. Subsequently, single-base primer extension was performed. The extension mixture contained nuclease-free water (0.62 µL), 10× iPLEX buffer plus (0.2 µL), iPLEX termination mix (0.2 µL), multiple extension primer mix (0.94 µL), and iPLEX enzyme (0.04 µL) (Agena Bioscience, San Diego, CA, USA). The extension reaction conditions were as follows: denaturation for 30 s at 95°C; 45 cycles of 95°C for 5 s, 52°C for 5 s, and 80°C for 5 s; and extension at 72°C for 3 minutes. In addition, the PCR products were desalted and purified by ion exchange and co-crystallized with the matrix on a 384 chip (Digena). Subsequently, the PCR products were separated according to m/z and their masses were detected by DP-TOF MS. DP-TOF software (Digena) was used to determine the corresponding allele in each sample based on the mass of the PCR product. Template-free and mutant controls were both included in each run.
The 998 retrospective DNA samples were analyzed using the cardiovascular drug response-associated gene detection kit and DP-TOF MS methods described above. Four samples containing heterozygous types of 11 genes and 17 loci were screened, then diluted to concentrations of 100, 50, 25, 10, 5, 2.5, 1, 0.5, 0.2, and 0.1 ng/μL, respectively. Each dilution was repeated three times to determine the limit of detection. Fourteen samples containing all common genotypes of 11 genes and 17 loci were screened. The inter-batch repeatability of each sample site was evaluated for 5 consecutive days, and each sample was repeated twice. The intra-batch repeatability was evaluated by repeating each sample 10 times on the same day. Two samples each containing wild-type and homozygous APOE rs429358 and MTHFR rs1801133 loci were screened to assess anti-interference performance. PCR products of homozygous samples were diluted to concentrations of 1013, 1011, 109, 107, 105, 103, and 101 copies/μL, respectively. Different concentrations of PCR products (0.5 μL each) from homozygous samples were added to the PCR products from wild-type samples. Each dilution was repeated three times.
Statistical analysis
The main evaluation indexes used to determine DP-TOF MS accuracy in the study group samples were the single nucleotide and overall concordance rates. All concordance rates were calculated with 95% confidence intervals (CIs). The concordance rate for a particular nucleotide was calculated as the number of times that the DP-TOF MS results matched the Sanger sequencing results divided by the number of times that the Sanger sequencing results displayed that nucleotide. The secondary evaluation index used to determine DP-TOF MS accuracy was the genotyping concordance rate based on the results obtained by DP-TOF MS and Sanger sequencing. The kappa statistic was used to assess the agreement in diagnostic value between the two platforms. For retrospective DNA samples, the DP-TOF MS results were compared with nine known results obtained using RT-qPCR or Sanger sequencing to calculate the concordance rate for each known locus, as well as the overall concordance rate.
Results
The results for the main evaluation indexes are summarized in Table 2. A, T, and G detection by DP-TOF MS and Sanger sequencing yielded 100% concordance at 95% CI, whereas the C nucleotide concordance was 99.86%. The overall concordance rate for the four nucleotides was 99.96% at 95% CI (detailed in Supplementary Table 2).
Single-nucleotide and total concordance rates.

Nucleotide frequency was multiplied by 2 if the locus was wild-type and homozygous.
Concordance rate is equal to the Study group count divided by the Control group count.
DP-TOF MS, dual-polarity time-of-flight mass spectrometry.
Results for the secondary evaluation index, genotyping concordance, are presented in Table 3. The genotypes identified by DP-TOF MS showed 99.96% concordance with those determined by Sanger sequencing for all 17 loci tested.
Genotype concordance rates.

Count, number of results obtained by DP-TOF MS or Sanger sequencing for this genotype; %, number of results obtained by DP-TOF MS or Sanger sequencing for this genotype divided by the total number of results for the gene locus.
MTHFR, methylenetetrahydrofolate reductase; VKORC1, vitamin K epoxide reductase complex subunit 1; CYP2D6, cytochrome P450 2D6; CYP2C9, cytochrome P450 2C9; AGTR1, angiotensin II receptor type 1; APOE, apolipoprotein E; ALDH2, aldehyde dehydrogenase 2; CYP2C19, cytochrome P450 2C19; SLCO1B1, solute carrier organic anion transporter family member 1B1; ACE, angiotensin-converting enzyme; ADRB1, adrenoceptor beta 1; DP-TOF MS, dual-polarity time-of-flight mass spectrometry.
For the 998 retrospective DNA samples, the results for the nine known loci and 17 loci from the DP-TOF MS results are detailed in Supplementary Table 3. The known loci and overall concordance rates are listed in Table 4. The overall concordance rate for the known loci was 99.91%. Notably, the results obtained by DP-TOF MS and RT-qPCR for APOE rs429358 were inconsistent at three loci, whereas the results for APOE rs7412 were inconsistent at one locus. In addition, the allele frequencies of the 17 loci in the retrospective samples were determined according to the DP-TOF results. The reference allele frequencies were consistent with the East Asian population frequencies reported in the National Center for Biotechnology Information (NCBI) database (Figure 1).
Concordance rates of nine known loci in retrospective DNA samples, obtained using RT-qPCR or Sanger sequencing.

VKORC1, vitamin K epoxide reductase complex subunit 1; CYP2C9, cytochrome P450 2C9; CYP2C19, cytochrome P450 2C19; MTHFR, methylenetetrahydrofolate reductase; APOE, apolipoprotein E; SLCO1B1, solute carrier organic anion transporter family member 1B1; RT-qPCR, quantitative reverse transcription polymerase chain reaction.

Allele frequencies of 17 loci in retrospective samples. Blue represents DP-TOF reference allele frequency; red represents NCBI reference allele frequency. DP-TOF, dual-polarity time-of-flight; NCBI, National Center for Biotechnology Information.
The detection limit results showed that at a concentration of 0.1 ng/μL, two samples of SLCO1B1 rs2306283 were not detected, whereas one sample each of ADRB1 rs1801253 and CYP2C19 rs12248560 were incorrectly detected. In addition, one sample of APOE rs7412 was not detected, and the other concentrations were normal. At a concentration of 0.2 ng/μL, all sites in the four samples could be accurately detected. At that point, the added DNA volume was 2 μL, indicating that the lower limit of detection was 0.4 ng DNA. The repeatability results showed that the inter-assay and intra-assay precision rates were both 100%. Anti-interference analysis revealed that homozygous samples at a concentration of 10^13 copies/μL could cause contamination whereby wild-type samples were falsely identified as heterozygous for the APOE rs429358 and MTHFR rs1801133 loci.
Discussion
Available technologies for gene detection have advantages and disadvantages. 30 For example, PCR-based detection technologies are rapid and simple, but they can only detect a few loci and are hindered by low sensitivity and limited application scope. 31 In contrast, NGS, a high-throughput sequencing technology, can reveal unknown mutations and pathogenic genes 32 ; however, the technology is expensive, turnaround time is slow, quality control is difficult, operation is complex, and repeatability is low, all of which impede large-scale clinical applications. 33 The present study showed that DP-TOF MS gene detection is a fast, sensitive, convenient, and medium-flux technique 34 combining the “MALDI” and “TOF MS” modules. A previous study demonstrated that the MS method could serve as an inexpensive, accurate, and dependable method for initial screening of gene mutations; it also constituted an adjunct to clinical diagnosis. 35 Nevertheless, performance validation is required before a new analytical procedure can be implemented in clinical settings. Our study revealed 99.96% concordance between the DP-TOF MS and Sanger sequencing results, supporting the notion that this technology can be applied to nucleotide detection in clinical laboratories.
Considering that DP-TOF MS has a detection range between 4000 and 10,000 Da, this study used a cardiovascular drug response-associated gene detection kit comprising 11 genes with uniform distributions for all 17 loci and extended primers between 4000 and 10,000 Da to validate the concordance of nucleotide detection between DP-TOF MS and Sanger sequencing. In addition, 150 clinical samples from participants across various regions of China were analyzed to capture regional genetic differences in the Chinese population. To statistically assess the capacity of DP-TOF MS to measure the rare CYP2C19 mutations rs12248560 and rs1799853, 20 DNA samples previously tested at Fuwai Hospital were selected. Among the 150 clinical samples, one no-signal sample was detected by DP-TOF MS, whereas Sanger sequencing showed a CC genotype corresponding to the CYP2D6 mutation rs1065852. We suspect that a whole-gene deletion or gene recombination of CYP2D6 occurred because the frequency of CYP2D6 gene deletion (CYP2D6*5 allele) is 4.82% in the Eastern Han Chinese population. 36 Furthermore, CYP2D6 can exchange genetic material with CYP2D7, CYP2D8, and other pseudogenes to form hybrid alleles. The DP-TOF MS technology is only designed to detect known nucleotide loci, leading to investigational limitations because unknown indels and instances of gene recombination cannot be identified.
Evaluations of DP-TOF MS accuracy among retrospective DNA samples supported the clinical feasibility of this technique. The overall concordance rate for the known loci was 99.91%. Notably, the DP-TOF MS and RT-qPCR results for APOE rs429358 were inconsistent in three samples, whereas the results for APOE rs7412 were inconsistent in one sample. The concordance rates for these results reached 100% after the DP-TOF MS experiment had been repeated once for the four inconsistent samples. According to the anti-interference analysis conducted during the study, the inconsistent results may have been caused by aerosol contamination of the PCR products. Thus, DP-TOF MS should be performed under controlled and standardized conditions during clinical testing.
The major limitations of this study were the inability to perform follow-up analyses of patient outcomes after intervention. Considerably more work is needed to explore the utility of DP-TOF MS in population genomics and epidemiology.
Conclusions
The DP-TOF MS platform exhibited good detection performance. Our results suggest that this technique can be used by clinical laboratories for nucleotide detection in applications such as pharmacogenomics and genetic disease testing.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605241255568 - Supplemental material for Detection accuracy and clinical applications of DP-TOF mass spectrometry
Supplemental material, sj-pdf-1-imr-10.1177_03000605241255568 for Detection accuracy and clinical applications of DP-TOF mass spectrometry by Zhaohui Liu, Juan Zhao, Kai Cui, Huimin Guo, Zhikai Li and Zhou Zhou in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605241255568 - Supplemental material for Detection accuracy and clinical applications of DP-TOF mass spectrometry
Supplemental material, sj-pdf-2-imr-10.1177_03000605241255568 for Detection accuracy and clinical applications of DP-TOF mass spectrometry by Zhaohui Liu, Juan Zhao, Kai Cui, Huimin Guo, Zhikai Li and Zhou Zhou in Journal of International Medical Research
Supplemental Material
sj-pdf-3-imr-10.1177_03000605241255568 - Supplemental material for Detection accuracy and clinical applications of DP-TOF mass spectrometry
Supplemental material, sj-pdf-3-imr-10.1177_03000605241255568 for Detection accuracy and clinical applications of DP-TOF mass spectrometry by Zhaohui Liu, Juan Zhao, Kai Cui, Huimin Guo, Zhikai Li and Zhou Zhou in Journal of International Medical Research
Footnotes
Acknowledgements
We thank Digena for providing instrumental and technical support regarding their DP-TOF MS system.
Author contributions
ZL and JZ: conceptualization, study design, data analysis, manuscript drafting, and manuscript revision;
KC: data curation and analysis;
HG: data analysis and investigation;
ZL: manuscript drafting and investigation;
ZZ: conceptualization, supervision, and manuscript revision.
Data availability statement
The data generated and/or analyzed in this study can be acquired from the corresponding author upon reasonable request.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
