Abstract
Chromosomal abnormalities involving 2q32q33 deletions are very rare and present with a specific phenotype. This case report describes a 37-year-old female patient with 2q32q33 microdeletion syndrome presenting with the characteristic features, but with the addition of secondary cognitive decline. Molecular karyotyping was performed on the patient and her parents. It revealed an 8.6 megabase deletion with the proximal breakpoint in the chromosome band 2q32.2 and the distal breakpoint in 2q33.1. The deletion encompassed 22 known genes, including the GLS, MYO1B, TMEFF2, PGAP1 and SATB2 genes. The observed deletion was confirmed using a paralogue ratio test. This case report provides further evidence that the SATB2 gene, together with GLS, MYO1B, TMEFF2 and possibly PGAP1, is a crucial gene in 2q32q33 microdeletion syndrome. The SATB2 gene seems to be crucial for the behavioural problems noted in our case, but deletion of the GLS, MYO1B and TMEFF2 genes presumably contributed to the more complex behavioural characteristics observed. Our patient is also, to our knowledge, the only patient with 2q32q33 microdeletion syndrome with secondary cognitive decline.
Keywords
Introduction
Individuals with deletions of 2q32q33 and who present with learning difficulties, growth retardation, dysmorphic features, thin and sparse hair, feeding difficulties, and high or cleft palate have been classified as having a new microdeletion syndrome. 1 A specific behavioural pattern, with hyperactivity and motor restlessness, chaotic behaviour, a happy personality but with periods of aggression and anxiety, sleeping problems and self-mutilation, has also been reported in some of these patients. 2
Until 2005, the only report of a similar behavioural phenotype with psychiatric problems in 2q deletion patients was presented by Vogels et al. 3 Subsequently, a report described a new recognizable pattern of abnormalities in four patients with an interstitial deletion of the chromosome region 2q32q33. 4 A behavioural phenotype associated with the 2q32q33 microdeletion was described in other clinical reports.1,5–10
We report on an adult patient with a de novo 8.6 megabase (Mb) deletion of 2q32.2q33.1 with the characteristic phenotype and behavioural problems, but whose prominent secondary cognitive decline was a key difference compared with other reported patients. To our knowledge, this feature has not previously been described in 2q32q33 microdeletion patients. This report aims to provide a new perspective on genotype–phenotype correlations in patients with 2q32q33 microdeletion syndrome, focusing on the behavioural phenotype and secondary cognitive decline, since these were the most prominent clinical features observed in this patient.
Case report
A 37-year-old woman, born in September 1977, first presented to our clinic in February 2015 because of severe aggressive outbursts. She was the only child of healthy, unrelated Caucasian parents with an unremarkable family history. She was born 3 days after the due date following an uneventful pregnancy with a birth weight of 3900 g (43rd centile) and a birth length of 52 cm (55th centile). Occipitofrontal circumference and Apgar scores were not reported. Cleft palate and craniofacial dysmorphic signs were noted at birth. She had problems with swallowing. She began sitting at 9 months and walked at 18 months. Her speech development was delayed; her vocabulary consisted of only a few words (mama, papa). She pointed with her hands to indicate what she wanted.
At the age of 6 years, the patient achieved a mental age of 3 years, with an intelligence quotient (IQ) in the mild mental retardation range (IQ 50–55) on the Wechsler Intelligence Scale for Children (WISC). She was hyperkinetic, with behavioural problems that were still manageable. She attended a special school, where she coped well with the programme. She learned to ride a tricycle. She had only a few, poorly-developed, protruding teeth.
At the age of 12 years, the patient’s mental state deteriorated. She regressed to the level of severe mental retardation, achieving an IQ of 25–30 points on WISC. Her speech deteriorated and she was no longer able to speak any words. Her height was 142 cm (10th centile) and her weight was 37 kg (26th centile). She understood and carried out only very simple instructions and tasks. She disliked the company of children of her age, preferring the company of adults. Her hyperkinetic and behavioural problems increased at this time, together with auto- and heteroaggression. Left facial hypotrophy was evident, and her walk was rigid and broad-based. She was admitted to a care home for mentally handicapped children.
In May 2015, when she was last examined for this report, the patient’s aggressive outbursts were even more frequent and persistent than they had been previously. Her behaviour was unpredictable, with shouting, throwing herself on the floor, hitting her head on the floor (causing nose bleeds), refusing to eat and being aggressive towards others. Swallowing problems were still present. Computed tomography scans, before and after the application of contrast media, revealed no abnormalities in the skull or brain matter. The patient was of normal height and weight; she had sparse hair, thin, pale skin, craniofacial dysmorphism (a long and asymmetric face, short and flat philtrum, and pointed chin), oligodontia and a left periorbital dermoid cyst (Figure 1).
Current appearance of a 37-year-old woman with a de novo 8.6 megabase deletion of 2q32.2q33.1 who presented with the characteristic phenotype and behavioural problems, but who also experienced prominent secondary cognitive decline. Characteristic features of 2q32q33 microdeletion syndrome noted in this patient are the sparse hair, thin, pale skin and craniofacial dysmorphism (including long and asymmetric face, short and flat philtrum, pointed chin, oligodontia, and periorbital dermoid cyst). The colour version of this figure is available at: http://imr.sagepub.com.
Molecular karyotyping was performed on this patient and her parents using DNA extracted from peripheral blood leukocytes with the QIAamp® DNA Blood Midi Kit (QIAGEN, Hilden, Germany). Analysis was carried out using the BlueGnome CytoChip ISCA 8×60K platform (Illumina UK, Cambridge, UK). The protocol was performed according to the manufacturer’s instructions and data were analysed using the BlueFuse™ Multi v3.1 software tool (Illumina UK). The deletion was confirmed using a paralogue ratio test (PRT) according to a previously described protocol. 11 For this, a pair of oligonucleotides (identification number: chr2ppn1p449839453) was selected, which coamplify a region within the deletion (chr2: 196,578,414–196,578,640; 226 base pairs [bp]) and a reference region from chromosome 17 (chr17: 58,027,459–58,027,690; 231 bp).
Molecular karyotyping in this patient revealed an 8.6 Mb interstitial deletion of the chromosomal region on the long arm of chromosome 2, between 191,711,722 bp and 200,347,340 bp (GRCh37/hg19). The deletion lies within the chromosomal bands 2q32.22q33.1 and encompassed 22 known genes listed in the Online Mendelian Inheritance in Man (OMIM) database,
12
including the GLS, MYO1B, TMEFF2, PGAP1 and SATB2 genes. The chromosomal position of the deletion in the present case is labelled red on Figure 2. Molecular karyotyping of the patient’s parents did not show any aberrations. The PRT analysis showed an ∼50% reduction in signal height for the chromosome 2 target locus (226 bp fragment) compared with the signal height from the reference 231 bp fragment from chromosome 17. Statistical analysis yielded a Z-score of 5.2 with a corresponding P-value <0.001. Thus, the PRT result confirmed the presence of copy number variation on chromosome 2 in this patient.
A schematic presentation of deletions in patients with reported 2q32q33 microdeletion syndrome and behavioural problems, including those of the present case (labelled red). Size and positions of deletions with implicated genes are presented in the University of California, Santa Cruz Genome Browser. The colour version of this figure is available at: http://imr.sagepub.com.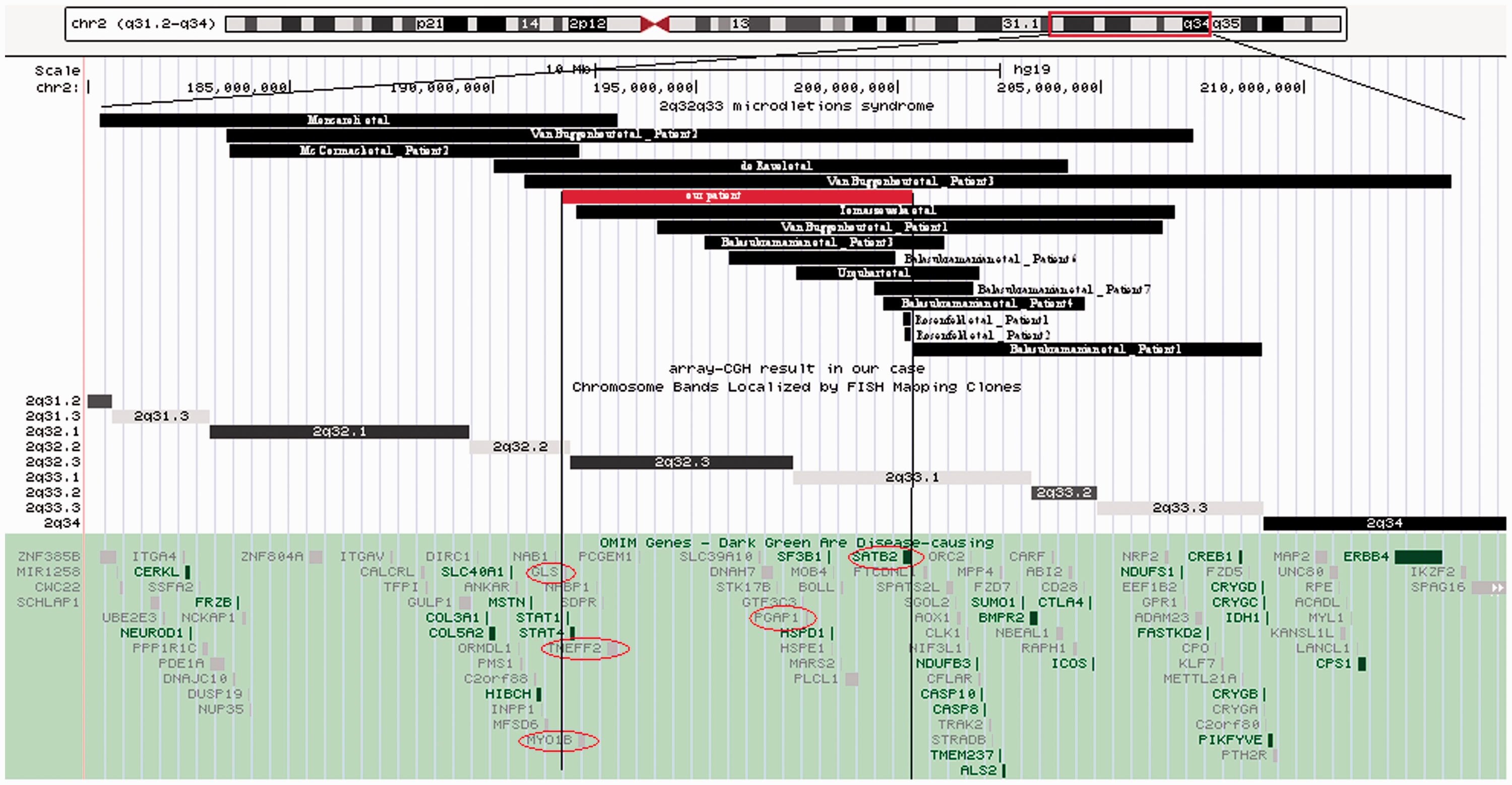
Ethics approval for this analysis was obtained from the Republic of Slovenia National Medical Ethics Committee (NMEC), Ljubljana, Slovenia (NMEC 89/01/11). Written informed consent was obtained from the parents of the patient for publication of this case report and its accompanying images.
Discussion
This present case report describes a patient with 2q32q33 microdeletion syndrome and secondary cognitive decline, which is, to our knowledge, a unique feature not previously described in people with this microdeletion syndrome. 2 The current proband reached a mental level in the mild mental retardation range by the age of 6, but deteriorated to the level of severe mental retardation by the age of 12. Her previously poor speech disappeared completely. The current proband is also the oldest patient with 2q32q33 microdeletion syndrome described in the literature, providing enough time for the emergence of secondary cognitive decline. Approximately 25 patients with deletions in 2q32q33 have been reported to date, the vast majority being <15 years of age.2,13 One patient from a previous report was 1 year younger than the current patient. 9 Another report described possible intellectual decline in a patient who developed normally until the age of 12 months, but in whom psychomotor retardation became evident by the age of 18 months. 1 However, it is premature to talk about secondary cognitive decline at the age of 18 months, so further analysis and longer follow-up are needed in order to confirm if the unique finding in the current case also occurs in other patients with this deletion. Behavioural problems in the current proband were manageable until the age of 6, after which bursts of self-mutilation, aggression and shouting dominated the clinical picture, leading to a constant need for antipsychotic medication, although her general mood was euthymic, happy and content. This specific behavioural pattern has already been observed in some patients with 2q32q33 microdeletion syndrome.1–10 Other phenotypic features observed in our patient (including facial dysmorphism, sparse hair, thin skin and oligodontia) were similar to those previously reported. 2
Molecular karyotyping in the current patient revealed an 8.6 Mb deletion with the proximal breakpoint in the chromosome band 2q32.2 and the distal breakpoint in 2q33.1. The deletion encompassed 22 known OMIM genes, including GLS, MYO1B, TMEFF2, PGAP1 and SATB2. These genes have already been implicated as being causative in the clinical outcome in patients with 2q32q33 microdeletion syndrome (Figure 2).4,5,7,8 The K-type mitochondrial glutaminase encoded by the GLS gene plays an important role in energy metabolism, ammonia trafficking and regeneration of the neurotransmitter glutamate, suggesting that the glutaminase is crucial for neuronal maturation. 14 The MYO1B gene is implicated in processes critical to neuronal development and function, such as cell migration, neurite outgrowth and vesicular transport. 9 The TMEFF2 gene encodes a survival factor for hippocampal and mesencephalic neurons. 15 PGAP1 encodes glycosylphosphatidylinositol inositoldeacylase, an enzyme involved in the Wnt signalling pathway. 16 PGAP1-deficient mice show a varied facial appearance, from normal to a complete lack of mouth and jaw. 17 SATB homeobox 2, encoded by the SATB2 gene, is a DNA-binding protein that participates in transcription regulation and chromatin remodelling. SATB2 knockout mice have been reported with craniofacial anomalies, and defects in osteoblast differentiation and function. 18 A de novo nonsense mutation of SATB2 has been described in an individual associated with cleft palate, osteoporosis and cognitive impairment. 19 Apparently balanced translocations within the SATB2 gene have also been described in individuals with some of the features seen in 2q32q33 microdeletions.20–22
Van Buggenhout et al. 4 suggested that haploinsufficiency of the SATB2 gene was crucial to the craniofacial abnormalities observed, including high or cleft palate. This gene plays an important role in craniofacial patterning and development, palate formation, and osteoblast differentiation and maturation. 23 A specific behavioural pattern was observed in three of four patients described by Van Buggenhout et al., 4 thus it has been suggested that other gene(s), and not SATB2, are implicated in the behavioural features. Considering that molecular karyotyping in this report was carried out using bacterial artificial chromosome array, it is difficult to define precisely how the boundaries of deletions in patients without behavioural problems differ from those with such problems. Two other reports described two additional patients with similar behavioural problems to those observed in patients 1, 2 and 3 reported by Van Buggenhout et al.4–6 By comparing their patients with patients 2 and 3 of Van Buggenhout et al., 4 both authors proposed that GLS, MYO1B and TMEF2 could be responsible for the behavioural phenotype observed in all patients,5,6 although the deletion in Van Buggenhout’s patient 1 did not include these three genes. 4 The importance of GLS, MYO1B and TMEF2 genes is probably even more evident in the report of Mencarelli et al., 5 since the deletion in their patient did not overlap the SATB2 gene. Based on the function of ZNF533 and MYO1B, the authors suggested that the aforementioned genes are good candidates for the neurological abnormalities noted in their patient. 5 Furthermore, the GLS gene encoding glutaminase could be responsible for the behavioural phenotype observed, since glutamate (which is a product of the enzymatic reaction of this enzyme) can act as a neurotransmitter.24,25 Interestingly, a high palate was described in the patient reported by Mencarelli et al., 5 indicating that other genes, in addition to SATB2, in this chromosomal region are probably also involved in palate formation. Further evidence for the importance of other genes in palate formation was gained from another report, 9 which described the proband’s mother as having an 8.6 Mb deletion of 2q32.1q32.3, which is fully covered by the deletion in Mencarelli's patient. 5 The FRZB gene (OMIM #605083), an antagonist of the Wnt8 signalling pathway involved in limb and craniofacial skeletogenesis, has been suggested as a possible candidate gene for the craniofacial abnormalities, while the GLS and MYO1B genes have once again been proposed as good candidates for the behavioural problems observed. 9 Moreover, the PGAP1 gene, together with the SATB2 gene, has been suggested as a plausible candidate for craniofacial abnormalities. 7
Patients reported by Urquhart et al., 7 Rosenfeld et al. (patients 1 and 2), 8 and Balasubramanian et al. 1 (patients 1, 3, 4, 6 and 7) were also described as having behavioural problems. In all of these patients, the deletion lies proximal to the GLS, MYO1B and TMEF2 genes and involves the SATB2 gene, which could be responsible for both the craniofacial features and behavioural problems.1,7,8 Doubts regarding the SATB2 gene were also raised by Rosenfeld et al., 8 who presented three patients with intragenic deletions of SATB2. Only two of the three had behavioural problems, suggesting that incomplete penetrance is possible. 8 This possibility is further supported by the balanced translocations of SATB2 reported, with one patient having only a learning disability 26 and the other having autistic spectrum disorder. 21 Behavioural problems were also not reported in Van Buggenhout’s patient 4 and Balasubramanian’s patient 2, where deletions in both patients encompassed the SATB2 gene.4,1
The proximal breakpoint of the deletion described in this current patient and that reported by Tomaszewska et al. 10 differ by <400 kilobases. In the current proband, the GLS gene was included in the deleted region, whereas in Tomaszewska’s patient the deletion begins with the MYO1B gene. 10 Since both patients showed behavioural problems (they were generally happy with bouts of aggression), we propose that TMEF2 and MYO1B could be the key genes for the observed behavioural features, as previously suggested.5,6 This current report also provides further evidence that GLS could be responsible for the behavioural phenotype, as proposed elsewhere.5,6 Deletion of the GLS gene in this current proband could have resulted in severe behavioural problems, such as self-mutilation and screaming, which were not observed in Tomaszewska's patients, 10 but were noted in a patient of Mencarelli et al. 5
In conclusion, considering the previously published findings regarding phenotype–genotype correlations in 2q32q33 microdeletion syndrome, this current case report presents further evidence that craniofacial features, including cleft palate and teeth abnormalities, are most likely the result of disruption of the SATB2 gene. The distal breakpoint of the deletion in this current proband was situated within the SATB2 gene, causing the loss of function of this gene. Previously described intragenic deletions have shown that dysfunction of the SATB2 gene can lead to the formation of cleft palate, with plausible incomplete penetrance. 8 The previously described patients with 2q32q33 microdeletions with palate malformations and deletions lying proximal 5 or distal 1 to the SATB2 gene indicate that other genes in this region are also involved in palate formation. Thus, haploinsufficiency of the PAGP1 gene, as previously outlined, 7 could be associated with palate anomalies in this current patient. Moreover, it is our opinion that the SATB2 gene is also crucial to the behavioural problems noted in this current proband and that deletion of the GLS, MYO1B, and TMEFF2 genes presumably contributes to the more complex behavioural characteristics observed. This current patient is also, to our knowledge, the only patient with 2q32q33 microdeletion syndrome with secondary cognitive decline documented by psychometric tests. This clinical finding cannot be explained only by the genes included in 2q32q33 microdeletion syndrome. There is a need for further observation and follow-up of other patients with 2q32q33 microdeletion syndrome in order to provide further information on possible phenotype–genotype associations.
Footnotes
Acknowledgments
We thank the patient’s family for their cooperation.
Declaration of conflicting interest
All authors declare that there are no conflicts of interest.
Funding
This work was supported by a PhD fellowship for young researchers and the research programme P4–0220, Comparative Genomics and Genomic Biodiversity, of the Slovenian Research Agency (ARRS).
