Abstract
Objective
To investigate whether single nucleotide polymorphisms (SNPs) in the 3′ untranslated region (UTR) of the matrix metallopeptidase 9 gene (MMP9) are associated with susceptibility to calcium oxalate stones.
Methods
A total of 428 patients with kidney stone disease (KSD) and 450 control individuals were enrolled. Three MMP9 SNPs (rs20544, rs9509, and rs1056628) were genotyped, and MMP9 mRNA and protein expression was determined in patients and controls. The dual luciferase reporter gene assay was conducted by transfecting HEK293 cells with miR-491-5p mimics and plasmids containing MMP9 with rs1056628 AA/CC genotypes.
Results
The rs1056628 CC genotype was significantly increased in KSD patients compared with controls (CC vs AA: odds ratio [OR] = 2.279, 95% confidence interval [CI] = 1.048–4.956). The rs1056628 C allele frequency was higher in KSD patients than controls. The increased KSD risks associated with rs1056628 were more evident in individuals aged <30 years (OR = 3.504, 95% CI = 1.102–11.139) and men (OR = 2.522, 95% CI = 1.004–6.334). mRNA and protein levels of MMP9 were significantly higher in KSD patients with the CC genotype than in those with the AA genotype.
Conclusion
This study demonstrates that MMP9 SNP rs1056628 is associated with a significant KSD risk in Chinese Han individuals.
Introduction
Nephrolithiasis, also known as kidney stone disease (KSD), is a common urinary system disease caused by an aggregation of crystals, typically composed of calcium oxalate, in the kidneys. The high medical expense associated with its treatment means that it is a major public health burden on national healthcare systems.1,2 Many factors increase the risk for KSD, such as a family’s KSD history, dietary intake,3,4 exposure to steel manufacturing, 5 hypercalciuria,6,7 and a low fluid intake. 8 However, the fact that not all individuals exposed to the same risk factors eventually develop KSD suggest that genetic factors play important roles in its development, and increasing evidence suggests that genetic variants are associated with KSD.
MicroRNAs (miRNAs) are a series of small, non-coding RNA molecules ∼22 nucleotides in length that regulate target gene mRNA levels by binding directly to 3′ untranslated regions (UTRs) and inhibiting gene expression. 9 miRNAs are therefore involved in a variety of physiological processes, including KSD. 10 Previous studies showed that single nucleotide polymorphisms (SNPs) within or near miRNA binding sites in target genes can disrupt or create new binding sites that affect the miRNA binding affinity. Thus, SNPs within miRNA binding sites modulate expression levels of their target mRNAs, and consequently contribute to the development of human diseases, including disease susceptibility. 11
Patients with kidney stones or urinary tract stones have higher blood uric acid concentrations than healthy individuals.12,13 Matrix metalloproteinase 9 (MMP9) plays a key role in uric acid metabolism 14 by regulating several pathological remodeling processes and degrading extracellular matrix proteins.15,16 There is some evidence that patients with kidney stones show MMP9 up-regulation, and MMP9 is hypothesized to be associated with kidney stone development.17,18 However, no study has investigated whether an association exists between MMP9 polymorphisms and calcium nephrolithiasis susceptibility in the Chinese Han population.
Using bioinformatics, we confirmed that the rs1056628 functional variant in the 3′-UTR of MMP9 influences miRNA–mRNA interactions, as previously described by Yuan et al. 19 We then investigated the relationship between MMP9 3′-UTR polymorphisms and calcium nephrolithiasis risk in the Chinese Han population and explored possible mechanisms. Our research will help understand the role of MMP9 in the development of KSD.
Patients and methods
Study design and study population
This case–control molecular epidemiological study was designed to explore the involvement of MMP9 SNPs in the formation of kidney stones. A total of 428 patients (276 men) with calcium oxalate stone disease were recruited from Danyang People’s Hospital and Lianshui County People’s Hospital between January 2014 and December 2017. A diagnosis of KSD was made by identifying kidney stones using abdominal X-ray or ultrasound. Exclusion criteria were: 1) taking medication that affects urinary calcium excretion, and 2) taking vitamin D and/or calcium supplements. The control group (n = 450; 288 men) comprised volunteers who regularly undergo health check-ups at our hospitals. They underwent imaging tests to confirm the absence of kidney stones and urinary tract stones. They also had normal serum creatinine and calcium concentrations, no history of kidney stones, no family history of kidney stones, and no other systemic diseases. The current study was approved by the Institutional Review Boards of Danyang People’s Hospital and Lianshui County People’s Hospital. Written informed consent was obtained from all participants.
Genotyping assay
Based on previous studies, three MMP9 genetic variants (rs20544, rs9509, and rs1056628) were selected to explore their relationship with KSD.19–21 Genomic DNA from patients and controls was isolated from ethylenediaminetetraacetic acid-anticoagulated peripheral blood samples using the Genomic DNA extraction kit (Tiangen, Beijing, China). Genotyping was performed with the ABI7500 Real-Time PCR System (Thermo Fisher Scientific, Waltham, MA, USA) using the TaqMan® Assay according to the kit’s protocol. Primers and probes (FAM- and VIC- labeled) were purchased from Applied Biosystems (Foster City, CA, USA) and PCR conditions were as described previously. 22 Briefly, all assays were carried out in 96-well plates with negative and positive controls provided by the kit. Plates were sealed and heated at 95°C for 5 minutes, then subjected to 40 cycles of 95°C for 30s and 60°C for 45 s. After amplification, 10% PCR products were selected for direct sequencing to confirm the genotype.
Detection of MMP9 expression
Serum MMP9 concentrations were measured by commercial kits as described previously. 23
MMP9 expression was also measured in the peripheral blood mononuclear cells (PBMCs) of 60 KSD patients (20 patients of each rs1056628 genotype). Mononuclear cells were isolated from peripheral blood through the density gradient centrifugation method using Ficoll™. Total RNA was isolated from PBMCs using the RNA Isolation Kit (Beyotime, Haimen, China). MMP9 mRNA expression was determined by real-time (RT)-PCR using previously described primers. 24 Reaction conditions were: 95°C for 3 minutes, then 30 cycles of 95°C for 45 s, 58°C for 45 s, and 72°C for 60 s. Ct values were generated by 7500/7500 Fast Real-Time PCR Software v2.0 (Applied Biosystems).
MMP9 protein levels were detected by western blot assay as described previously. 25 MMP9 and GAPDH primary polyclonal antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA).
Plasmid construction
MMP9 rs1056628 luciferase reporter plasmids were constructed by amplifying the MMP9 region containing rs1056628 from participants carrying the rs1056628 AA or CC genotype. Amplification primers were: 5′-CCTGAGGACTAGGGTTCCCG-3′ and 5′-ATACCGTTTGTATCCGGCA-3′. Amplification was performed under the following conditions: 95°C for 3 minutes followed by 30 cycles of 95°C for 10 s, 60°C for 30 s, and 72°C for 30 s. PCR products were digested with Xba I (Takara, Dalian, China) and inserted into a pGL3-control vector (Promega, Madison, WI, USA). The plasmids were named pGL3-A and pGL3-C, respectively, and were confirmed by sequencing.
Dual luciferase reporter gene assay
HEK293 cells (ATCC, Manassas, VA, USA) were grown in RPMI 1640 media (Gibco, Waltham, MA, USA) supplemented with 10% fetal bovine serum (Gibco) at 37°C with 5% CO2. Cells were seeded into six-well plates at a density of 2 × 105 cells/well. They were then co-transfected with 30 nM miR-491-5p mimics (synthesized by Shanghai GENEPHARMA Co., Ltd., [Shanghai, China]) or water without DNA as a negative control, followed by secondary transfection of pGL3-blank, pGL3-A, or pGL3-C. After 48 hours, relative luciferase activities were detected using the dual luciferase assay kit (Solarbio, Beijing, China) according to the manufacturer’s instructions. 26
Bioinformatics Analysis
TargetScan (http://www.targetscan.org/cgi-bin/targetscan/vert_71/) was used to predict the binding of miRNA to target genes. miRSNP was used to analyze the effect of SNPs on miRNA binding. RNAhybrid was used to analyze the effect of SNPs on minimum free energy (MFE).
Statistical analysis
GraphPad Prism 7 software was used for statistical analysis. The goodness-of-fit χ2 test was used to detect Hardy–Weinberg equilibrium (HWE) among controls. Quantitative data are shown as means ± standard deviation, and were compared using the independent Student’s t-test to test for significant differences between two means. Pearson’s χ2-test was used to compare differences of categorical variables. Multiple logistic regression analysis was used to assess the relationship between MMP9 SNPs and the risk of KSD. A value of P<0.05 was considered statistically significant.
Results
Clinical characteristics of study subjects
Demographic and clinical characteristics of patients and controls are shown in Table 1. The average age for KSD patients was 38.7±10.9 years (range, 18–67 years) and 64.4% were male. The incidence of kidney stones was higher in men than in women. The average body mass index in patients was 23.2 ± 1.4 kg/m2. None of the participants were overweight or obese. Seventy-four patients (21.3%) had a family history of kidney stones. Biochemical examinations showed that serum creatinine and urinary calcium concentrations were significantly higher in patients than in controls (P ≤ 0.05; Table 1). Differences of other biochemical tests were not significant between groups.
Clinical characteristics of the study subjects.
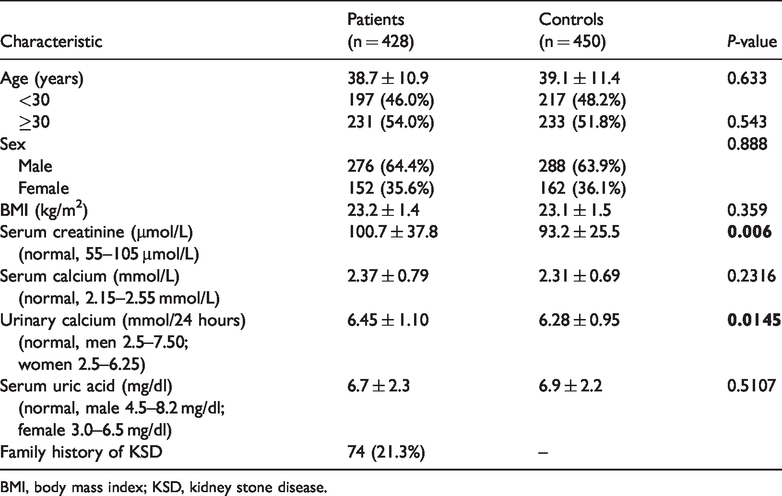
BMI, body mass index; KSD, kidney stone disease.
Associations between MMP9 polymorphisms and risk of KSD
MMP9 polymorphism frequencies in KSD patients and controls are shown in Table 2, and the distribution of MMP9 polymorphisms in controls was in accordance with the HWE (Table 3). Genotype frequencies of rs1056628 were 61.9% (AA), 33.4% (AC), and 4.7% (CC) in KSD patients, and 67.1% (AA), 30.7% (AC), and 2.2% (CC) in controls (Table 3). When the rs1056628 AA homozygote genotype was used as the reference group, the CC genotype was associated with a significantly increased risk of KSD (CC vs AA: odds ratio [OR] = 2.279, 95% confidence interval [CI] = 1.048–4.956, P = 0.039). However, the AC genotype was not associated with a KSD risk (AC vs AA: OR = 1.181, 95% CI =0.887–1.572). rs1056628 C allelic frequencies were significantly higher in KSD patients than in controls (P<0.05). We found no significant differences in genotype frequencies of rs20544 or rs9509 between KSD patients and controls.
MMP9 genotypes and risk of KSD.

MMP9, matrix metalloproteinase 9; KSD, kidney stone disease; OR, odds ratio; CI, confidence interval.
Hardy–Weinberg equilibrium in the control group.
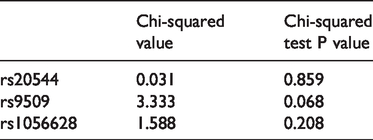
Stratification analysis of rs1056628 and the risk of KSD
We next investigated the interactions between rs1056628 and potential risk factors of KSD including age and sex (Table 4). Increased risks of KSD associated with rs1056628 were significantly more evident in individuals aged <30 years (OR=3.504, 95% CI =1.102–11.139, P = 0.037) and men (OR = 2.522, 95% CI =1.004–6.334, P = 0.049). These results suggest that MMP9 rs1056628 plays an important role in the occurrence of KSD in young men.
Stratification analysis of MMP9 rs1056628 and risk of KSD.

KSD, kidney stone disease; OR, odds ratio; CI, confidence interval.
Functional relevance of rs1056628
To examine the effect of rs1056628 on kidney stone formation, we examined whether it affected MMP9 expression in HEK293 cells. TargetScan predicted that miR-491-5p may bind the 3′ UTR of MMP9 (Figure 1a), while miRSNP showed that the CC genotype of rs1056628 destroyed a binding site of miR-491-5p (Figure 1b). RNAhybrid predicted that miR-491-5p has a higher MFE with the C allele of rs1056628 than with the A allele (MFE = −13 kcal/mol vs. MFE =−18.1 kcal/mol, respectively). Thus, we hypothesized that individuals with the rs1056628 C allele show up-regulated MMP9 expression.

Modification of rs1056628 on the expression of MMP9. a: TargetScan predicted that miR-491-5p binds the 3′-UTR of MMP9. b: rs1056628 destroyed an miRNA-mRNA binding site. c: miR-491-5P significantly decreased luciferase activities in HEK293 cells transfected with rs1056628 A allelic reporter plasmids compared with those transfected with rs1056628 C allelic reporter plasmids. P = 0.045. d: MMP9 mRNA levels were detected in PBMCs of KSD patients with various genotypes (20 per genotype). Each experiment was repeated three times. e, f: MMP9 protein levels were detected in PBMCs of KSD patients with various genotypes (20 per genotype). Each experiment was repeated three times. * P < 0.05. UTR, untranslated region; MMP9, matrix metalloproteinase 9; PBMCs, peripheral blood mononuclear cells; KSD, kidney stone disease.
Luciferase activity assays showed that miR-491-5P significantly decreased luciferase activities in HEK293 cells transfected with rs1056628 A allelic reporter plasmids compared with those transfected with rs1056628 C allelic reporter plasmids (P = 0.04, Figure 1c). Similarly, mRNA and protein levels of MMP9 were significantly higher in PBMCs from KSD patients with the CC genotype than the AA genotype (P < 0.05, Figure 1d–f).
Discussion
The main features of KSD include pain, persistent kidney damage, and acute or chronic kidney disease. 27 Several studies found that SNPs in the calcium-sensing receptor gene (CASR) function in recurrent nephrolithiasis formation, 28–30 while Orai1 polymorphisms were also reported to play important roles in nephrolithiasis. 31 Additionally, a significant association was documented between the vitamin D receptor gene (VDR) genotype and idiopathic nephrolithiasis.32,33
Increasing evidence has shown that SNPs located in mRNA 3′-UTRs are associated with the modulation of miRNA target genes, thus influencing susceptibility to various diseases.34,35 For example, Chen et al. 36 recently found that the rs2507800 TT genotype of the angiopoietin-1 gene might be associated with a lower risk of stroke by influencing the miR-211 binding efficiency. Additionally, the progestin and adipoQ receptor 6 gene variant rs759330, located at the binding site of hsa-miR-424*, may increase the risk of KSD. 37 In the current study, we found that MMP9 SNP rs1056628 appears to alter the miR-491-5p binding efficiency and influence the risk of KSD in a Chinese Han population. Furthermore, logistic regression analyses showed that individuals with the rs1056628 C allele had an increased risk of developing KSD. Our research will help early-stage disease prevention in high-risk groups and reduce the incidence of kidney stones.
Human MMP9 is located on chromosome 20q11.2-q13.1. Through bioinformatics analysis, we identified several SNPs. The most common MMP9 variant is rs3918242 which is located in the promoter 38 and has been shown to influence MMP9 expression. A previous luciferase activity assay revealed higher luciferase activity from plasmids with the rs3918242 T allele than plasmids with the rs3918242 C allele. 39
To date, few studies have investigated the association of MMP9 SNPs and kidney stones. Mehde et al. 18 reported that subjects with the rs3918242 TT genotype had an increased risk of KSD. Our current findings suggest that SNP rs1056628 influences the binding affinity between MMP9 mRNA and miR-491-5p and is involved in the development of KSD. Pirooz et al. 40 reported that an A→C transition in the miR-491-5p recognition site of MMP9 enhanced gene expression, potentially through the inhibition of miR-491-5p binding. rs1056628 plays a pivotal role in the predisposition of carriers to the development of atherosclerosis and ischemic stroke. 19 Furthermore, the relationship between MMP9 3′-UTR SNPs with the risk of cancers has been investigated. 21
In summary, we demonstrate for the first time that MMP9 SNP rs1056628 plays an important role in the occurrence of KSD in Chinese populations. rs1056628 has not previously been identified as a kidney stone risk factor in any population. The mechanism of action of MMP9 in calcium metabolism and kidney stone formation should be further investigated, and a larger sample size should be used to determine whether rs1056628 can be used as a biomarker of KSD.
Footnotes
Author contributions
Qiang Bu and Yu Zhu performed the RT-PCR, calculated the combined medication index, collected and analyzed the data, and prepared and wrote the paper; Qiaoyun Chen contributed to cell culture and the luciferase activity assay; Hao Li was responsible for case collection and patient consultations; Yan Pan contributed to the research design, data analysis, and manuscript writing. All authors critically reviewed the manuscript and approved it for submission.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This study was financially supported by The Jiangsu Health and Family Planning Commission Fund (No. z201625).
