Abstract
Antibodies are indispensable tools in biomedical research, yet the widespread use of animal-derived antibodies contributes to poor reproducibility and raises serious ethical concerns. Many commercial antibodies fail to bind their intended targets, leading to irreproducible results, wasted resources and stalled progress. In Europe alone, over a million animals are used annually for antibody production. Non-animal-derived recombinant antibodies and antibody mimetics offer a scalable, scientifically valid alternative. Endorsed by the European Union Reference Laboratory for alternatives to animal testing (EURL ECVAM), these reagents deliver greater consistency, transparency and ethical alignment. Despite their advantages, they are currently estimated to make up less than 5% of research reagents, held back by limited awareness, access barriers and persistent misconceptions about performance. To address these challenges, the Centre for Human Specific Research developed the open-access Recombinant Antibodies & Mimetics Database. This platform links researchers to ethically sourced, sequence-defined affinity reagents and features a unique colour-coded classification system to clearly indicate the extent of animal involvement. By improving transparency, the database empowers informed decision-making and supports alignment with best practices. More than a sourcing tool, the platform drives advocacy for systemic change, encouraging funders, institutions, publishers and regulators to prioritise recombinant antibodies and antibody mimetics. Its ultimate aim is to make high-quality animal-free recombinant and mimetic antibodies the global standard, supporting a more reproducible, ethical and forward-looking future for antibody science.
Antibodies in research: A call for reliable and ethical alternatives
Animal use in antibody production: An ongoing concern
The research-use-only (RUO) antibody market comprises over 6 million antibodies from more than 300 suppliers, 1 and is projected to grow from US$1.6 billion in 2023 to US$2.2 billion by 2030 (a compound annual growth rate (CAGR) of 4.8%). 2 Growing demand is being fuelled by advances in genomics and proteomics, which continue to uncover new proteins, isoforms and post-translational modifications, each representing a potential new target for antibody development. 3 To meet these demands, large numbers of animals are still used to generate antibodies, sustaining reliance on traditional animal-based production methods. The European Union Reference Laboratory for alternatives to animal testing (EURL ECVAM) estimates that ∼1 million animals are used annually in Europe alone for antibody production. 4 Globally, the number will be significantly higher, though a lack of transparency makes accurate estimations difficult.
Quality and reproducibility: A systemic failure
Despite the scale of use, there is still no industry-wide standard for validating antibodies, contributing to what has been termed the ‘reproducibility crisis’. 5 A 2008 study found that only 49% of 5436 commercial animal-derived antibodies were specific to their intended target, 6 and a 2014 analysis of 13,000 antibodies used in Western blotting found that only 45% produced supportive staining. 7 This irreproducibility leads to publication retractions, wasted resources and terminated research projects, with estimates suggesting up to US$800 million annually is lost due to poor-quality animal-derived antibodies. 8
A call to the scientific community for a validated solution: Non-animal-derived antibodies
In response to these challenges, the EURL ECVAM Scientific Advisory Committee (ESAC) reviewed the evidence supporting animal-free antibody technologies. ESAC concluded that non-animal-derived antibodies are not only scientifically valid, but often superior, offering better-defined reagents that enhance reproducibility and research efficiency. In 2020, EURL ECVAM published a recommendation stating: “Animals should no longer be used for the development and production of antibodies for research, regulatory, diagnostic and therapeutic application… EU countries should no longer authorise the development and production of antibodies through animal immunisation, where robust, legitimate scientific justification is lacking.”
4
This position aligns with the requirements of EU Directive 2010/63/EU, which states that “wherever possible, a scientifically satisfactory method or testing strategy, not entailing the use of live animals, shall be used instead of a procedure”. 9 Despite these advantages, uptake of recombinant, non-animal-derived antibodies remains limited. They are currently estimated to make up less than 5% of research reagents, 10 underscoring the urgent need for greater awareness, adoption and policy alignment within the scientific community.
The Recombinant Antibodies & Mimetics Database
Developed by the Centre for Human Specific Research, the Recombinant Antibodies & Mimetics Database 11 is an open-access platform that connects researchers to global suppliers of sequence-defined reagents. While all listed products meet this sequence-defined standard, they differ in origin, discovery methods and whether animal-derived biomaterials (ADBs) were used at any point in development or production. These distinctions are critical to transparency and are clearly detailed for each product. By combining sourcing with advocacy, the database aims to remove barriers to adoption, and highlights the scientific, ethical and cost-effective benefits of next-generation affinity reagents, supporting a global move toward more reproducible and ethical research.
Built-in transparency: A colour-coded classification system
To empower informed decision-making, the database features a colour-coded classification system that highlights key product attributes such as: discovery method; production technique; and degree of ADB involvement. This system allows users to quickly assess how well each product aligns with both scientific standards and ethical best practices.
The classification framework includes seven categories, developed through extensive consultation with stakeholders across academia, industry, NGOs, policy, and the European Commission’s knowledge services. Categories A and B reflect full alignment with the EURL ECVAM recommendations, avoiding any use of active animal immunisation: — Category A includes products that are entirely non-animal-derived. These may originate from naïve human B-cell libraries, synthetic or consensus frameworks, or in silico-designed scaffolds. — Category B covers reagents from pre-2020 historical immunisation or from naïve B-cell libraries from non-immunised animals. This category also includes sequence-defined hybridomas that are no longer maintained in animals, consistent with EURL ECVAM guidance. — Categories C to F include products developed through active animal immunisation (post-2020), or involving primary ADBs during any stage of their development or manufacture.
A seventh category is reserved for reagents with insufficient data for full classification. This structured approach ensures clarity for researchers seeking to avoid animal-derived products entirely or to understand the ethical nuances of each product’s origin.
Importantly, antibodies produced by using traditional methods, such as polyclonal and hybridoma-derived monoclonal antibodies (including those made by using the ascites method), are excluded from the database due to significant ethical and reproducibility concerns. 12 It is also worth noting that some suppliers may market recombinant antibodies as ‘animal-free’ despite them having originated from immunised animals or hybridomas — an ambiguity that the database aims to clarify.
Manufacture of non-animal-derived antibodies
Non-animal-derived recombinant antibodies (also known as rAbs) are monoclonal, sequence-defined antibodies produced in vitro by using recombinant DNA technology. They can be generated without the need to immunise animals and are suitable replacements for animal-derived antibodies in the vast majority of research applications. However, it should be noted that recombinant antibodies are also sourced from antibody gene sequences derived from immunised animals (see section on Libraries derived from immunised animals, below).
Library display technologies
Non-animal-derived recombinant antibodies are generated from diverse antibody libraries using in vitro selection techniques. These libraries can be displayed on various platforms, such as phage, bacteria, yeast, mammalian cells or ribosomes, with phage display being one of the most established and widely used methods.
In phage display, antibody gene segments are fused to a phage coat protein gene and inserted into bacteria. These bacteria produce phages displaying unique antibody fragments on their surfaces. Through a selection process called panning, phages that bind to a specific immobilised antigen are captured, while non-binders are washed away. The bound phages are eluted and amplified over several rounds to enrich for high-affinity binders. Final candidates are sequenced, cloned and expressed as purified antibodies. In a process termed ‘affinity maturation’, the affinity of the candidate antibody may be further improved by introducing random mutations and performing further rounds of selection.
This method supports the production of multiple antibody formats, such as single-chain variable fragments (scFv), Fab fragments or full-length IgGs, and can theoretically generate libraries with up to 1011 distinct clones 13 — equivalent to a lifetime of antibodies from immunised animals.
The effectiveness of these display platforms depends on the quality and diversity of the antibody libraries they use, which can originate from several different sources, each offering different levels of diversity, control and ethical impact.
Sources of antibody sequences
Naïve, non-immunised B-cell libraries
These are derived from B-cells collected from non-immunised human or animal donors. Antibody gene segments are amplified and cloned into phagemid display vectors to create recombinant libraries. Naïve libraries reflect the natural immune repertoire, with each donor contributing millions of unique gene sequences due to the hypervariable regions of antibodies. Pooling donors enhances diversity and increases the chances of finding high-affinity binders. While these libraries offer substantial diversity, not all antibodies are stable, so synthetic approaches may be preferred in some cases.
Framework/synthetic-based libraries
Framework sequences, derived from conserved regions of antibody genes, act as stable backbones for constructing synthetic or naïve antibody libraries. To achieve the necessary diversity and antigen specificity, complementarity-determining regions (CDRs) — the portions of the antibody responsible for recognising and binding to antigens — are designed and inserted into the variable domains of the antibody’s heavy (VH) and light (VL) chains. These regions are critical for the antibody’s target-binding function.13,14 Two main types of framework sequences are used: — Synthetic frameworks: These are computationally engineered and optimised for properties such as solubility, stability and efficient expression in host systems (e.g. E. coli, yeast or mammalian cells). They are fully animal-free and allow the precise tailoring of antibody characteristics, as well as the elimination of problematic sequences from the outset. — Natural frameworks: Historical (legacy) antibody sequences generated by immunisation, copied from existing databases, published research or expired patents. These sequences originate from a range of species, including human, mouse, rabbit, camel and shark. Their use does not require new animal immunisation or tissue harvesting. In some cases, they are designed as consensus sequences compiled from multiple natural frameworks to enhance performance and diversity.
Libraries derived from immunised animals
Some recombinant antibodies are derived from new immunisations, where animal use is not historical — typically mice, rabbits or camelids. In these cases, B-cells are harvested from organs such as the spleen or lymph nodes, and the extracted mRNA is used to construct antibody libraries. Although this method yields antigen-specific antibodies, it conflicts with the EURL ECVAM recommendation issued in May 2020, which discourages further animal immunisation where scientifically robust, non-animal alternatives exist.
To align with this guidance and evolving ethical and regulatory expectations, researchers should prioritise naïve or synthetic/framework-based libraries, which offer a level of antibody diversity and specificity comparable to that obtained through immunisation-based methods, without relying on either current or historical animal use.
The use of animal-derived biomaterials
Animal-derived biomaterials (ADBs) are still commonly used in the production or formulation of antibodies, raising concerns about reproducibility, contamination and ethics. However, a growing number of alternatives are now available, many of which are catalogued in resources like the Animal-free Biomaterial (AFB) Providers database. 15 These alternatives support greater standardisation, batch consistency and ethical research. ADBs on the Recombinant Antibodies & Mimetics Database have been placed into two categories: primary (directly sourced from animal tissues or fluids) and secondary (derived indirectly from animal components or systems).
Primary animal-derived biomaterials
Primary ADBs are obtained directly from animal tissues, organs or fluids. Common examples include:
Fetal bovine/calf serum (FBS/FCS), newborn calf serum, serum protein or tissue hydrolysates
Widely used in cell culture media, FBS is collected from bovine fetuses during slaughter, raising ethical concerns and contributing to batch variability and contamination risks.
16
Serum-free media formulations are increasingly available, supported by detailed protocols for transitioning to humanised or fully defined systems (see the Fetal Calf Serum-Free Database).
17
Common alternatives include: — Human serum, which offers clinical relevance and eliminates interspecies issues, though supply limitations and batch variability remain. — Human platelet lysate, a serum-free supplement derived from blood donations, useful especially for clinical-grade applications.
18
— Plant- and microbial-derived hydrolysates (e.g. from soy, wheat, pea or yeast), which are being actively tested to identify optimal combinations for specific applications.19,20 Artificial intelligence (AI) tools in development are expected to support this by modelling cell growth across different conditions and predicting the best-performing formulations.
21
Bovine serum albumin (BSA)
Commonly used as a blocking agent to prevent non-specific binding and as a protein stabiliser. BSA is a source of inconsistency due to variations in herds, processing methods, and contamination with residual animal proteins such as IgG or proteases. These issues contribute to assay failures and irreproducible results. 22 An alternative is recombinant, animal-free albumin, which eliminates these variables and offers more consistent performance, while also reducing regulatory complications.
Trypsin
Traditionally derived from porcine pancreas, it is a proteolytic enzyme used to detach adherent cells in culture. It carries risks of contamination and porcine virus transmission. Animal-free alternatives include TrypZean, a recombinant version expressed in corn that eliminates these concerns.23,24
Gelatin and extracellular matrix proteins (e.g. collagen, fibronectin, laminin)
Extracted from animal connective tissues, these components are sometimes used as stabilisers or surface coatings. Alternatives include: plant-based or recombinant collagens (e.g. expressed in tobacco or yeast); synthetic peptide hydrogels; fibrin from human plasma; alginate from brown algae; and chitin/chitosan-based scaffolds. 25
Keyhole limpet haemocyanin (KLH)
Used to conjugate small molecules (haptens) to boost immunogenicity in antibody production, KLH is derived from marine molluscs. Synthetic carriers, such as polymers based on polyethylene glycol (PEG), are being explored as ethical and scalable alternatives. 26
Secondary animal-derived biomaterials
Secondary ADBs are not directly sourced from animal tissues, but originate from animal products or byproducts. These are not categorised in the Recombinant Antibodies & Mimetics Database, which currently focuses on primary ADBs. Examples include: — Tryptone, a pancreatic digest of casein, produced using trypsin (which is usually animal-derived), and commonly used as a nutrient in bacterial culture media. — Hen egg white lysozyme (HEWL), which is employed in bacterial cell lysis to release intracellular recombinant proteins, including antibodies.
Alternatives to these ADBs include recombinant proteins, plant-based proteins and chemically defined media components.
Animal-derived cell lines
Although not considered ADBs themselves, many commonly used production cell lines — such as CHO (Chinese Hamster Ovary) cells — are of animal origin. Their use does not inherently involve ‘new’ animal use, but efforts to develop equivalent human-origin or plant and stem cell-derived expression systems are ongoing and represent an additional route toward fully animal-free antibody production.
Advantages of recombinant antibodies over those produced through traditional methods
Some advantages of recombinant antibodies over those produced through traditional methods.

Limitations and practical considerations for recombinant antibodies
While recombinant antibodies present compelling advantages over those produced through traditional animal-based methods, their adoption is not without challenges. The following limitations and practical considerations highlight areas where further progress or caution may be needed: — Intellectual property (IP): Barriers still exist around sequence and format protection among the companies that generate antibodies by display methodologies. Very few make the DNA sequences available for free, though access is improving with public libraries and development kits. — Library type constraints: Regarding libraries from immunised animals, these require a new library for each antigen, which is time-consuming and incompatible with non-immunisation policies. — Fragment conversion: Therapeutic applications may require conversion of antibody fragments to full-length IgGs or humanisation to reduce immunogenicity.
While recombinant antibodies offer many benefits, they are not the only animal-free affinity reagents available. Another class of alternatives, antibody mimetics, offers unique advantages in both research and therapeutic applications.
Antibody mimetics: Engineered, animal-free binding molecules
Antibody mimetics are molecules that mimic the antigen-binding function of antibodies, but are not structurally related to immunoglobulins. They are typically smaller, structurally simpler, and more stable under a range of conditions compared to antibodies produced by using traditional methods.
Examples of antibody mimetics include ankyrons, 28 aptamers, 29 affimers 30 and DARPins. 31 Aptamers, for instance, are short, single-stranded DNA or RNA oligonucleotides that bind their targets with high specificity and affinity. Ankyrons are engineered short repeat proteins designed for high-affinity, thermal stability and ease of manufacturing. Their manufacture does not involve live animal use.
Design of antibody mimetics
The development of a new antibody mimetic begins with the selection of an appropriate (protein or non-protein) scaffold to form a well-defined, three-dimensional framework. A suitable scaffold must maintain structural integrity and have sufficient flexibility in its primary structure to accommodate modifications such as mutations and insertions, without compromising secondary structure or overall stability.
Key advantages over full-length (IgG) antibodies
Recent advancements in computational design, library construction and selection technologies have enabled the development of antibody mimetics with strong selectivity and affinity. It is now possible to engineer antibody mimetics with binding affinities in the femtomolar range (10–15 moles per litre (mol/l) range), indicating exceptionally high affinity between two molecules rivalling or even surpassing that of full-length antibodies.
Like recombinant antibodies, antibody mimetics offer multiple scientific and technical advantages. They can, for example, be engineered to possess desirable properties such as enhanced solubility, protease resistance, and stability at high temperatures or extreme pH values. Their simpler structure facilitates efficient production in microbial expression systems, often resulting in significantly higher yields. In contrast, full-length antibodies are large and complex, making them prone to degradation under extreme temperatures and pH. Their size also limits their ability to efficiently penetrate tissues, which can hinder their effectiveness in certain applications. 28
Current use and limitations
A review by Yu et al. 32 highlights the growing use of antibody mimetics in analytical and diagnostic applications, and Chaves et al. 33 provide a detailed comparison of their characteristics relative to antibodies. 23
Antibody mimetics are not without challenges. Their development can be labour-intensive and costly, and some existing mimetic structures and sequences are protected by patents so are not readily accessible to the research community. Traditional methods of antibody production still dominate biomedical research. Nonetheless, the expanding range and functionality of these next-generation binders are steadily shifting the landscape of affinity reagents.
Future directions for antibody mimetics
Reducing the cost and complexity of early-stage design and production would help mitigate these barriers. Expanding access to antibody mimetics could significantly benefit the scientific community, particularly in non-animal-based research approaches.
The EURL ECVAM Scientific Advisory Committee (ESAC) working group briefly addressed the emergence of alternative binders entering the market, and recommended a more comprehensive review of these affinity reagents once additional performance data becomes available.
This aligns with the goals of the Recombinant Antibodies & Mimetics Database, which brings together information on the sources, characteristics and potential applications of antibody mimetics. By consolidating emerging evidence and real-world use cases, the database aims to raise awareness, support wider adoption, and encourage the collection and sharing of performance data, particularly in comparison to antibodies produced by using traditional methods.
Raising the standard: Validation, reliability, and the case for recombinant antibodies and mimetics
The importance of validation and standardisation
Recombinant antibodies and antibody mimetics offer a potentially powerful solution to address the reproducibility crisis in biomedical research. Unlike polyclonal antibodies, which are heterogeneous and bind multiple epitopes, recombinant antibodies are monoclonal and defined by their precise amino acid sequences. In contrast, antibodies produced by using traditional methods, such as monoclonal antibodies derived from hybridomas, often lack this definition; studies have shown that up to one-third contain rogue B-cell sequences that can cause cross-reactivity. 34
Because recombinant antibodies are sequence-defined, the risk of hidden or contaminating clones is eliminated. However, knowing the sequence is only the starting point. To ensure reliability, each antibody’s functional performance — including binding specificity, activity and application suitability — must be rigorously validated. Even well-engineered antibodies can fail due to off-target effects, particularly in complex biological systems. Robust, standardised validation is essential to build confidence in the use of recombinant antibodies and mimetics across research applications.
Common pitfalls of poorly validated antibodies
Poor antibody validation remains a major driver of irreproducible results in scientific research. Common issues include cross-reactivity with related proteins, inconsistent performance between batches, use in unvalidated applications, low or variable affinity, and poor specificity across species or model systems. These problems can lead to misleading data, wasted resources and even retracted publications. 8
Despite growing awareness, uptake of rigorous validation practices remains limited. Many researchers are not trained in proper validation methods, and the lack of universal standards continues to hinder widespread adoption. 35
What should antibody validation include?
Antibodies are not universally compatible across all applications or species. To ensure reliability, each must be validated for the specific model system and intended use. A robust validation strategy should assess several key factors: — Target specificity: Confirming that the antibody binds exclusively to the intended protein and not to structurally similar off-targets. — Application specificity: Demonstrating that the antibody performs reliably in the desired techniques (e.g. Western blot, immunofluorescence, IHC, ELISA, immunoprecipitation), as sample processing can alter epitope accessibility. — Reproducibility: Ensuring consistent results across different production lots and experimental conditions. — Functional activity: Evaluating binding affinity, kinetics, or neutralisation capacity to understand how well the antibody functions in context. — Purity: Verifying the absence of contaminants or degradation using SDS-PAGE, chromatography, or mass spectrometry.
Validation data should be transparently reported for each intended application, and ideally for each species or cell type in which the antibody is to be used.
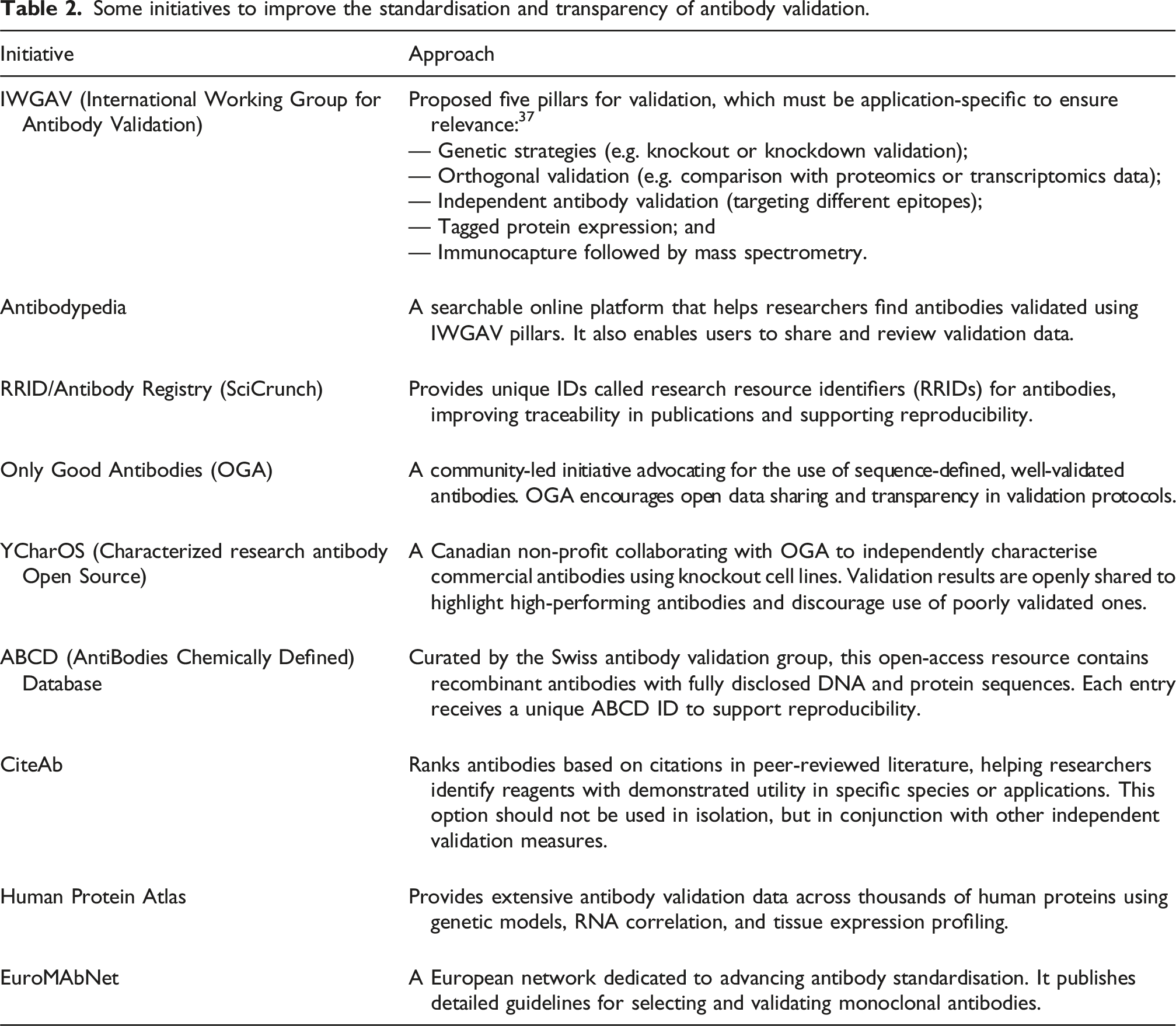
Community and global initiatives driving better antibody validation
Some initiatives to improve the standardisation and transparency of antibody validation.
Innovative education and training models
Education and training are critical to embedding ethical, reproducible practices in the next generation of scientists. One innovative example is the collaboration between ABCD Antibodies (a recombinant antibody manufacturer) and the University of Geneva, which equips students with practical skills in antibody validation while also contributing to the scientific literature. As part of the programme, students evaluate different antibodies targeting the same protein, using techniques such as Western blotting, immunofluorescence, ELISA and flow cytometry. The results are compiled into peer-reviewed Antibody Reports publications, with students credited as co-authors — providing them with both hands-on experience and academic recognition.
This initiative recently expanded through a partnership with the PETA Science Consortium, which funds the development of sequence-defined antibodies targeting extracellular matrix proteins. These antibodies are validated by a specialist laboratory under real-world experimental conditions, with the data made publicly available in Antibody Reports. This collaboration, linking funders, academic institutions and validation experts, provides a replicable model for other disciplines and research funding programmes.
Commercial validation strategies
Examples of company-led initiatives to validate antibodies.

The future of antibody science
As awareness continues to grow, the scientific community is well-positioned to benefit from the broader adoption of recombinant antibodies and antibody mimetics. These next-generation reagents offer enhanced reproducibility, consistent batch performance, and long-term renewability, while also aligning with ethical standards and increasing demands for cost-effective, reliable research tools.
However, realising their full potential requires more than individual purchasing decisions. While researchers play a vital role through informed choices, broader and more sustainable progress depends on coordinated action from funders, regulatory bodies, publishers, institutions and suppliers. For example: — Funding bodies could require grant recipients to use non-animal-derived antibodies or mimetics that are sequence-defined and free from ADBs, such as those in Category A or B, unless a compelling scientific justification is provided. — Universities and research institutions could update procurement policies to prioritise recombinant, non-animal-derived antibodies for all new purchases, permitting exceptions only with clear scientific rationale. — Scientific publishers could mandate full disclosure of antibody origin and sequence information, and promote the use of sequence-defined reagents in studies that depend on antibody-based detection. — Regulators and ethics committees could integrate antibody sourcing into review processes, encouraging or requiring the use of non-animal alternatives wherever suitable. — Suppliers could increase transparency in product labelling, clearly identifying whether antibodies are animal-derived, recombinant or mimetic, and commit to phasing out poorly characterised or unvalidated animal-derived reagents.
Embedding these expectations into policy and evaluation frameworks would send a strong institutional signal, accelerating the shift toward reproducible, ethical and future-ready antibody technologies.
The Recombinant Antibodies & Mimetics Database supports systemic change by providing a practical tool for funders, regulators, publishers and institutions, to identify sequence-defined, ethically sourced reagents, enabling alignment with best practices in reproducibility and more ethical animal-free science.
Ultimately, the goal is for recombinant antibodies and mimetics to become the default: reliable, affordable, ethically sourced and universally accessible. By supporting decision-makers across the research ecosystem, the platform helps build a more reproducible, responsible and humane future for antibody science.
Footnotes
Acknowledgements
The author thanks Dr Ross Dobie (Centre for Human Specific Research) for his valuable review and feedback on the draft. Sincere gratitude is also extended to our collaborators, whose expertise and insights were instrumental in developing the antibody categorisation system and ensuring its scientific robustness. Appreciation is further given to the steering group members for their important role in shaping the foundation of the database, and to ABCD Antibodies, Abcalis GmbH, Aptamer Group, Bio-Rad and ProImmune for their valuable input.
Funding
No funding was received by the authors in connection with this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
