Abstract
Objective
To measure levels of microRNA (miR)-21 and its target gene, programmed cell death 4 (PDCD4), in samples of human cutaneous malignant melanoma and normal non-malignant control skin.
Methods
Relative levels of miR-21 and PDCD4 mRNA were measured using a quantitative real-time reverse transcription–polymerase chain reaction. Correlations between the levels of the two molecules and the clinicopathological characteristics of malignant melanoma were analysed.
Results
A total of 67 cases of human cutaneous malignant melanoma were analysed and compared with 67 samples of normal nonmalignant control skin. Compared with normal skin samples, the relative level of miR-21 was significantly higher and the relative level of PDCD4 mRNA was significantly lower in the melanoma specimens. A significant negative correlation between PDCD4 mRNA and miR-21 was demonstrated in malignant melanoma (r = −0.602). Elevated miR-21 and reduced PDCD4 mRNA levels were both significantly correlated with increased tumour size, a higher Clark classification level and the presence of lymph node metastases in malignant melanoma.
Conclusions
These findings suggest that miR-21 and PDCD4 might be potential biomarkers for malignant melanoma and might provide treatment targets in the future.
Introduction
Malignant melanoma, a tumour derived from naevus cells and melanocytes, has a rapidly increasing incidence and is the leading cause of skin cancer deaths, worldwide. 1 The main reasons for the poor prognosis and high mortality rate associated with malignant melanoma include the diagnosis of late-stage melanoma, its early metastasis via blood vessels and lymph nodes and resistance to chemotherapy.2,3 There is an urgent need to determine the detailed cellular and molecular mechanisms underlying the development and progression of this disease, which are complex and remain poorly understood.
MicroRNAs (miRs) belong to a class of small, highly conserved noncoding RNAs (14–24 nucleotides) that are widely present in mammalian genomes, which can regulate gene expression at the post-transcriptional level by the degradation of target mRNA or the inhibition of mRNA translation. 4 Studies reveal that several miRs play important roles in the tumourigenesis, progression and metastasis of malignant melanoma, such as miR-137, miR-101, let-7, miR-26a, miR-221/222, miR-155, miR-21, miR-182, miR-30b/30d, and miR-214. 5 The overexpression of miR-21 has already been shown to be associated with progression, metastasis and poor prognosis in a variety of cancers, such as colon cancer, pancreatic cancer, and breast cancer.6–8
Programmed cell death 4 (PDCD4), a tumour suppressor gene, has been shown to be downregulated or absent in different human cancers and this downregulation has been associated with a poor cancer outcome.9,10 Through the regulation of p21, cyclin-dependent kinase 4, ornithine decarboxylase, carbonic anhydrase II, urokinase receptor and JNK/c-Jun/AP-1, PDCD4 performs many functions in neoplastic transformation, apoptotic regulation, and invasion. 11 Recent studies have shown that PDCD4 is the downstream target gene of miR-21 in colon cancer, gastric cancer and hepatocellular carcinoma.6,12,13 However, a possible relationship between miR-21 and PDCD4 remains to be investigated in malignant melanoma.
The aim of the present study was to measure miR-21 and PDCD4 mRNA levels in human cutaneous malignant melanoma tissues, and to analyse the association between their levels and clinicopathological characteristics of malignant melanoma.
Patients and methods
Patients and tissue samples
Paraffin wax-embedded malignant melanoma tissue specimens were sequentially collected from patients with malignant melanoma by the Department of Dermatology, Qilu Hospital, Shandong University, Jinan, China, the Department of Pathology, Medical School, Shandong University, Jinan, China, the Department of Dermatology, The Traditional Chinese Medicine Hospital of Jinan City, Jinan, China and the Department of Dermatology, Weifang People’s Hospital, Weifang, China between January 2010 and December 2012. Paired nonmalignant normal control skin specimens were sequentially collected from healthy volunteers from the Department of Plastic Surgery, Qilu Hospital, Shandong University, Jinan, China. Sex and age were matched between patients and healthy control subjects between January 2010 and December 2012. The location and size of the tumour and the presence of lymph node metastases were recorded for every patient with malignant melanoma. Tissue sections derived from the specimens were routinely stained with haematoxylin and eosin, and were reviewed by two pathologists to assess the Clark classification. 14 Patients diagnosed with primary cutaneous malignant melanoma who had not undergone chemotherapy, radiotherapy or surgical treatment were included into this study. Written informed consent was obtained from all patients and healthy control subjects. This study was approved by the Ethics Committee of Qilu Hospital (reference number: QL2009079).
Extraction of total RNA and miRNA
Paraffin wax surrounding the tissue samples was removed and serial sections (10µm) were cut from the block specimens. Four sections were loaded into a 1.5 ml microcentrifuge (BT620-NS; Sangon Biotech, Shanghai, China) and 1 ml xylene was added to the sample. The sample was vortexed vigorously for 10 s and then centrifuged at 13 400 g for 2 min at room temperature. The supernatant was discarded and 1 ml 100% ethanol was added to the pellet, which was centrifuged at 13 400 g for 2 min at room temperature. The supernatant was discarded and the pellet was air dried at room temperature. Subsequent extraction procedures were undertaken according to the manufacturer’s instructions for the total RNA extraction kit and the microRNA rapid extraction kit (BioTeke, Beijing, China). An ultraviolet spectrophotometer (SMA4000; Merinton, Beijing, China) was used to detect the purity and concentration of total RNA.
Measurement of miR-21 and PDCD4 mRNA using qRT–PCR
Real-time quantitative reverse transcription–polymerase chain reaction (qRT–PCR) for miR-21 and PDCD4 mRNA was performed on cDNA generated from the above total RNA according to the manufacturer’s instructions described below, using U6 noncoding small nuclear RNA and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as the internal controls, respectively. Primer sequences were designed as follows (Sangon Biotech): miR-21 forward 5′-GAAATGCCTCACAGCTATCGT-3′ and reverse 5′-CCTCCACAAAGAGCCACC-3′; U6 forward 5′-ATCATGTTTGAGACCTTCAACA-3′ and reverse 5′-CATCTCTTGCTCGAAGTCCA-3′; PDCD4 forward 5′- CCTGAATTAGCACTGGATACTCCT-3′ and reverse 5′- CTAGCCTGCACACAATCTAAGTT-3′; GAPDH forward 5′- CGGAGTCAACGGATTTGGTCGTAT-3′ and reverse 5′- AGCCTTCTCCATGGTGGTGAAGAC-3′. The RT reaction system (20 µl; All-in-one First Strand cDNA Synthesis Kit; FulenGen, Guangzhou, China) contained 5 µl RNA, 0.4 µl 10 mM dNTP, 0.8 µl N6 random primer, 4 µl 5 × Reverse buffer, 0.3 µl Moloney murine leukaemia virus reverse transcriptase, 0.4 µl RNasin and 9.1 µl diethylpyrocarbonate water. The reaction conditions were as follows: 37℃ for 60 min and 94℃ for 3 min. After the RT reaction, qRT–PCR reaction was performed as follows using an ABI7500 real-time PCR detection system (Applied Biosystems, Foster City, CA, USA). The qRT–PCR reaction system (20 µl) contained 2.0 µl cDNA, 10 µl 2 × SYBR Premix ExTaq™ II (TaKaRa Biotechnology, Shiga, Japan), 1.6 µl of the specific primers, 0.4 µl 50 × ROX Reference Dye (TaKaRa Biotechnology) with additional triple-distilled water up to a final volume of 20 µl. The PCR cycling conditions comprised of preliminary denaturation at 95℃ for 30 s, followed by 40 cycles of denaturation at 95℃ for 5 s, annealing and elongation at 60℃ for 34 s, and then a dissociation stage at 60℃ for 15 s. Each sample was measured in triplicate and the amount of miR21 and PDCD4 mRNA relative to the internal controls were determined using the 2−ΔΔC
Statistical analyses
All statistical analyses were performed using the SPSS® statistical package, version 17.0 (SPSS Inc., Chicago, IL, USA) for Windows®. Relative levels of miR-21and PDCD4 mRNA are presented as mean ± SD. The difference between the two groups was analysed using Student’s t-test. The correlation between the relative levels of miR-21 and PDCD4 mRNA was assessed using Spearman’s rank correlation coefficient analysis. Fisher’s exact test was used to assess the correlation between the relative levels of miR-21 and PDCD4 mRNA and the clinicopathological characteristics of malignant melanoma. A P-value < 0.05 was considered statistically significant.
Results
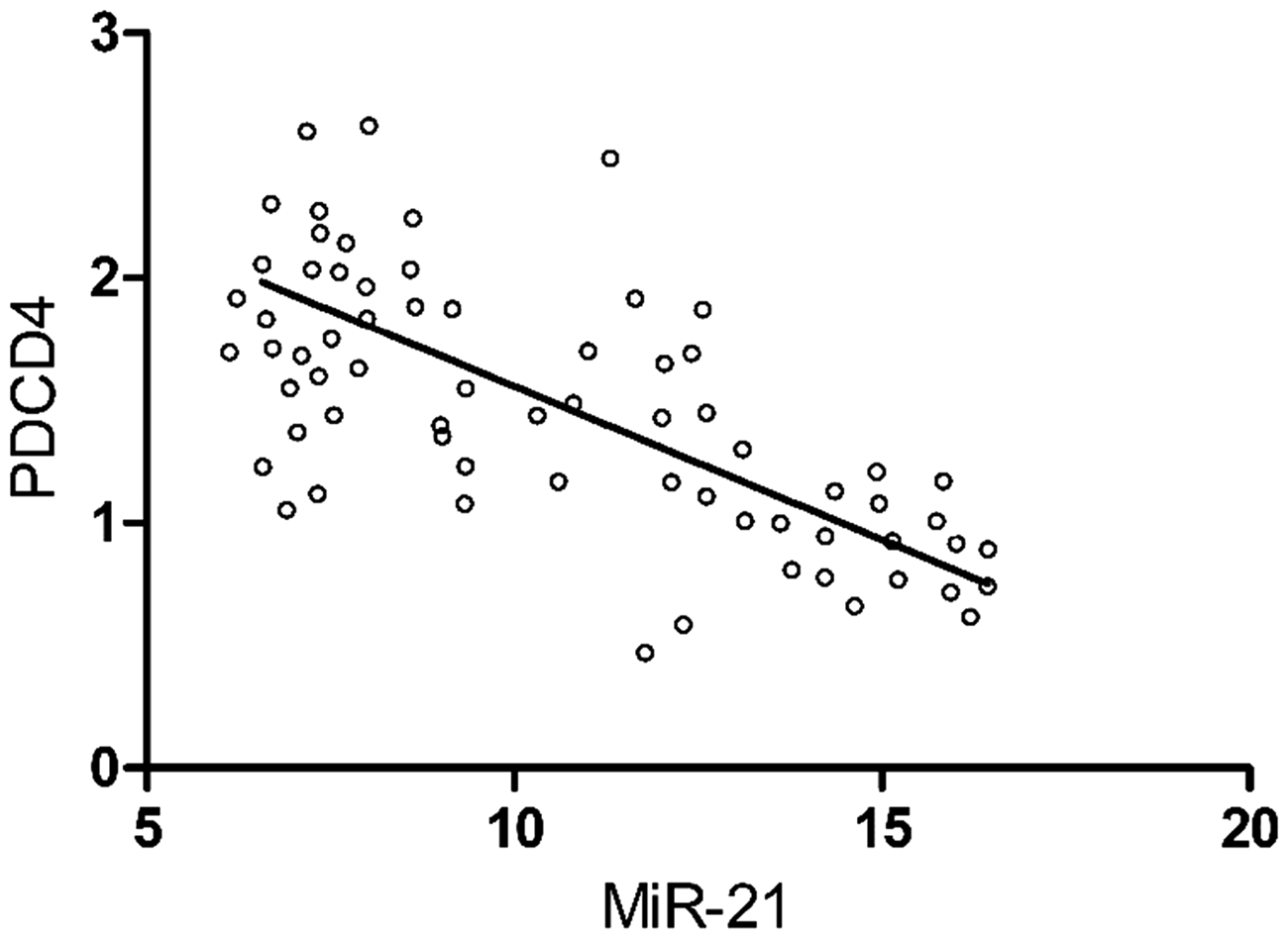
This study used 67 malignant melanoma tissue specimens from 43 male and 24 female patients (median age, 50.7 years [range 40–73 years]) and 67 paired nonmalignant normal control skin specimens. Among the 67 malignant melanoma specimens, 59 specimens were from the limbs, six from the face an two from the chest. The mean ± SD relative level of miR-21 was significantly higher in malignant melanoma specimens compared with control samples (9.853 ± 3.136 versus 1.107 ± 0.113, respectively; P < 0.05) (Figure 1). The mean ± SD relative level of PDCD4 mRNA was significantly lower in malignant melanoma specimens compared with control samples (1.385 ± 0.429 versus 9.341 ± 1.252, respectively; P < 0.05) (Figure 2). As shown in Figure 3, there was a significant negative correlation between the relative levels of miR-21 and PDCD4 mRNA in malignant melanoma tissues, as shown by Spearman’s rank correlation coefficient analysis (r = −0.602, P < 0.05).
Relative levels of microRNA (miR)-21 in human cutaneous malignant melanoma tissue specimens (malignant melanoma) and human normal nonmalignant control skin samples (normal). Data presented as mean ± SD. *P < 0.05; normal skin versus malignant melanoma; Student’s t-test. Relative levels of programmed cell death 4 (PDCD4) mRNA in human cutaneous malignant melanoma tissue specimens (malignant melanoma) and human normal nonmalignant control skin samples (normal). Data presented as mean ± SD. *P < 0.05; normal skin versus malignant melanoma; Student’s t-test. Correlation between relative levels of microRNA (miR)-21 and programmed cell death 4 (PDCD4) mRNA in human cutaneous malignant melanoma specimens (n = 67). Spearman’s rank correlation coefficient analysis (r = −0.602, P < 0.05).

Clinicopathological characteristics and relative levels of microRNA (miR)-21 and programmed cell death 4 (PDCD4) mRNA in 67 malignant melanoma tissue specimens.

Data presented as n patients and mean ± SD.
Fisher’s exact test.
Discussion
Similar to most other tumour types, the genesis, progression and invasion of malignant melanoma is a complex biological process that involves the abnormal regulation of multiple genes. 15 MiRNAs play important functions in cell differentiation, cell proliferation, apoptosis, metabolism, and immune regulation by promoting the degradation of their target mRNA or inhibiting mRNA translation. 16 Upregulated miRNA levels, for example miR-182 and miR-149, and downregulated levels of miRNAs, for example miR-26a and miR-101, have been shown to be involved in the development and progression of malignant melanoma.5,17
Overexpression of miR-21 is associated with the progression, metastasis and poor prognosis of many types of tumours via its regulation of the phosphatase and tensin homologue (PTEN), the tissue inhibitor of metalloproteinase 3 and PDCD4 genes.6–8 A study of 127 primary melanoma tissues demonstrated that levels of miR-21 were significantly increased compared with benign naevi. 18 Increased expression of miR-21 was also demonstrated in 86 primary cutaneous melanomas and expression levels were correlated with Breslow thickness and advanced clinical stage. 19 This present study showed that consistent with previous studies,19,20 the upregulation of miR-21 in malignant melanoma was correlated with clinicopathological characteristics such as an increased tumour size, a higher Clark classification level and the presence of lymph node metastases.
At present, searching for the downstream target genes of miR-21 and clarifying the regulating mechanisms have become important areas of research in malignant melanoma. PDCD4, a tumour suppressor gene, plays multiple important roles in transformation, tumourigenesis, progression and invasion. 11 The downregulation of PDCD4 was associated with tumour stage, nodal metastasis, and with both vascular and perineural invasion in oesophageal cancer. 10 This present study showed that the levels of PDCD4 mRNA were significantly lower in malignant melanoma specimens than in normal control skin samples. The downregulation of PDCD4 mRNA in malignant melanoma was also correlated with clinicopathological characteristics such as an increased tumour size, a higher Clark classification level and the presence of lymph node metastases in the present study. Research has demonstrated that the PDCD4 gene is a direct downstream target for miR-21. 20 Overexpression of miR-21 regulated the metastatic behaviour of B16 melanoma cells by inhibiting PDCD4, PTEN and B-cell translocation gene 2. 20 This present study also found a negative correlation between PDCD4 mRNA and miR-21 in malignant melanoma tissue, which suggests that the miR-21-modulated downregulation of the PDCD4 gene might be a crucial oncogenic mechanism in malignant melanoma. To our knowledge, this is the first published article demonstrating a correlation between PDCD4 mRNA and miR-21 in malignant melanoma tissue.
This present study was limited by the fact that it simply measured the levels of miR-21 and PDCD4 mRNA in malignant melanoma specimens compared with normal nonmalignant control skin. Further research is required to provide a detailed understanding of the molecular mechanisms responsible for the association between PDCD4 mRNA and miR-21in malignant melanoma tissue.
In conclusion, elevated miR-21 and reduced PDCD4 mRNA levels, which were negatively correlated with each other, were associated with clinicopathological characteristics in malignant melanoma. These findings suggest that miR-21 and PDCD4 mRNA levels might be potential biomarkers for malignant melanoma, and might provide treatment targets in the future.
Footnotes
Acknowledgements
We would like to thank Drs Weiyuan Ma and Yukun Wang for assessing the Clark classification of the sections.
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
