Abstract
Objective
To investigate whether the Notch–Hif-1α signaling pathway is involved in liver regeneration.
Methods
Rats were divided into two groups and treated with daily intraperitoneal injections of saline (control) or the gamma-secretase inhibitor, Fli-06, for 2 days. Two-thirds of the rat livers were resected and rats were later euthanized at specific time points post-resection to analyze the remnant livers. Each group's liver/body weight ratio was calculated, and immunostaining and western blotting were used to determine the cell proliferation marker, PCNA and Ki-67 expression. Real-time PCR and western blotting were used to compare the mRNA expression of Notch homolog-1 (Notch1), hairy and enhancer of split-1 (Hes1), and vascular endothelial growth factor (Vegf), and the protein expression of NICD and HIF-1α, respectively.
Results
The liver/body weight ratios and number of Ki-67- and PCNA-positive cells were significantly lower in the experimental group than the control group, indicating lower levels of liver regeneration following the disruption of Notch signaling by Fli-06. The Hes1 and Vegf mRNA levels and NICD and HIF-1α protein expression levels were all down-regulated by Fli-06 treatment.
Conclusion
Notch–Hif-α signaling pathway activation plays an important role in liver regeneration, where it may contribute toward liver cell proliferation.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is one of the most common malignancies of the digestive system, and the incidence of HCC is increasing year by year. 1 With the development of advanced surgical techniques, liver resection has become a common treatment for this type of cancer. 2 Indeed, hepatectomy is now the first choice of treatment for hepatic malignancies. For this reason, elucidating the mechanism of liver regeneration after hepatectomy is particularly important.
Partial hepatectomy (PHx) is a classical model for studying liver regeneration in rats. 3 In this model, the left and the middle lobes of the liver (approximately two-thirds) undergo surgical resection, inducing liver regeneration after acute liver injury. The various cells affected, including the liver cells, bile duct epithelial cells, and liver sinusoidal endothelial cells (LSEC) among others, are all involved in the process of liver regeneration after liver resection. 4
Previous reports have shown that activation of the Notch signaling pathway is involved in the process of liver regeneration after liver injury. 5 , 6 In particular, the proliferation of liver and biliary duct cells is controlled by this pathway. 7 In the adult liver, Notch receptors and ligands are widely distributed in the bile duct and parenchyma cells. 8 When the liver is injured, the expression of Jag1 significantly increases, activating the downstream Notch signaling pathway and inducing liver regeneration. 9 The resulting Notch–Jag1 complexes undergo two phases of hydrolysis, the first of which is catalyzed by a metal protease family, while the second is catalyzed by a specific gamma-secretase that releases the Notch intracellular domain (NICD). The NICD binds to the CSL-binding protein in the nucleus, forming a transcriptional activation complex that activates cell differentiation and proliferation. Thus, gamma secretase plays a crucial role in the activation of the Notch signaling pathway.
Hepatic resection is often associated with hepatic ischemia and hypoxia damage, which activate the HIF-1 signaling pathway involved in the regulation of downstream products such as vascular endothelial growth factor (VEGF).10,11 Thus, the HIF-1α signaling pathway plays a positive regulatory role in the process of liver regeneration. There is, therefore, a regulatory relationship between the Notch signaling pathway and multiple other signaling pathways, and the mechanisms by which these pathways act together to promote regeneration in the liver still need to be explored.
In this study, we used the gamma-secretase inhibitor, Fli-06, to construct a Notch signaling inhibition model of liver regeneration after partial hepatectomy in rats. Molecular biology techniques were then applied to investigate the role of the Notch signaling pathway in liver regeneration. This model provides a basis for the further validation of the mechanism of Notch signaling in vivo and in vitro.
Materials and Methods
Animals
Healthy male Sprague-Dawley rats were selected with a 200- to 220-g body weight at 6 to 8 weeks old (Bangyao bio, Shanghai, China). The rats were reared in a clean, ventilated environment at a constant temperature of 26°C. This study conformed with the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health. The study protocol was approved by the Animal Ethics Committee of Sun Yat-sen Memorial Hospital (approval no. 18100212).
Partial hepatectomy model in rats
The Sprague-Dawley rats were divided randomly into two groups, namely the Fli-06 group and the control group, with 24 rats in each group. The Fli-06 group was given a daily intraperitoneal injection of 3 mg/kg/day Fli-06 (MedChemExpress, Monmouth Junction, NJ, USA) prepared in a 0.9% (w/v) sodium chloride solution (Kinghunt Bio, Nanjing, China) with a total volume of 1 mL per injection. The control group was given a daily 1-mL intraperitoneal injection of 0.9% sodium chloride solution alone. After 2 days, two-thirds of the left and middle hepatic lobes of the liver of each rat were resected according to the Higgins–Anderson method. 12 At each of the selected time points, six rats in the control group and six in the Fli-06 group were euthanized (on Days 1, 3, 7, and 28 after the initial resection) to extract the remnant liver. The weight of each rat was recorded before anesthesia. The remnant livers were frozen at −130°C (Olympus, Tokyo, Japan) in preparation for paraffin sectioning.
Immunohistochemistry
The remnant liver tissues were subjected to paraffin sectioning (3–4 µm). The paraffin sections were dewaxed and dewatered, incubated with 3% (v/v) H2O2 (Kinghunt bio) for 15 minutes, followed by incubation with undiluted sheep serum (Beijing Zhongzuo Jinqiao Co., Ltd., Beijing, China) for 20 minutes at 37°C. The sections were then incubated overnight at 4°C with antibodies against PCNA or Ki-67 (mouse anti-rat PCNA monoclonal antibody, 1:50 dilution; rabbit anti-rat Ki-67 polyclonal antibody, 1:200 dilution) sourced from Bohai Bioengineering (Wuhan, China). The sections were then washed in phosphate-buffered saline (PBS) three times. Next, the sections were incubated with a sheep anti-mouse secondary antibody (Bohai Bioengineering) at room temperature for 30 minutes, washed in PBS three times, incubated with horseradish peroxidase for 15 minutes, exposed to 3,3ʹ-diaminobenzidine (DAB) for 40 s, then washed in water and stained with hematoxylin for 15 to 20 s at room temperature. Finally, the samples were observed under a microscope (Olympus CX31-LV320; Olympus). Positive expression was determined based on the number of cells stained and the intensity of staining in the cytoplasm and nucleus.
RT-PCR
Liver tissues were selected from the Fli-06 and control groups and 100 mg each were dissolved in 1 mL TRIzol. The traditional three-step method of chloroform, isopropanol, and absolute ethanol extraction was used to isolate the total RNA, which was then dissolved in diethyl pyrocarbonate (DEPC). The extracted RNA was quantified by spectrophotometry and converted to cDNA with a reverse transcription cDNA kit (Bao Bioengineering, Dalian, China). Using cDNA as the template, the mRNA expression of Notch homolog-1 (Notch1), hairy and enhancer of split-1 (Hes1), and Vegf was detected by real-time PCR using a SYBR Green PCR Master Mix (Bao Bioengineering). The primers are listed in Table 1. The amplification conditions were as follows: pre-denaturation at 95°C for 2 minutes, followed by 40 cycles of denaturation at 94°C for 20 s, annealing at 59°C for 45 s, and extension at 72°C for 20 s. The PCR reactions were conducted in 25-µL reactions containing 2 µL cDNA (20 ng), 12.5 µL SYBR Green (12.5 µmol/L), 1 µL forward primer (0.4 µmol/L), 1 µL reverse primer (0.4 µmol/L), and 8.5 µL DEPC water.
The primers used for the RT-PCR of Notch1, Vegf, and Hes1.
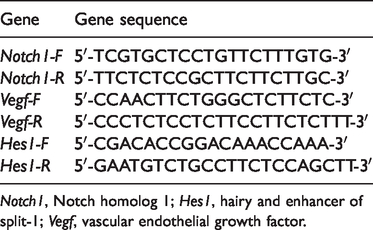
Notch1, Notch homolog 1; Hes1, hairy and enhancer of split-1; Vegf, vascular endothelial growth factor.
Western blotting
The HIF-1α and NICD protein samples were separated by 8% polyacrylamide gel electrophoresis, transferred to a polyvinylidene difluoride membrane (Thermo Fisher Scientific, Rockford, IL, USA) for 2 hours, blocked with skimmed milk powder for 30 minutes, and incubated with rabbit-derived anti-HIF-1α (1:1,000 dilution) or anti-NICD (1:500) for 12 hours at 4°C. Next, the samples were incubated with sheep anti-rabbit secondary antibody (1:5,000) at room temperature for 2 hours, followed by luminescence detection to determine the protein expression levels of HIF-1α and NICD.
Statistical analysis
Statistical analysis was conducted using IBM SPSS Statistics for Windows, version 19.0 (IBM Corp., Armonk, NY, USA). Measurement data were expressed as the mean ± standard deviation (SD) and compared with the two-tailed t-test, with P < 0.05 regarded as being statistically significant.
Results
Fli-06 inhibits liver regeneration after partial hepatectomy
The partial hepatectomy model was successfully established in rats by two-thirds liver resection. The remnant liver weights were measured on Days 1, 3, 7, and 28 after resection (Figure 1a) and used to calculate the liver weight ratio: sacrificed liver weight (g)/rat weight (g) × 100%. The results showed that the liver weight ratio in the control group was significantly higher than that in the Fli-06 group on Day 1 (P < 0.001), Day 3 (P < 0.01), Day 7 (P < 0.01), and Day 28 (P < 0.01) (Figure 1b, Table 2).

Effect of Fli-06 inhibition of Notch signaling on liver regeneration in rats after partial liver resection. (a) Photographs of the remnant rat livers on Days 1, 3, 7, and 28 after partial liver resection. (b) The remnant liver-to-body weight ratios were calculated to determine the effect of Fli-06 on gains in tissue mass after resection. Data represent the mean ± SD of at least three independent experiments, n = 6 in each group. *P < 0.05, **P < 0.01 vs. the respective control group.
Comparison of the liver weight ratio of each group.

Elevated expression of Ki-67 in the regenerating liver
Residual liver Ki-67 immunohistochemical staining was performed (Figure 2a, b). The results showed that the livers of both the control and Fli-06 groups were positive for cell proliferation after hepatectomy. However, the proportion of Ki-67-positive cells in the control group was significantly higher than that in the Fli-06 group on Days 1 and 2 (P < 0.05 for both), supporting the results of the western blotting (Figure 2c, d). Thus, compared with the control group, the liver regeneration capacity of the Fli-06 group was reduced after partial hepatectomy, with a lower proportion of Ki-67-positive cells observed.

The expression of Ki-67 after hepatectomy was detected by the immunohistochemical staining and western blotting of remnant liver samples from Days 1, 3, 7, and 28 after partial liver resection. (a) Ki-67 immunohistochemical staining of paraffin sections. (b) Quantification of the number of Ki-67-positive cells in the paraffin sections. (c) Western blot analysis of Ki-67 expression in the remnant liver. (d) Relative quantification of Ki-67 expression based on the western blot data in (c). Data represent the mean ± SD of at least three independent experiments, n = 6 in each group. *P < 0.05, **P < 0.01 vs. the respective control group.
Elevated expression of PCNA in the regenerating liver
The expression of PCNA in the liver after hepatectomy was similarly detected by immunohistochemistry (Figure 3a, b) and western blotting (Figure 3c, 3d). The results showed that the expression of PCNA was significantly higher in the control group than in the Fli-06 group on Days 1, 3, 7, and 28 after hepatectomy (P < 0.01 for all). Thus, compared with the control group, the liver regeneration capacity of the Fli-06 group was reduced after partial hepatectomy, with a lower proportion of PCNA-positive cells observed.

The expression of PCNA after hepatectomy was detected by the immunohistochemical staining and western blotting of remnant liver samples from Days 1, 3, 7, and 28 after partial liver resection. (a) PCNA immunohistochemical staining of paraffin sections. (b) Quantification of the number of PCNA-positive cells in the paraffin sections. (c) Western blot analysis of PCNA expression in the remnant liver. (d) Relative quantification of PCNA expression based on the western blot data in (c). Data represent the mean ± SD of at least three independent experiments, n = 6 in each group. *P < 0.05, **P < 0.01 vs. the respective control group.
Fli-06 inhibits the expression of HIF-1α and NICD
Western blotting was used to detect changes in the protein expression of HIF-1α and NICD in the residual liver tissues. The results showed that after partial hepatectomy, the expression of NICD in the remnant livers of the control group was significantly higher than that in the Fli-06 group on Days 1, 3, 7, and 28 after resection (P < 0.01 to < 0.05, Figure 4a, 4b). The relative expression levels of HIF-1α in the remnant liver tissue of the Fli-06 group on Days 1, 3, 7, and 28 were also significantly lower than that of the control group (P < 0.01 for all, Figure 4c).
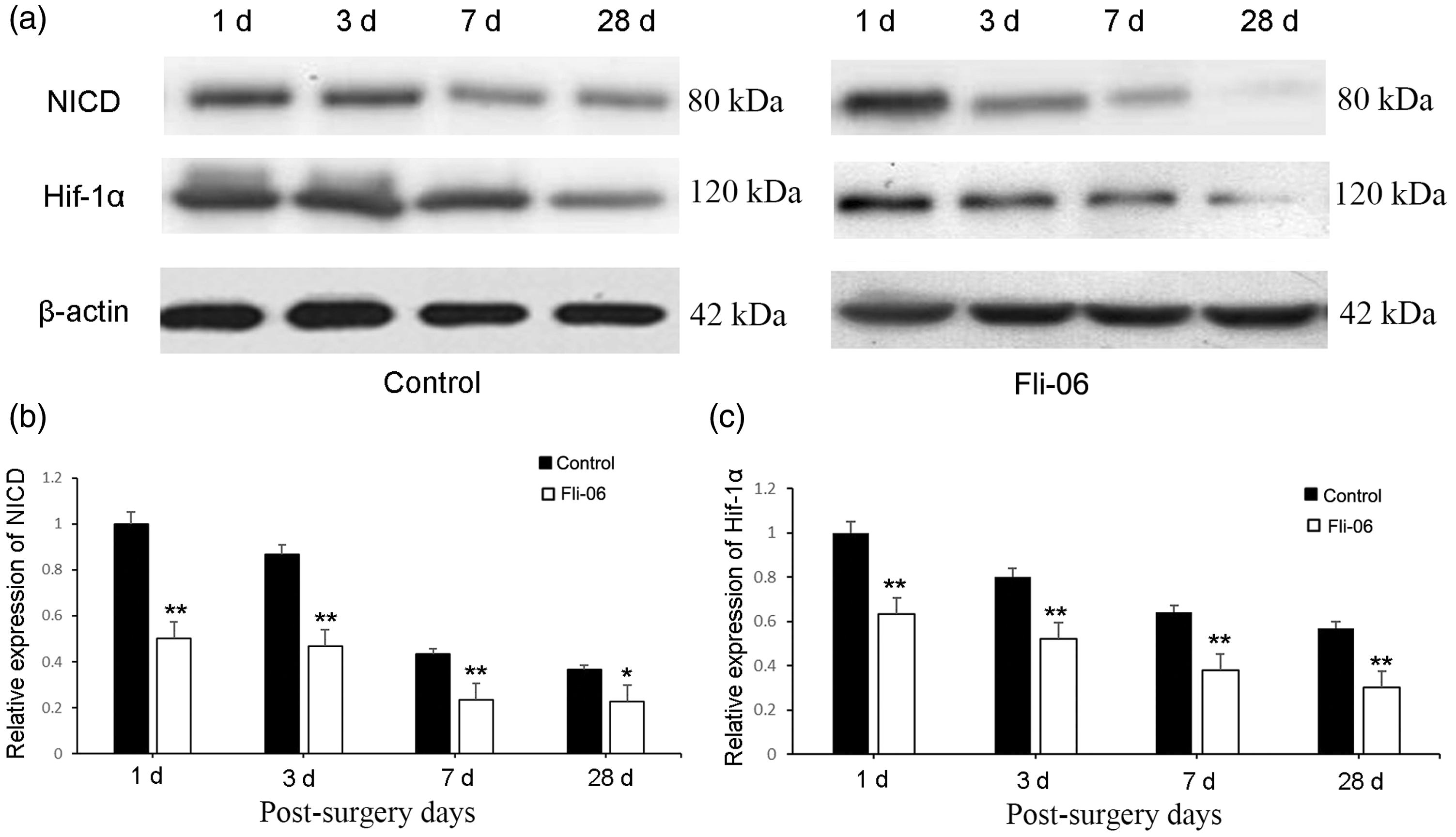
The expression of HIF-1α following the inhibition of Notch signaling was detected by the western blotting of remnant liver samples from Days 1, 3, 7, and 28 after partial liver resection. (a) Western blot analysis of NICD and HIF-1α expression in the control and Fli-06 groups. (b, c) Relative quantification of NICD and HIF-1α expression, respectively, based on the western blot data in (a). Data represent the mean ± SD of at least three independent experiments, n = 6 in each group. *P < 0.05, **P < 0.01 vs. the respective control group.
Expression levels of Notch1, Hes1, and Vegf in the regenerating liver
To further validate our findings, we analyzed the mRNA expression changes of Notch1, Hes1, and the downstream molecule, Vegf, by RT-PCR. There were no statistically significant differences between the control and Fli-06 groups in the level of Notch1 mRNA expression on any of the days tested following resection, indicating that the inhibitor was unable to interfere with the expression of Notch1. However, the expression levels of Hes1 and Vegf mRNA in the Fli-06 group were significantly lower than those in the control group (P < 0.01 to < 0.05, Figure 5). Both Notch1 and Hes1 showed a decreasing trend in mRNA expression over time post-resection in both the control and Fli-06 groups, while Vegf showed an increasing trend in mRNA expression.

RT-PCR analysis of the mRNA expression levels of Notch1, Hes1, and Vegf in remnant liver samples from the control and Fli-06 groups on Days 1, 3, 7, and 28 after partial liver resection. Data represent the mean ± SD of at least three independent experiments, n = 6 in each group. *P<0.05, **P<0.01 vs. the respective control group.
Discussion
Liver regeneration is a complex physiological process. A variety of signaling pathways, such as Notch and Wnt, are involved in liver regeneration after liver injury. 13 The Notch signaling pathway is involved in regulating the proliferation of parenchymal and non-parenchymal cells such as hepatocytes, bile duct cells, and vascular endothelial cells during liver regeneration. 14 Morell et al. 15 showed that the process of liver regeneration after partial hepatectomy in rats activates the Notch signaling pathway and that the Notch ligand, Jagged1, promotes liver cell proliferation, suggesting that Notch signal is a positive regulator of liver cell proliferation. After hepatectomy, in the early stage of liver regeneration, activation of the Notch–HIF-1α pathway can be found, and the expression of its downstream target molecules, such as Hes1, rapidly increases. Yang et al. 16 reported that after partial hepatectomy, the expression of hepatocyte growth factor activator inhibitor type 1 (Hai-1) mRNA in both the control and cirrhotic groups of rats increased first and then decreased, although the expression of Hai-1 mRNA in the cirrhotic rats was higher than that in the control group. However, these studies looked only at the short-term production of hepatocyte growth factor activators, and no long-term records were included.
Gamma-secretase inhibitors inhibit the activity of gamma secretase, thereby inhibiting the production of NICD and, in turn, signal transmission along the Notch signaling pathway. 17 We used Fli-06 in the present study, as it is a gamma-secretase inhibitor that suppresses Notch signaling. 18 The Groeneweg et al. 19 study found that γ-secretory enzyme inhibitors inhibit tumor growth in a tumor-bearing mouse model of uterine serous carcinoma. Mizugaki et al. 20 found that the combination of a gamma-secretase inhibitor and radiotherapy in the treatment of nude mice with lung cancer significantly enhances the anti-tumor effect of radiotherapy and inhibits tumor growth. The Wang 21 study showed that DAPT (a γ-secretory enzyme inhibitor) inhibits tumor growth in nude mice with esophageal adenocarcinoma.
To date, investigations of the role of Notch signaling in liver regeneration have mostly been carried out using conditional gene knockout of RBP-J in mice, a key molecule in a Notch downstream pathway. 22 The present study investigated the possibility of using gamma-secretase inhibitors to block Notch activation in vivo to construct a simple and feasible observation model. Our experiments showed that Fli-06 could significantly inhibit liver regeneration after partial hepatectomy, as evidenced by the significantly lower liver-to-body weight ratios in the Fli-06 group compared with those in the control group.
Through Ki-67 and PCNA staining, we found that the proliferation of hepatocytes in the Fli-06 group was significantly decreased, indicating that the Notch pathway promotes the proliferation of hepatocytes during liver regeneration. By examining the protein levels of NICD in the control and Fli-06 groups and the mRNA expression of Vegf and Hes1, it was apparent that the Notch signal showed marked activation after surgery, consistent with earlier reports. 23 However, the mRNA expression levels of Notch1 did not change significantly in the Fli-06 group, indicating that Fli-06 did not affect the expression of Notch. Overall, our in vivo experiments confirmed that the Notch signaling pathway plays an important role in liver regeneration.
Our findings indicated that the HIF-1α signaling pathway is also involved in the process of liver regeneration following liver injury. The protein expression levels of HIF-1α and the mRNA expression of its downstream target, Vegf, were significantly lower in the Fli-06 group than those in the control group. Thus, our study confirmed previous results showing that the in vivo inhibition of Notch signaling can inhibit the expression of HIF-1α and its downstream target, VEGF, in the regenerating liver. 24 It has been reported that HIF-1α can combine with intracellular NICD to increase the stability of NICD in hypoxic environments. 25 Further, Lee et al. 26 showed that the Notch inhibitor, DAPT, inhibits Notch signaling in mouse embryonic stem cells under hypoxic conditions, leading to the down-regulation of VEGF protein expression in these cells, which is also consistent with our experimental results.
Conclusion
The capacity for liver regeneration in rats following hepatectomy decreased significantly after treatment with a gamma-secretase inhibitor, Fli-06, which inhibited Notch signaling and reduced the levels of HIF-1α and its target, Vegf. These findings suggest that both the Notch and Hif-1α signaling pathways are involved in the regulation of rat liver regeneration after partial hepatectomy. However, the regulatory relationship between these two signaling pathways is complex and requires further investigation to fully understand the mechanisms involved in liver regeneration.
