Abstract
Objective
This prospective clinical study comparatively investigated the effects of tobacco smoking on global methylation and hydroxymethylation in oral epithelial cells.
Methods
Buccal cells from the inside of the cheeks were collected from 47 individuals, including smokers, former smokers, and never smokers. DNA was extracted using dedicated kits. Methylated and hydroxymethylated DNA fractions were measured using assays similar to enzyme-linked immunosorbent assays. The levels of methylation and hydroxymethylation were compared among groups using unpaired two-tailed t-tests or the Mann–Whitney U test; P < 0.05 was considered statistically significant.
Results
There was no statistically significant difference in the average number of cigarettes between smoker and former smoker groups. Although methylation levels were lower for smokers (3.1%) and former smokers (2.16%), compared with never smokers (4.16%), these differences were not statistically significant. There was a two-fold increase in hydroxymethylation level in never smokers, compared with smokers.
Conclusions
Our findings suggest that smoking leads to global reductions in both methylation and hydroxymethylation levels in oral epithelial cells in a manner influenced by the intensity and length of exposure to tobacco smoke.
Keywords
Introduction
Tobacco use is the leading cause of cardiovascular, respiratory, and cancer morbidity and mortality. Relatively recent studies have shown modifications in DNA methylation patterns due to tobacco exposure. 1 , 2 To the best of our knowledge, there is considerably less published information regarding the direct effects of tobacco smoke on the oral mucosa, compared with respiratory mucosal epithelium; most analyses have been focused on sputum, blood, cell lines, or tumor tissue. Tobacco comes into contact with the oral mucosa in a direct manner (i.e., through smoke exposure) and indirectly through dissolution in saliva and subsequent absorption by the oral epithelium. 3
Exposure of oral mucosa to tobacco smoke produces structural/macroscopic effects that have been correlated with the severity and length of exposure. 4 The cellular and molecular mechanisms underlying these changes have not been fully elucidated and involve various degrees of inflammation-related alterations in growth rates, as well as enhancement of the nucleus/cytoplasm ratio with associated pyknosis, karyorrhexis, and karyolysis. 5 , 6 The cytotoxic effects lead to epigenomic and transcriptomic changes, followed by apoptosis. 7
The epigenetic changes exerted by tobacco use on oral mucosa are not yet fully understood. DNA methylation, an epigenetic mechanism that plays key roles in various biological processes, is influenced by smoking. Multiple analyses of blood samples have revealed significant changes in genome methylation status in smokers, compared with non-smokers, which may persist for years after smoking cessation. 8 , 9 The reversal of smoke-induced methylation changes depends on the time elapsed since smoking cessation. 10
Recent studies have shown that demethylation is associated with elevated levels of 5-hydroxymethyl-cytosine (5-hmc) in euchromatic regions, which is produced by ten-eleven translocation enzyme-induced conversion of 5-methyl-cytosine (5-mc). 11 However, 5-hmc has also been shown to possess distinct regulatory functions as both a stimulatory and an inhibitory transcriptional regulator. 12 , 13 Ex vivo and in vivo experiments have linked oxidative stress with a global reduction in 5-hmc levels, thus providing a mechanistic link between smoking (known to directly deplete systemic antioxidants) and genome hydroxymethylation.14–16
The effects of tobacco smoking on the degree and patterns of global genome hydroxymethylation in oral mucosa have been far less studied compared with its effects on genome methylation. This study aimed to comparatively investigate the effects of tobacco smoking on global methylation and hydroxymethylation in oral epithelial cells.
Methods
Participants
This prospective clinical study was approved by the Ethics Committee at Victor Babes University of Medicine and Pharmacy (Timisoara, Romania) (approval no. 17/17.08.2018). This study enrolled volunteers without oral diseases who presented for their regular dental checkup; all provided written informed consent to participate and completed a questionnaire regarding their smoking habits. All participants were instructed not to eat, drink coffee or juices, or smoke for at least 30 minutes before sample collection.
Sample collection and DNA isolation
Buccal cells were collected using sterile OmniSwab buccal swabs (GE Healthcare, Chicago, IL, USA); buccal cells from the inside of the cheeks were collected on two swabs from each patient and dried before DNA extraction. DNA was extracted using a QIAmp DNA Mini kit (Qiagen, Hilden, Germany), in accordance with the manufacturer’s instructions. After isolation, DNA concentration and quality were assessed using a NanoDrop Microvolume Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA); all samples were stored at −20°C until further use.
Methylated and hydroxymethylated DNA quantification
Quantifications of global DNA methylation and hydroxymethylation were performed using the MethylFlash Methylated DNA Fluorometric Quantification Kit (Epigentek, Farmingdale, NY, USA) and MethylFlash Hydroxymethylated DNA Fluorometric Quantification Kit (Epigentek), respectively, in accordance with the manufacturer’s protocols. In these assays, similar to enzyme-linked immunosorbent assays, the methylated and hydroxymethylated fractions of DNA are detected using capture and detection antibodies, then quantified fluorometrically by reading the relative fluorescence units with a fluorescence spectrophotometer (GloMax Discover Multimode Microplate Reader; Promega, Madison, WI, USA). Data are represented as proportions of methylation and hydroxymethylation, respectively. All assays/tests were run in duplicate; average values were used for calculation.
Statistical analysis
The levels of methylation and hydroxymethylation were compared among groups using unpaired two-tailed t-tests or the Mann–Whitney U test (as appropriate, after data were assessed for normality of distribution using the D'Agostino & Pearson omnibus normality test). P < 0.05 was considered statistically significant.
Results
This study enrolled 47 individuals (25 men and 22 women), aged 26 to 89 years. The detailed clinical and demographic characteristics of the study participants are presented in Table 1. The average number of cigarettes did not significantly differ between the smoker and former smoker groups.
Study participants’ demographic characteristics and DNA methylation data.
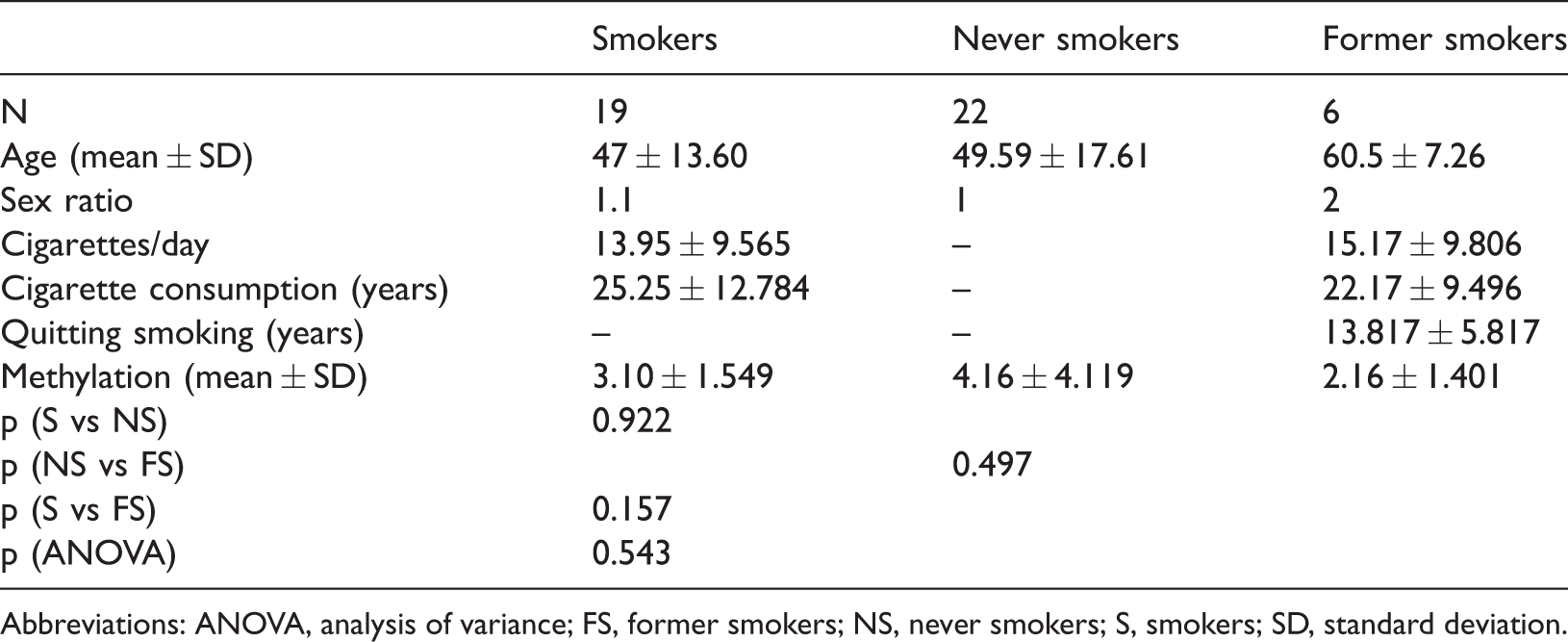
Abbreviations: ANOVA, analysis of variance; FS, former smokers; NS, never smokers; S, smokers; SD, standard deviation.
Although the methylation levels were lower for smokers (3.1%) and former smokers (2.16%), compared with never smokers (4.16%), these differences were not statistically significant. With respect to smoking intensity (i.e., number of cigarettes per day and duration of smoking), the heavy smokers (>20 cigarettes/day) and long-term smokers (over 20 years) exhibited significantly higher levels of DNA methylation (P = 0.0294 and P = 0.0273, respectively; Table 2 and Figure 1). This stratified analysis could not be performed in the former smoker group because it included a small number of participants.
Global DNA methylation in relation to smoking intensity and duration.
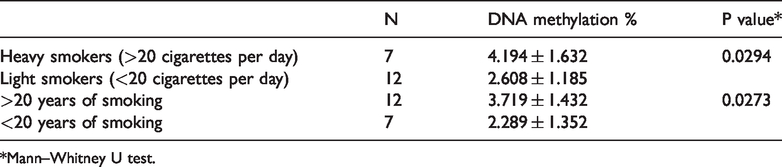
*Mann–Whitney U test.

Differences in DNA methylation according to smoking behavior. a. DNA methylation (%) and smoking intensity (mean number of cigarettes per day) (P = 0.0294 for comparison between groups); b. DNA methylation (%) and duration of smoking (mean number of years of smoking) (P = 0.0273 for comparison between groups).
Given the small size of the former smoker group, hydroxymethylation status was only compared between the smoker and never smoker groups. There was a two-fold increase in hydroxymethylation level in never smokers, compared with smokers (P = 0.0312; Table 3).
Study participants’ demographic characteristics and DNA hydroxymethylation data.

Abbreviations: NS, never smokers; S, smokers; SD, standard deviation.
Discussion
This pilot study investigated the effects of smoking on DNA methylation and hydroxymethylation. Smoking-associated modifications of methylation and hydroxymethylation patterns have been shown to influence the risks of several diseases, including cancer and chronic obstructive pulmonary disease. 17
Recently, several studies have revealed associations between tobacco smoking and widespread changes in genomic DNA methylation patterns (i.e., global hypomethylation and site-specific hypermethylation of multiple gene promoters). Zeilinger et al. 2 identified both hypo- and hypermethylation of multiple CpG islands in smokers compared with non-smokers using whole blood cells; however, they did not identify a clear direction or magnitude of changes in global DNA methylation due to tobacco smoking. Furthermore, in an analysis of oral epithelial cells, Teschendorff et al. 18 identified 1501 CpG islands impacted by smoking, of which 912 were hypermethylated and 589 were hypomethylated. Finally, Wan et al. 19 showed that the differences in methylation status were greater in oral epithelial cells than in whole blood cells of smokers, indicating that analysis of the oral epithelium may be more appropriate for assessment of the impact of tobacco smoking on methylation status.
Our results are consistent with the findings in these previous reports in that tobacco smoking reduced the global methylation status by approximately 25%. Notably, the reduction in methylation status was even more abrupt in former smokers (approximately 50%); however, this might represent the cumulative influence of age 20 in this group. Moreover, some CpG sites reportedly maintain their hypomethylated statuses after smoking cessation. 2
To identify a correlation between the intensity of exposure to tobacco smoke and the genome-wide level of DNA methylation, we classified study participants with >20 years of smoking and/or >20 cigarettes per day as heavy smokers, based on published data collected in a neighboring country. 21 As reported by De Araújo Costa et al., 22 we found a statistically significant, two-fold enhancement of methylation level in the heavy smoker group, compared with the light smoker group. The alteration of global DNA methylation levels was also studied in blood lymphocytes by Sabi et al., 23 who observed a significant enhancement of global DNA methylation, compared with the control group. The differences in levels of methylation and hydroxymethylation between the present study and the investigation by De Araújo Costa et al. could be explained by the different methods used for the analysis. De Araújo Costa et al. 22 used mouthwash for collection of biological samples, which yielded heterogeneous cell populations from the oral cavity (i.e., lymphocytes and epithelial cells); in contrast, our method comprised collection of oral epithelial cells on cotton swabs. Furthermore, we used a fluorescence-based technique similar to that of enzyme-linked immunosorbent assays; our technique was more sensitive than the classical enzyme-linked immunosorbent assay method that measures absorbance values, which was used by De Araújo Costa et al. 22
Our results showed a much lower level of hydroxymethylation, compared with methylation, in oral epithelial cells. This is consistent with the findings in previous studies regarding 5-mc and 5-hmc levels in human tissues, 8 and is presumably because 5-hmc represents an intermediate of 5-mc demethylation. 24 In our study, smokers exhibited significantly lower levels of hydroxymethylation, compared with non-smokers; this difference did not appear to be influenced by participant age.
Our results should be interpreted with caution because this study had several limitations. First, our investigation had a low number of samples and corresponding pilot study design; larger studies are needed to obtain more conclusive results as well as to explore the relationship between hydroxymethylation and methylation statuses and their correlations with the intensity and length of exposure to tobacco smoke. Second, our statistical analysis was based on participants’ responses regarding their smoking habits; therefore, the findings are subject to recall bias. Third, no passive smokers were included in our study. Finally, considerable age variation was observed among the participants, so no analyses that involved stratification of age and sex were performed.
Conclusions
This study aimed to quantify the genome-wide methylation and hydroxymethylation levels in oral epithelial cells exposed to tobacco smoke. Our findings suggest that smoking may influence the levels of DNA methylation, but larger studies are needed to confirm these results.
Supplemental Material
sj-pdf-1-imr-10.1177_0300060520954677 - Supplemental material for Changes in global DNA methylation and hydroxymethylation in oral mucosa according to tobacco smoke exposure
Supplemental material, sj-pdf-1-imr-10.1177_0300060520954677 for Changes in global DNA methylation and hydroxymethylation in oral mucosa according to tobacco smoke exposure by Gabriela-Emilia Avram, Anca Marcu, Alexandra Moatar, Corina Samoila, Angela Podariu, Edward Seclaman and Catalin Marian in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_0300060520954677 - Supplemental material for Changes in global DNA methylation and hydroxymethylation in oral mucosa according to tobacco smoke exposure
Supplemental material, sj-pdf-2-imr-10.1177_0300060520954677 for Changes in global DNA methylation and hydroxymethylation in oral mucosa according to tobacco smoke exposure by Gabriela-Emilia Avram, Anca Marcu, Alexandra Moatar, Corina Samoila, Angela Podariu, Edward Seclaman and Catalin Marian in Journal of International Medical Research
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
