Abstract
Antimony trioxide (AT) is used as a flame retardant in fabrics and plastics. Occupational exposure in miners and smelters is mainly through inhalation and dermal contact. Chronic inhalation exposure to AT particulates in B6C3F1/N mice and Wistar Han rats resulted in increased incidences and tumor multiplicities of alveolar/bronchiolar carcinomas (ABCs). In this study, we demonstrated Kras (43%) and Egfr (46%) hotspot mutations in mouse lung tumors (n = 80) and only Egfr (50%) mutations in rat lung tumors (n = 26). Interestingly, there were no differences in the incidences of these mutations in ABCs from rats and mice at exposure concentrations that did and did not exceed the pulmonary overload threshold. There was increased expression of p44/42 mitogen-activated protein kinase (MAPK) (Erk1/2) protein in ABCs harboring mutations in Kras and/or Egfr, confirming the activation of MAPK signaling. Transcriptomic analysis indicated significant alterations in MAPK signaling such as ephrin receptor signaling and signaling by Rho-family GTPases in AT-exposed ABCs. In addition, there was significant overlap between transcriptomic data from mouse ABCs due to AT exposure and human pulmonary adenocarcinoma data. Collectively, these data suggest chronic AT exposure exacerbates MAPK signaling in ABCs and, thus, may be translationally relevant to human lung cancers.
Keywords
Introduction
Antimony trioxide (AT), or Sb2O3, is used as a flame retardant in textiles, paper, and plastics. AT is also used in combination with some chlorinated or brominated flame retardants on commercial furniture, draperies, wall coverings, and carpets.1,2 Workers are exposed to antimony in the metal ore smelting and mining industries mainly through inhalation and dermal contact with dusts. 3 In the National Toxicology Program’s rodent cancer bioassay, 4 chronic inhalation exposure of Wistar Han rats and B6C3F1/N mice to AT particulates for 2 years resulted in increased incidences of alveolar/bronchiolar adenomas (ABAs) and carcinomas (ABCs) (Figure 1, Tables 1 and 2).
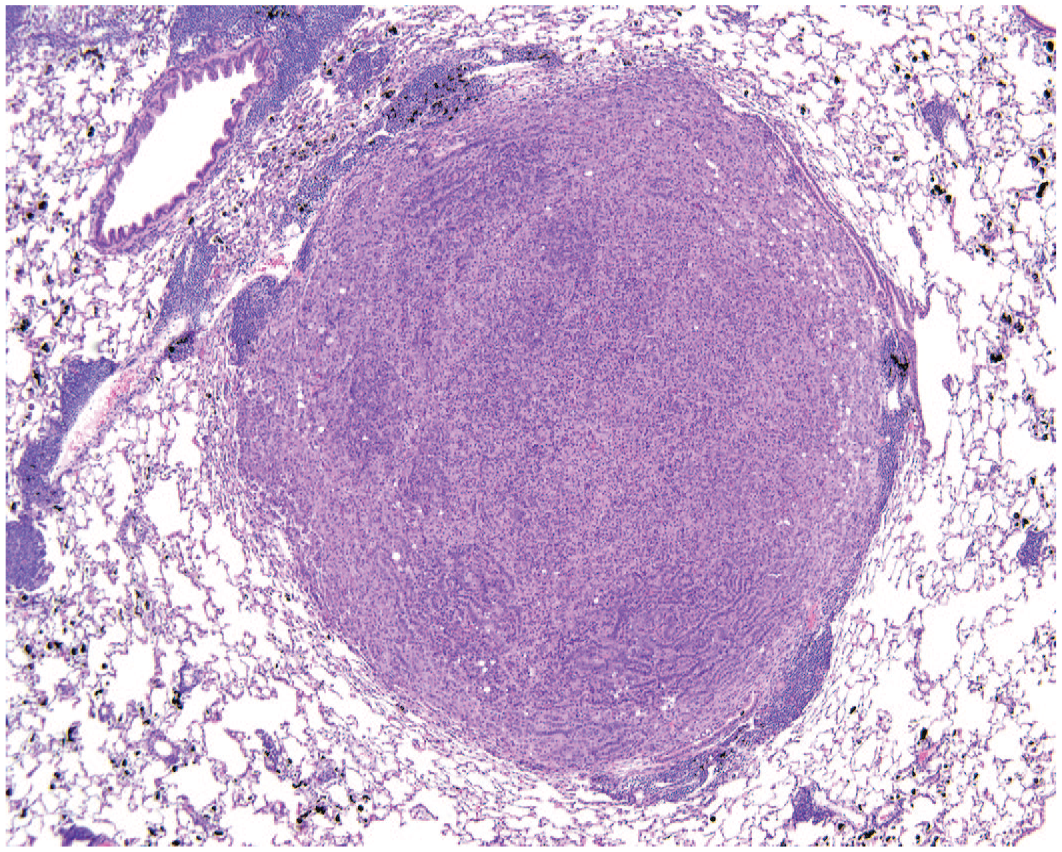
Alveolar/bronchiolar carcinoma, hematoxylin and eosin. A solid, densely cellular mass in the lung of a female B6C3F1/N mouse exposed to 30 mg of antimony trioxide/m3 by inhalation for 2 years. Please note the scattered black aggregates of antimony trioxide dust within the parenchyma. To access the whole-slide image provided with this figure, scan the QR code on cover page 3 or visit slide 1 .
Incidences of alveolar/bronchiolar tumors in Wistar Han rats after 2-year inhalation exposure to antimony trioxide (NTP TR 590).
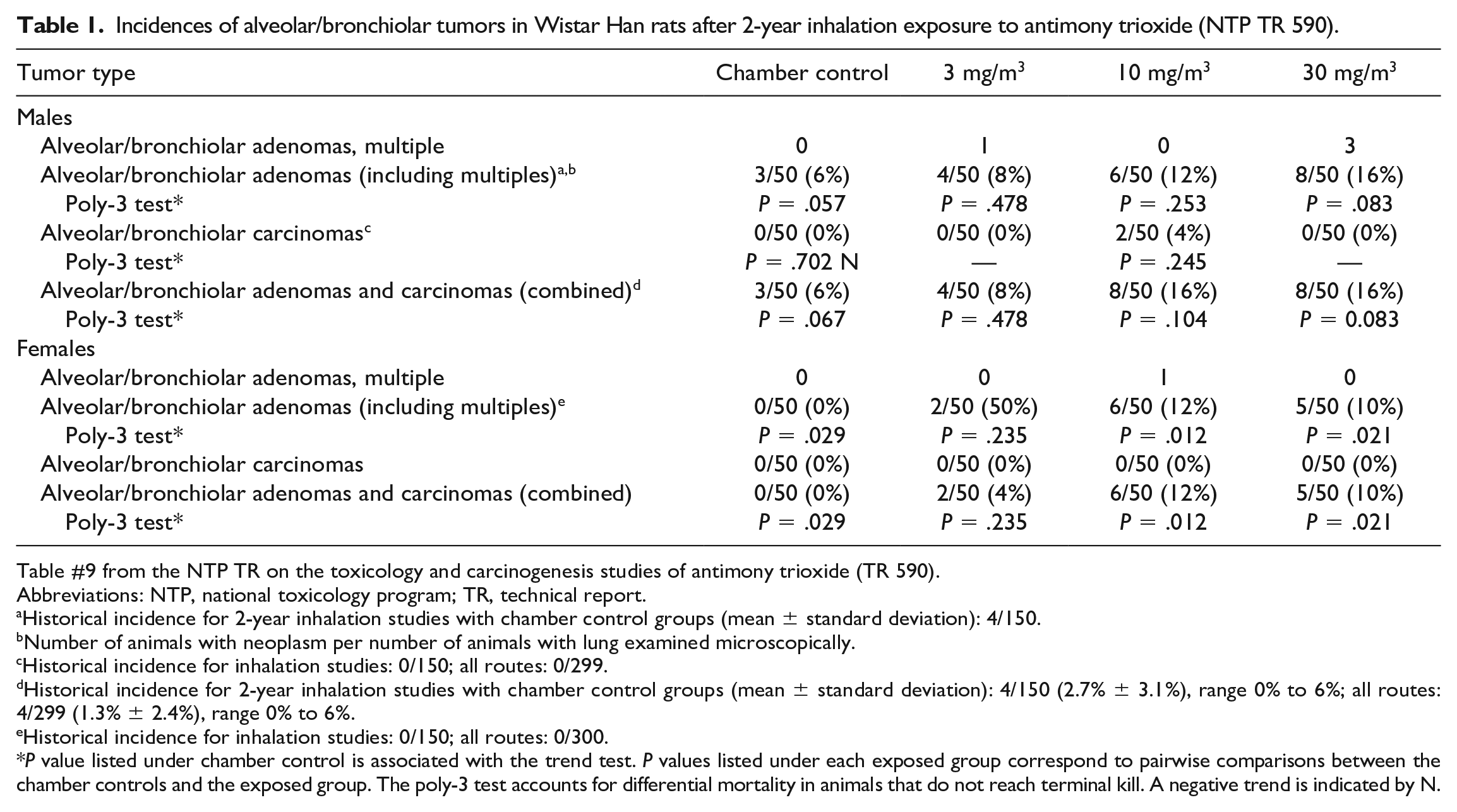
Table #9 from the NTP TR on the toxicology and carcinogenesis studies of antimony trioxide (TR 590).
Abbreviations: NTP, national toxicology program; TR, technical report.
Historical incidence for 2-year inhalation studies with chamber control groups (mean ± standard deviation): 4/150.
Number of animals with neoplasm per number of animals with lung examined microscopically.
Historical incidence for inhalation studies: 0/150; all routes: 0/299.
Historical incidence for 2-year inhalation studies with chamber control groups (mean ± standard deviation): 4/150 (2.7% ± 3.1%), range 0% to 6%; all routes: 4/299 (1.3% ± 2.4%), range 0% to 6%.
Historical incidence for inhalation studies: 0/150; all routes: 0/300.
P value listed under chamber control is associated with the trend test. P values listed under each exposed group correspond to pairwise comparisons between the chamber controls and the exposed group. The poly-3 test accounts for differential mortality in animals that do not reach terminal kill. A negative trend is indicated by N.
Incidences of alveolar/bronchiolar tumors in B6C3F1/N mice after 2-year inhalation exposure to antimony trioxide (NTP TR 590).
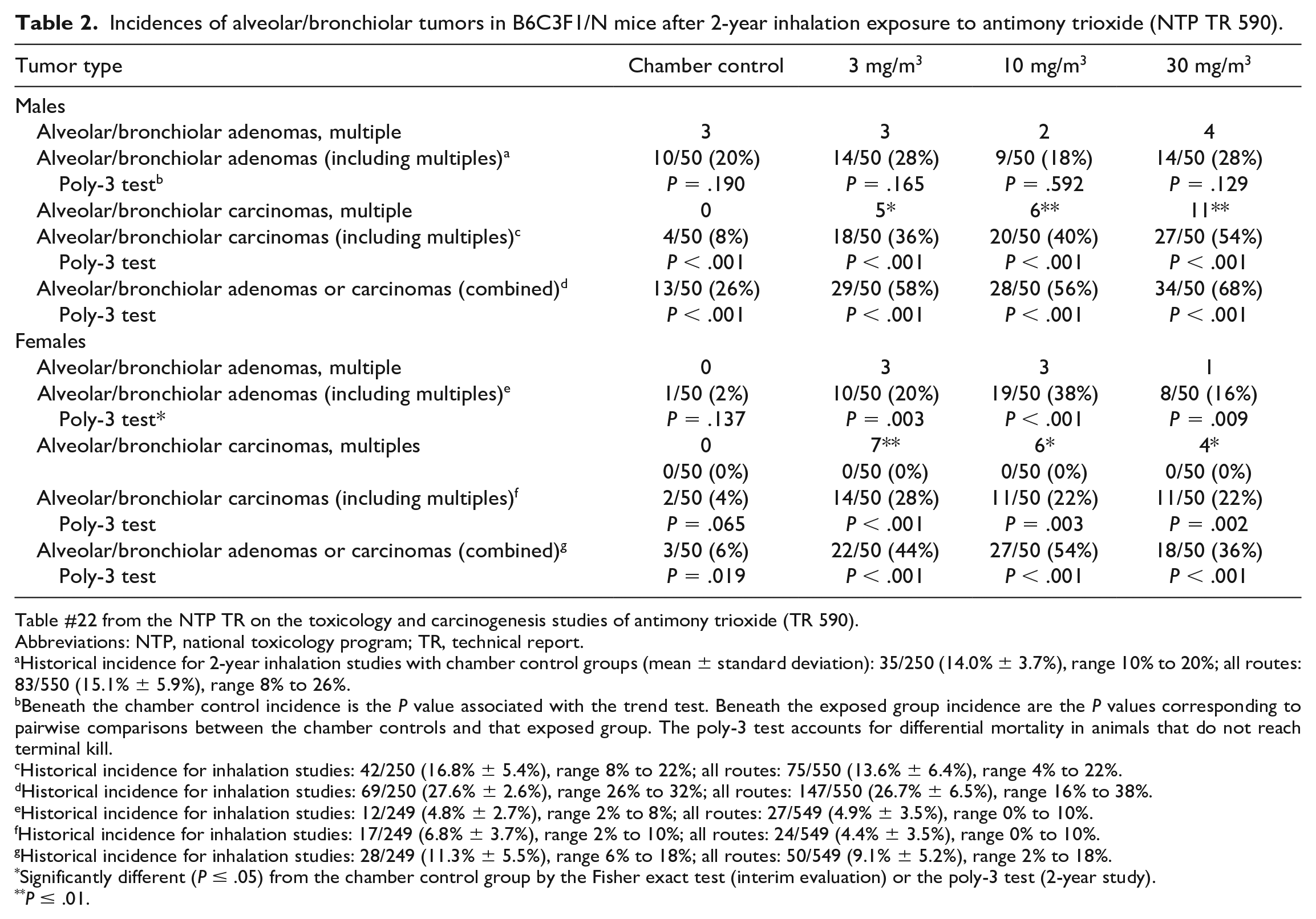
Table #22 from the NTP TR on the toxicology and carcinogenesis studies of antimony trioxide (TR 590).
Abbreviations: NTP, national toxicology program; TR, technical report.
Historical incidence for 2-year inhalation studies with chamber control groups (mean ± standard deviation): 35/250 (14.0% ± 3.7%), range 10% to 20%; all routes: 83/550 (15.1% ± 5.9%), range 8% to 26%.
Beneath the chamber control incidence is the P value associated with the trend test. Beneath the exposed group incidence are the P values corresponding to pairwise comparisons between the chamber controls and that exposed group. The poly-3 test accounts for differential mortality in animals that do not reach terminal kill.
Historical incidence for inhalation studies: 42/250 (16.8% ± 5.4%), range 8% to 22%; all routes: 75/550 (13.6% ± 6.4%), range 4% to 22%.
Historical incidence for inhalation studies: 69/250 (27.6% ± 2.6%), range 26% to 32%; all routes: 147/550 (26.7% ± 6.5%), range 16% to 38%.
Historical incidence for inhalation studies: 12/249 (4.8% ± 2.7%), range 2% to 8%; all routes: 27/549 (4.9% ± 3.5%), range 0% to 10%.
Historical incidence for inhalation studies: 17/249 (6.8% ± 3.7%), range 2% to 10%; all routes: 24/549 (4.4% ± 3.5%), range 0% to 10%.
Historical incidence for inhalation studies: 28/249 (11.3% ± 5.5%), range 6% to 18%; all routes: 50/549 (9.1% ± 5.2%), range 2% to 18%.
Significantly different (P ≤ .05) from the chamber control group by the Fisher exact test (interim evaluation) or the poly-3 test (2-year study).
P ≤ .01.
Somatic mutations in oncogenes such as Kras and Egfr (both major components of the MAP kinase signaling pathway) are common driver mutations in human lung cancers.5 -8 In rodent lung tumors, these mutations may arise either spontaneously or due to chronic chemical exposures. The mutation spectra observed in rodent tumors arising from certain chronic chemical exposures are often conserved across species and are also observed in certain human cohorts. For example, G to T transversions in codon 12 of Kras gene are commonly noted in lung cancers resulting from tobacco smoke exposure and also in rodent lung tumors resulting from chronic exposure to cobalt, which is mechanistically attributed to metal-induced oxidative stress.9,10
In human non-small-cell lung cancer (NSCLC), the incidence of Kras mutations is approximately 26%, with the majority of the point mutations seen in codon 12 followed by fewer mutations in codons 13 and 61.5,11 Point mutations in codons 12, 13, and 61 of Kras are activating mutations that result in constitutive activation of the KRAS protein, making it refractory to the inhibitory GTPase-activating proteins, which results in stimulus-independent, persistent activation of downstream effectors, in particular, the RAF-MEK-ERK cascade. Constitutive activation of this kinase cascade results in cellular proliferation and transformation.6,7
The overall incidence of Egfr mutations in human NSCLC, a subtype of adenocarcinoma, differs by region with the highest in the Asia-Pacific region (47%) and lowest in the Oceania region (12%). 12 The majority (70%) of Egfr mutations are observed within exons 19 and 21. 5 Epidermal growth factor receptor (EGFR) is a transmembrane receptor, which upon ligand binding, dimerizes and activates the cytosolic kinase domain of the receptor tyrosine kinase, resulting in activation of signaling pathways, such as the aforementioned mitogen-activated protein kinase (MAPK)/extracellular signal-regulated kinase (ERK) pathway, that support cancer development and progression. Other signaling pathways that can be activated include the phosphatidylinositol 3-kinase (PI3K) pathway, which, when activated, leads to AKT activation and apoptosis inhibition.
In the current study, we have examined the hotspot mutations in Kras and Egfr (rats and mice) and transcriptomic alterations (mice only) in ABCs resulting from chronic inhalation exposure to AT for 2 years to better understand the translational mechanistic relevance of these rodent tumors to human lung cancers.
Materials and Methods
Animal Tissue Samples
Frozen lung samples and formalin-fixed, paraffin-embedded (FFPE) lung tissues were obtained from male and female Wistar Han rats (Charles River Laboratories, Raleigh, North Carolina) and from male and female B6C3F1/N mice (Taconic Farms, Germantown, New York) used in the AT cancer bioassay. The lung tumors were diagnosed ABAs and ABCs based on microscopic morphologic criteria described in the INHAND (International Harmonization of Nomenclature and Diagnostic Criteria) document on rat and mouse respiratory system. The diagnostic terminology “alveolar/bronchiolar” is the preferred NTP terminology and is slightly different from the “bronchiolo-alveolar” terminology described in the INHAND document. All the diagnoses were confirmed by an NTP pathology working group. The ABAs and ABCs that arose either spontaneously in chamber controls or due to chronic inhalation exposure to AT had similar morphologic features with the exception of increased tumor multiplicity and the test material accumulation in the exposed lungs. Animal care and use were in accordance with the Public Health Service Policy on Humane Care and Use of Animals. Animal studies were in compliance with the IACUC (Institutional Animal Care and Use Committee) protocol approved by the Battelle Toxicology Northwest Animal Care and Use Committee and were conducted in an animal facility accredited by the Association for the Assessment and Accreditation of Laboratory Animal Care International.
Alveolar/Bronchiolar Neoplasms for Mutation Analysis
Morphologic evaluation based on the INHAND criteria was performed to diagnose and select alveolar/bronchiolar neoplasms. Alveolar/bronchiolar adenomas (n = 23) and ABCs (n = 3) from Wistar Han rats and ABCs (n = 80) from B6C3F1/N mice were used from AT-exposed groups. Rat ABAs (n = 2) and mouse ABCs (n = 9) obtained from the chamber controls were considered to have arisen spontaneously. Nontumor lung tissues from chamber control rats (n = 11) and mice (n = 10) were used as negative controls. A mutation analysis was performed on FFPE tissues. The size of the selected mouse tumors was generally greater than 5 mm in diameter. Rat tumors were smaller than 5 mm in diameter and scattered throughout the pulmonary parenchyma. The alveolar/bronchiolar tumors selected for molecular biology analysis were based on their overall size and tissue viability (minimal to no necrosis, autolysis, or hemorrhage microscopically) in order to maximize the amount and quality of DNA obtained from the FFPE sections. DNA quantity was measured on a NanoDrop spectrophotometer (Thermo Fisher Scientific, Wilmington, Delaware). DNA sample quality was estimated by 260/280-nm absorbance ratio; samples with an absorbance ratio range between 1.7 and 2.0 were used for analysis. Samples outside this DNA purity range were re-isolated from FFPE sections until a suitable purity measure was obtained.
DNA Extraction, Polymerase Chain Reaction, Autosequencing, and Mutation Analysis
The lung tumors were evaluated for hot spot mutations within specific codons or exons in Kras and Egfr genes that are relevant in human lung cancer. Ten-micron-thick FFPE sections were collected on charged glass slides, and the tumor-only tissue was grossly dissected using a sharp microtome blade and collected into screw-top tubes. In some cases (especially in rats) where the alveolar/bronchiolar tumors had a miliary distribution throughout the pulmonary parenchyma, the entire lung section was used for DNA isolation. DNA was isolated from these FFPE sections using a DNeasy Tissue Kit (Qiagen, Valencia, California) following the manufacturer’s protocols. Amplification reactions were carried out by semi-nested polymerase chain reaction (PCR) using primer sets designed for Kras (exons 1 and 2) and Egfr (exons 18 to 21). 9 Controls without template DNA were run with all sets of reactions. PCR products were purified using a QIAquick Gel Extraction Kit (Qiagen). The purified products were cycled with Terminal Ready Reaction Mix-Big Dye (PerkinElmer, Foster City, California), and the extension products were purified with the DyeEx 2.0 SpinKit (Qiagen). The lyophilized PCR products were sequenced with an automatic sequencer (PerkinElmer ABI Model 3100). 13 The resulting electropherograms were compared to identify mutations in alveolar/bronchiolar neoplasms that either arose spontaneously or were due to AT exposure using CLC Genomic Workbench (Redwood City, California).
Statistics for Mutation Analysis
A one-sided Cochran-Armitage trend test was conducted to test for significance of exposure concentration-related trends in the incidences of mutations. One-sided Fisher’s exact tests were conducted to test for significant differences in the proportions of mutations between the chamber control and various exposed groups.
RNA Isolation
Mouse ABCs arising either spontaneously (n = 6) in chamber controls or due to chronic AT exposure (n = 6) were used for transcriptomic analysis. RNA was extracted from tumor-only tissue obtained using laser-capture microdissection of the frozen mouse ABCs collected from the AT cancer bioassay. RNA from mouse nontumor lung tissue (n = 6) was used as negative controls. Invitrogen PureLink Mini kit (Invitrogen, Carlsbad, California) was used for RNA extraction following manufacturer’s protocol. On-column deoxyribonuclease (DNase) treatment was performed using the Invitrogen PureLink DNase kit (Invitrogen) to purify RNA samples. The concentration and quality of the extracted RNA were measured on a Bioanalyzer (Agilent Technologies, Santa Clara, California). Samples were aliquoted and stored at −80°C until they were analyzed. All the RNA samples used for the microarray study had an RNA integrity number value of at least 7.
Microarray Protocol
A global differential gene expression analysis was done using the Affymetrix Mouse Genome 430 2.0 GeneChip arrays (Affymetrix, Santa Clara, California). Twenty nanograms of total RNA was amplified as directed in the WT-Ovation Pico RNA Amplification System (Nugen, San Carlos, California) protocol and labeled with biotin following the Encore Biotin Module. Amplified biotin-aRNAs (4.6 μg) were fragmented and hybridized to each array for 18 hours at 45°C by rotating hybridization. Array slides were stained with streptavidin/phycoerythrin utilizing a double-antibody staining procedure and then washed for antibody amplification according to the GeneChip Hybridization, Wash, and Stain Kit user manual following protocol FS450-0004. Arrays were scanned in an Affymetrix Scanner 3000, and data were obtained using the GeneChip Command Console and Expression Console Software (AGCC, Version 3.2, and Expression Console, Version 1.2) using the MAS5 algorithm to generate .CHP files. Preliminary analyses were performed with OmicSoft Array Studio (Version 7.0) software (Qiagen, Germantown, MD).
Microarray Data Analysis
Gene expression data were normalized across all samples using the robust multiarray analysis (RMA) methodology 14 in the R statistical software environment. RMA analysis fits a linear additive model to the log-transformed perfect match values using median polish to produce the gene expression measures used in the statistical analyses. RMA-normalized data were used for statistical analysis to determine differentially expressed genes. For each probe set, a pairwise t-test was performed between groups of samples (either control vs spontaneous ABC or control vs AT-exposed ABC). In each case, the null distribution of the t-statistic was derived using 10,000 bootstrap samples obtained by resampling the residuals in the ORIOGEN (Shyamal Peddada at NIEHS, https://www.niehs.nih.gov/research/resources/software/biostatistics/oriogen/index.cfm.) software package. 15 A false discovery rate (FDR) threshold of 0.05 was used to establish statistical significance.
Partek Genomics Suite, version 7.0 (release date November 25, 2019; Partek, St. Louis, Missouri), was used to perform principal components (PC) analysis (PCA) on the complete catalog of probe sets from the normalized gene expression data and to generate heat maps to compare the control lung samples, spontaneous ABC tumor samples, and AT-exposed ABC tumor samples for differentially expressed probe sets. PCA uses linear transformation to reduce the dimension of the data from p variables to k PCs. The first three PCs capture about 50% of the variation in the data and were used to visualize the spatial relationship of the control, spontaneous ABC, and AT-exposed ABC tumor samples. For hierarchical clustering, gene expression measures were first standardized across samples in a probe-set-specific manner in which the mean expression level of a transcript is subtracted from each gene expression value and the quantity is divided by the standard deviation of values across all samples. After standardization, the probe sets (rows) and samples (columns) of the data matrix holding gene expression measures for significant genes were subjected to agglomerative hierarchical clustering, where Pearson’s dissimilarity was used as the distance measure and Ward’s method was used to evaluate the distance between clusters. 16 Microarray data files (.cel) and associated annotations have been submitted to the GEO database: accession GSE152629 (reviewer token “wbubysqchpknnsd”).
Determination of Overrepresented Canonical Pathways
Ingenuity Pathway Analysis (IPA), application building_utopia, version 47547484, was used to evaluate the most statistically significant overrepresented or enriched canonical pathways included in the Ingenuity knowledge base. The significant biological canonical pathways were derived from the entire IPA library, and P < 0.01 (Fisher’s exact test) was used to indicate statistical significance.
Comparison of AT-Exposed ABCs from Mice to Human NSCLCs
Illumina BaseSpace Correlation Engine (previously known as NextBio; Illumina, Santa Clara, California) was used to compare the transcriptomic data sets of AT-exposed mouse ABCs from this study to human NSCLCs and identify the common pathways enriched in both species. Illumina BaseSpace Correlation Engine is a curated and correlated repository of experimental data derived from an extensive set of public sources (e.g., ArrayExpress and GEO) that allows the user to compare patterns of gene expression in their experiment to the published data sets. Statistical analysis is carried out using rank-based enrichment analysis to compute pairwise correlation scores of the uploaded data set and all studies in the Illumina BaseSpace Correlation Engine. The statistical analysis method used by the Illumina BaseSpace Correlation Engine is referred to as a “Running Fischer” algorithm and is very similar to the gene set enrichment analysis. 17 A detailed explanation of the analysis protocol used by the Illumina BaseSpace Correlation Engine is described elsewhere. 18
The Illumina BaseSpace Correlation Engine software was utilized to compare the differential transcriptomic changes common to human NSCLCs (curated by the Illumina BaseSpace Correlation Engine) and the AT-exposed mouse ABCs. The top-ranked upregulated and downregulated genes, as well as functional categories (termed biogroups by Illumina BaseSpace Correlation Engine) and canonical pathways that are common to AT-exposed mouse ABCs and human NSCLC, were compared. A biogroup’s score is based on the overall statistical significance and consistency of the enrichment, or overlap, between the set of genes that make up the biogroup and each of the queried biosets. The most significant biogroup receives a score of 100. All other biogroup scores are normalized to the top-ranked biogroup. 18
Validation of Transcriptomic Data
Quantitative gene expression levels of relevant targets were confirmed using the QuantStudio 3 Real-Time PCR System (Thermo Fisher Scientific, Waltham, Massachusetts). Fold increases or decreases in gene expression were determined by quantitation of cDNA from tumor samples relative to control samples using the 2−(ΔΔCt) method.
Immunohistochemistry for p44/42 MAPK (Erk1/2)
An immunohistochemical staining for p-MAPK was performed on sections of normal age-matched control lung tissue (n = 3) and AT-exposed ABCs (n = 7) from B6C3F1/N mice using the standard avidin-biotin-peroxidase technique. Formalin-fixed, paraffin-embedded tissue sections were deparaffinized in xylene and rehydrated through graded ethanol solutions. Heat-induced epitope retrieval was then performed using a citrate buffer (pH 6.0; Biocare Medical, Concord, California) in the Decloaker pressure chamber for 15 minutes at 110°C. Sections were then immersed in 3% H2O2 for 15 minutes to block endogenous peroxidase. Nonspecific sites were blocked by incubating slides for 20 minutes with 10% normal donkey serum (Jackson ImmunoResearch, West Grove, Pennsylvania), and endogenous biotin and avidin binding sites were blocked using an avidin-biotin blocking kit (Vector Laboratories, Burlingame, California). The sections were then incubated with rabbit polyclonal p-MAPK antibody (Cell Signaling Technology, Danvers, Massachusetts) at a 1:1000 dilution for 60 minutes at room temperature. For negative control tissue sections, normal rabbit IgG (MilliporeSigma, Burlington, Massachusetts) diluted to match the protein concentration of the p-MAPK antibody was utilized. Secondary incubation was done using a biotinylated donkey antirabbit IgG antibody (Jackson ImmunoResearch) at a dilution of 1:500 for 30 minutes at room temperature. Labeling incubation was done with the Vectastain RTU Kit Label (Vector Laboratories) for 30 minutes at room temperature. The antigen-antibody complex was visualized by staining with 3-diaminobenzidine chromogen (Agilent Dako, Santa Clara, California) for 6 minutes and counterstained with modified Harris hematoxylin.
Results
Kras and Egfr Mutations in AT-Exposed Mice and Rats
The incidences of Kras and Egfr mutations in alveolar/bronchiolar tumors in AT-exposed rats were 4% (1/26) and 50% (13/26), respectively (Table 3). The single Kras mutation was a non-hot-spot mutation on codon 16 with no established functional significance. The Egfr mutations were localized within exons 18 to 21 (G to A or C to T transitions). The incidence of Egfr mutations in alveolar/bronchiolar tumors from AT-exposed male rats was 62% compared to 39% in AT-exposed female rats.
Summary of Kras and Egfr mutations in nontumor lung tissue and alveolar/bronchiolar adenomas and carcinomas from Wistar Han rats after 2-year inhalation exposure to antimony trioxide.
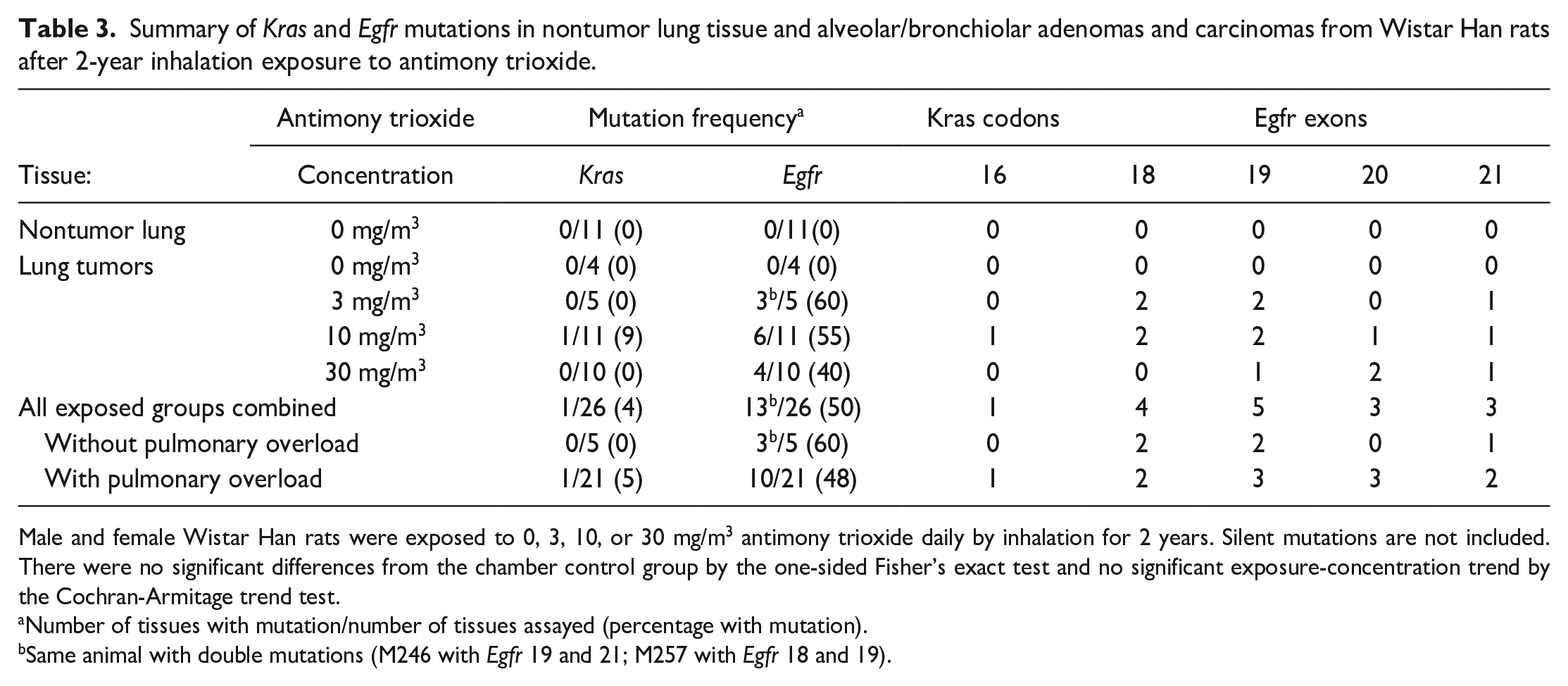
Male and female Wistar Han rats were exposed to 0, 3, 10, or 30 mg/m3 antimony trioxide daily by inhalation for 2 years. Silent mutations are not included. There were no significant differences from the chamber control group by the one-sided Fisher’s exact test and no significant exposure-concentration trend by the Cochran-Armitage trend test.
Number of tissues with mutation/number of tissues assayed (percentage with mutation).
Same animal with double mutations (M246 with Egfr 19 and 21; M257 with Egfr 18 and 19).
The incidence of Kras and Egfr mutations in alveolar/bronchiolar tumors in AT-exposed mice was 43% (34/80) and 46% (37/80), respectively (Table 4). A majority of the Kras mutations were located in codon 12 (G to A transitions). The Egfr mutations were mainly localized within exons 18 to 20 (G to A or C to T transitions). In alveolar/bronchiolar tumors from AT-exposed male mice, the incidence of Kras and Egfr mutations was 38% and 47%, respectively, compared to 49% and 46% in AT-exposed female mice.
Summary of Kras and Egfr mutations in nontumor lung tissue and alveolar/bronchiolar carcinomas from B6C3F1/N mice after 2-year inhalation exposure to antimony trioxide.
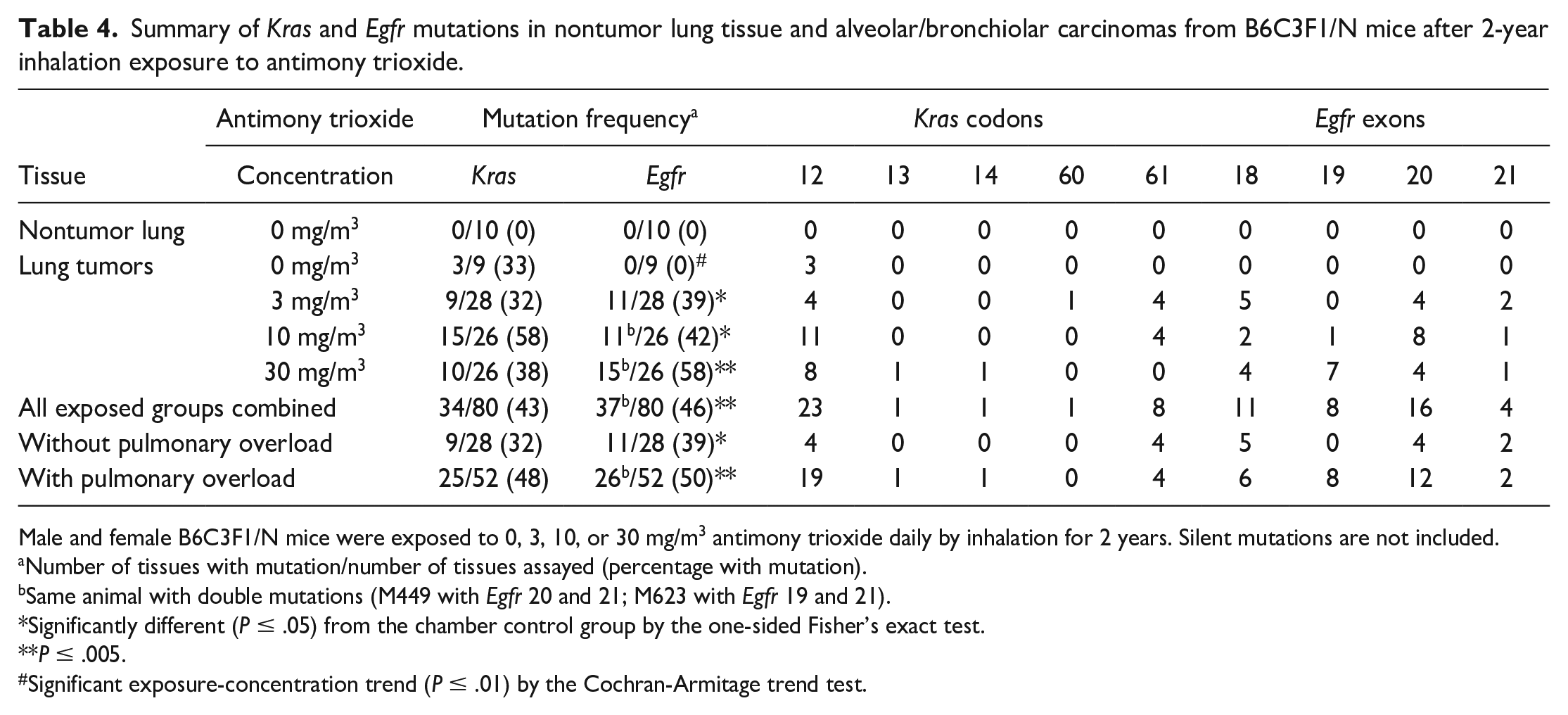
Male and female B6C3F1/N mice were exposed to 0, 3, 10, or 30 mg/m3 antimony trioxide daily by inhalation for 2 years. Silent mutations are not included.
Number of tissues with mutation/number of tissues assayed (percentage with mutation).
Same animal with double mutations (M449 with Egfr 20 and 21; M623 with Egfr 19 and 21).
Significantly different (P ≤ .05) from the chamber control group by the one-sided Fisher’s exact test.
P ≤ .005.
Significant exposure-concentration trend (P ≤ .01) by the Cochran-Armitage trend test.
Microarray Analysis
Principal component analysis was performed on all samples and all probes to characterize the variability present in the data. There was variability within the groups (control, spontaneous, AT-treated). The control group separated from the tumors, but the spontaneous and AT-treated tumors clustered together (Figure 2). Three samples (1 control [#5] and 2 spontaneous ABCs [#5 and #6]) that labeled poorly during microarray hybridization compared to the other samples were treated as outliers and removed from the analysis. Hierarchical cluster analysis (HCA) comparing standardized global gene expression profiles of all groups also showed clustering based on treatment, with some overlap between the spontaneous and AT-treated groups (Figure 3).
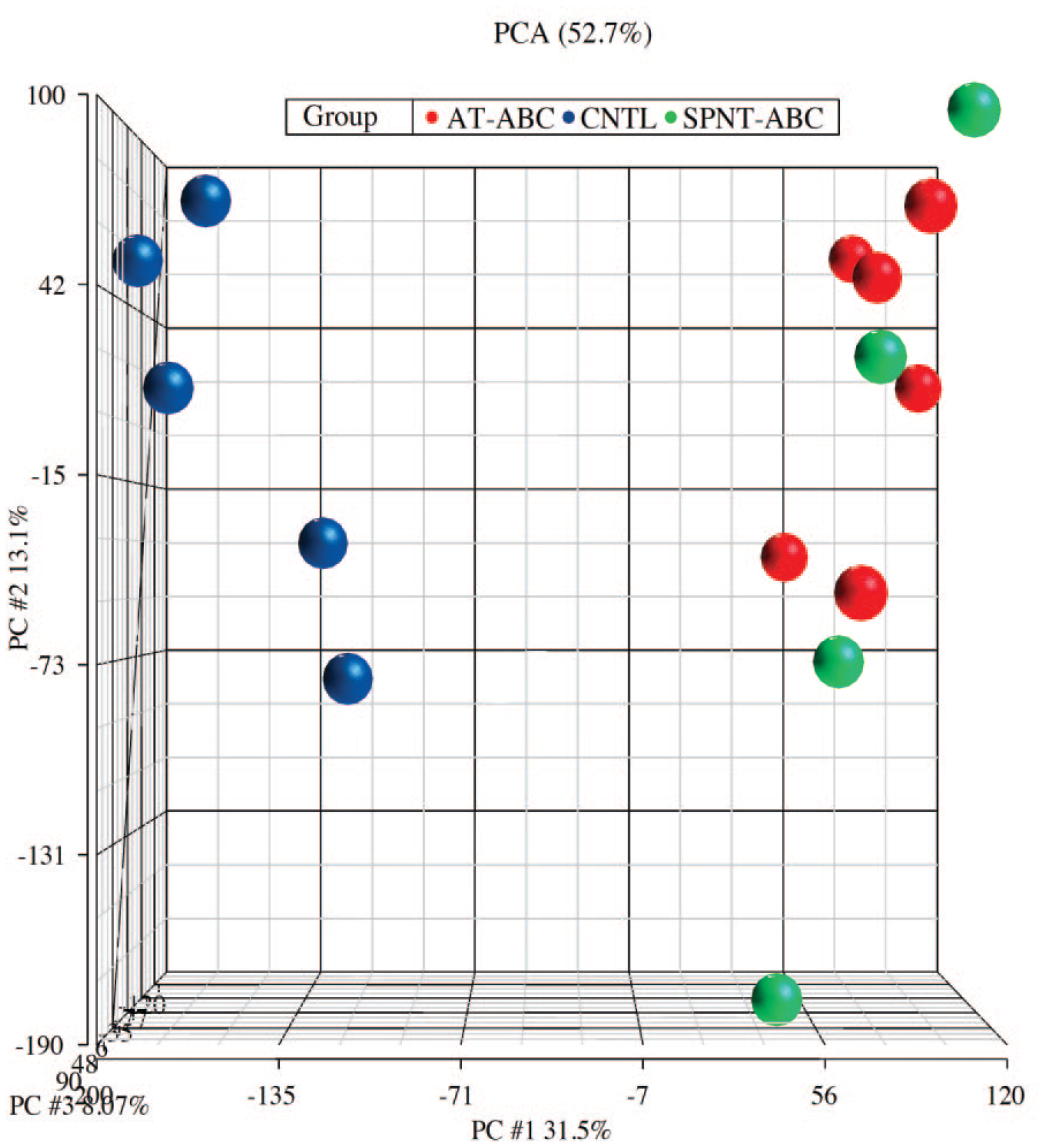
Principal component analysis (PCA) showing the distribution of various sample types (control, spontaneous, AT-treated). Three samples (1 control and 2 spontaneous) that labeled poorly compared to the other samples were treated as outliers and removed from the analysis. ABC, alveolar/bronchiolar carcinomas (ABCs); AT, antimony trioxide; CNTL, control; PC, principal components; PCA, principal component analysis; SPNT, spontaneous ABCs.
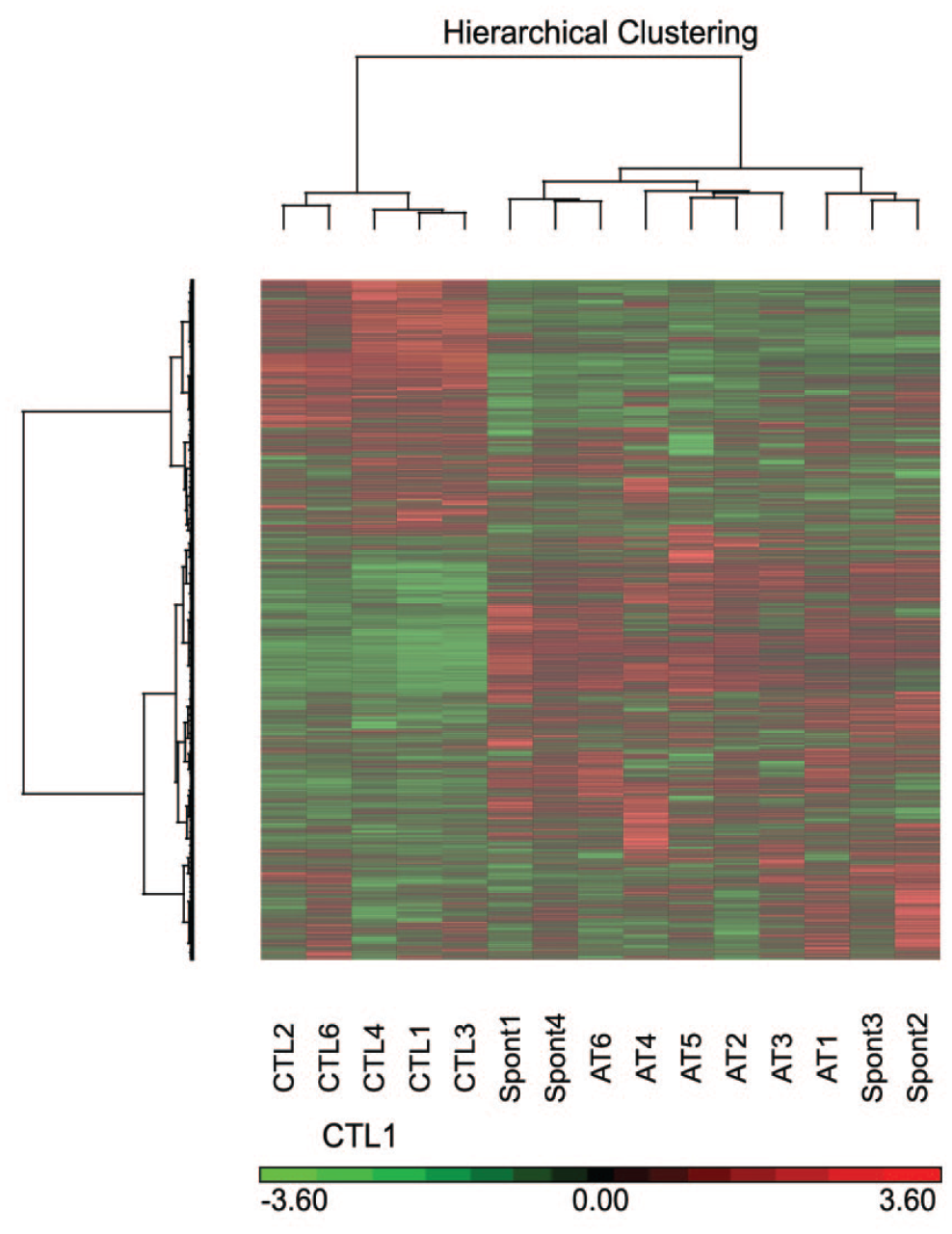
Hierarchical cluster analysis (HCA) comparing standardized global gene expression profiles (all 45,101 probe sets on the array) of normal lungs from control group, alveolar/bronchiolar carcinomas from the spontaneous group, and alveolar/bronchiolar carcinomas (ABCs) from the AT-treated group, in B6C3F1/N mice. AT, Antimony trioxide induced ABCs; SPNT, Spontaneous ABCs; CTL, Controls.
A total of 9,941 transcripts were differentially expressed in spontaneous tumors compared to control samples while 12,441 transcripts were altered in AT-exposed tumors versus control (FDR < 5%). Of this number, 7,716 transcripts were altered in both spontaneous tumors and AT-exposed tumors compared to control. In addition, 2,225 transcripts were unique to spontaneous tumors while 4,725 transcripts were unique to AT-exposed tumors (Figure 4).
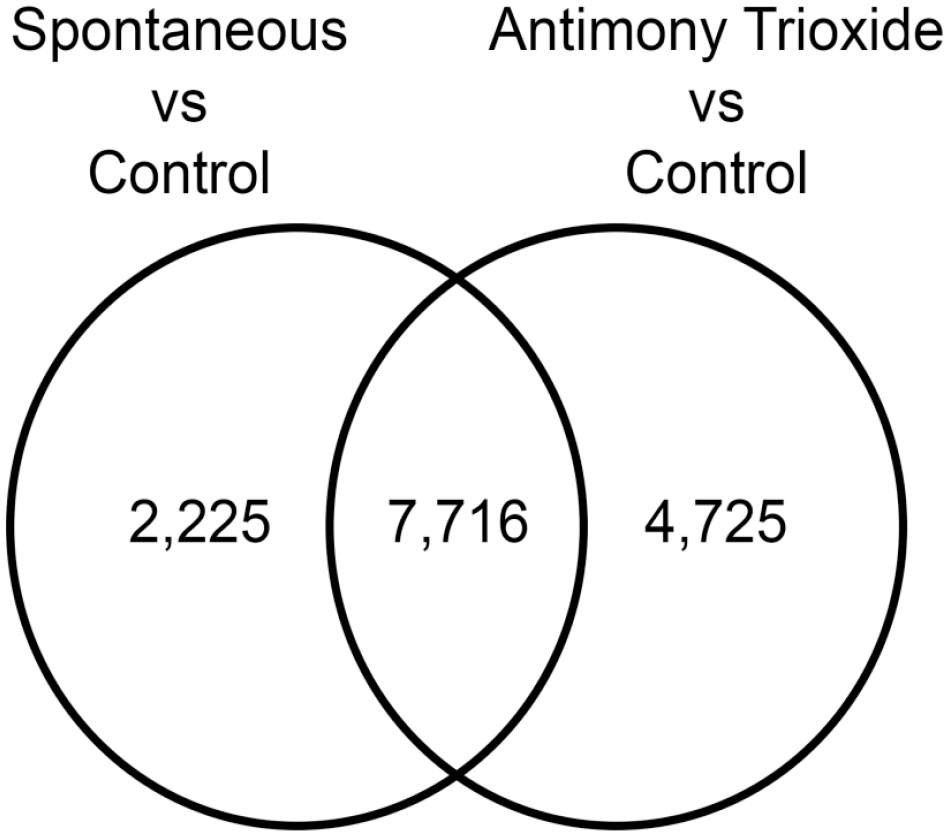
Venn diagram showing transcripts differentially expressed in spontaneous alveolar/bronchiolar carcinomas (ABCs) compared to control samples and antimony trioxide–induced ABCs compared to control samples and the corresponding significant overlap of differential expressed transcripts between the two tumor types.
Validation of Microarray Analysis (Real-Time PCR)
A subset of genes identified by microarray as dysregulated in AT-treated lung tumors were examined by TaqMan qPCR. In general, the expression of the genes examined by real-time PCR correlated well with results from microarray analysis.
Significantly Overrepresented Canonical Pathways in Mice AT-Exposed ABCs
Canonical pathways significantly altered in mouse AT-ABCs compared to controls (but not in spontaneous ABCs vs controls) as determined by Ingenuity pathway analyses of transcripts unique to AT-exposed ABCs are shown in Table 5. The ratio of number of genes within the differentially expressed gene list compared to the cataloged genes in that particular canonical pathway within the IPA database are shown, as well as −log(P value) as determined by Fisher’s exact test. Some relevant cancer pathways include cell cycle control of chromosomal replication (−log(P value) = 3.52), Ephrin receptor signaling (2.86), phospholipase C (PLC) signaling (2.79), signaling by Rho-family GTPases (2.58), paxillin signaling (2.27), and integrin-linked kinase (ILK) signaling (2.06).
Most significantly altered canonical molecular pathways in alveolar/bronchiolar carcinomas (ABCs) due to chronic antimony trioxide exposure in B6C3F1/N mice as determined by Ingenuity Pathway Analysis (IPA).
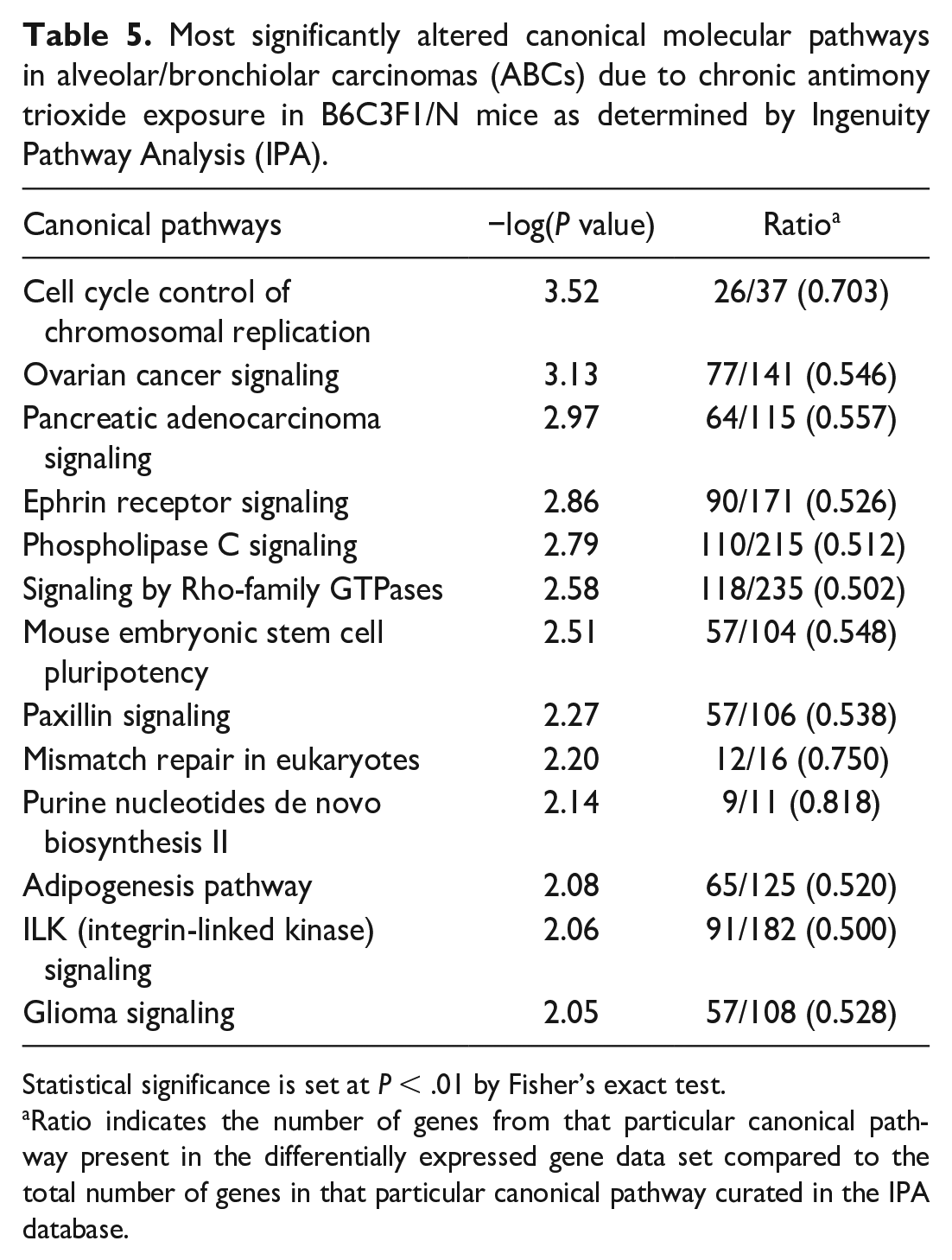
Statistical significance is set at P < .01 by Fisher’s exact test.
Ratio indicates the number of genes from that particular canonical pathway present in the differentially expressed gene data set compared to the total number of genes in that particular canonical pathway curated in the IPA database.
Significantly Upregulated and Downregulated Genes in AT-ABCs
Table 6 shows the top significantly (P < .001) upregulated and downregulated genes in AT-ABCs compared to controls. These genes were not significantly differentially expressed in spontaneous ABCs compared to controls. Top significantly upregulated genes include Dlk1 (fold change = +209.86), Tmem27 (+203.92), Rian (+66.39), Ctse (+44.96), Npy2r (+36.51), and Mycn (+12.83). Top significantly downregulated genes include Cyp2b6 (−112.085), Cbr2 (−23.29), Scgb1a1 (−21.67), Kcnk2 (−16.60), and Ripply3 (−13.77). Also, Ereg (P = .0019, fold change = +13.23) and Areg (P = .0053, fold change = +5.02), although not statistically significant at P < .001, were differentially expressed only in AT-exposed ABCs vs controls.
Top significantly upregulated and downregulated genes in alveolar/bronchiolar carcinomas (ABCs) from B6C3F1/N mice exposed to antimony trioxide for 2 years as determined by Ingenuity Pathway Analysis (IPA).
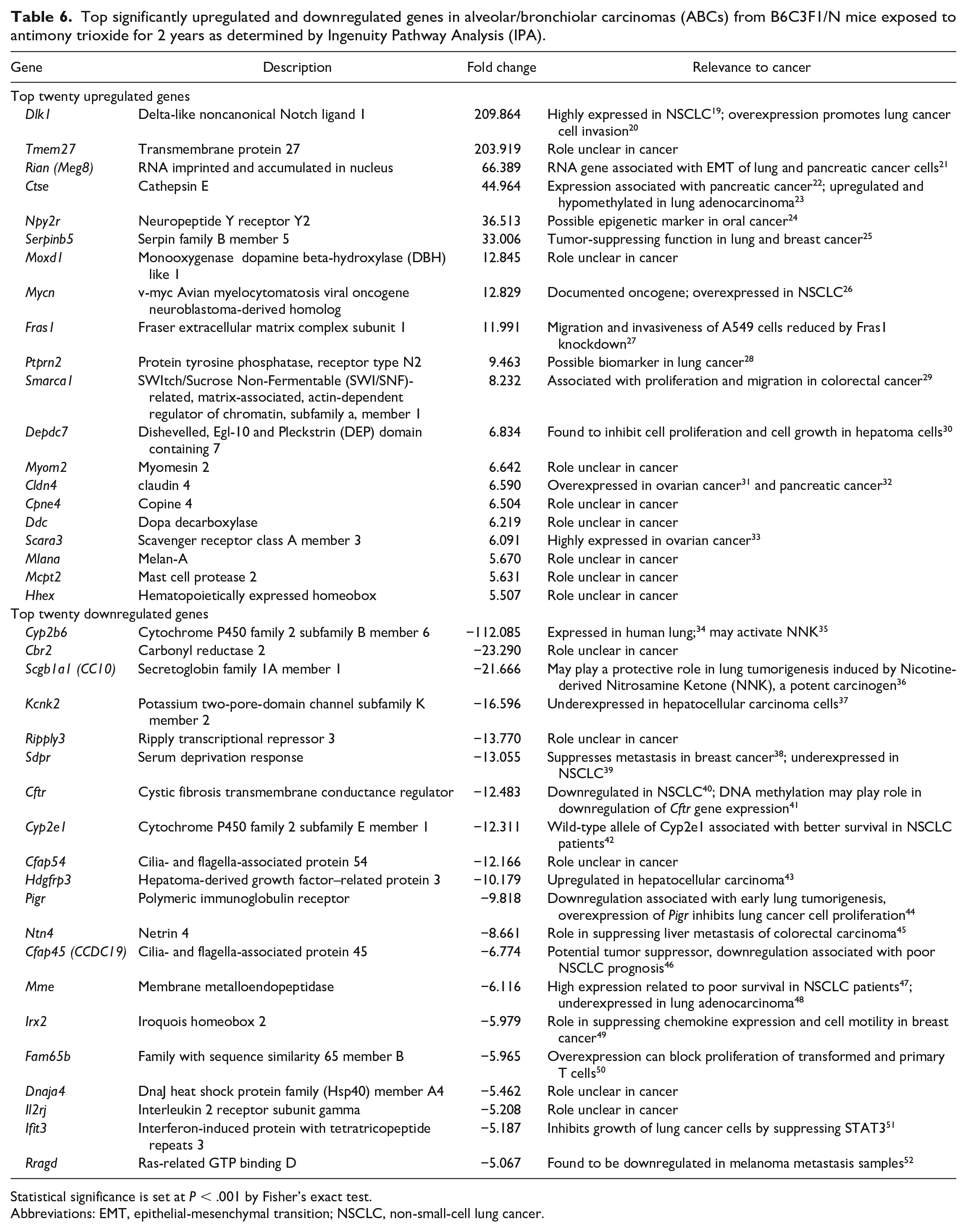
Statistical significance is set at P < .001 by Fisher’s exact test.
Abbreviations: EMT, epithelial-mesenchymal transition; NSCLC, non-small-cell lung cancer.
Genes and Pathways in Mouse AT-Exposed ABCs Similar to Those in Human NSCLC (Illumina BaseSpace Correlation Engine)
Using Illumina BaseSpace Correlation Engine, the differentially expressed transcripts data set of mouse AT-ABCs was compared to the most similar human NSCLC data set (from the Illumina BaseSpace Correlation Engine curated data sets as well as the National Center of Biotechnology Information’s GEO database). The human NSCLC data set (GSE27262) that was most concordant with the mouse AT-exposed ABC tumor data set was from stage-I lung adenocarcinoma from patients compared to adjacent nontumor tissue using Affymetrix Human Genome U133 Plus 2.0 Array.
Comparisons of the 8,067 differentially expressed transcripts in AT-ABCs in mice to the 12,667 differentially expressed human NSCLC genes resulted in an overlap of 3,953 genes (P = 1.0E-48), with 1,735 upregulated genes (P = 3.9E-23) and 1,181 downregulated genes (P = 1.6E-78) (Figure 5). Top concordant upregulated genes in AT-ABCs in mice and human NSCLC include Dlk1, Tmem27, Bpifa1 and Ros1, and the top concordant downregulated genes include Ogn, Gpm6a, Mfap4, and Scn7a (Table 7). Top concordant canonical pathways in mouse AT-ABCs in mice and human NSCLC include Nop56p-associated pre-rRNA complex, 60S ribosomal subunit (cytoplasmic), 55S ribosome (mitochondrial), 40S ribosomal subunit (cytoplasmic), and oxidative phosphorylation (Table 8).
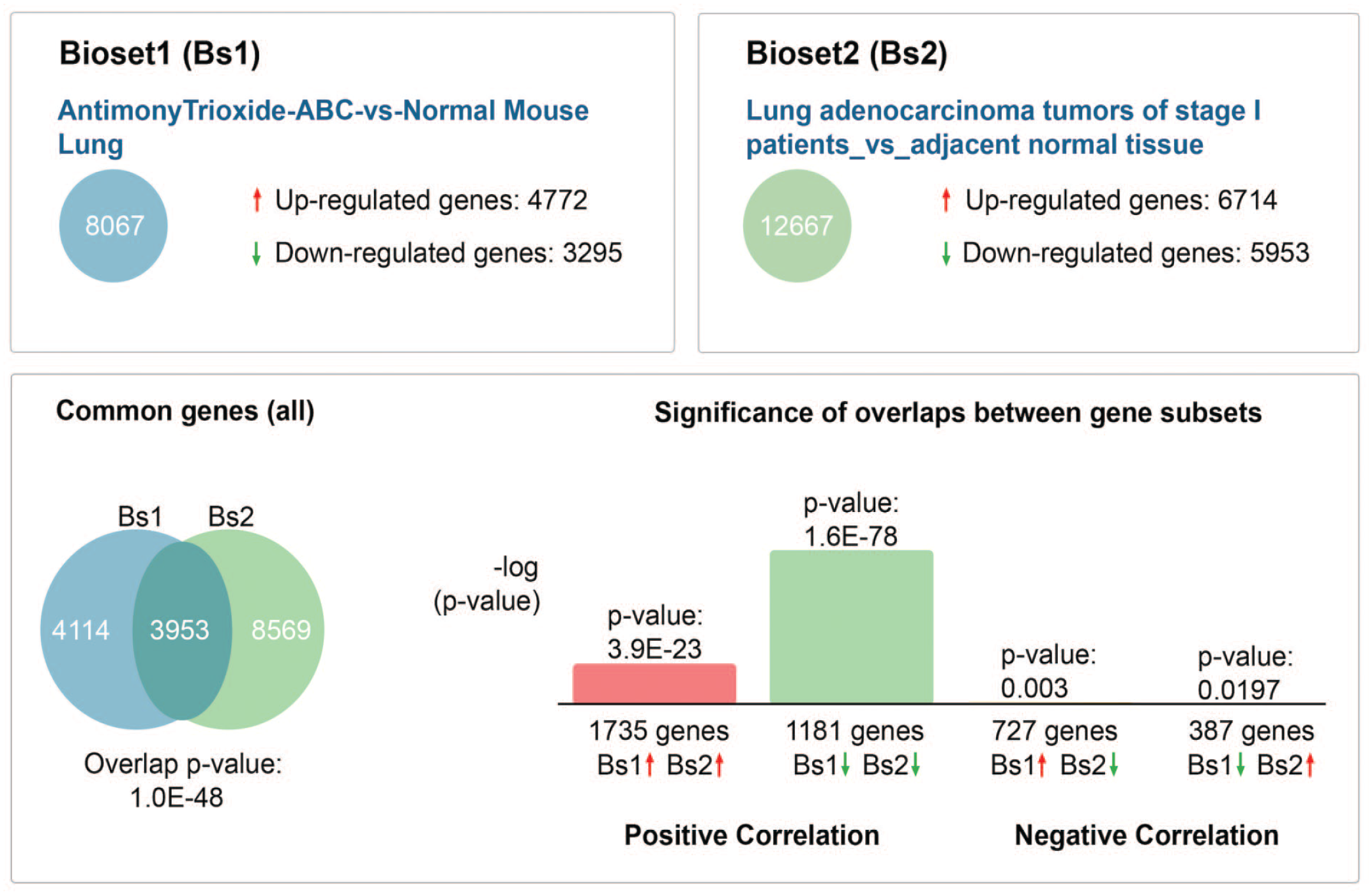
Illumina BaseSpace Correlation Engine was used to compare the differentially expressed mouse antimony trioxide (AT)-exposed ABCs tumor data set to the most similar human NSCLC data set (from the Illumina BaseSpace curated data sets as well as National Center for Biotechnology Information’s Gene Expression Omnibus (GEO) database). The human NSCLC data set (GSE27262) that closely matches the mouse AT-ABC tumor data set was from stage-I lung adenocarcinoma from patients compared to adjacent nontumor tissue using Affymetrix Human Genome U133 Plus 2.0 Array. ABC, alveolar/bronchiolar carcinoma; NSCLC, non-small-cell lung cancer.
The top concordant upregulated and downregulated genes in ABCs from B6C3F1/N mice exposed to antimony trioxide for 2 years and human non-small-cell lung cancer as determined by Illumina BaseSpace Correlation Engine.
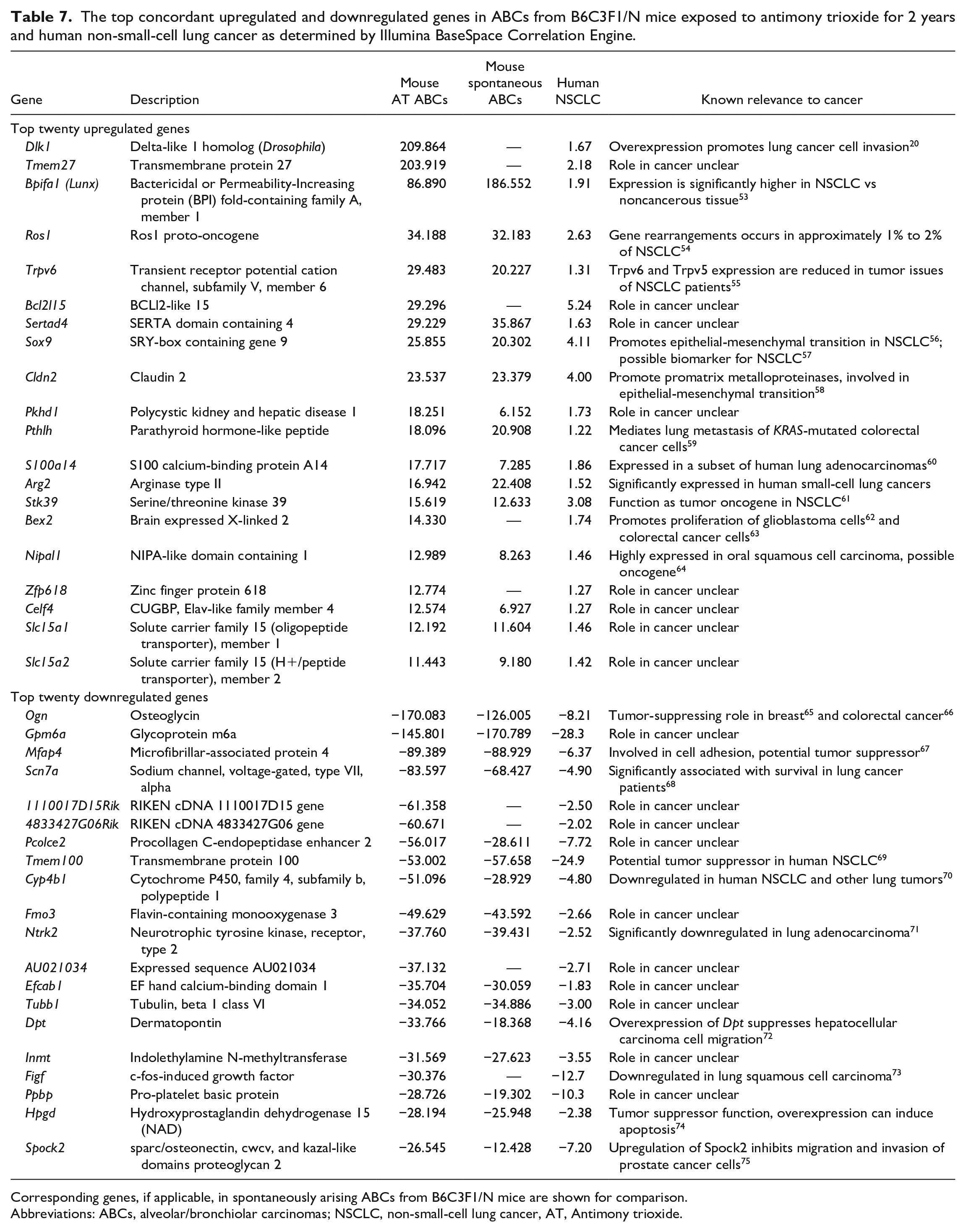
Corresponding genes, if applicable, in spontaneously arising ABCs from B6C3F1/N mice are shown for comparison.
Abbreviations: ABCs, alveolar/bronchiolar carcinomas; NSCLC, non-small-cell lung cancer, AT, Antimony trioxide.
Illumina BaseSpace Correlation Engine generated top canonical pathways common to alveolar/bronchiolar carcinomas (ABCs) from B6C3F1/N mice exposed to antimony trioxide (AT) for 2 years and human non-small-cell lung cancer as predicted by the Broad Institute’s Molecular Signatures Database (MSigDB) using the respective transcriptomic data.
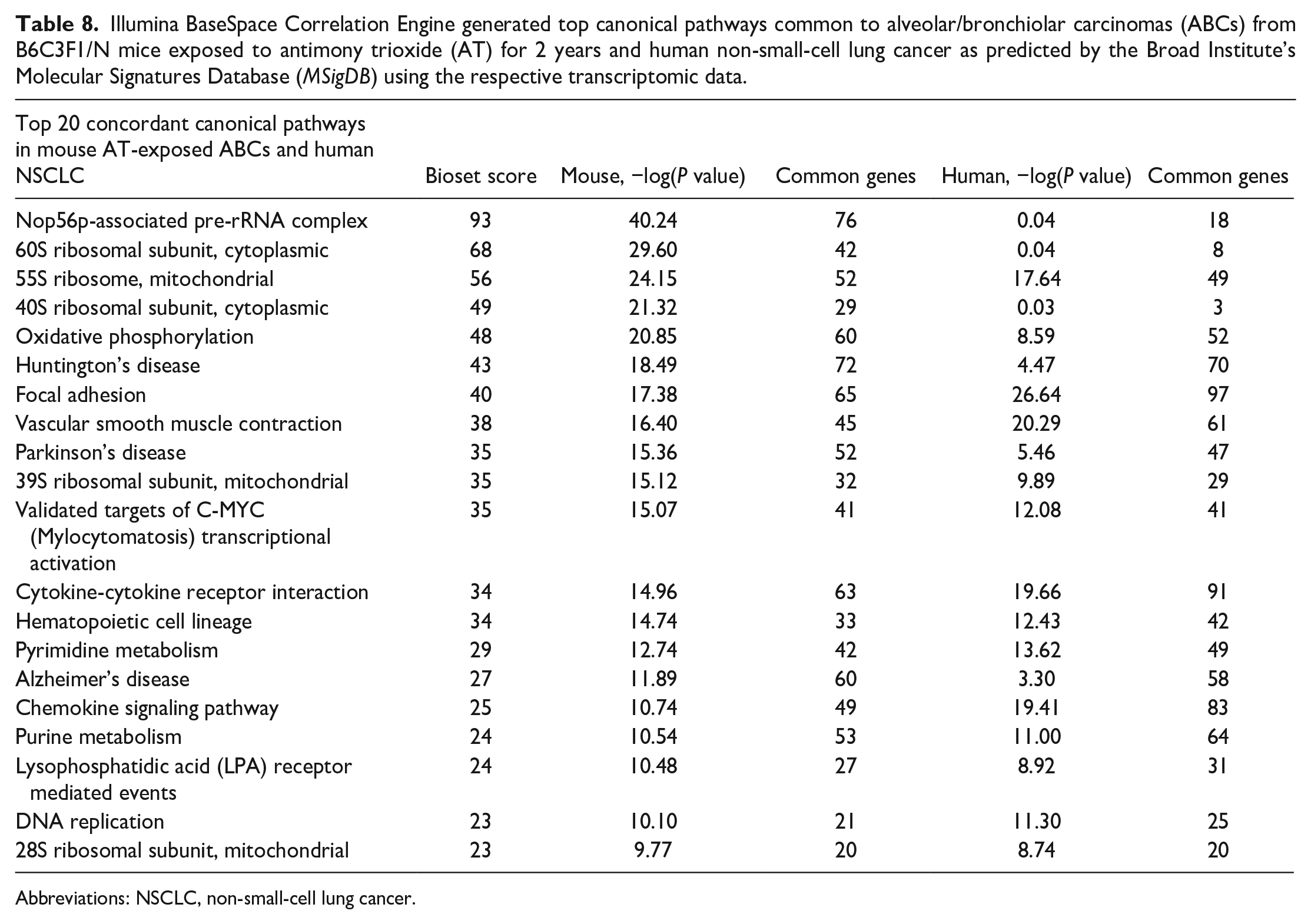
Abbreviations: NSCLC, non-small-cell lung cancer.
Immunohistochemistry for p44/42 MAPK (Erk1/2)
In normal control lungs, nuclear expression of p44/42 MAPK was present in the endothelium, but the pulmonary epithelial cells did not exhibit immunoreactivity for pErk1/2 (Figure 6A and B). In ABCs with Kras mutations, pErk1/2 expression was predominantly nuclear with some cytoplasmic expression. Expression was mainly in the neoplastic cells at the periphery of the neoplasm and in the neoplastic cells that infiltrated the surrounding parenchyma (Figure 6C and D).
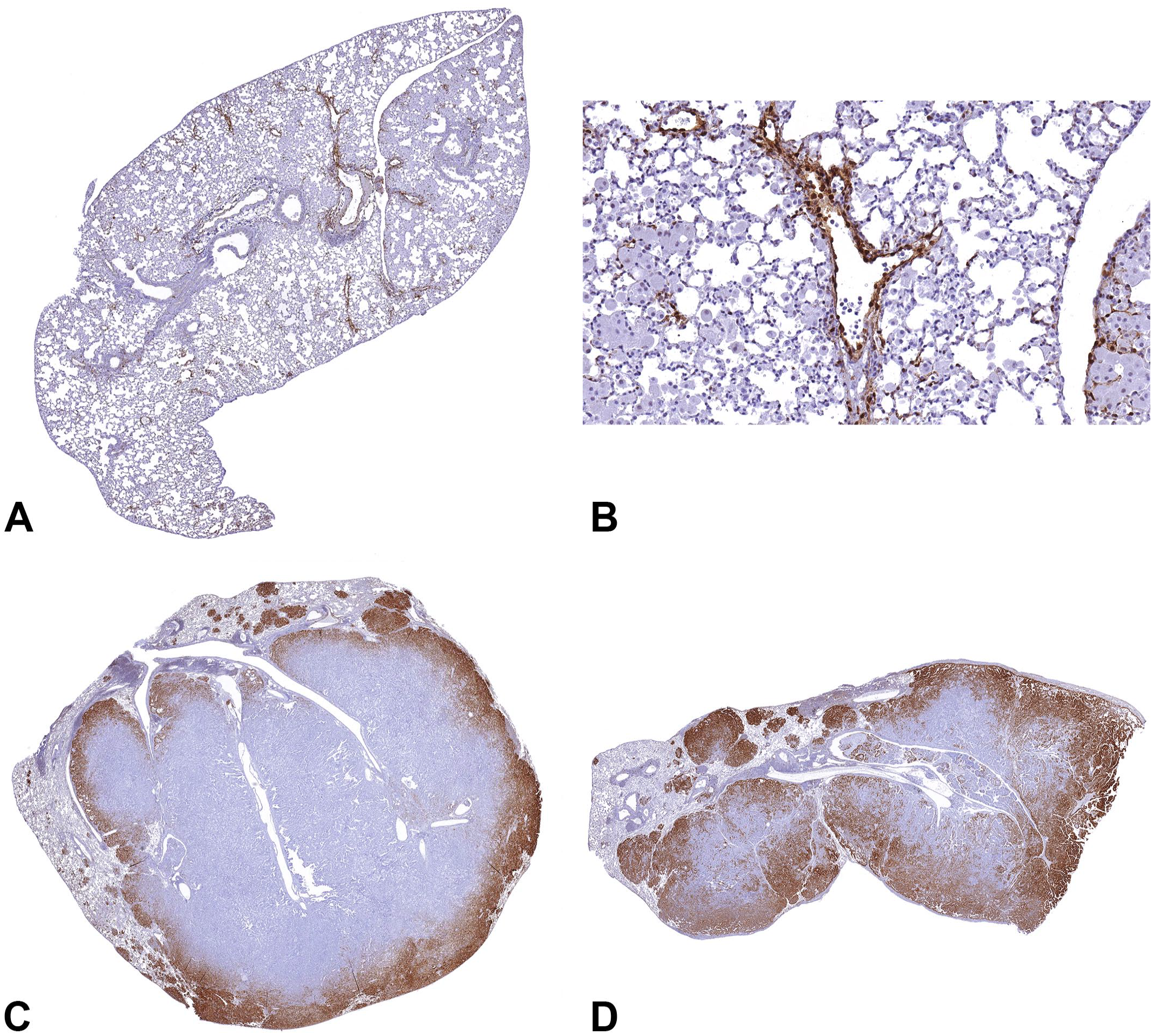
Expression of p44/42 MAPK in B6C3F1/N mouse lung. In normal mouse lungs, p44/42 MAPK expression is predominantly in the endothelium (A and B). However, the neoplastic alveolar/bronchiolar cells stain positive for p44/42 MAPK in the mouse lung tumors. The distribution pattern of p44/42 MAPK expression in neoplastic cells was predominantly nuclear and mainly in neoplastic cells at the periphery of the neoplasm and those infiltrating the surrounding parenchyma (C and D). MAPK, mitogen-activated protein kinase. To access the whole-slide images provided with this figure, scan the QR code on cover page 3 or visit slide 1 , slide 2 , slide 3 .
Discussion
Alveolar/bronchiolar tumors resulting from exposure to AT in both mice and rats harbored mutations in the Egfr gene. However, Kras mutations (mainly G to A transitions in codon 12) were observed only in alveolar/bronchiolar tumors in B6C3F1/N mice, but not in rats, exposed to AT. These Kras mutations, however, were also common in spontaneous ABCs, 13 suggesting AT exposure may have enhanced proliferation of neoplastic clones harboring the spontaneous Kras mutations in mice.
KRAS mutations in NSCLC are seen more frequently in smokers, while EGFR mutations in lung cancers are most frequently observed in never-smokers, Asians, and women. 5 In general, mutations within EGFR and KRAS are considered to be mutually exclusive events in human lung and colon cancers.76 -79 Interestingly in this study, 32% (15/47) of the ABCs from AT-exposed mice harbored both Egfr and Kras mutations. In AT-exposed rats, only one ABC harbored mutations in both Egfr and Kras (a non–hot spot mutation on codon 16 with unknown functional significance). The occurrence of concurrent mutations was also noted in ABCs in rats and mice resulting from chronic exposure to cobalt metal dust. 9 These data suggest that chronic chemical exposure may lead to multiple simultaneous mutational events leading to heterogenous clonal evolution.
Another interesting feature of this study is the occurrence of pulmonary overload in rats and mice at 10 and 30 mg/m3 but not at 3 mg/m3 (Tables 3 and 4). The concept of pulmonary overload implies that the exposure to high levels of poorly soluble particles exceeding the clearance capacity of the lung leads to a carcinogenic response unrelated to the intrinsic toxicity of the exposure, that is, lung overload due to inert particles such as talc and carbon black cause pulmonary inflammation, fibrosis, epithelial hyperplasia, and eventually lung tumors. 80 In this study, the incidences of lung tumors as well as the incidences or types of Kras or Egfr mutations in lung tumors from mice and rats were not significantly different in exposures that did and did not approach the pulmonary overload threshold. This suggests that the mutation spectra of these neoplasms were not affected by pulmonary overload and are probably related to the intrinsic toxicity of AT exposure.
Immunochemical demonstration of p44/42 MAPK expression in FFPE sections of ABCs arising from AT-exposure in mice that harbored Kras and/or Egfr mutations showed upregulation of the downstream MAPK signaling. The upregulation seems to be predominantly in the neoplastic cells at the periphery of the neoplasm or in the infiltrating neoplastic cells in the surrounding parenchyma. Moreover, the staining pattern was similar in ABC sections with a Kras mutation or both Kras and Egfr mutations. The role of ERK 1/2 or MAPK pathway activity in epithelial-mesenchymal transition (EMT) was previously demonstrated by several studies both in transformed and nontransformed epithelial cells.81 -84 The expression pattern of phosphorylated Erk1/2 in mouse ABCs harboring mutations in Kras or both Kras and Egfr is predominantly restricted to the periphery of the neoplastic mass, supporting possible involvement in EMT and tumor cell migration.
Differential transcriptomic analysis of B6C3F1/N ABCs in this study showed several significantly dysregulated canonical pathways relevant to carcinogenesis. Alveolar/bronchiolar adenomas/carcinomas that arise spontaneously or from chronic chemical exposures are morphologically indistinguishable. 13 In this study, the PCA and HCA plots showed overlap between the ABCs arising spontaneously or due to chronic AT exposure, suggesting shared molecular events leading to tumorigenesis and tumor progression between these etiologies. However, there were some significantly altered molecular changes and canonical pathways unique to AT-exposed ABCs.
Several canonical pathways were significantly altered in AT-exposed ABCs but not in spontaneous ABCs (Table 5), including cancer-relevant pathways such as ephrin receptor signaling (−log(P value) = 2.86), PLC signaling (2.79), signaling by Rho-family GTPases (2.58), paxillin signaling (2.27), and ILK signaling (2.06). Eph receptors are transmembrane proteins (TMEMs) involved in organismal growth and development, including processes such as axon guidance, neural development, and vascular development. 85 They are activated by a family of cell surface–associated ligands called ephrins. Aberrant expression of ephrin receptor signaling is found to contribute to various cancers. 85 Several Eph receptor genes were altered in ABCs of AT-exposed mice. EphA2 is overexpressed in NSCLC, 86 and its expression also correlates with brain metastasis in NSCLC patients. 87 The silencing of EphA7 is shown to inhibit proliferation and metastasis of A549 human lung cancer cells. 88 Eph receptors are also negative regulators of MAPK signaling. 89 EphrinA1 can suppress MAPK signaling and the proliferation of prostatic cancer cells. 90 It is possible that aberrant ephrin receptor signaling plays a role in alteration of MAPK signaling in AT-exposed ABCs. Phospholipase C signaling plays a role in intracellular and second messenger signaling. It consists of six classes of PLC. One of these classes, PLC-β, is found to mediate the proliferation of NSCLC and SCLC cells. 91 In the canonical pathway signaling by Rho-family GTPases, three subfamilies of small GTP-binding proteins, RHO, RAC, and CDC42, play the primary roles in mammals in the diverse biological processes regulated by the pathway, including cytoskeleton formation and transcription regulation. 92 RAC and CDC42 can also activate the PI3K/AKT signaling pathway, which is dysregulated in many cancers. Rho GTPases are aberrantly expressed in various human cancers, including breast, lung, and colon cancers. 92 RhoA is found to be overexpressed in lung cancers, 93 and RhoE is overexpressed in NSCLC. 94 Transformation of cells by oncogenic Ras also requires crosstalk between Ras and Rho-family GTPases signaling. 95 Rho signaling can block the growth inhibitory effects of ERK-MAPK signaling. In turn, ERK-MAPK signaling can block RhoA signaling to the cytoskeleton, effectively promoting motility and proliferation of transformed cells. 95 Paxillin is a focal adhesion-associated protein that regulates cell spreading and motility. 96 In NSCLC cell lines, paxillin overexpression was found to inhibit cell motility. 97 Integrin-linked kinase was shown in vitro to promote NSCLC development and metastasis through increased cell migration and invasion by inducing EMT. 98 In patients with EGFR-mutation-positive NSCLC, high levels of mRNA expression for ILK point toward shorter overall survival. 99
Several genes were significantly differentially expressed only in AT-exposed ABCs (Table 6). Dlk1, delta like noncanonical Notch ligand 1, is markedly upregulated (+209.86 fold change) in AT-ABCs. DLK1 is highly expressed in NSCLC, 19 and its overexpression can promote lung cancer cell invasion. 20 Transmembrane proteins are a group of 310 proteins that are components of cellular membranes. TMEMs are noted to function as both tumor suppressors and oncogenes in various cancers. 100 Tmem27 is markedly upregulated (+203.92) in AT-exposed ABCs. The role of TMEMs in AT-induced carcinogenesis is unclear. Rian (+66.39), RNA imprinted and accumulated in nucleus (also known as Meg8), encodes for a long noncoding RNA expressed in the nucleus. It is associated with EMT in lung and pancreatic cancer cells 21 and the tumor progression of NSCLC. 101 Ctse (+44.96), cathepsin E, is a member of the peptidase A1 proteases and is found in various cell types, such as cells of the gastrointestinal and immune systems and also in cancer cells. 22 Its expression is associated with pancreatic cancer, 22 and it is hypomethylated and upregulated in lung adenocarcinoma. 23 Mycn, v-myc avian myelocytomatosis viral oncogene neuroblastoma-derived homolog, is a well-documented oncogene and is upregulated in AT-exposed ABCs (+12.83). Mycn is overexpressed in NSCLC and was found to promote NSCLC cell proliferation. 26 Scgb1a1 (−21.67), secretoglobin family 1A member 1, is a secreted protein and member of the secretoglobin family. It is associated with anti-inflammation, and defects in Scgb1a1 are associated with asthma susceptibility. Mice with this gene knockout and exposed to the carcinogen 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone (NNK) showed markedly increased frequency of Kras mutations and alterations of MAPK/Erk1 signaling in lung tumors. 36
Commonly altered canonical pathways between mice AT-exposed ABCs and human NSCLC (as determined by Illumina BaseSpace in Table 8) include 55S ribosome, mitochondrial (-log(P value) = 24.15 in AT-ABCs and 17.64 in human NSCLC), oxidative phosphorylation (20.85 in AT-ABCs and 8.59 in human NSCLC), focal adhesion (17.38 in AT-ABCs and 26.64 in human NSCLC), and vascular smooth muscle contraction (16.40 in AT-ABCs and 20.29 in human NSCLC). Several differentially expressed genes that were statistically significant and highly altered (±fold changes) in mouse AT-ABCs were also altered in human NSCLCs (Table 7). Dlk1 and Tmem27 were already discussed previously in genes significantly differentially expressed only in AT-exposed ABCs as determined by IPA. Osteoglycin (Ogn) is part of the small leucine-rich proteoglycans in the extracellular matrix and is abundantly expressed in the mouse lung. Ogn is markedly downregulated both in AT-exposed ABCs (−170.083) and in human NSCLC (−8.21). Deficiency in OGN is related to defects in collagen fibril morphology. 102 OGN plays a tumor suppressor role in human breast cancer where it inhibits cell proliferation and invasiveness by suppressing PI3K/Akt/mTOR signaling. 65 It was shown that decreased OGN expression allowed tumor cells to induce EGFR activity to sustain proliferative signaling in colorectal cancer. 66 Microfibrillar-associated protein 4 (Mfap4) is an extracellular matrix protein the binds collagen and contains a C terminal fibrinogen-like domain and an N-terminal integrin-binding motif. Mfap4 is involved in cell adhesion and intercellular interactions and is downregulated in both murine and human lung tumors. Mfap4 is downregulated in AT-exposed ABCs (−89.389) and human NSCLC (−6.37). The mRNA level of MFAP4 is significantly downregulated in cancer tissues compared with normal tissues, indicating that MFAP4 may potentially be a tumor suppressor gene. 67
There were over twice as many unique genes (4,725 transcripts) that were significantly differentially expressed in ABCs in mice exposed to AT compared with those (2,225 transcripts) from spontaneously arising ABCs. Several noted cancer genes such as Ereg (+13.2), Areg (+5.0), and Mycn (+12.9) related to MAPK signaling and lung cancer were significantly differentially altered in the AT-exposed ABCs but not in ABCs arising spontaneously. It has been shown that EREG, a ligand of EGFR, is required for the promotion of lung tumors. 103 It has also been shown that oncogenic mutations in the Kras and Egfr genes activate the MEK/ERK pathway and induce overexpression of Ereg. Consequently, Ereg overexpression plays an essential role in oncogenic KRAS-mediated tumorigenesis. 104 It is, therefore, possible that Ereg also plays a role in the tumorigenesis of ABCs of mice exposed to AT through oncogenic activation of the MEK/ERK pathway. Another ligand of EGFR, AREG, is secreted by NSCLCs. AREG has been shown to promote autonomous growth of tumor cells and to provide resistance to apoptosis. 105 Areg is significantly overexpressed in ABCs resulting from AT exposure but not differentially expressed in spontaneous ABCs compared to normal lung. Therefore, Areg, like Ereg, may also play a role in the progression of ABCs in AT-exposed mice. Mycn is overexpressed in AT-treated ABCs. It has been shown that MYCN is overexpressed in NSCLC in humans and may also play a role in promoting the proliferation of these cancer cells. 26 Myc/Mycn is a target molecule in the MAPK signaling pathway. Ingenuity Pathway Analysis identified Mycn as a predicted upstream regulator in AT-treated and spontaneous alveolar/bronchiolar tumors.
Recently, Riva et al 106 examined the whole genomes of a subset of mouse ABCs arising spontaneously or due to chronic exposure AT obtained from this study and demonstrated that the mutation burden and mutation signatures (single-base substitutions [SBS], doublet mutations, and small insertions and deletion) are comparable. The main difference being slightly increased SBS18 signature in ABCs arising from AT exposure, indicating increased oxidative stress. Differential transcriptomic analysis of these ABCs suggests shared molecular events leading to tumorigenesis and tumor progression in both groups of ABCs, with the exception of some significantly differentially expressed genes and altered canonical pathways unique to AT-exposed ABCs. Based on the data from the study by Riva et al 106 and this study, it appears that AT-exposure potentiates some of these pathways in spontaneous ABCs as observed by increased tumor incidences and tumor multiplicities in ABCs after chronic AT-exposure.
In summary, this study shows that mice and rats exposed to AT develop alveolar/bronchiolar tumors that harbor Egfr mutations (mice and rats) and Kras mutations (mice). Hot spot mutation data and transcriptomic analysis of these lung tumors show that alterations in the MAPK signaling are important in AT-induced pulmonary carcinogenesis. Alterations in MAPK signaling are also important in human lung cancers, demonstrating the translational relevance of these tumors in human health risk.
Footnotes
Acknowledgements
We would like to dedicate this manuscript to the memory of Dr. Gordon Flake, our trusted and knowledgeable colleague who passed away after completing this technical report (NTP TR 590). We appreciate the help provided by the National Toxicology Program Tissue Archives, the National Institute of Environmental Health Sciences (NIEHS) pathology core laboratories, as well as the NIEHS Microarray core laboratory. In addition, we express our gratitude to Drs. Jian-Liang-Li and Alex Merrick for reviewing the manuscript and providing feedback.
Abbreviations
ABA, alveolar/bronchiolar adenoma; ABC, alveolar/bronchiolar carcinoma; AT, antimony trioxide; FFPE, formalin-fixed paraffin-embedded; HCA, hierarchical cluster analysis; IPA, ingenuity pathway analysis; NTP, national toxicology program; PCA, principal component analysis; RMA, robust multiarray analysis; SPNT, spontaneous.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research has been supported by the DNTP/NIEHS intramural program (ES103319-06 [2021], ES102505-14 [2021], and ZIA ES103383-01).
