Abstract
Objectives
To study the pathogenic bacterial profile and drug resistance in older patients with community-acquired pneumonia (CAP) in outpatients with fever, and provide evidence to diagnose and treat CAP timely and accurately.
Methods
We studied older (>60 years) patients with CAP in Beijing Shijitan Hospital from 2016 to 2017. Pathogenic bacteria from sputum of patients were isolated and identified and their resistance to antibiotics was tested. Risk factors for multidrug-resistant CAP (MDR-CAP) and clinical outcomes were analyzed.
Results
A total of 5563 outpatients with fever were recruited and 391 had CAP. A total of 117 isolates of pathogenic bacteria were obtained from 176 CAP cases. The main pathogenic bacteria were Klebsiella pneumoniae (27.4%), Escherichia coli (17.9%), Staphylococcus aureus (12.0%), Pseudomonas aeruginosa (10.3%), and Streptococcus pneumoniae (9.4%). A drug sensitivity test (DST) showed that K. pneumoniae, E. coli, and P. aeruginosa had good sensitivity to imipenem, cefoperazone/sulbactam, piperacillin/tazobactam, and amikacin. Staphylococcus aureus and Streptococcus pneumoniae had strong sensitivity to vancomycin, linezolid, and levofloxacin. Previous multiple antibiotic treatment was an independent risk factor for MDR-CAP.
Conclusions
Gram-negative bacteria are the main pathogenic bacteria in older patients with CAP. Identification and DSTs of pathogens could enable accurate diagnosis and treatment of CAP.
Keywords
Introduction
Although community-acquired pneumonia (CAP) is a common infectious disease, it threatens human health worldwide. CAP is also an leading cause of death in adults, especially older people with underlying comorbidities and who are frail, in developing countries. 1 In developed countries, the estimated incidence of CAP is 0.2% to 1.1% in adults and the mortality is 2% to 14% in patients with CAP.2–6 Some studies have shown that the overall incidence of CAP in Mediterranean coastal areas is 0.12%, 7 it is 0.35% in Shanghai city, 8 and the reported mortality rate in Asia is 7.3%. 9 In clinical practice, the pathogens that cause CAP are rarely identified. Antibiotic treatment of CAP is usually empirical, and this delays the diagnosis and treatment of CAP. 10 Appropriate selection of antibiotics depends on the identification of pathogens and the results of a drug sensitivity test (DST). Therefore, a timely and accurate etiological diagnosis and DST help to treat CAP and reduce the mortality of patients with CAP. In the present study, we isolated and identified bacterial pathogens from older patients with CAP. We performed a DST of pathogens to provide scientific evidence for accurate diagnosis and treatment of CAP.
Patients and methods
Study population
This retrospective analysis included data collected between 1 May 2016 and 30 October 2017 at the Department of Infectious Diseases, Beijing Shijitan Hospital, Beijing, China. We enrolled patients who were at least 60 years old with fever (axillary temperature ≥37.5°C) and who underwent a chest X-radiography (CXR) examination. Information on sex, age, days of fever, and respiratory symptoms (e.g., cough and sputum) of cases was recorded by nurses. The study was approved by the ethics committee of Beijing Shijitan Hospital and all patients provided written informed consent.
Diagnostic criteria of CAP
The patients were diagnosed according to the Guidelines of Diagnosis and Treatment for CAP in China. 11 The criteria included the following. Patients were infected in the community and had the presence of new patchy infiltrate, consolidation of a lobe or segment, ground-glass opacity, or interstitial changes on CXR, and had at least one of the following symptoms: (1) recently emerging cough, sputum, or deterioration of the original respiratory symptoms; (2) fever; (3) pulmonary consolidation and/or moist rales; and (4) a white blood cell (WBC) count >10 × 109/L or < 4 × 109/L, excluding pulmonary tuberculosis, pulmonary tumor, non-infectious pulmonary interstitial disease, pulmonary edema, pulmonary atelectasis, pulmonary embolism, pulmonary eosinophilia, pulmonary vasculitis, and acquired immunodeficiency syndrome.
Detection of pathogens
Sputum sample collection
Sputum samples were collected by nurses before the patients had received antibiotics. Smears were prepared from all specimens and examined for squamous epithelial cells and WBCs. All smears were treated with gram stain and examined via light microscopy. Under a low-power field in microscopy, if the number of squamous epithelial cells was < 10, the WBC count was > 25, or the number of squamous epithelial cells/WBCs was less than 1/2.5, the specimens of sputum were considered to qualify for analysis. The specimens were re-collected if the sputum did not meet these criteria.
Isolation, identification, and DSTs of pathogenic bacteria
The sputum collected from patients was inoculated on a blood agar plate, chocolate plate, and MacConkey plate, and cultured at 35°C with 5% carbon dioxide for 18 to 24 hours. Identification of pathogens and the DST were carried out in a clinical microbiology laboratory using the Vitek 2 system (bioMerieux, St. Louis, MO, USA). The Clinical and Laboratory Standards Institute document M100-S24 (2014) was used for interpretation of the DST. 12
Analysis of clinical outcomes
The treatment information of patients was analyzed. Patients with pathogenic bacteria that were isolated from sputum were treated according to the results of the DST of pathogens and were included in the targeted therapy group. Patients with no pathogenic bacteria that were isolated from sputum were treated empirically and were included in the empiric therapy group. The alleviating time of clinical symptoms and the rate of hospitalization were compared between the two groups.
Statistical analysis
Univariate analysis of all potential risk factors for MDR-CAP was performed using SPSS 19.0 (Armonk, NY, USA). Variables with significant differences between two groups in univariate analysis were analyzed using multivariate logistic regression. Comparison between the targeted therapy and empiric therapy groups was performed using the χ2 test for categorical variables and the two-sample t-test for continuous variables. P < 0.05 indicated statistical significance for all statistical analyses.
Quality controls
Quality control of each batch test was performed with the reference strains Escherichia coli (ATCC 25922), Klebsiella pneumoniae (ATCC 700603), Pseudomonas aeruginosa (ATCC 27853), and Staphylococcus aureus (ATCC 25923).
Results
Characteristics of the patients
The study enrolled a total of 5563 outpatients with fever and 391 were diagnosed with CAP. Among the patients, 176 were older than 60 years old, 94 were men, and 82 were women. The mean age of the patients was 68.3 ± 4.3 years.
Isolation and identification of pathogens
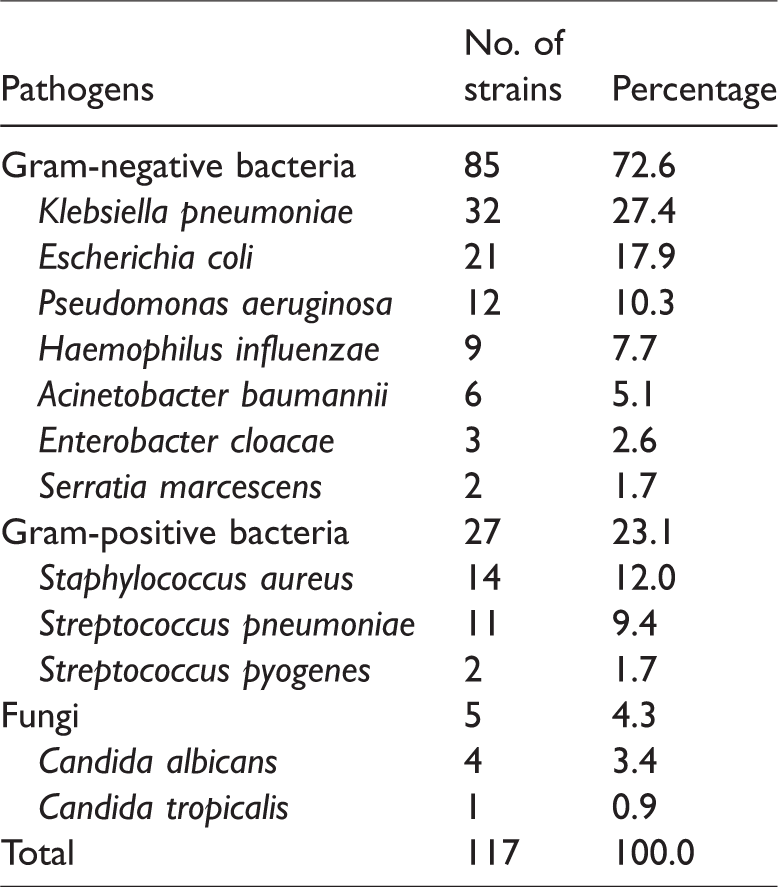
A total of 117 isolates of pathogens were obtained from sputum specimens of 97/176 (55.1%) patients. Of all the isolates, 85/117 (72.6%) were gram-negative bacteria, 27/117 (23.1%) were gram-positive bacteria, and 5/117 (4.3%) were fungi. Additionally, 8/176 (4.5%) patients had more than one pathogen. The main pathogens identified were K. pneumoniae (32 strains, 27.4%), E. coli (21 strains, 17.9%), P. aeruginosa (12 strains, 10.3%), Staphylococcus aureus (14 strains, 12.0%), and Streptococcus pneumoniae (11 strains, 9.4%) (Table 1).
Distribution of the major pathogens in sputum samples of 176 patients.
DST of the pathogens
Drug susceptibility of major gram-negative bacteria to commonly used antibacterial agents.

Values are n (%). S: susceptible; I: intermediate; R: resistance; ESBL: extended‐spectrum beta‐lactamase; −: not tested; SMZ-TMP: sulfamethoxazole-trimethoprim.
Drug susceptibility of major gram-positive bacteria to commonly used antibacterial agents.

Values are n (%). S: susceptible; I: intermediate; R: resistance; −: not tested; SMZ-TMP: sulfamethoxazole-trimethoprim.
Risk factors for MDR-CAP
A total of 29 MDR-CAP cases were identified in this study. Univariate analysis showed that MDR-CAP was significantly associated with age ≥ 75 years (odds ratio [OR]: 2.643; 95% confidence interval [CI]: 1.080–6.472, P = 0.045), comorbidities (OR: 3.986; 95% CI: 1.544–10.288, P = 0.004), prior multiple antibiotic treatment (OR: 6.593; 95% CI: 2.366–18.373, P < 0.001), previous episode of pneumonia (OR: 2.714; 95% CI: 1.098–6.712, P = 0.045), and previous hospitalization (OR: 2.597; 95% CI: 1.066–6.329, P = 0.044). The risk factors in univariate analysis were included in multivariate logistic regression analysis. Prior multiple antibiotic treatment was the only independent risk factor for MDR-CAP (OR: 3.542; 95% CI: 1.141–14.827, P = 0.002) (Table 4).
Risk factors for MDR-CAP in univariate and multivariate analyses.

MDR: multidrug-resistant; CAP: community-acquired pneumonia; OR: odds ratio; CI: confidence interval.
Analysis of clinical outcomes
In our study, 97 patients were included in the targeted therapy group and 79 patients in the empiric therapy group. The alleviating times of coughing, high fever, and lung rales in the targeted therapy group were significantly shorter than those in the empiric therapy group (t = 4.422, 7.862, and 6.848, all P < 0.05). The rate of hospitalization in the targeted therapy group was significantly lower than that in the empiric therapy group (χ2 = 4.262, P < 0.05) (Table 5).
Clinical outcomes in the different patient groups.

Discussion
In the present study, 117 pathogens were isolated and identified from sputum samples of 176 outpatients with CAP. Gram-negative bacteria were the main pathogens (72.6%), including K. pneumoniae, E. coli, and Pseudomonas aeruginosa. The pathogenic bacterial profile of CAP is complicated and varies in different populations and regions. Our findings in this study are different to those of some previous studies as follows. Peto et al. 13 showed that Streptococcus pneumoniae was the main bacterial pathogen isolated from patients with CAP, followed by Legionella pneumophila and Haemophilus influenzae. Furthermore, Gadsby et al. 14 showed that H. influenzae and Streptococcus pneumoniae were the main bacteria in patients with CAP, and their rates of detection were 40.2% and 35.6%, respectively. In one Chinese study, Mycoplasma pneumoniae (20.7%) and Streptococcus pneumoniae (10.3%) were the main pathogens, followed by H. influenzae (9.2%), K. pneumoniae (6.1%), Staphylococcus aureus (3.8%), and E. coli (1.6%%). 15 In our study, the detection rates of H. influenzae and Streptococcus pneumoniae were low, at only 7.7% and 9.4%, respectively. In China, some older people, especially those with chronic diseases, are often treated with antibiotics or even prolonged use of antibiotics. In particular, antibiotics against gram-positive coccus are used by people outside the hospital when they feel slightly uncomfortable. Additionally, some primary community doctors lack the professional knowledge to choose antibiotics rationally, and cause overuse of antibiotics, especially penicillins and cephalosporins. 16 This may be the reason why more gram-negative bacteria are isolated than gram-positive bacteria. The growth conditions of H. influenzae and Streptococcus pneumoniae are more demanding in vitro, which may also be the cause of their low detection.
Our study showed that resistance of K. pneumoniae to cefuroxime, piperacillin, and ampicillin was high. The sensitivity of E. coli to antibiotics was low, including cephalosporins, levofloxacin, and ampicillin. However, the resistance of the main gram-negative bacteria to carbapenems, aminoglycosides, and β-lactamase inhibitor complex was low, which is similar to the studies of Xie et al. 17 and Liu et al. 18 in China. Resistance of the main gram-negative bacteria to cephalosporins in our study was high, which may be related to their widespread use. Although the sensitivity of the major gram-negative bacteria to amikacin was high, amikacin has a higher toxicity to the kidney and poor activity against some drug-resistant strains. Therefore, amikacin is not usually preferred for treating CAP. In the current study, the detection rate of E. coli-producing ESBLs was higher than that of K. pneumoniae-producing ESBLs. This finding could explain why the resistance of E. coli to antibiotics was higher than that for K. pneumoniae because ESBL production increases bacterial resistance. 19
In our study, vancomycin, linezolid, levofloxacin, amikacin, and teicoplanin had strong antimicrobial activity against Staphylococcus aureus (sensitivity rate, ≥78.6%). Streptococcus pneumoniae showed high sensitivity to penicillin, amoxicillin/clavulanate, azithromycin, erythromycin, vancomycin, linezolid, clindamycin, and levofloxacin (sensitivity rate, ≥72.7%). However, Liu et al. 15 and Yang et al. 20 showed that, in macrolides, antibiotics and clindamycin had low activity against Streptococcus pneumoniae strains in pulmonary infection (resistance rate, >75.0%). This difference in sensitivity among studies may be related to the smaller sample size in this study and less use of these two types of antibiotics in clinics in this region. We also found 29 MDR-CAP cases in this study and these cases were associated with prior multiple antibiotic treatment. Although use of multiple antibiotics might significantly inhibit bacterial growth, it also might lead to frequent bacterial mutation and drug resistance. When more than one antimicrobial agent is present in the microorganism environment, pressure from these antimicrobial agents results in selection of bacteria using multiple or polyvalent resistance mechanisms. Therefore, bacteria optimize one resistance mechanism to survive in variable environments or increase mutational events during situations of bacterial stress.21,22
Analysis of clinical outcomes showed that the clinicians’ targeted therapy according to results of identification and DST of pathogenic bacteria could significantly shorten the alleviating time of clinical symptoms and hospitalization rate of patients. The results of identification and DST of pathogenic bacteria can help clinicians select optimal targeted antibiotics, and can eliminate pathogenic bacteria. Although current methods can isolate and identify a pathogen from only 30% to 40% of patients,23–25 analysis of the pathogenic bacterial profile and drug resistance in the region can help clinicians carry out reliable empiric therapy protocols to treat patients with no isolation of pathogenic bacteria. Our study was retrospective and the sample size was limited, with some possible selection bias. However, our findings are still meaningful for providing a foundation for future improvements in microbiological diagnostic measures and an antibiotic regime for bacterial pathogens in patients with CAP.
Footnotes
Acknowledgements
The authors would like to thank the patients enrolled in the study, and Dr Hao Zhu and Dr Jiyong Jian for the identification and DSTs of the pathogens.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not for-profit sectors.
