Abstract
Assessment and communication of toxicology data are fundamental components of the work performed by veterinary anatomic and clinical pathologists involved in toxicology research. In recent years, there has been an evolution in the number and variety of software tools designed to facilitate the evaluation and presentation of toxicity study data. A working group of the Society of Toxicologic Pathology Scientific and Regulatory Policy Committee reviewed existing and emerging visualization technologies. This
Data visualization tools create representations (also known as graphical displays) of data that can take many forms, ranging from single-parameter graphs and scatterplots to complex multistudy heat maps. Most pathologists are familiar with graphical displays as a means of presenting study data to an audience. However, in recent years, several factors (e.g., common data formatting standards and advances in software variety and capability) have converged to facilitate the use of data visualization software for analysis and communication of study data. A working group of the Society of Toxicologic Pathology (STP) Scientific and Regulatory Policy Committee (SRPC) was formed to review some of the current data visualization tools and practices (with emphasis on their use in the context of toxicologic pathology), potential strategies for software configuration, and some of the potential pitfalls of their use.
One of the primary functions of data reporting systems used in toxicology studies is to create individual and summary tables from large anatomic and clinical pathology data sets in a format that is appropriate for submission to regulatory agencies. However, such tables tend to be cumbersome due to the frequently large volume of data, often confounding the ability to succinctly communicate study findings to other stakeholders. Therefore, in addition to tabular reports, pathologists increasingly use visualization tools to review, analyze, and present data.
Nonclinical toxicology studies (single, repeat dose, and carcinogenicity) initiating after December 18, 2017, and intended for Investigational New Drug submission must conform to the terminology of Standard for Exchange of Nonclinical Data (SEND; U.S. Food and Drug Administration 2014; Keenan and Goodman 2014). Providers of toxicology data management programs and companies supporting drug development have implemented software and process upgrades to conform to these new requirements. The standardized data structure and terminology required by SEND provides a basis for creating data warehouses without requiring extensive reconciliation or conversion of data and has spurred the growth of applications designed to mine and visualize SEND-compliant data. SEND-controlled terminology standards related to pathology end points are being developed and implemented through collaborations of the International Harmonization of Nomenclature and Diagnostic Criteria (INHAND) Global Editorial Steering Committee, SEND Controlled Terminology subteam, goRENI, and global STPs.
Software companies have taken a variety of approaches to create visualization software for nonclinical data sets. An informal survey of STP members involved in toxicologic pathology indicated that nearly all have used some form of graphical display within their organizations. Some companies are adapting current software used in chemical development or clinical trial patient data review to visualize SEND-compliant data, others are using off-the-shelf products focused specifically on nonclinical SEND data, and others are developing their own in-house solutions. User interface options range from allowing the user to select from predefined visualizations to giving the user the ability to write their own highly customizable queries each with a different learning curve dependent on complexity of the software user interface.
The type of graphical display used depends on the size of the data set being analyzed and the questions that are being asked. Graphical programs can be customized as a front-end viewer to analyze data across single or multiple studies using data from single or multiple sources. Graphical displays of data from single toxicology studies evaluating changes by dose group, by time point, and among types of data (e.g., organ weights, histopathology) are commonly used. Published examples of these types of graphical display include using SigmaPlot to evaluate the relationship of kidney biomarkers and histopathology to nephrotoxicants (Vlasakova et al. 2014), Spotfire to evaluate the maturity-related changes in cynomolgus thymus parameters (Snyder et al. 2016), and R for the evaluation for miR-122 as a marker for hepatotoxicity (Sharapova et al. 2016). Large genomics and metabolomics data sets can be integrated into these evaluations using heat map–style hierarchical data clustering, phenotranscriptograms, or principal component analysis (PCA) displays (Hamadeh et al. 2002; Lobenhofer et al. 2006; Raddatz et al. 2017; Xu et al. 2008). Comparison of individual study data against large historical data sets has also been demonstrated using KNIME, a free open-source data mining tool (Briggs 2017).
System Structure and Configuration
Data from toxicology studies go through a series of steps. Data are generated and populate the files of a toxicology data management program. These programs can export data files (e.g, *.txt, *.csv) and/or can connect to a data warehouse. Data from toxicology data management programs can also be used to generate SEND-compliant data files using integrated functionality or third-party solutions.
Data repositories hold data from multiple studies and use a standard data structure. Data repositories can be an “off-the-shelf” product supplied by vendors or a custom-built system. Data repositories (or warehouses) generally contain data from numerous studies sponsored by a single company. These data repositories may include only data from reported studies or may also include all data released by the performing laboratory. Since data repositories impose a standard data structure, they offer an efficient means to evaluate data across studies or to explore historical data to put current data into context.
To ensure that meaningful comparisons can be made across studies, data brought into a data repository require reconciliation. This involves transformation of synonymous terminology to a single term for user-generated terms (e.g., in-life findings, histology terminology) and for test names and units. Reconciliation may require significant effort for data originating from multiple facilities or data generated prior to SEND implementation. Even if SEND-controlled terminology is utilized, terms used at the time of data collection (prior to availability of SEND nomenclature) may require further analysis to map to the standard lexicon. This mapping increases the value of future cross-study comparisons.
Graphical representations of data sets can be created by manually (i.e., case by case) importing data in a recognizable format into a visualization tool, by mapping the visualization tool to one or more data sources, or by pointing the visualization tool to a database or data warehouse where the data have been optimized for querying and reporting.
Data can be visualized during any step in this process, but methods may be different depending on study status. Data from a subset of unreported or ongoing studies may reside only within the source data systems (e.g., toxicology data management systems) or may also be present in a data warehouse (see Figure 1). For those data that only reside within a toxicology data management system (source data system), the data are generally accessible via the extraction of files (*txt, *.csv) for a single study or by direct connection to the data collection system database. These files can be imported into Excel or another graphing program (e.g., GraphPad Prizm, Spotfire) either manually or through an automated process. Visualization of these data can be automated via macros or web-based viewers or can require users to customize their own visualizations as needed.
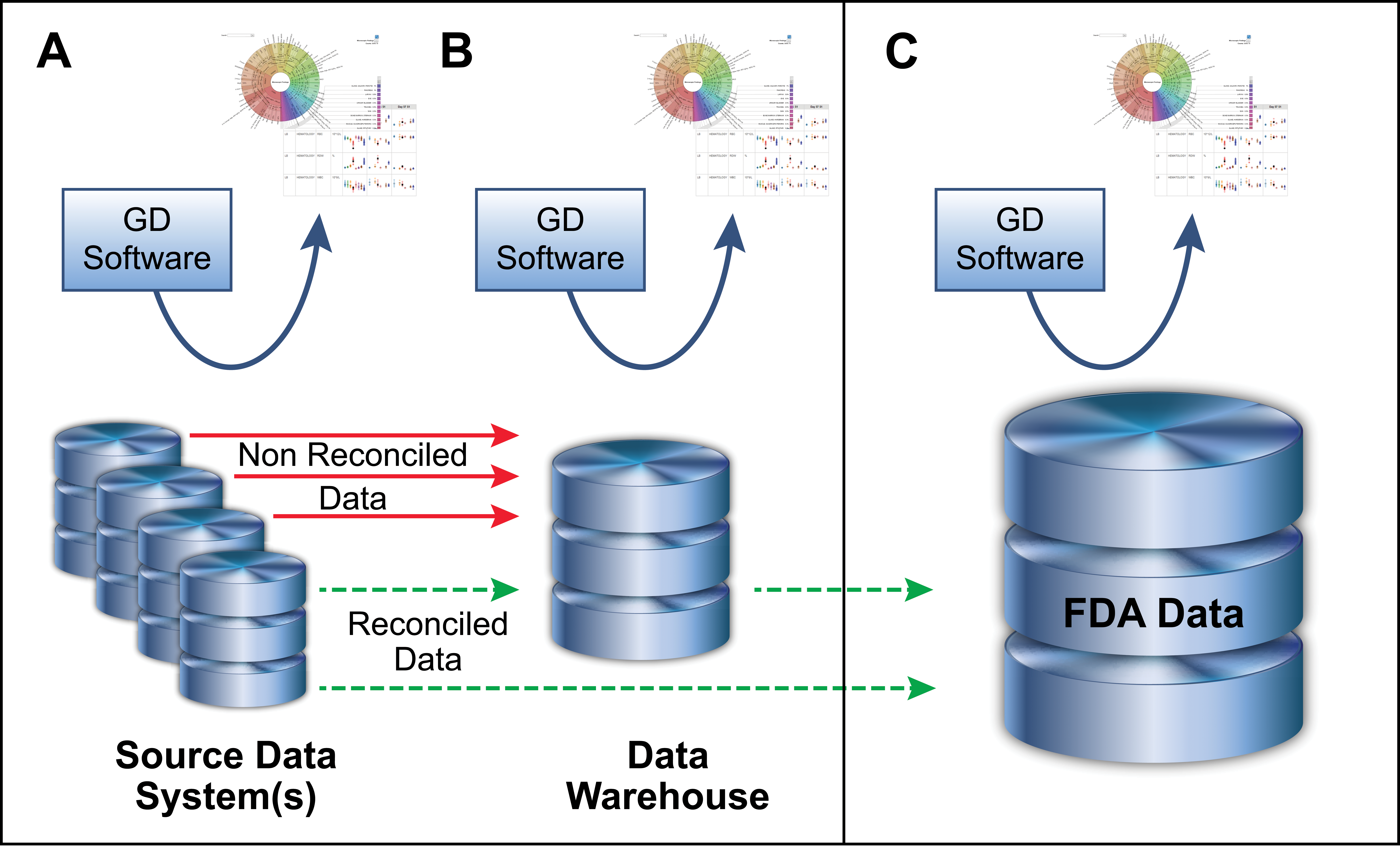
Multiple options for graphical display (GD) software implementation. Depending on the configuration options and capabilities of the GD software, data can be viewed directly from the source data system (A) and/or from a downstream data warehouse (B) prior to submission to the agency (C). These data may be queried before (red arrows) or after data reconciliation (green arrows) to standardized terminology. Querying the source data system(s) allows for more real-time access to study data; however, data may not be converted to a common terminology. Querying transformed data (e.g., from a downstream data warehouse) can facilitate cross-study evaluations but may involve a lag time for data transformation and transmission.
A potential downside to analyzing data real time is that the data visualization is of nontransformed data that may not match what is ultimately generated in a SEND-compliant output; however, this may be an acceptable trade-off particularly for companies with source data lexicons closely matching SEND terminology. For SEND-compliant data sets, visualization is more straightforward due to the standardized data structure and terminology required by SEND. Some visualization software systems are constructed specifically to accommodate the SEND-compliant data. More attention to mapping may be required to incorporate findings that do not use controlled terminology into the data visualization software. At the current time, data sets are generally not transformed to SEND format until after completion and reporting of a study, which limits the value of SEND-based data visualization tools for ongoing studies and for studies that are never transformed to SEND format.
Graphs and other data displays are effective when they clearly communicate results of interest. The best graph to use for a data set is the simplest one that adequately conveys information that addresses the question being asked. The same data set can be viewed in various ways to answer different questions, so systems used to visualize data should be flexible and customizable to the questions specific to the study, scientist, or reviewer. For example, in a study set with 4 groups with changes in alanine aminotransferase (ALT) activity, organ weight, and histology data from 3 time points, a pathologist might want to use a graphical display to visualize and understand the relationship among any of these parameters, or the relationship to metadata, such as dose group, time point, or study phase. In order to select a display format that effectively communicates the data set(s) of interest, it is important to consider the type of data being evaluated and the potential advantages or limitations of various data visualization strategies for different data types.
Selection of an Appropriate Display Format
Categorical and Ordinal Data Displays
Categorical data include pathology end points with a finite number of values that have no intrinsic order (e.g., yes/no, male/female). Ordinal data is a subcategory of categorical data where a ranked value is assigned to a finding (either a word or a number; small, medium, large; or 1, 2, 3). With ordinal data in pathology, the biological difference in meaning interval between categories assigned 1 and 2 may be larger or smaller than the difference between categories 2 and 3. Examples of pathology end points that produce these types of data include urinalysis dipstick results (e.g., 1+, 2+, or 3+ urine protein) and microscopic data (blood smear morphology, urine microscopic evaluations, and histomorphologic evaluations). Categorical data’s graphical displays often utilize a format similar to a standard data table but can include enhancements such as color coding to differentiate attributes (e.g., severity, dose groups, or individual data points) and the ability to “drill down” within a summary display into individual data points of interest. For example, color-coded reports of blood smear morphology data (Figure 2) or histopathology data (Figure 3) have a familiar format and data density but can be reviewed more efficiently than one without color coding. Use of (stacked) bar graphs such as those presented in figures 4 and 5 can be a useful alternative to the more commonly used tabular presentation for categorical toxicology data.
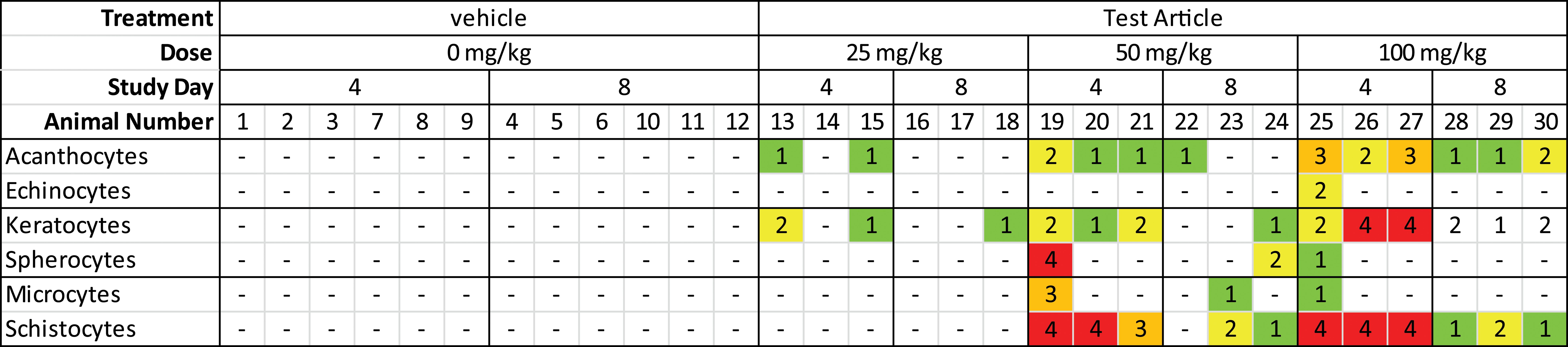
Blood smear morphology data with color-coded severity scores.

Histopathology data table with color-coded severity scores.
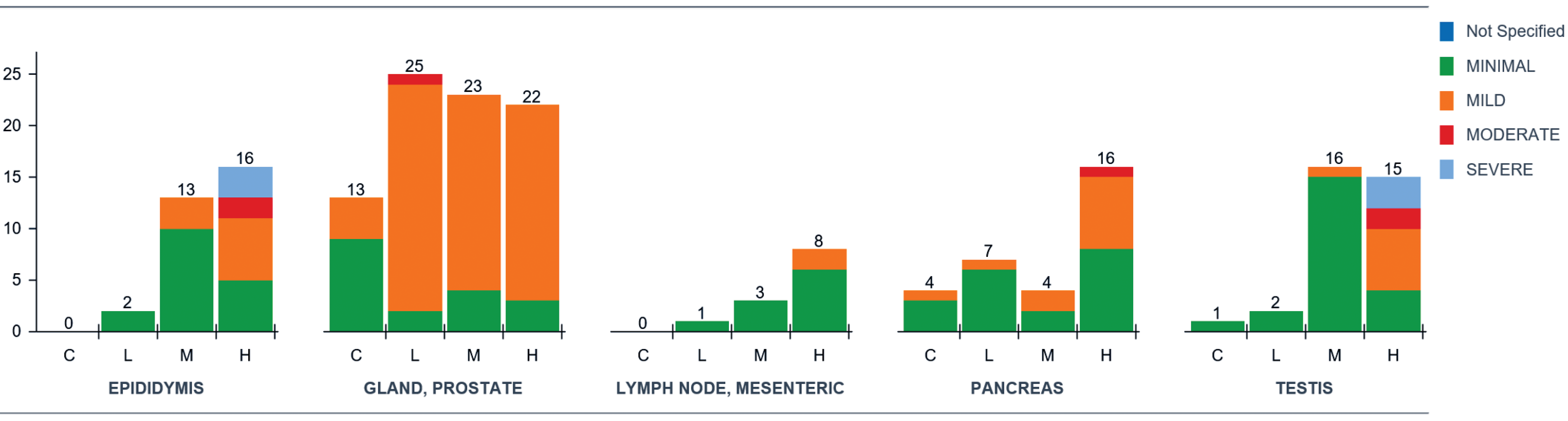
Histopathology bar graph with color-coded severity scores representing the same histopathology data presented in Figure 3 (dose levels are abbreviated as C = control, L = low, M = mid, and H = high;
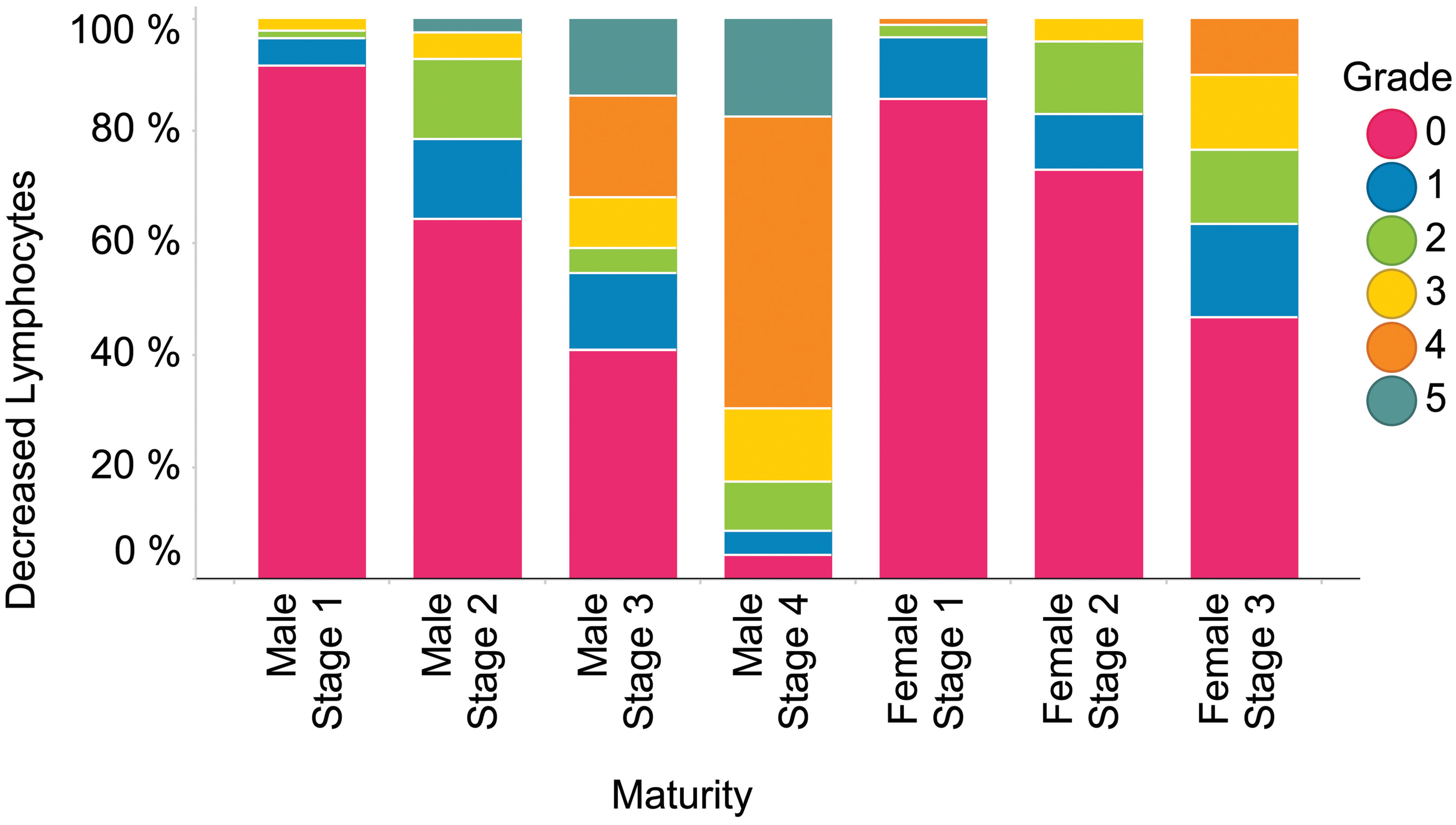
Stacked bar graph of thymus histologic findings as a function of maturity stage.
Continuous Data Displays
Pathology end points that produce continuous data include most instrument-generated clinical pathology results (including biomarker results), organ weights, and image analysis measurements. These data may be analyzed repeatedly (repeated measures) on the same subject (e.g., clinical pathology for large animals) or only at one time point (e.g., organ weights, clinical pathology for rodents). Line graphs are useful for visualizing repeated measures for individual animals, allowing the visualization of changes over time in an individual animal while also understanding differences among groups (Figure 6A). Depending on what software is used, these data can be stacked or trellised (e.g., lines for each group on a different row or column) or can be constructed as a pivot graph (lines for each group clustered together). Lines can be further delineated by color coding. Line graphs are particularly useful to evaluate data as change from pretest (percent change or difference in absolute values, depending on which approach is most appropriate) as well as absolute values; this can be accomplished rapidly with pivot charts in Excel. While traditional line graphs and pivot charts excel in demonstrating trends in a single data set over time, it is challenging to use a line-graph format to effectively demonstrate relationships between a continuous data end point and other study end points (e.g., reticulocyte counts vs. body weight or ALT activity vs. histologic scores for hepatocellular necrosis). For these comparisons, scatterplots are generally more informative and user-friendly.
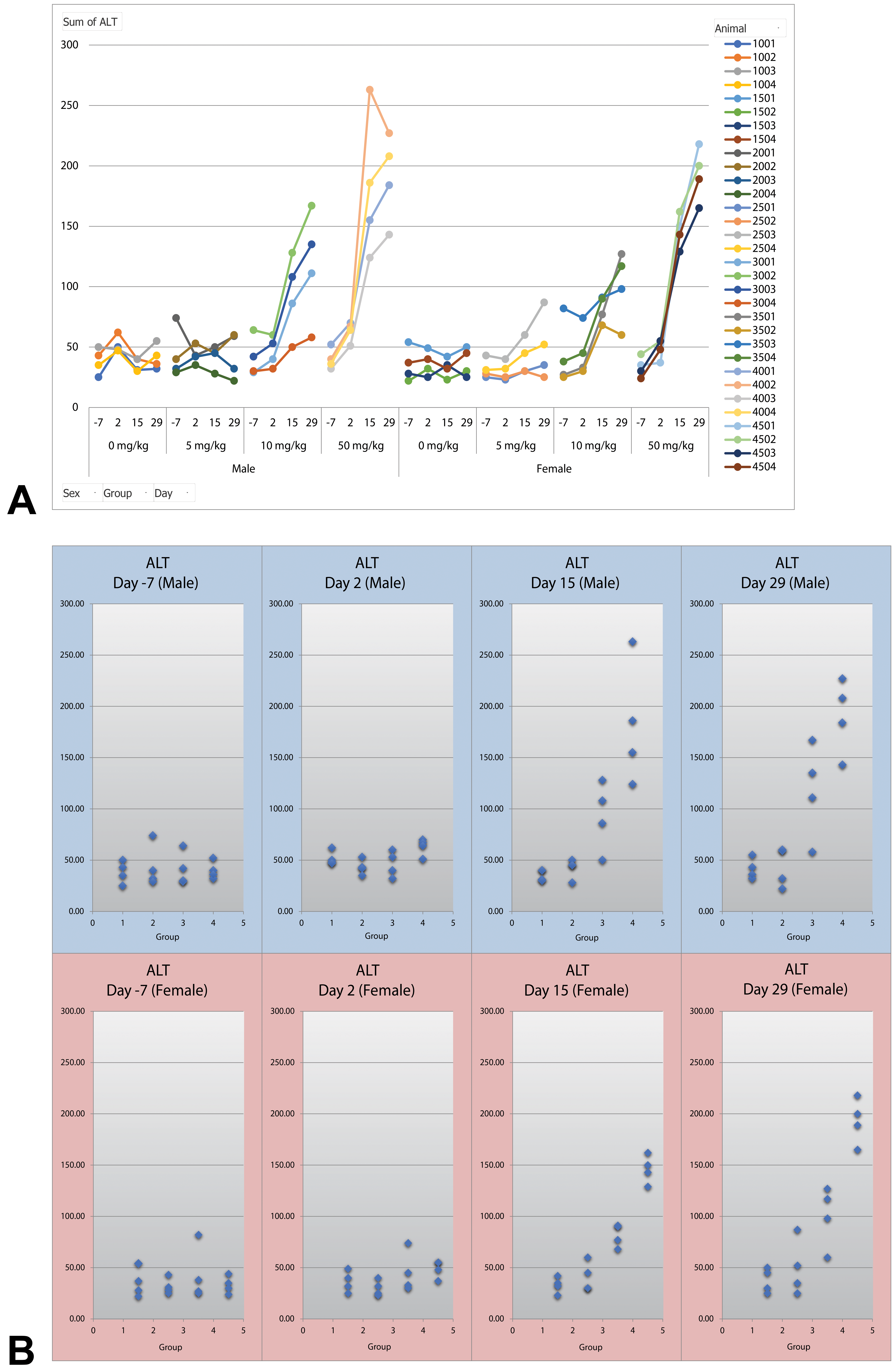
(A and B) Basic graphing options for visualizing continuous data collected at multiple intervals. (A) Serum alanine aminotransferase (ALT) data for nonhuman primates are shown at multiple collection time points, with metadata (time point, dose group, and sex) plotted on the
For data that are only collected at one time point per animal, scatterplots with group designation on the
Scatterplots can also be useful for demonstrating the relationship between two continuous parameters. For example, in a rat study with a dose-related increase in reticulocyte counts, mean cell volume and reticulocyte count in rats might be positively correlated. These scatterplots colored by dose or group, with one continuous variable on the
Categorical data can also be graphed against continuous data in order to understand relationships. For example, relationship of ALT activities of animals with grade 0, 1, 2, or 3 for necrosis of hepatocytes may be graphed using a number of different types of graphs (including but not limited to scatterplots, bar graphs, whisker plots, heat maps, or PCA). An example of a visualization combining categorical data with continuous data is shown above (Figure 7), illustrating that animals with similar ALT activities can have a range of histologic grades for hepatocellular necrosis. The metadata associated with a data point (dose, sex, group designation, and time point) are used to graph either categorical or continuous data. Additionally, this concept can be extended to understand the relationship of similar data between two or more studies. For example, this graphical approach can be used to demonstrate the potential of sorbitol dehydrogenase (SDH) activity as a biomarker of hepatocellular necrosis in the liver. A bar and whisker plot of terminal SDH activity (Figure 8) demonstrates the relationship between the severity of the hepatocyte necrosis and increased serum levels of SDH.
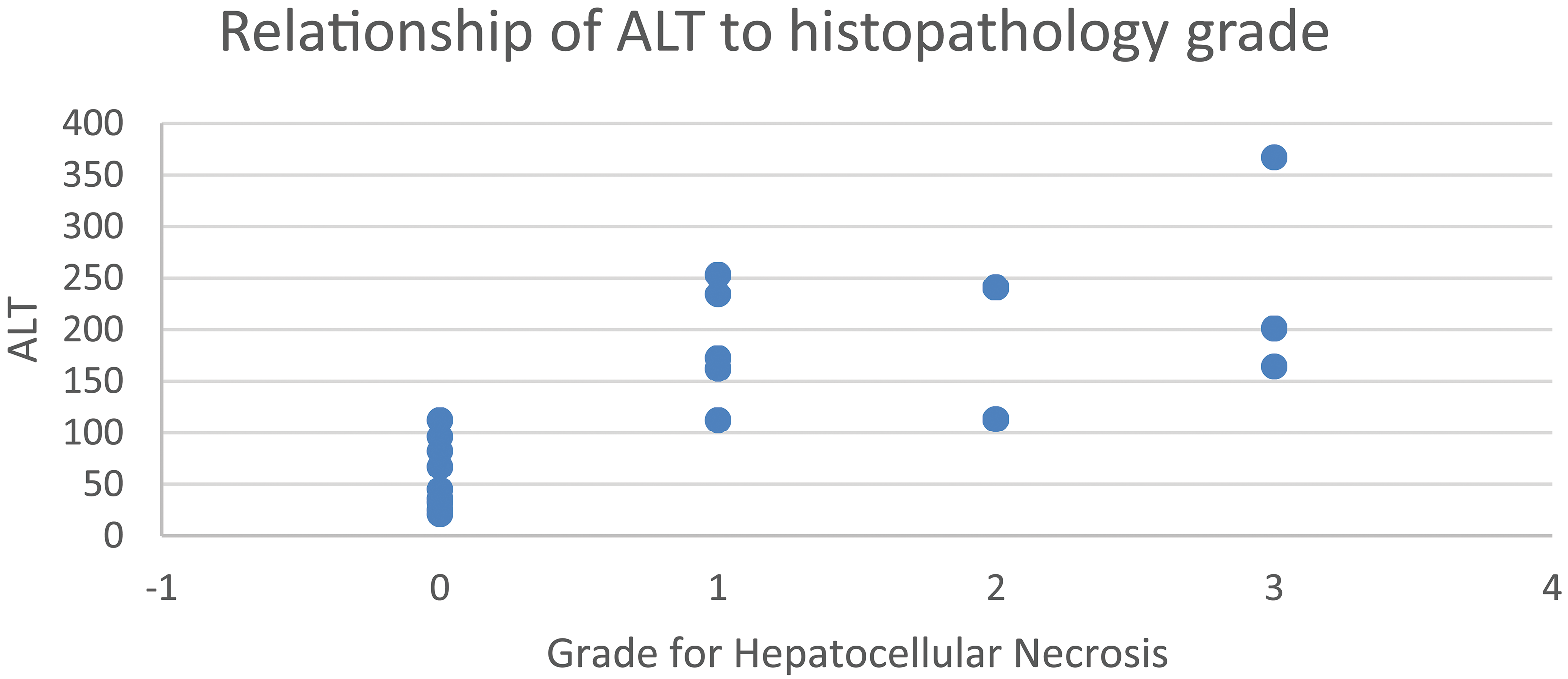
A graphical display plotting histopathology severity grade (necrosis) versus ALT activity in a single study.
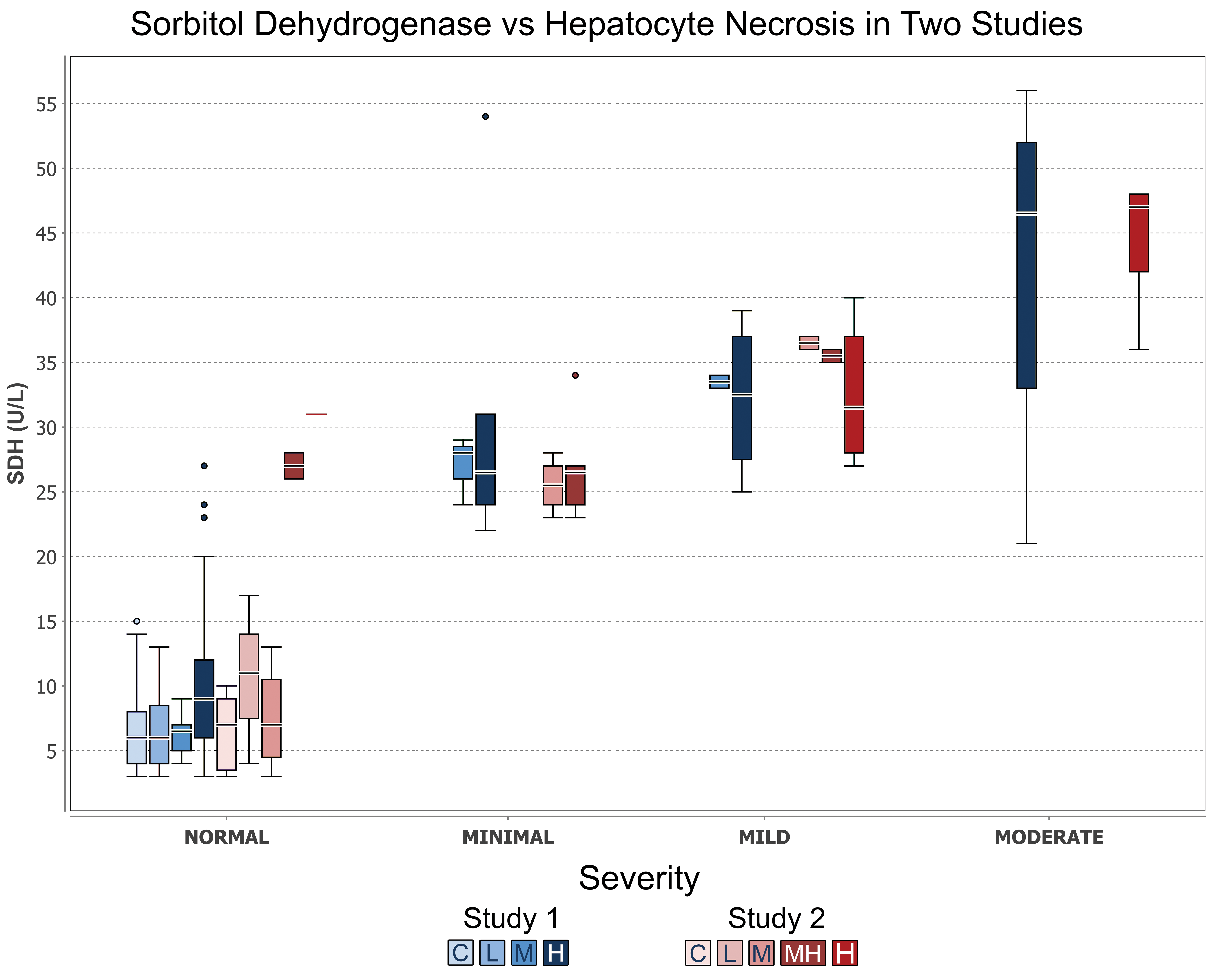
A bar and whisker plot for visualizing similar data collected from two different studies in Wistar Han rats. Terminal serum sorbitol dehydrogenase (SDH) is plotted on the
Treemap and sunburst displays (Figures 9 and 10) can be useful in providing visual cues for specifically defined parameters (e.g., incidence, severity) using broad graphical representations of entire studies or collections of historical toxicology data. With a treemap display (Figure 9), findings within a study are represented by a rectangle with descriptors such as incidence or severity influencing the area of the box. This type of display can also be organized with hierarchical ordering of findings by box area. The “sunburst” display (Figure 10) was originally developed for metagenomics evaluation and has since been adapted for visualization of other data types. The sunburst display uses a pie chart format with each slice representing a finding and its size influenced by the incidence or severity. These types of displays are useful when investigating trends within a study or across multiple studies.

Multistudy treemap view of background histopathology findings in control rats. Treemaps use nested rectangles colored by category with area proportional to outcome (e.g., incidence/severity). Such displays can be particularly useful for compact representation of large hierarchical data sets. In this example, color is used to specify the organ, each rectangle represents a separate histomorphologic diagnosis, and the area of the rectangle is proportional to incidence.
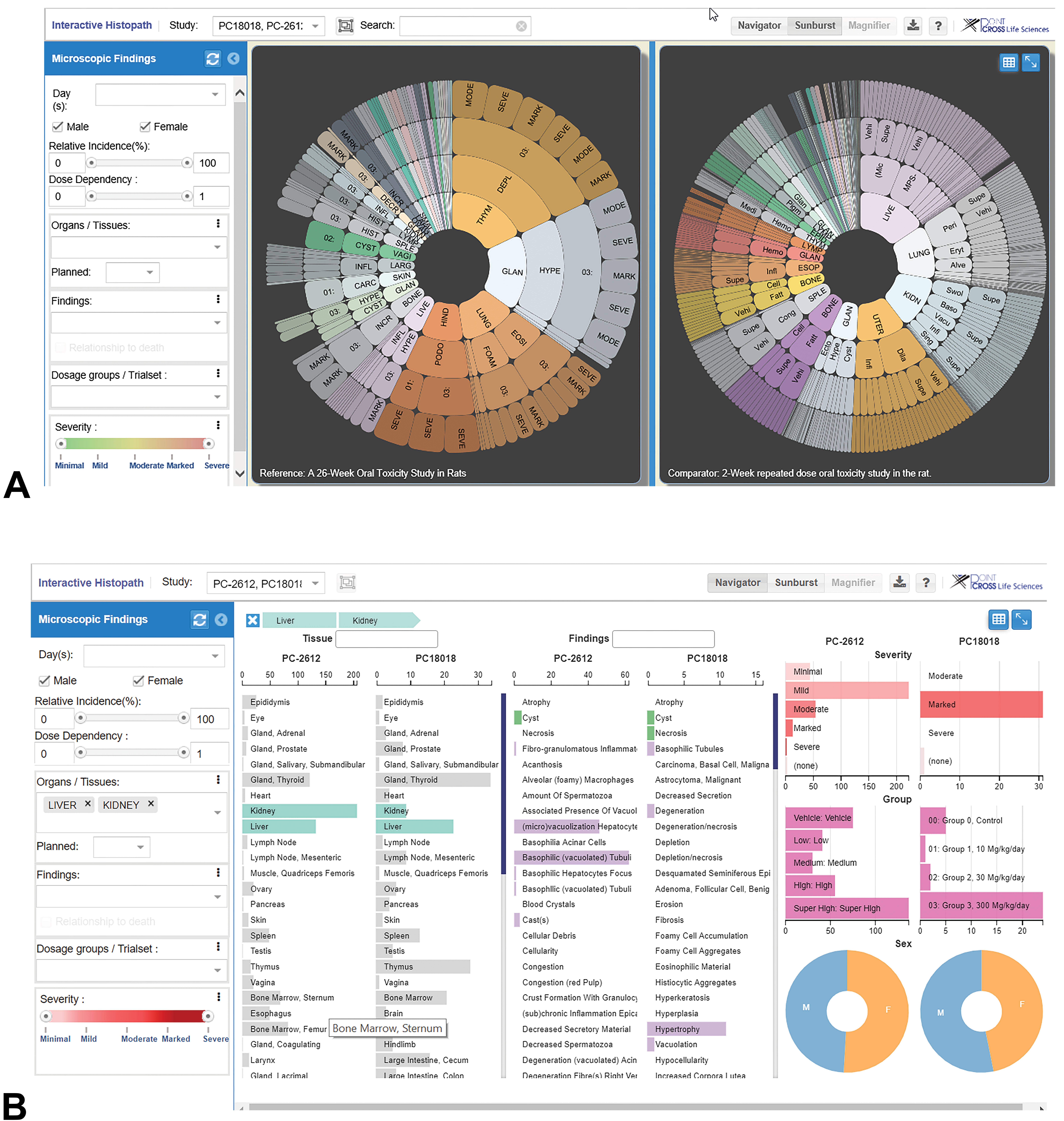
Sunburst displays can provide dynamic hierarchical representations of data (microscopic findings in this example) from an entire study with the highest order at the center of the circle. Users can quickly implement changes to the order of the hierarchical view and can drill down to explore parameters in more detail. (A) A comparison of histopathology findings across 2 rat studies PC2612 (2-week repeated dose oral toxicity study in rats) and PC18018 (26-week oral toxicity study in rats). (B) The same data set using a combination of bar graphs and pie charts to compare incidence and severity of histomorphologic findings across in males and females with focus on dose-related kidney and liver changes.
Use of Data Visualization Tools for Real-time versus Retrospective Data Analysis
During the course of toxicology studies, a variety of in-life study data are frequently monitored to determine animal suitability for assignment to studies, to assess changes in toxicology or pathology end points that may require further monitoring, and to identify potential critical animal health issues that may necessitate clinical intervention or modification of study design. Such real-time assessments are needed to maintain acceptable study conduct. However, the extent to which data visualization tools can be used for this purpose may be limited by degree of mapping of visualization software to source database and/or frequency of data transmission to a mapped data warehouse while a study is in progress. Therefore, depending on system configuration, scientists evaluating data during the conduct of a study may be limited to the use of native tabular outputs from the source data system or more basic visualization tools. Interim data are generally released on a preliminary basis and, therefore, assessments made during the course of the study should be verified at the completion of the study to ensure that the interpretation is accurate for the final SEND-compliant data sets used for regulatory submission.
Once a study or series of studies has been completed and “finalized” the data from these studies can be further visualized or analyzed retrospectively. Retrospective data visualization and analysis presents fewer technical challenges, given the availability of complete, reconciled, and “finalized” data sets that can be particularly helpful in enabling complex cross-study data comparisons (although such cross-study evaluations are not without caveats as described below).
Use of Data Visualization Software for GLP Study Evaluation
Most data visualization systems generate graphic displays that facilitate viewing of existing study data. Because these systems are not creating new data, they are not subject to the rigorous validation and documentation requirements required by the Code of Federal Regulations, Volume 21, Part 58 (good laboratory practices [GLP]) and Part 11 (electronic records/signatures) of other GLP “data capture” systems. For this reason, developers of data visualization software may avoid the inclusion of statistical packages or other capabilities that may be construed as data generating. For GLP studies, data visualization graphics included in the report are accompanied by the raw data in tabular format and the graphics are considered illustrative, in the same manner that a photomicrograph might be used to communicate the nature of a microscopic finding. If the interpretation of data is conducted with a non-GLP-compliant data visualization system, it should be confirmed with evaluation of GLP-compliant data tables.
Selection of Optimal Data Display Formats and Potential for Misrepresentation of Data
While graphical displays represent a valuable tool for conveying data in a succinct manner, it is important to recognize that incomplete or inaccurate conclusions can easily be drawn from a poorly designed display or from a type of display that is not well suited to the data being presented. Therefore, particular care should be taken in the selection of visualization tools and graphical display types to ensure that the graphical presentation of the data is free of misleading visual perceptions, such as visual distortion associated with 3-dimensional graphs, incorrect use of scale or error bars, truncated graphs, or graphs with inappropriate intervals (Hickey 2013; Duke et al. 2015; Gallery of Data Visualization 2000). Consideration should also be given to the type of data being presented relative to the type and amount of information that various graphical types can inherently demonstrate. For example, Weissgerber et al. (2015) highlight the fact that bar graphs are commonly used to visualize intergroup comparisons of continuous data, although such displays convey only mean values (often with standard error or standard deviation) and do not provide any information regarding data distribution within and among groups that can be critical for complete scientific assessment of important intergroup differences (such as could be conveyed using a scatterplot of the same data set). This concept becomes even more important in presentations of cross-study data relationships where, in addition to avoiding the visual misperceptions, continuous numerical data and categorical grading scales may require standardization to ensure that potential variability in data reporting among studies does not unintentionally influence the visual interpretation of trends in a particular end point or correlations among end points.
Use of Data Visualization Tools for Indiscriminant Large-scale Analyses
In an effort to identify potential test article–related effects or data relationships within a study, it would be theoretically possible to set up a large query to plot every data end point within a study against another in a search for potential treatment effects or cross-domain relationships; however, such an approach would come with several downsides. Not only would it likely be impractical to review such a mass of data, many of the resultant (random) comparisons would not make sense scientifically and the sheer number of assessments would predispose to more false positive “flags” due to random variability. A more efficient use of data visualization tools involves initial assessment using a core set of display (with or without modifications based on anticipated or known effects associated with a compound class or mechanism of action) to provide an overview of the data with additional hypothesis-driven queries (e.g., specific cross-domain displays) conducted on a case-by-case basis based on the initial findings.
Cross-study Data Displays
Cross-study (or multistudy) analyses should be performed with caution and with full knowledge of the limitations, quality, and assumptions surrounding the source data. Although the use of SEND-compliant data using consistent terminology should theoretically facilitate comparison of histopathology findings across studies/programs, it is probable that those studies were reviewed by different pathologists with an associated interobserver variability in thresholds and grading criteria, thus limiting any direct comparison. Attaining high precision in the comparison of clinical pathology results generated from different instrument platforms may also be difficult due to instrument bias and/or analytical differences. Similar caution is warranted with cross-domain analyses (e.g., comparison of histopathology and clinical pathology), where interpretation is predicated upon a basic understanding of the normal relationships between the parameters in each domain in order to avoid misleading or erroneous associations. Figure 11 provides a multistudy comparison of histopathology data using bar graphs.
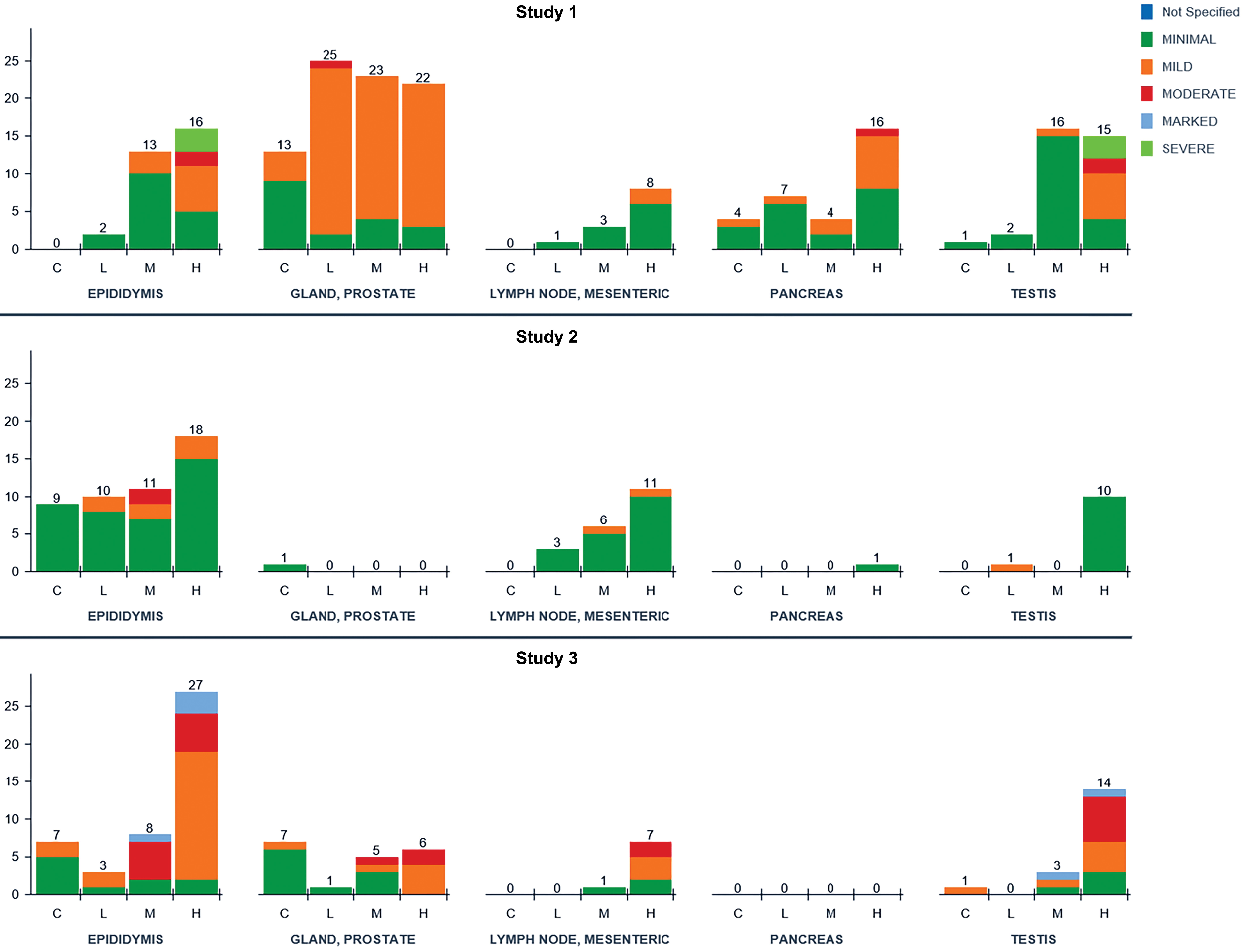
A histopathology bar graph display can be used for multistudy comparisons (dose levels are abbreviated as C = control, L = low, M = mid, and H = high;
Application of Statistics to Data Displays
While it may be tempting to supplement data visualization software with statistical analysis capabilities, caution should be used when considering such an approach, given the implications for the potential generation of “new data” particularly in the context of a GLP study. While many data visualization systems can graph displays of basic descriptive statistics (e.g., mean, median, standard deviation, and other aggregated measurements), statistical analysis and interpretation should be limited to quantitative data (organ weights, numeric clinical pathology end points) and are best performed using a dedicated GLP-compliant system rather than within data visualization software. The use of statistical analysis for analysis of ordinal data such as histology scores is generally not recommended in routine toxicology studies (Mann et al. 2012).
Discussion
We have provided a review of some of the many types of data visualizations that can be used to supplement data review and presentation by scientists involved in toxicology assessments. All data visualization systems come with unique trade-offs in terms of level of customization, complexity, and ease of use and implementation. Likewise, there are additional trade-offs to consider when contemplating strategies for accessing/querying source data with data visualization software.
Data visualization tools are available to improve agility, speed, and clarity during data review or presentation for assessment of nonclinical toxicology data. These tools do not generate data and are generally not subjected to GLP computer system validation requirements. Verification of study conclusions should always be made against the raw data. Analysis of the clinical and anatomic pathology data directly from the slides, source application, or tabular data is the foundational requirement for toxicity study data interpretation, and visualization tools are not intended to be a substitute for analytical ability. Since evaluation preferences can vary by the study, scientist, or reviewer, the tools used to facilitate data visualization need to be flexible and customizable and users should be cognizant of subjectivity and other variables involved in these evaluations.
Along with their many benefits, graphical display software also has the potential to be used inappropriately. To safeguard against potential misinterpretation of data, it is important that users have a basic understanding of toxicology, pathology, and the normal relationships between the parameters being evaluated. Scientists using these tools should also be aware of and guard against pitfalls such as indiscriminant or irrelevant cross-domain analyses, inappropriate use of statistics, or misleading use of graphical scale.
The use of dedicated data visualization software has been limited by cost, up-front investment in system configuration, and the effort involved in preparing queries. However, the establishment of a standardized data format (SEND) coupled with availability of more intuitive and user-friendly query tools and well-designed predefined data views have made implementation somewhat easier than in the past. It is likely that the use of such tools will increase among pathologists as the technology and graphics continue to improve and as the visualizations are further embraced by data reviewers. The Food and Drug Administration has been actively collaborating with scientists and software developers through the Pharmaceutical Users Software Exchange (PhUSE) consortium (Brown et al. 2016; Kropp et al. 2013; PhUSE 2015) to evaluate the use of graphical display software for review of nonclinical data submissions. Pathologists and other scientists involved in nonclinical toxicology studies continue to play an important role in providing feedback to enable the continued development and improvement of these tools.
Footnotes
Acknowledgments
The authors would like to thank Joyce Zandee, Mathew Mohit, and Kurien Abraham for their assistance in generating graphics and sample data and reviewers from the SRPC Committee and PhUSE graphical display working group for their critical comments on this article.
Author Contributions
All authors (ST, NE, WS, BK, ML, JH) contributed to conception or design; data acquisition, analysis, or interpretation; drafting the manuscript; and critically revising the manuscript. All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential, real, or perceived conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
