Abstract
Objective
To compare hepatitis C virus (HCV) RNA levels from serum and explanted native liver samples from patients undergoing living donor liver transplantation (LDLT).
Methods
This was a prospective observational study. Serum and liver samples were collected from consecutive serum anti-HCV-positive transplant recipients between February 2016 to August 2019. HCV RNA was extracted from liver samples and subjected to one-step reverse-transcription qPCR. using the TopScript One Step qRT-PCR Probe Kit with HCV qPCR probe assay and human
Results
Among the 80 patients, 36% (29/80) were HCV RNA positive in serum and 85% (68/80) had positive hepatic HCV RNA. Post-liver transplantation, 4% (3/80) patients were serum positive.
Conclusions
Our study suggests that pre-transplant serum HCV RNA levels may give an underestimate of the number of positive HCV RNA cases and that hepatic HCV RNA data may be more accurate.
Introduction
Chronic hepatitis C virus (HCV) infection, which leads to liver cirrhosis and hepatocellular carcinoma, is a worldwide problem. 1 Despite the widespread use of direct-acting antiviral agents (DAAs), their impact remains controversial. Some authors suggest there is a link between the use of DAAs and the occurrence of HCC.2–4 Indeed, guidelines suggest that irrespective of a sustained viral response (SVR) in patients with HCV-infection, patients treated with DAAs remain at risk of HCC development and require continued HCC surveillance.1,5
Living donor liver transplantation (LDLT) plays an important role in advanced liver disease. 6 Patients with a positive serum HCV-antibody test and a negative HCV RNA polymerase chain reaction (PCR) test are considered to have no evidence of current (active) HCV infection. 7 However, they are not protected from re-infection. Therefore, quantitative HCV RNA testing using serum samples is recommended prior to liver transplantation or initiation of anti-viral therapy to document the baseline viral load, and following liver transplantation to monitor HCV reactivation. 7 In this study, we compared HCV RNA levels from explanted native liver tissues and serum from patients undergoing LDLT.
Methods
Study population
Consecutive patients who underwent LDLT at Chang Gung Memorial Hospital, Kaohsiung, from February 2016 to August 2019, were eligible for this prospective, cohort study. To be included in the study, patients were ≥18 years of age, and had a positive serum HCV-antibody test. Patients excluded from the study had the following: a positive serum hepatitis B surface antigen (HBsAg) test result; primary biliary cirrhosis; alcohol-related liver disease; underlying psychological illness. The study was approved by the Medical Ethics Committee of Chang Gung Memorial Hospital, Kaohsiung (February, 2016; ethical approval number: 201701633B0) and written informed consent was obtained from each patient.
RNA extraction
Total RNA was extracted from liver tissues using Total RNA Isolation Kit (Vazyme Biotech., CN) according to the manufacturer’s instructions. Absorbance values of 260nm (A260), 280 nm (A280) and 230nm (A230) represent absorbance of nucleic acids, proteins, and organic salts, respectively. The ratio of A260/A280 was used as a reference for detecting nucleic acid quality. The A260/A280 ratio and concentration of total RNA were determined using an Epoch spectrophotometer system (Biotek, USA). The A260/A280 ratio of RNA of liver specimens was between 1.8 to 2.0.
One step reverse-transcribed qPCR for HCV and human GAPDH
An RNA sample from each liver specimen was subjected to one-step quantitative reverse transcription polymerase chain reaction (qPCR) using the TopScript One Step qRT-PCR Probe Kit with HCV qPCR probe assay and human glyceraldehyde phosphate dehydrogenase (GAPDH) qPCR probe assay (Topgen Biotech., TW) on a ViiA7 Real-Time PCR System (Applied Biosystems, USA) following the manufacturer’s instructions. The reaction mixture contained: 2 µl RNA (10 ng/ul); 2 µlnuclease-free water; 5 µl one-step RT-PCR master mix; 0.5µl enzyme; 0.5 ul HCV or GAPDH qPCR probe assay. The thermal cycling conditions were as follows: 50°C for 20 min; 95°C for 1 min; 40 cycles of 95°C for 3 s; 60°C for 40 s. Data collection occurred after 60°C at every cycle step.
Absolute quantification of HCV to determine copy number and normalize with human GAPDH
To generate a standard curve for the absolute quantification of HCV, we performed one-step reverse-transcription qPCR with arbitrary copies (1E7, 1E5, 1E3, and 1E2 copies) of custom-made HCV 5′UTR RNA prime (Topgen Biotech., TW). The liver specimens were amplified in triplicate with appropriate non-template controls. Amplification data were quantified using the HCV standard curve and normalized to GAPDH expression. Quantification of relative expression (reported as arbitrary units (copy number)) was performed using the 2−ΔΔCt relative quantification method. 7 The datasets used and/or analysed during the study are available he corresponding author [K-W. C.] on request.
Serum RNA was extracted automatically using COBAS AmpliPrep/COBAS TaqMan HCV Test followed by fluorescent probes for RT-qPCR amplification and detection of HCV RNA.
Statistical analyses
Data were analysed using SPSS version 22.0 (SPSS Inc., Chicago, IL, USA) and SAS software, version 9.4 (SAS Institute, Inc., Cary, NC, USA). All tests were two-sided and a
Results
In total 80 patients (40 men, 40 women) underwent LDLT at our center between February 2016 and August 2019. Their mean age was 60 years (range, 38–70 years) and 46 patients were diagnosed with HCC. Serum screening showed, 19 patients had HCV genotype 1, 15 patients had HCV non-genotype 1, and 46 patients had uncertain genotypes. Of the 80 patients, 34 (43%) had received pre-transplant DAAs for three months including
HCV RNA identification
Clinical profiles of the 80 recipients with HCV are shown in Table 1. Before LDLT, 36% (29/80) were serum positive and 64% (51/80) were serum negative for HCV RNA. Although there was a statistically significant difference (
Clinical profile of patients with chronic hepatitis C (HCV) undergoing living donor liver transplantation according to HCV RNA detected in serum.
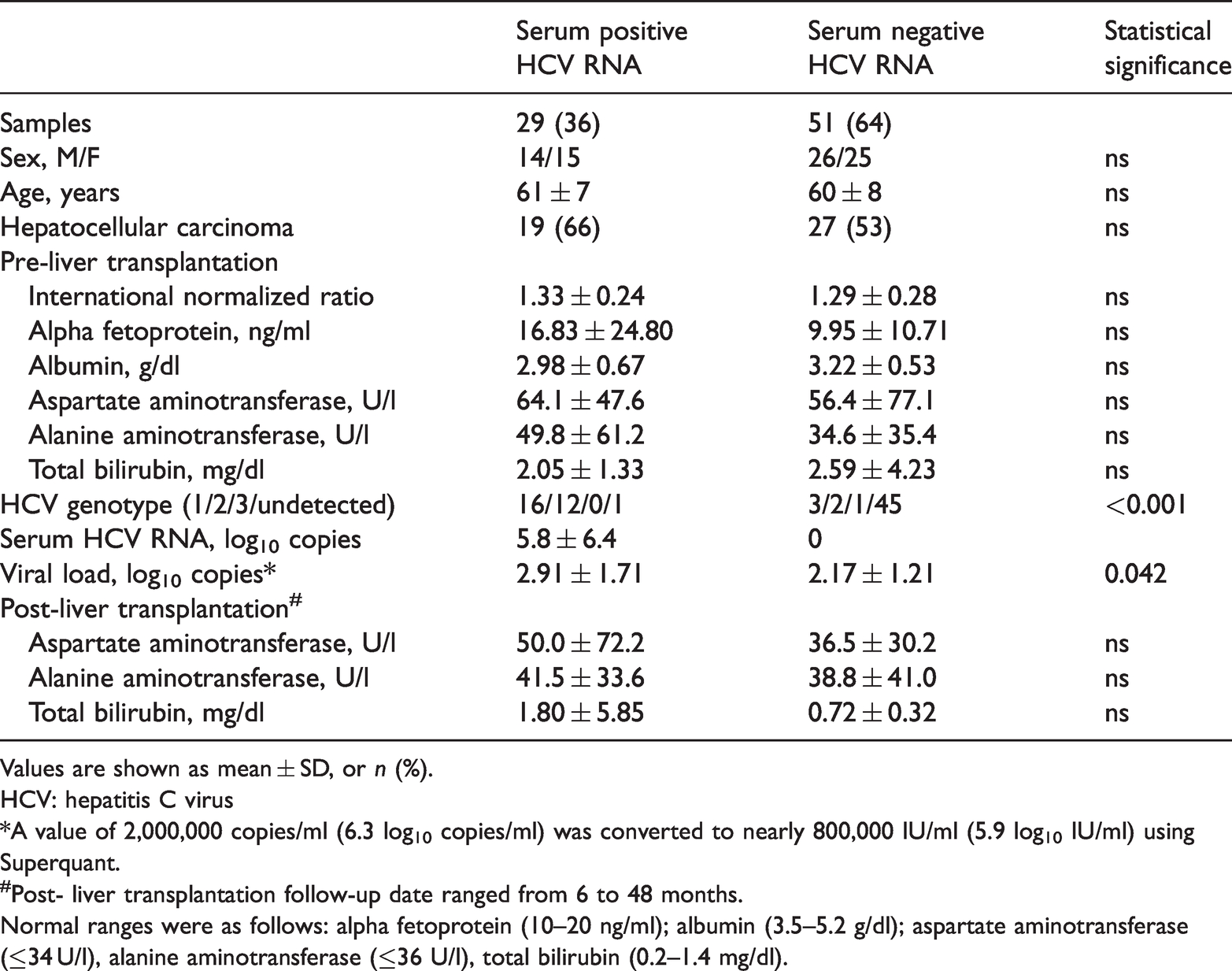
Values are shown as mean ± SD, or
HCV: hepatitis C virus
*A value of 2,000,000 copies/ml (6.3 log10 copies/ml) was converted to nearly 800,000 IU/ml (5.9 log10 IU/ml) using Superquant.
#Post- liver transplantation follow-up date ranged from 6 to 48 months.
Normal ranges were as follows: alpha fetoprotein (10–20 ng/ml); albumin (3.5–5.2 g/dl); aspartate aminotransferase (≤34 U/l), alanine aminotransferase (≤36 U/l), total bilirubin (0.2–1.4 mg/dl).
Analysis of native liver samples showed 85% (68/80) were positive for hepatic HCV RNA, and 15% (12/80) were negative (Table 2). Post-liver transplantation, 4% (3/80) patients were serum positive and 96% (77/80) were serum negative for HCV RNA (Table 3).
Clinical profiles of patients with chronic hepatitis C (HCV) undergoing living donor liver transplantation according to HCV RNA detected in native liver samples.

Values are shown as mean ± SD, or
HCV: hepatitis C virus
*A value of 2,000,000 copies/ml (6.3 log10 copies/ml) was converted to nearly 800,000 IU/ml (5.9 log10 IU/ml) using Superquant.
#Post- liver transplantation follow-up date ranged from 6 to 48 months.
Normal ranges were as follows: alpha fetoprotein (10–20 ng/ml); albumin (3.5–5.2 g/dl); aspartate aminotransferase (≤34 U/l), alanine aminotransferase (≤36 U/l), total bilirubin (0.2–1.4 mg/dl)
HCV RNA detected in serum and in corresponding liver samples in 80 patients undergoing living donor liver transplantation.

Values are shown as
HCV: hepatitis C virus.
According to International Consensus, in our quantitative HCV RNA assay, we used a value of 2,000,000 copies/ml (6.3 log10 copies/ml) converted to nearly 800,000 IU/ml (5.9 log10 IU/ml) using Superquant.
10
There was a statistically significant difference (
Hepatic HCV RNA and pre-treatment with DAA
Among the 68 recipients with positive hepatic HCV RNA, 40% (27/68) had been treated with DAA before LDLT, of which 51% (15/27) were positive for serum HCV RNA. The three cases that remained positive for serum HCV RNA after the LDLT had received DAA prior to the transplant. Following DAA treatment post-transplant, only one patient remained HCV RNA positive.
Discussion
We found that measurement of HCV RNA in serum identified far fewer positive cases than measurement of hepatic HCV RNA from the same patients. Similar to the mechanisms of antiviral treatment for hepatitis B infection, 11 patients with negative serum HCV RNA may not have undergone extensive HCV clearance in the liver. The high percentage of positive hepatic HCV RNA found in this study suggests that HCV returns to the hepatocyte as well as a target organ for its replication.
Following LDLT, three patients had positive serum HCV RNA. These patients had received DAAs pre-transplantation and had achieved a sustained viral response. Moreover, studies have reported that patients with detectable HCV RNA at the time of liver transplantation can experience recurrent HCV infection. 12 Although some studies have suggested that pre-emptive or prophylactic antiviral treatment is effective at preventing post-transplant HCV recurrence, 12 other studies have shown mixed results. 13 Impairment of bioavailability has been suggested as a possible explanation for an inferior DAA treatment response, although the exact mechanism for suboptimal activity has yet to be defined. 14 In many cases, DAAs may have a poor response in transplant patients because the population will have some level of decompensated liver disease.12,14 As a consequence, the timing of the DAA treatment before or after liver transplantation increases the complexity of the procedure when donor feasibility is taken into account. 15 Interestingly, a real-world experience of the transplantation of HCV-viraemic organs into HCV negative recipients found that in carefully selected patients, the use of HCV-viraemic grafts and careful selection of post-transplant treatment strategies appears to be efficacious and well tolerated. 16 We found that although serum HCV RNA was present in three patients after LDLT, following DAA treatment post-transplant, only one patient remained HCV RNA positive.
The study had several limitations. For example, our sample size was small and we did not include control groups. Additionally, there were no post-LDLT hepatic HCV RNA data available because biopsies on the transplanted tissues were precluded. Furthermore, we did not investigate differences in viral load.
In conclusion, our study suggests that pre-transplant serum HCV RNA levels may give an underestimate of the number of positive HCV cases and that hepatic HCV RNA data may be more accurate. In addition, careful selection and administration of antiviral therapy pre- and post-transplant may be beneficial.
Footnotes
Acknowledgments
We gratefully acknowledge all the participants who participated in the study and the study team for their support.
Declaration of conflicting interests
The authors declare that there are no conflicts of interest.
Funding
This work was supported by the grant number CMRPG8F1541 and CMRPG8H1131 from the Chang Gung Memorial Hospital of Taiwan
