Abstract
How do experts stitch together seemingly irreconcilable scientific objects? This paper examines the way “endophenotypes” and “deep phenotyping” are deployed to bridge the infamously recalcitrant genotype–phenotype divide. The endophenotype concept emerged in mid-20th-century evolutionary biology/entomology and was briefly adopted in schizophrenia genetics in 1972 before virtually disappearing for a generation. Decades later, in the “postgenomic” 2000s, the concept exploded in medical—and especially psychiatric—genetics in the United States and United Kingdom. Endophenotypes now refer to phenotypic observations like subclinical biomarkers or traits that are both more fine-grained than disease categories and yield stronger associations with genetics. We argue that the endophenotype concept functions as an “epistemic quilting device” that allows experts to forge research programs across disjunct scientific ontologies and fields, creating new wholes that respond to historically specific needs. But they are also destabilizing existing medical categories and foundational concepts in genetics like penetrance and recessivity, with radical implications for “precision medicine.” Today, endophenotypes and the more loosely defined “deep phenotyping” have been integrated into the infrastructures of postgenomic research in both rare disease and big-data genomics, quilting together genetics and other fields interested in human illness and difference even as it disrupts them.
Biomedical experts have developed an expansive toolkit for bridging the infamously recalcitrant chasm between our knowledge about genotypes and phenotypes. It is the quintessential “postgenomic” project: finding ways to turn data about our DNA into meaningful information about human disease and difference (Reardon 2017; Rheinberger and Müller-Wille 2018; Richardson and Stevens 2015). To meet this challenge, researchers have built up massive databases of genomic and phenotypic information, with an army of bioinformaticians, algorithms, and artificial intelligence tools to make sense of it all (Stevens 2013). Others have turned the problem on its head, using genetic variants to delineate and diagnose new categories of disease and difference (Navon 2019). Polygenic scores are the flavor of the moment (their millions of meagre ingredients and poor explanatory kick notwithstanding). Clinical geneticists often engage in a complex semiotic matching process between genotype and phenotype in order to make calls about a DNA variant's pathogenicity (Timmermans 2017). More broadly, the field has increasingly pivoted toward a narrative of “enlightened geneticization” (Hedgecoe 2001) that accommodates environmental factors and complexity even as it centralizes genetics. To that end, there is a seemingly endless parade of multiomics approaches that integrate various combinations of epigenomics, proteomics, metabolomics, environmental “exposomics” and early childhood development with the genomics mothership (Landecker 2011; Landecker and Panofsky 2013; Pickersgill et al. 2013; Shostak 2013; Meloni 2019; Lock 2015; Penkler 2022). 1 These and other postgenomic research programs have attracted widespread social scientific critique.
Yet the social studies of science and medicine have mostly overlooked an increasingly influential pair of strategies that both cut across these various postgenomic approaches and help tie them together. 2 Consider the titles of a just few influential papers: “Endophenotypes: Bridging Genomic Complexity and Disorder Heterogeneity” (Insel and Cuthbert 2009 in Biological Psychiatry); “Deep Phenotyping for Precision Medicine” (Robinson 2012 in Human Mutation); “From Genotype to EEG Endophenotype: A Route for Post-Genomic Understanding of Complex Psychiatric Disease?” (De Geus 2010 in Genome Medicine); “Deep Phenotyping: The Details of Disease” (Delude 2015 in Nature); and “Deep Phenotyping Unveils Hidden Traits and Genetic Relations in Subtle Mutants” (San Miguel et al. 2016 in Nature Communications).
But what are these things—“endophenotypes” and “deep phenotyping”—that promise to suture the ontological gap between genomics and medical classification? How can we explain their dramatic rise in recent years, and what lessons might their uneven history hold for the social studies of science more generally?
A phenotype, of course, is the observable features of an organism which result from the interaction of its genotype and the many relevant environments (cellular, organic, nutritional, social, etc.) that shape the organism's development. In practice, phenotypes tend to be the traits and diseases that one might discuss independent of genetics—in lay parlance, medicine, or any other expert field. From a genetic perspective, phenotypes are usually beguilingly complex: apart from rare diseases and certain cancers, it is quite unusual for any form of genetic analysis—be it pedigree charts, heritability estimates, or some sort of chromosomal, gene, or DNA test—to yield a straightforward explanation of their presence. In fact, experts usually struggle to find any clear-cut matches between a given phenotype and a genotype: the postgenomic riddle in a nutshell.
That is where endophenotypes come in. The prefix endo points to something that lies within or makes up a part of a larger whole. But endophenotyping has never been a simple matter of parceling up phenotypes into more fine-grained components. Instead, endophenotyping is all about identifying novel forms of phenotypic specificity to enhance the explanatory power of genetics. But how? When we disaggregate traits and medical conditions into more subtle, specific phenotypes, a couple of important things are supposed to happen. First, we should find phenotypic elements that lend themselves to much stronger associations with heritability estimates or genotypes. Second, we may discover features that are also often present in family members who share the same genetic background but were thought to be either unaffected or mere carriers. In this way, endophenotypes help to marry knowledge about genetics and human difference even as they destabilize long-standing modes of classification.
This paper shows how endophenotyping and its conceptually looser offspring, “deep phenotyping,” became commonplace, powerful, and disruptive strategies in British and US human genetics—though they are now circulating globally as well. 3 After gaining traction in genetics research on schizophrenia, endophenotyping was adopted by a variety of fields—above all in psychiatric and behavior genetics, but now also across medical genetics and even beyond. Researchers can develop an understanding of a genetic variant or mutation that is more fine-grained and nuanced than “x percent of carriers have autism,” “y percent have intellectual disability,” or “z percent have schizophrenia.” Instead, they can identify a variety of autistic behaviors and differences, particular cognitive strengths and weaknesses, and varying forms of “schizotypy.” They can also look for a variety of “intermediate” phenotypes—biomarkers from molecular or metabolic assays, imaging findings, and so on—that bolster the connection between heredity and pathology. Whether it is so much convenience experimentation to enhance the statistical power of a genome-wide association study (GWAS) analysis, an effort to flesh out the phenotype of a genetic disorder, or many things in between, endophenotypes and deep phenotyping are now a core part of the postgenomic project.
But we will also see how the term “endophenotype” has metamorphosed since it was introduced into the scientific literature over half a century ago. The first significant appearance came in a 1966 Science article on species differentiation in entomology (John and Lewis 1966). The endophenotype concept then mutated into something akin to its contemporary form when prominent psychiatric geneticists Irving Gottesman and James Shields repurposed it in a book about schizophrenia to identify a series of subtle biological and behavioral similarities present in pairs of identical twins who were discordant for schizophrenia itself. Yet, even as Gottesman and Shields grasped its radical implications from the very beginning, the endophenotype concept was hemmed in by prevailing approaches to human genetics, and almost completely abandoned for a generation. It was only in the early 2000s that endophenotypes began to make a real mark—the start of a dramatic upswing of publications across a growing range of fields. The reason, we argue, was not some breakthrough in endophenotype research, but the burgeoning realization that vast investments in the field headlined by the Human Genome Project (Hilgartner 2017) were yielding precious few insights when it came to common disease. Endophenotypes represented a tantalizing route through the postgenomic impasse.
Today, endophenotyping has morphed into a verb and been largely supplanted by deep phenotyping, shedding much of the methodological stringency stressed by Gottesman and others. Crucially, endophenotypes are no longer just things, but the core component of a widely deployed strategy in research programs that seek to salvage the promise of genomic medicine. Endophenotypes and “subclinical traits” are often seen as the corollary of more biologically precise diagnosis, and researchers are designing major (endo)phenotype ontologies and databases to accommodate the massive influx of genomic data seen in recent years. In short, endophenotypes and deep phenotyping have emerged as central tools, built into the very infrastructures of postgenomic research. At the same time, we argue that they are increasingly destabilizing established medical categories and foundational concepts in genetics like penetrance and recessivity, with potentially radical implications for the rollout of “precision medicine.” Deep phenotyping is even taking off in clinical research and epidemiology—changing the classification of human difference in fields far-removed from genetics.
So, this paper outlines the surprising history of the endophenotype concept and tries to explain how it came to play an increasingly central role in contemporary human, medical, and especially psychiatric genetics.
But more than that, we suggest that the endophenotype concept and deep phenotyping can be usefully thought of as a pair of epistemic quilting devices that allow experts to sew together otherwise discordant ontologies of human difference. A quilt, after all, is made from two layers of fabric that were never meant to be sewn together, and an intermediary, mostly unseen layer that helps stitch them together in a new whole. The endophenotype concept is akin to this middle layer of “batting”—a hidden material that allows researchers to sew together pieces from a top layer of medical classification and a bottom of genomic data. Just as batting is there to join two outer layers, an epistemic quilting device is created to fulfill a historically specific need to bridge disjunct scientific ontologies. Thinking about the endophenotype concept using the metaphor of epistemic quilting helps us understand its postgenomic resurgence.
Even though the endophenotype concept may be intrinsically relational, we will see how it allows experts to sew together new wholes that are destabilizing the meaning of human difference in the postgenomic era. This, in turn, raises a series of issues for both modern human genetics and science and technology studies (STS). On the one hand, it challenges biomedical experts and STS scholars alike to consider to what extent the move to deep phenotyping is driven by patient needs and clinical utility, as opposed to the political need to salvage the epistemic, statistical, and sociopolitical power of genomics after much investment yielded few insights or treatments for common diseases. On the other, the concept of epistemic quilting device offers a new analytic tool to help STS scholars better understand how researchers bridge disjunct scientific ontologies—and the disruptive effects that can ensue.
The Strange Career of the Endophenotype
Endophenotypes and deep phenotyping have decisively arrived as key tools in contemporary medical genetics, but they took a winding road to get here. The concept of the endophenotype can be traced back to midcentury developments in the seemingly far-flung fields of entomology and evolutionary biology. Stranger still, the very definition of an endophenotype underwent a dramatic mutation when it made the jump to its new environment in psychiatric genetics in the 1970s. It then lay mostly dormant for a generation, only to be revived just as the Human Genome Project was coming to an end and the field found itself desperately needing to connect genotype and phenotype. This section tells the uneven story that saw the endophenotype travel from its confinement inside the nuclei of several insect species to a blossoming universe of fine-grained human traits, biomarkers, and dis/abilities.
Insecta and the “Chromosome Phenotype”
The term “endophenotype” dates to the 1963 book Chromosome Marker, by Lewis and John (1963). However, it was their article in Science a few years later that made a lasting impact. The paper, “Chromosome Variability and Geographic Distribution in Insects,” built on the pioneering work of Theodosius Dobzhansky to make a comprehensive case for using what John and Lewis (1966, 711) called the “chromosome phenotype” to untangle tricky issues in population genetics.
4
They contrasted this to the “exophenotype”—the phenotype found outside of the nucleus in other parts of the cell or organism (what we would usually simply call its phenotype). John and Lewis (1966, 714, our emphasis) explain that there is: a second component to the phenotype, a component which includes, among other things, the behavior of the chromosomes themselves. And all chromosome mutations affect this aspect of the endophenotype, for they all interfere to some extent with the course, and hence the consequences, of meiosis.
The endophenotype was therefore a matter of the observable structure of the chromosomes. John and Lewis lamented that chromosomal variation was mostly neglected or ignored. This was especially important, they argued (1966, 711), because “The chromosome complement is not just another character.” Instead, “The material of the genotype itself forms part of the structure of the chromosomes.” That had two important implications: “the chromosome phenomena is far less influenced by external factors than is the morphological or the physiological phenotype. Moreover, the chromosome phenotype is often a much more sensitive indicator of biological change and biological distinctiveness” (John and Lewis 1966, 711).
In other words, the very first appearance of the “endophenotype” concept in the scientific literature situates it as nothing less than a second and distinct “component” or dimension of the foundational notion of the phenotype. Distinct from the genotype (even though the endophenotype contains its material) and from the received notion of the phenotype (which they call the “exophenotype”), John and Lewis had theorized a fundamentally intermediary dimension of research on genetics and biological difference.
This intermediary status made endophenotypes extraordinarily useful for natural history: unlike the exophenotype, they are barely influenced by their environment; unlike gene mutations, endophenotypes cannot be ascribed to adaptive selection because they “cannot be detected in the exophenotype” (John and Lewis 1966, 714). The endophenotype “has no meaning for the individual in which it occurs…adaptation is a job for genes” (John and Lewis 1966, 720, our emphasis). It was a uniquely pure signal. So here we have it: the endophenotype operating solely within the nucleus, on a different temporality, and according to different forces—a singular object for natural history research.
John and Lewis’s (1966) paper is fascinating, and there is hardly a more high-profile venue than Science. Nevertheless, its impact appears to have been modest. It has been cited around 240 times over the past fifty-seven years, mostly as a precursor to endophenotype research in human genetics. While the term “endophenotype” did circulate in its target fields for a time, it does not appear in the titles or abstracts of any contemporary papers in entomology, evolutionary biology, or the other disciplines John and Lewis were speaking to. The endophenotype had a long way to go before it would gain lasting traction as a scientific concept.
Endophenotypes in Schizophrenia Research, but Before Their Time
How, then, did endophenotypes mutate and migrate into behavior and psychiatric genetics? The leap came in Irving Gottesman and James Shields's 1972 book, Schizophrenia and Genetics: A Twin Study Vantage Point (see also Irving I. Gottesman and Shields 1973), where they discuss the lessons and challenges of twin studies on schizophrenia and their implications for both schizophrenia research and psychiatric genetics more broadly. 5 The notion of endophenotypes only appears, at least explicitly, in a handful of passages toward the end of the book's 400-plus pages.
Gottesman and Shields present a wealth of evidence that schizophrenia is at once heritable and (like any phenotype) the product of genotype–environment interaction. More provocatively, however, they argue that some of the complexity we see with a condition like schizophrenia might be the product of our own conceptual frameworks for making sense of genetics and psychiatric diagnoses. As they recap, “Monogenic theories can be divided into recessive…and dominant.” But Gottesman and Shields (1972, 319-20 our emphasis) argue that those are theories rather than natural laws. What concept, then, could get closer to the underlying biological reality? Genes themselves are neither dominant nor recessive; the terms only have meaning with respect to a particular phenotypic characteristic. John and Lewis (1966) introduced the useful distinction between exophenotype (external phenotype) and endophenotype (internal), with the latter discernable only after aid to the naked eye, e.g., a biochemical test or a microscopic examination of chromosome morphology. As endophenotypes have become more available, the distinction between recessivity and dominance has become blurred; in a sense all genes are “dominant” (cf. sickle-cell anemia vs. sickling trait) when we have a way of detecting gene action molecularly.
Reimagined as any form of “internal” or subtler observation, endophenotypes help us see “genes” exerting an impact even when they do not cause the associated phenotype, at least in the way it is commonly understood.
This had far-reaching implications. Researchers could look for molecular traces or, as they explain a little later, other sorts of phenotypes that fall well short of standard diagnostic criteria but may still be associated with underlying genetic risk factors. As an example, Gottesman and Shields (1972, 320) point to inborn errors of metabolism like phenylketonuria—which had recently become the posterchild for national newborn screening—where carrier phenylalanine levels were known to be reduced but functional. Crucially, they also explained how one can sometimes identify more subtle physical, behavioral, or cognitive differences than the ones that qualify a person for a particular psychiatric diagnosis, that is, nonmolecular endophenotypes. This is the core set of ideas that would see the endophenotype concept take off in 21st-century human genetics.
But what made the endophenotype concept—originating as it did in entomology and natural history—so compelling to Gottesman and Shields back in the early 1970s? How did they even come across such far-flung research? John and Lewis's paper on endophenotypes was published in Science, arguably the top generalist journal in the world. Its title, and especially its blurb (“Chromosome rather than gene variations provide the key to differences among populations”) could easily grab the attention of an ambitious psychiatric geneticist looking for threads that might strengthen the association between schizophrenia and heritability findings. At precisely the same time, major studies on XYY (a chromosome abnormality that was infamously associated with aggression, crime, and mental “subnormality” by both researchers and mass media in the 1960s and 1970s) and psychiatric problems were being funded by the National Institute of Mental Health and other leading organizations, and cytogenetics was revolutionizing human genetics more broadly (De Chadarevian 2020; Hogan 2016; Lindee 2008; Navon 2019). In short, there is every reason to think that Gottesman and Shields would have been intrigued by an approach that used chromosome structure to get at problems of population and classification.
At a deeper level, the original endophenotype concept promised to fulfill a historically specific need for genetics researchers to rethink population categories—a simultaneously scientific and political need that was visible to a pair of innovative schizophrenia genetics researchers in the early 1970s and would eventually become widespread in human genetics in the 21st century. For John and Lewis with Insecta, Gottesman and Shields with schizophrenia twins, and for human genetics writ large today, what made the endophenotype concept so compelling is precisely its intermediate status and potential for reconciling discrepancies between genetic data and population classification. For as much as Gottesman and Shields changed what an endophenotype is at a material level, it retained its relational core. They hoped that endophenotypes would help solve long-standing riddles about the genetics of schizophrenia (e.g., discordant monozygotic twins) and help answer deep questions about the genetics of complex disorders. 6
So, the endo of the endophenotype concept had gone from something that resides “within the nucleus” to a universe of fine-grained observations that lie in between genotypes and clinical diagnoses. With hindsight, Gottesman and Shields's adaptation of the endophenotype concept is probably the book's most important contribution, but while the book appears to have made an impact in the field in the few decades after it came out, endophenotypes did not.
To put it bluntly, the endophenotype concept did not catch on. It was almost completely ignored in psychiatric and medical literatures for a generation until Gottesman and colleagues revived it for a postgenomic world. Figure 1 draws on a targeted ISI Web of Science search. 7 It illustrates how only a handful of papers used the term endophenotype (or a derivation) in their titles in the 20th century. Turning to a topic search, which includes terms from abstracts and keywords but is only reliable in a Web of Science search for papers published from around 1991, Figure 2 gives us confidence that the title search is not missing a large well of papers. 8 In short, despite a few passing references in schizophrenia research in the decades after Gottesman and Shields’s (1972) book, 9 the endophenotype concept gained minimal traction prior to the turn of the millennium. The first indexed paper about humans to deploy “endophenotype” in its title or abstract was Sharma et al.’s (1992) Schizophrenia Research article, “The Search for an Endophenotype Correlated with a Subtype of Schizophrenia: Magnetic-Resonance-Imaging in 2 Large British Pedigrees.” Then, from 1992, we begin to see a small trickle of papers using the term in human genetics and allied fields.
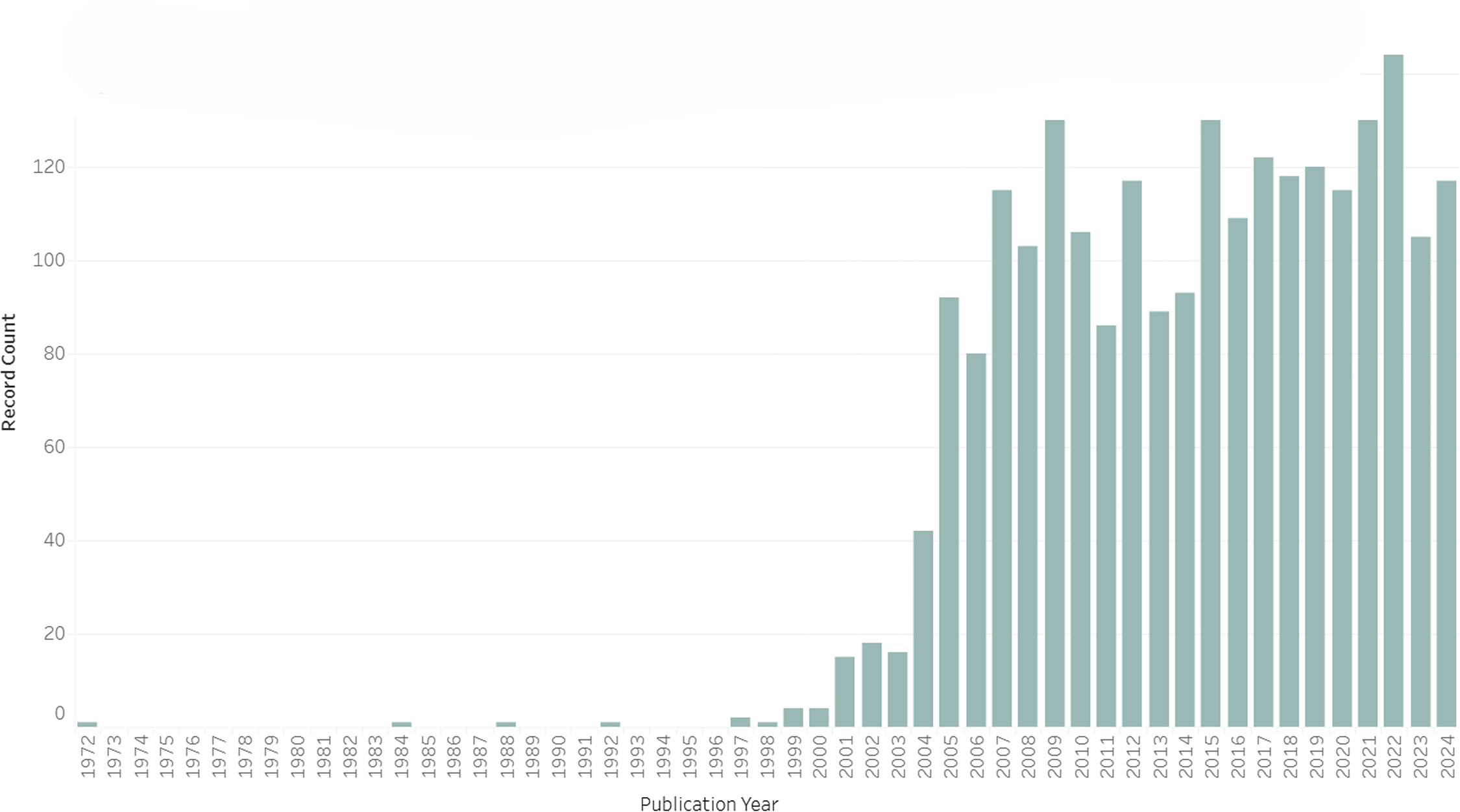
Papers Published Per Year with “Endophenotype” or a Related Term in Their Title Through 2024 (n = 2,339).
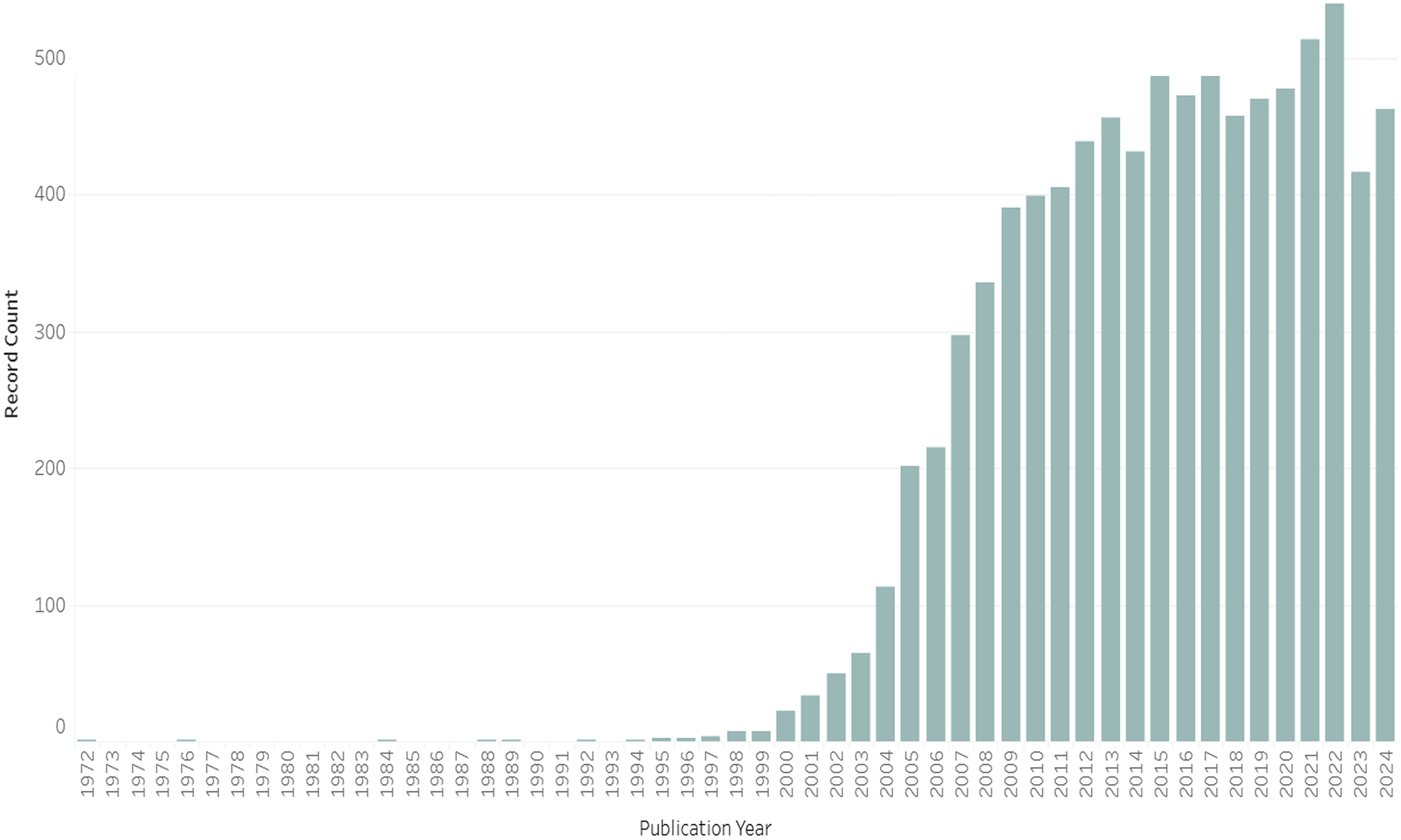
Papers Published Per Year with “Endophenotype” or a Related Term in Their Title Through 2024 (n = 8,748).
Turn-of-the-Century Resurgence
That trickle of papers discussing endophenotypes in the 1990s gave way to a flood following the turn of the millennium. A 2003 paper by Gottesman and Todd Gould in the American Journal of Psychiatry was especially pivotal. Entitled “The Endophenotype Concept in Psychiatry: Etymology and Strategic Intentions,” this agenda-setting piece takes the 1972 concept and updates it for the postgenomic era. Gottesman and Gould (2003, 637) pointed out that many new techniques (“neurophysiological, biochemical, endocrinological, neuroanatomical, cognitive, or neuropsychological”) for discovering endophenotypes had become available, and outlined criteria for identifying valid endophenotypes. 10 The long-term hope was that endophenotype analysis would “be useful for establishing a biological underpinning for diagnosis and classification” (Gottesman and Gould 2003, 641). While their rigorous criteria would be increasingly ignored as part of the dilution and diffusion of the concept, the paper helped bring endophenotype analysis to the fore some thirty years after Gottesman first introduced it into psychiatric genetics.
Gottesman and Gould (2003, 637) situated the explosion of interest in endophenotypes squarely in the postgenomic humbling the field was just beginning to reckon with: Despite the inherent advantages of the concept of endophenotype, the term and its promise lay dormant for a number of years. However, now that multiple genetic linkage and association studies using current classification systems and the development of practical animal models, have all fallen short of success, the term and its usefulness have reemerged… Endophenotypes are being seen as a viable and perhaps necessary mechanism for overcoming the barriers to progress.
Indeed, as an increasingly wide array of advanced genomics techniques and projects continued to flounder, endophenotypes were deployed ever more broadly. The need for an epistemic quilting device that could unite genotype and phenotype was becoming widespread in postgenomic research.
Gottesman and Gould's paper has been cited almost 7,400 times, compared with just over 1,200 for Gottesman and Shields's much broader 1972 book on schizophrenia genetics, and only 254 for John and Lewis's original paper in Science, according to Google Scholar. While “endophenotype” was appearing in paper titles or abstracts a few dozen times a year by 2002, as opposed to just a handful even in the late-1990s, its ascent was only just beginning (see Figures 1 and 2). In the years following Gottesman and Gould's paper, “endophenotypes” took off in the human genetics literature, leveling off at around 100 paper titles and 400-500 paper abstracts and keywords throughout the 2010s.
But those figures likely just scratch the surface. For one thing, authors may not include the term in their title or abstract. A 2003 proposal in Nature Genetics to create a “Human Phenome Project” to parallel the Human Genome Project puts it like this: Disease diagnoses are the tip of the iceberg of phenotypic variation in the population. The HPP can facilitate understanding of diseases (its main goal) by investigating the full range of such variation. For some disorders this task is facilitated by previously identified intermediate phenotypes, or endophenotypes, that are more objectively defined than the disease diagnosis…Therefore, an important task for the HPP will be to promote investigation of the heritability of endophenotypic measures. (Freimer and Sabatti 2003, 16 our emphasis)
Yet, this paper would not appear in Figure 1 or Figure 2. Even in 2003, when the concept was still relatively novel in medical genetics writ large, it could be invoked in the pursuit of far-reaching goals without appearing in a paper's indexed terms. For another, endophenotypes have now been integrated into experts’ taken-for-granted background assumptions. The term has spawned a variety of synonyms and related terms. Already by 2003, Gottesman and Gould (2003, 637) noted that “Other terms with patently synonymous meaning, such as ‘intermediate phenotype,’ ‘biological marker,’ ‘subclinical trait,’ and ‘vulnerability marker,’ have been used interchangeably.” Today, many papers employ the language of “imaging phenotype” (e.g., magnetic resonance imaging), “molecular phenotype,” “disease trait,” or simply “trait.”
Finally, just in the last decade we have seen the concept morph into the verb form—not just something a researcher might discover, but a strategy or mode of discovery that they might use. Thus, endophenotyping and the now-dominant verb form “deep phenotyping” have gained increasing traction. As a recent paper puts it, endophenotypes help address the problem of a “dilut[ed] genetic signal,” and “the application of endophenotypes…has become known as ‘deep phenotyping’” (Greenwood, Shutes-David, and Tsuang 2019, 7, our emphasis).
A Postgenomic Quilt
Today, as experts throughout human genetics routinely use endophenotypes and deep phenotyping, they almost invariably eschew the rigorous criteria laid out in Gottesman's 1972 and 2003 work. As Le Goff, Allard, and Landecker (2021) astutely point out in their historical survey of “epimutation,” scientific terms often lose their solidity as they are adopted by new fields, and they are sometimes held together by their relational status to other objects. For epimutations, Le Goff et al. show how it is held together by a negative definition—it is a mutation not in DNA. In our case, whatever solidity the endophenotype concept had in 1972 or even 2003 has given way to a functional definition as a liminal phenotype that allows researchers to quilt together genotype and phenotype.
Endophenotypes and deep phenotyping have become uncontroversial, even commonplace postgenomic strategies for bridging the genotype/phenotype gap. But they take many different forms. Endophenotypes are now intrinsically relational rather than a certain kind of thing: their power lies precisely in their capacity to sew together previously disjunct categories of human difference. And what's more, that relational logic has been so broadly adopted that the term “endophenotype”—and even the looser “deep phenotyping” or other related terms—need not be invoked at all. So, while endophenotypes are usually highly differentiated, the endophenotype concept has become increasingly amorphous and clearly intermediary, making it a versatile tool that can help address a pressing political need to salvage postgenomic research.
But even as endophenotypes and deep phenotyping are intrinsically relational, they allow for the creation of qualitatively new wholes. As we will see, deep phenotyping might help show that a genetic variant is in fact always penetrant, changing our understanding of its phenotype and the clinical profile of the rare disorder it causes. An endophenotype might help identify more robust associations between genomic variants and a psychiatric condition, but it can also get caught up in broader processes of diagnostic change and “looping” (Hacking 2007) that transform the condition itself. It might help show that distinct disorders share both genomic roots and subtle phenotypic traits, further challenging existing classification systems. Increasingly, an endophenotype target might allow a trial of a pharmaceutical compound or therapy to demonstrate efficacy in a way the disease itself would not. In this way, endophenotypes allow experts to create a postgenomic quilt out of objects of knowledge that had been impossible to conjoin—new wholes that transform the meaning of the parts.
Epistemic Quilting Devices
How can we explain the ascendance of endophenotypes and deep phenotyping in human genetics? Scholars in STS and the sociology of science have long been interested in knowledge production and exchange across fields. Foundational work by Kuhn (1996), Bourdieu (1975), and many others treated normal scientific fields as unitary—bound by shared frameworks, norms, and incentives. This idealized picture has given way to a range of more complex theoretical accounts that focus, inter alia, on the disunity of science (Dupré 1983), “trading zones” (Collins, Evans, and Gorman 2007; Galison 1997) and spaces between fields (Eyal 2010), how certain scientific fields modify “auxiliary hypotheses” to salvage the core of a research program (Lakatos 1970) or subsume opposition to maintain hegemony (Decoteau and Daniel 2020), diverse “situated knowledges” (Haraway 1988), lay expert interventions in research (Epstein 1995), and many other things besides. We will see these and many other dynamics at play in the history of the endophenotype concept. However, none of them can fully explain or unpack its development or postgenomic resurgence.
Perhaps the most apt tool from STS theory is the concept of “boundary objects.” Susan Leigh Star and James Griesemer introduced the term in their groundbreaking 1989 paper to explain how objects of knowledge can facilitate scientific cooperation and exchange even in the face of divergent interests and epistemic frameworks. They explained how boundary objects are “both plastic enough to adapt to local needs and constraints of the several parties employing them, yet robust enough to maintain a common identity across sites” (Star and James Griesemer 1989, 393). Researchers from different fields can then connect and collaborate around an object of knowledge/practice even when they have different ideas about what it is, what it means, and what might be accomplished by studying it. It is easy to imagine how any given endophenotype might function as a boundary object. As we will see, they clearly can enable collaboration and exchange across fields.
Yet, not all endophenotypes are boundary objects. Some are mostly confined to single fields or even single unidisciplinary studies. Others, by contrast, might have a coherent meaning and value across fields. Whether any given endophenotype is a boundary object is therefore an empirical question. As Star herself argued (2010), we should not apply the concept too loosely or broadly. More importantly, we will show how the concept of the endophenotype is another thing altogether. It is more like a device for creating entities that can bridge ontological gaps between genotypes and phenotypes. If anything, it is a conceptual apparatus for (sometimes) producing boundary objects. Likewise, one could also say that endophenotypes play a key role in the “trading zones” that span medical genetics and other fields, or even that they are part of the interlanguage or “parole” that develops between fields (Collins, Evans, and Gorman 2007; Galison 1997; see Navon and Eyal 2014 on trading zones in psychiatric genetics). But that is certainly not always the case, and endophenotypes can never constitute a trading zone by themselves. In sum, we need a different conceptual tool to understand endophenotypes and deep phenotyping in postgenomic knowledge production.
At their core, endophenotypes and deep phenotyping are about bridging objects of knowledge, not fields. We need to explain how they stitch together things—genotypes and phenotypes—that do not easily fit together within a single research program. That is why we call the endophenotype concept and deep phenotyping a pair of epistemic quilting devices. To flesh out our analogy with literal quilting techniques, we argue that the endophenotype concept is akin to the mostly hidden padding or “batting” that, concealed between two outer layers, satisfyingly holds them together even though they were not designed to be adjoined. It makes it possible to sew robust threads back and forth between genotype and phenotype ontologies, allowing for research programs that are internally coherent even as they destabilize their constituent fields. Epistemic quilting devices like endophenotypes are created to forge robust connections within a research program, not for their own sake. They are intrinsically relational. And yet, they allow for the creation of new and qualitatively different wholes.
Textile metaphors have a long history in science and technology studies, especially in feminist STS where they have been used to analyze the inherent relationality of knowledge practices. 11 Recently, a paper in this journal by Thomer and Rayburn (2024) explored how museum and archive staff use quilting metaphors to describe their database curation practices, and how quilting can serve as an analytic lens to understand layered database development. While we are indebted to this scholarship, we use the quilting metaphor in a very specific way, not just to explore the fact that different forms of knowledge intersect with one another, but to highlight a particular sort of intermediary object and explain how it helps tie divergent scientific ontologies together in a single research program without reducing one to the other.
In Figure 3, we see how epistemic quilting devices allow experts to produce novel objects that mediate between two otherwise incommensurable ontologies, A and B, where reduction or unification has failed but the drive to bridge fields within a given research program is strong. Then, in Figure 4 we see how endophenotypes stand in between observations of heritability or the genome on the one hand, and established categories of human disease and difference on the other, making it possible to quilt them together by creating mediated associations between them.
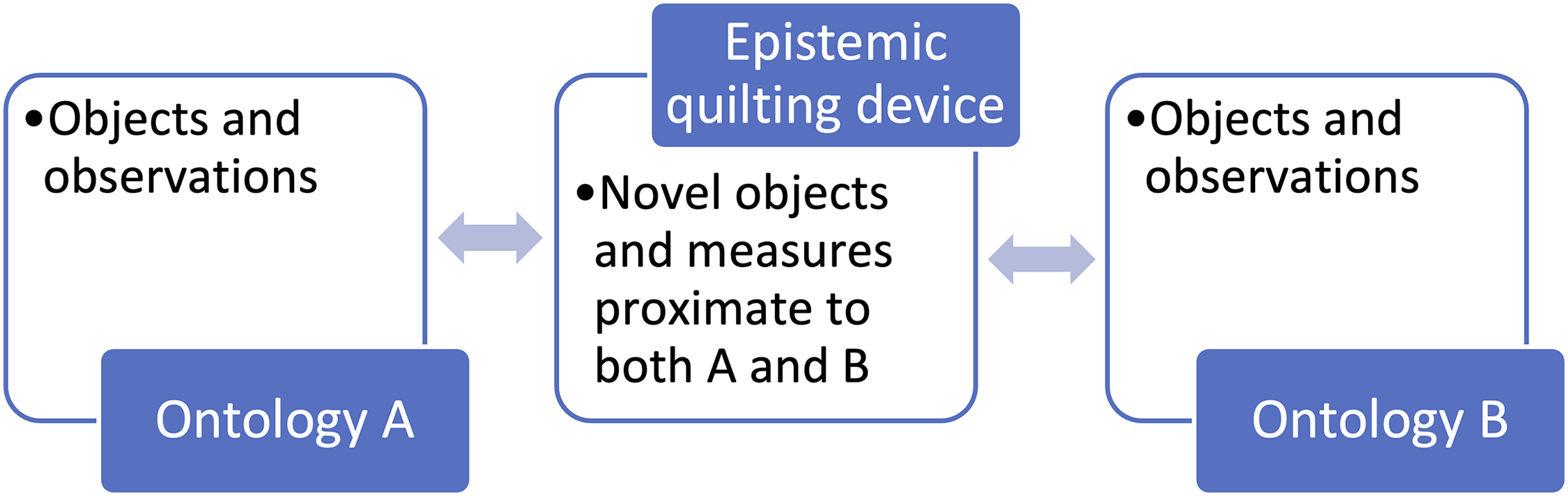
Mediating Role of Epistemic Quilting Devices.
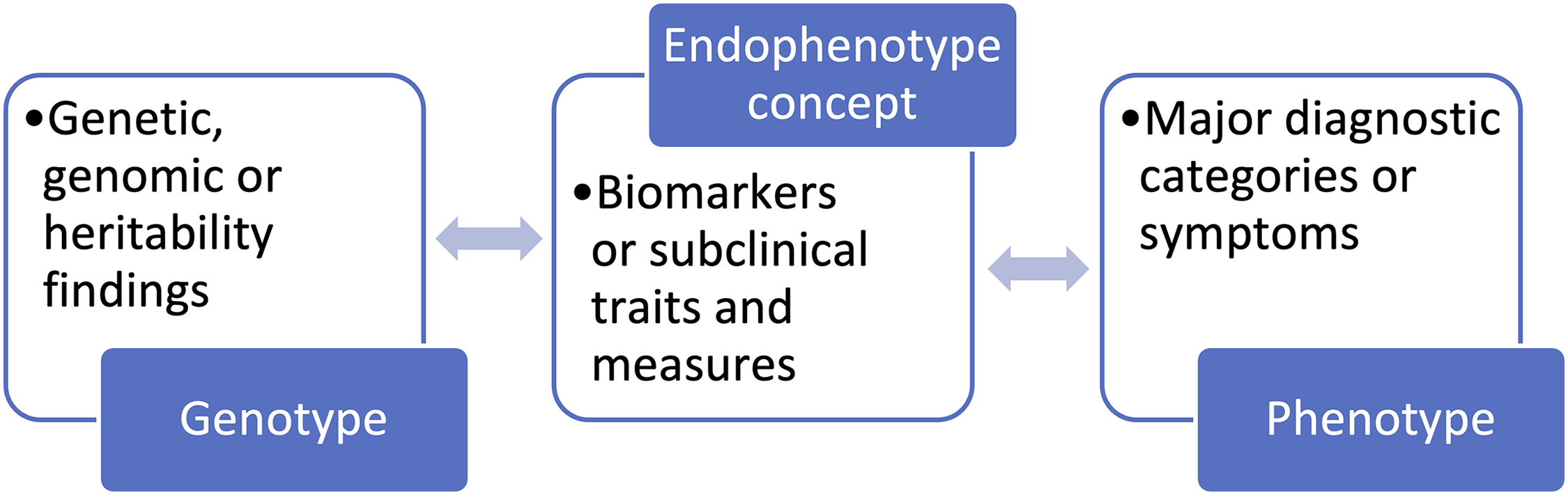
Mediating Role of the Endophenotype Concept.
Epistemic quilting devices are only likely to come into their own when there is an apparent need to bring two disjunct objects or ontologies together. As we saw, the endophenotype concept was mostly ignored in the decades after 1972, only to explode in the postgenomic era as it became clear that most common medical conditions would remain impervious to genetic explanation despite the growing mountain of sequenced genomes at researchers’ disposal. Reduction and unification were both doomed to fail, and a calamitous disunity loomed large. At the same time, too much money, expertise, and hope had been invested in human genomics to accept this kind of partial defeat. Whatever one thinks about the results, endophenotypes provide points of contact through which the pent-up desire to connect genotypes and phenotypes can readily flow. In this way, epistemic quilting devices can respond to historically specific needs and open up expansive new research programs.
Finally, it is important to clarify that it is the endophenotype concept and the practice of deep phenotyping that are epistemic quilting devices. Specific endophenotypes or subclinical traits are highly differentiated, unlike a quilt's batting. However, the endophenotype concept has become so amorphous and so clearly intermediary that it does resemble a quilt's batting, allowing one to sew together elements that would prove too thin or quickly fray and fall apart in its absence. To be sure, just as one does not always need batting to make a quilt, many scientific ontologies do not need epistemic quilting devices to create meaningful ties between them. Indeed, not all genotype–phenotype associations require endophenotypes. Nevertheless, just like batting and the various fabrics one might seek to conjoin in a quilt, the endophenotype concept opens up a universe of new possibilities for tying genotypes and phenotypes together. Indeed, as with a real quilt, endophenotypes can help stitch together different fragments from the same layer that were not easily conjoined—in our case when multiple phenotypes or multiple genotypes are associated with a single endophenotype. Like literal batting, the endophenotype concept is mostly invisible to nonexperts and helps us forget that two outward-facing layers—themselves composed of numerous different pieces—were not supposed to go together, even as we wrap ourselves in the comfort they provide. The quilting device is there to hold it all together.
So, how do endophenotypes and deep phenotyping function as epistemic quilting devices in contemporary psychiatric genetics and allied fields?
Endophenotypes and Deep Phenotyping in Psychiatric Genetics
It is no coincidence that the modern endophenotype concept emerged in Anglo-American psychiatric research. On the one hand, psychiatric classification in the American Psychiatric Association's Diagnostic and Statistical Manual of Mental Disorders (DSM) has always been based on symptoms. 12 On the other, leaders in the field have famously tried and failed to move toward biological classification in psychiatry (Regier et al. 2009; Whooley and Horwitz 2013). Yet despite their efforts, the DSM still specifically excludes diagnoses that are tied to biological markers, be it abnormal brain scans, hormone levels, genes, and so on. 13 So, while psychiatry's vanguard has long sought to turn it into a bona fide field of biomedicine, the way we classify mental illness remains “too transparently socially constructed” (Aronowitz 1999, 31) to avoid withering criticism on multiple fronts. Psychiatry has been wrestling with this old ghost since the dawn of clinical medicine at the end of the 18th century (Davidson 2004). Ever since the late-1970s, the DSM has been framed as a kind of “neo-Kraepelin” wet-nurse that will see diagnostic reliability to one day give way to diagnostic validity, that is, a new biological classification system (Horwitz 2002). But decades of research later, there remains an enormous epistemic gulf between human biology and psychiatric classification. Nevertheless, the proponents of a new biological psychiatry have regrouped from the infamous failure of the DSM-5 Task Force—casting it as one battle in a much longer campaign. In his much-discussed 2010 announcement, Thomas Insel (then director of the US National Institute of Mental Health [NIMH]) bemoaned the failure to establish a biological system of psychiatric classification and charted a new (if long) path forward. Insel was announcing the NIMH's controversial Research Domain Criteria (RDoC), a “project to transform diagnosis by incorporating genetics, imaging, cognitive science, and other levels of information to lay the foundation for a new classification system…patients and families should welcome this change as a first step toward ‘precision medicine’” in psychiatry (2010).
Our goal in this paper is not to rehash scholarly and public debates about RDoC and DSM-5 (Insel 2013; Whooley and Horwitz 2013). Instead, we point to how modern champions of biological psychiatry plan to use endophenotypes and deep phenotyping to stitch together discordant knowledge about human genomes and our understandings of mental illness into a coherent postgenomic quilt. Consider the first of the three references cited in Insel's controversial 2013 blog post: “Endophenotypes: bridging genomic complexity and disorder heterogeneity” (Insel and Cuthbert 2009). The paper notes how “endophenotypes have enjoyed accelerating popularity in recent years due to their potential associations with genomic data to provide a way forward for parsing complex, heterogeneous disease phenotypes such as schizophrenia” (Insel and Cuthbert 988, our emphasis). As Insel put it to the New York Times, “biology never read [the DSM]” (quoted in Belluck and Carey 2013). To extend the analogy, biology inscribes neuropsychiatric phenotypes in an entirely different language than the ones we humans have used to describe ourselves, and endophenotypes allow us to decipher it now that straightforward translation has proven illusory.
A recent perspective piece in Nature Neuroscience captured the apparent promise of these approaches in its abstract (Sanchez-Roige and Palmer 2020, 475): By studying endophenotypes it is possible to both dissect psychiatric disorders (“splitting”) and to combine multiple phenotypes (“clumping”), which can either reinforce or challenge traditional diagnostic categories. Such endophenotypes may also permit a deeper exploration of the neurobiology of psychiatric disorders. A coordinated effort to fully exploit the potential of endophenotypes is overdue.
By moving away from dichotomous clinical diagnoses that “dilute[e] the genetic signal” to phenotypic spectra that can be measured and quantified, researchers are using the endophenotype concept to find much more statistically powerful threads with which to sew together psychiatric and genetic ontologies (Greenwood, Shutes-David, and Tsuang 2019, 7).
It is easy to see, then, why endophenotypes and deep phenotyping came to us via psychiatric genetics. The need for endophenotypes and deep phenotyping is particularly acute. While it first took root in schizophrenia genetics, the endophenotype concept quickly expanded to autism (Abrahams and Geschwind 2008; Sucksmith, Roth, and Hoekstra 2011) and ADHD (Castellanos and Tannock 2002; Pinto et al. 2016), and then onto Alzheimer's disease (Deming et al. 2017) and many other psychiatric disorders. But to be clear, psychiatry is unique in degree, not in kind: there are very few genomic silver bullets to be had outside of fields like cancer and rare disease research. That is why researchers are now turning to endophenotypes and deep phenotyping in medical genetics writ large, touching on questions from dentistry (Wright and Herzberg 2021) to cardiology (Kapur and MacRae 2020) and many other fields besides.
As we will see, psychiatric endophenotypes and deep phenotyping are now deployed by researchers working on radically different scales, from big-data genomics to small-n rare disease research.
Endophenotypes in Big-Data Research
After countless failures to shed light on the genetics of common illnesses, today's big-data methods like GWAS and polygenic risk or index scoring routinely deploy endophenotypes to amplify weak genetic signals. 14 From the “chronotype” or sleep timing preference (Lane et al. 2016) to brain structure (Elliott et al. 2018) and far beyond, these sorts of studies are appearing in leading journals and receiving hundreds of citations precisely because they deploy intermediary objects to stitch together genotype and phenotype more durably than previous approaches had allowed. In this way, the vast complexity of population-level genomics is increasingly mediated via endophenotypes and deep phenotyping.
Endophenotype research is enabled by large biobanks and patient registries. While some are specific to neuro- and psychiatric genetics—or even specific conditions like autism or Rett syndrome (e.g., iPSYCH, the Simons Simplex Collection, and InterRett 15 )—others like 23andMe, All of Us, and the UK Biobank combine data from millions of people and cover a broad range of medical conditions and fields. They are all increasingly replete with endophenotype data. Take a recent paper in Nature on the UK Biobank titled “The UK Biobank resource with deep phenotyping and genomic data,” which describes a dizzying list of traits including imaging, dozens of biomarkers, heart and lung function measures, cognitive and behavioral tests, and physical activity monitoring (Bycroft et al. 2018). It has already been cited over 8,800 times. Across these sorts of programs, comparative approaches relying on shared endophenotypes are used to bolster sample sizes, increase statistical power, and make sense of the ever-vexing relationship between diagnosis and etiology.
These are significant undertakings that entail a twofold clinical reclassification. As Peter Robinson (2012, 777) argued in an agenda-setting piece on the Human Phenome Project in Human Mutation, “Deep phenotyping for precision medicine” will require comprehensive databases in which “disease subtypes” can be identified against the expanse of human diversity. However, as Robinson and others (see Haendel, Chute, and Robinson 2018, for a review) have argued, standardizing such measures and ontologies poses a significant challenge when “Many phenotypic descriptions in medical publications describe the phenotype in sloppy or imprecise ways” (Robinson 2012, 777). Deep phenotyping therefore requires a wholesale rethinking of clinical data inputs, where lump diagnosis has been valued over granular symptoms and idiosyncratic descriptions abound. These kinds of proposals led to the creation and broad adoption of the Human Phenotype Ontology (HPO) for annotating the clinical profiles of patients with genetic disorders and facilitating big-data genomics across a wide range of academic disciplines and medical fields. As HPO's website explains at the top of its “WHO WE ARE!” page: The HPO is being used for computational deep phenotyping and precision medicine as well as integration of clinical data into translational research. Deep phenotyping can be defined as the precise and comprehensive analysis of phenotypic abnormalities in which the individual components of the phenotype are observed and described.
16
In sum, the endophenotype concept has been integrated into the infrastructures that make big-data genomics research possible. Researchers rely on massive datasets containing a variety of subtle traits and differences to amplify weak signals that reverberate between genotype and phenotype. Developing even larger datasets with ever more fine-grained variables has, in turn, become a common priority for funders, researchers, and biobanks alike. Across these projects, endophenotypes and deep phenotyping are widely understood as key to disentangling the genotypic muddle of complex conditions at the population level.
Endophenotypes in Rare Disease Research
Another major line of research takes an almost mirror-opposite approach: it starts with the genotype and then uses endophenotypes and deep phenotyping to flesh out its implications. When it comes to the growing number of cases of “genomic designation,” where the conditions are delineated and diagnosed strictly according to genetic mutations or variants (Navon 2019), the genetic specificity of a disorder may be the very thing that drew researchers to it in the first place—making it “low-hanging fruit” for getting at the genetics of human difference. However, these variants often do not cause the key phenotype of interest: Fragile X might be the genetic variant most strongly tied to autism in the literature (Krueger and Bear 2011), but many or most people with Fragile X do not have autism per se (depending on the diagnostic instrument); the 22q11.2 microdeletion might be the schizophrenia variant (Drew et al. 2011), but most people with 22q11.2 Deletion Syndrome never develop schizophrenia.
Endophenotypes provide the batting researchers need to sew newly robust threads between genetic variants and their most high-profile phenotypes. Just as Gottesman and Shields (1972, 319) used endophenotypes to challenge a key concept in genetics when they wrote “Genes themselves are neither dominant nor recessive,” so researchers today use endophenotypes to challenge notions of incomplete penetrance. A variant might always be penetrant for some sort of spectrum of difference or disability, even if its (endo)phenotype does not always cross the thresholds that govern medical classification (Finucane et al. 2016; Navon 2020). For example, scoring significantly higher than an unaffected parent or sibling on an autism spectrum diagnostic instrument can count as penetrance even if the person with the genetic variant is nowhere near the cutoff for an autism spectrum disorder (ASD) diagnosis per se. The same goes for IQ, even when a person's score leaves them well above the threshold for a diagnosis of intellectual disability, or when their performance IQ indicates a real impairment even as their verbal or full IQ score does not. These kinds of subclinical traits and other endophenotypes can end up playing a key role in the neurodevelopmental and clinical profiles of a genetic condition.
Take the Fragile X mutation: a CGG repeat expansion on the FMR1 gene. Fragile X syndrome (FXS) happens when the expansion exceeds 200 trinucleotide repeats, silencing FMR1 and leading to an FMRP protein deficit. FXS is characterized by very high rates of intellectual disability (especially in males), facial dysmorphism, a range of associated medical issues, and increased risk of autism spectrum disorders (Kniffin 2022, 202). Indeed, the association with autism, alongside a pioneering patient advocacy movement, has made Fragile X syndrome one of the most intensely studied rare genetic disorders in the world (Navon 2019). But Fragile X is increasingly associated with a richly complex clinical/cognitive/behavioral profile and strongly differentiated from idiopathic autism, mostly thanks to endophenotype and deep phenotyping research (e.g., Kniffin 2022; Turk 2011). What's more, endophenotypes have also emerged as the main target of drug development for Fragile X—allowing for the development of an animal model and opening a path “Toward Fulfilling the Promise of Molecular Medicine in Fragile X Syndrome” as the title of an influential Annual Review of Medicine piece puts it (Krueger and Bear 2011, 416-19).
Perhaps even more strikingly, people who have the much more common (around one in 150-300 females) “premutation” expansion of 55-200 CGG repeats in their FMR1 gene and were long thought to be mere “carriers” for Fragile X are now understood as having a mild genetic disorder whose symptoms are visible from birth. But what kinds of symptoms? The answer is written in endophenotypes. While there are also slightly elevated risks of neurodevelopmental disorders and high risks of two late-onset conditions, the Fragile X premutation phenotype is composed almost entirely of mild or subtle cognitive and behavioral differences that resemble FXS itself, but would not even be visible were it not for the language of endophenotypes (e.g., Cornish et al. 2005; Gallego, Burris, and Rivera 2014; Wang et al. 2012).
So, as important as endophenotypes and deep phenotyping may be for highly penetrant genetic differences like Fragile X syndrome, it may be even more transformative for milder variants like the premutation. Indeed, these sorts of approaches challenge the very notions of penetrance and recessivity. A child with a pathogenic variant may appear unaffected by the usual clinical standards and thresholds, only for deep phenotyping to reveal differences that would have otherwise been overlooked. As the senior author of one of the premutation papers cited above told us in an interview about neurodevelopmental evaluation and intervention for people with complex genetic disorders like Fragile X, 22q11.2 Deletion Syndrome, and so on, “It is all about endophenotyping as widely and broadly, as interdisciplinarily, as wide as you can possibly get.”
Discussion
Endophenotypes and deep phenotyping are simultaneously disrupting and suturing. They are at the vanguard of a major shift in medical genetics—one that challenges core Mendelian concepts like dominant versus recessive variants, carrier status, penetrance, and more, by turning toward “subclinical” traits, biomarkers, and spectra of variation. In this way, the endophenotype concept helps refocus the clinical genomic gaze, rendering genetic differences more complex, penetrant, and potentially relevant to a far greater proportion of the population. The postgenomic research program is sewn together, however tenuously, even as its constituent objects and fields are destabilized in the process.
The turn to endophenotypes and deep phenotyping is playing out very broadly in clinical genetics, from the HPO and other infrastructures through to cutting-edge research on rare disorders and pathogenic variants. Take the Clinical Sequencing Exploratory Research Consortium, or CSER (pronounced like the emperor): a massive, National Institutes of Health (NIH)-funded consortium of sites across the United States investigating the rollout of clinical genomic sequencing. As a team of 279 CSER authors puts it in a mission statement in the American Journal of Human Genetics, studies had found that apparently healthy people with pathogenic variants had been “followed up with in-depth phenotyping to reveal that roughly half of the population carrying such variants had subtle phenotypes of underlying genetic disease but were unaware of this” (Green et al. 2016, 1058). In other words, the variants were highly penetrant if you examined their bearers for endophenotypes. They explained how this had profound implications for our thinking about variant classification, penetrance estimation, and future treatments for both rare and common conditions. As the final line of the paper puts it: “with iterative and more in-depth phenotyping… there is an opportunity to define disease and diminished function in entirely new ways” (Green et al. 2016, 1061). Though they never use the precise term per se (and thus would not show up in the charts provided in Figures 1 and 2), it could hardly be clearer that deep phenotyping plays a key role in their vision for genomic medicine. In sum, endophenotypes and deep phenotyping allow researchers to see clinically relevant signs that were invisible to traditional modes of medical classification, strengthening the ties between genotype and phenotype in rare disease research, big-data genomics, and everything in between.
It will be interesting to see if endophenotypes and deep phenotyping can truly take on a life of their own, shorn of their originating intermediary role as a research tool. It will also be important to track how they circulate beyond Western human genetics into postcolonial settings and international research programs—a process that is clearly underway but beyond the scope of this paper. Either way, endophenotypes have traveled a long way from the insect nuclei where they debuted as scientific facts.
This brings us back to the endophenotype concept as an epistemic quilting device. A quilt, of course, is much more than the sum of its parts—the independently unsatisfying batting and the various scraps or offcuts that might otherwise go in the dustbin. We hope that other STS scholars will find the notion of epistemic quilting devices helpful for analyzing how intermediary objects can help to create ties across scientific ontologies and fields. But they can do more than just build connections. Like the batting of a quilt, endophenotypes only have relational value, yet by allowing experts to create new wholes they can be revolutionary. Indeed, they can enable what Foucault (1977, 194-5, emphasis in original) called an apparatus: the assemblage or “system of relations” between various discursive and material elements, “which has as its major function at a given historical moment that of responding to an urgent need.” As we saw, endophenotypes and deep phenotyping are now integral to the postgenomic apparatus, simultaneously suturing and disrupting genotype and phenotype ontologies.
And yet, they did not take off by sheer force of reason. The endophenotype concept was available in something close to its mature form from 1972, when it helped fulfill a need in a small corner of schizophrenia research. But then it lay dormant for a generation, only to explode in the postgenomic era as it became clear that most common medical conditions would remain impervious to genetic explanation despite the rapidly growing availability of DNA data. Reduction or unification was doomed to fail, threatening a calamitous disunity. There was no turn-of-the-millennium discovery or innovation that propelled endophenotypes to become so widespread, important, and disruptive. If anything, the opposite was true: the endophenotype concept shed much of its rigor and specificity. Endophenotypes and deep phenotyping were only taken up in response to broad failure; when there was a compelling need to sew together divergent ontologies and suture the postgenomic tear. That need was scientific and intellectual, to be sure, but also borne of a distinctly ideological urge to salvage the staggering amount of money, expertise, hope, and hype that had been invested in human genomics. As Howard Becker (1982, 249) puts it, “People made quilts, then, because they needed them.”
But whose needs do endophenotypes and deep phenotyping respond to? Just because the endophenotype concept has emerged triumphant in postgenomic research and beyond 17 does not mean we should be pollyannish about it. Moving forward, it will be important to critique the clinical, social, and ethical implications of the turn to endophenotypes and deep phenotyping. When does deep phenotyping provide genuine insight about genetics and human difference, and when is it an attempt to move the goal posts and enhance the statistical power of big-data genomics? What happens when we let associations with genomic variants reshape clinical classification in the name of “precision medicine”? How might mild differences and impairments that might have never risen to the attention of a doctor, teacher, parent, or patient be medicalized, and at what costs to patient wellbeing, healthcare systems, and health equity? Will endophenotypes and deep phenotyping truly help improve patient prognosis, treatment, and care? And for whom? Why are questions of race, ethnicity, and ancestry largely absent in this subfield, especially given their growing salience in human genetics research more broadly? Given the well-documented underrepresentation of BIPOC populations in genetics research, databases, and testing (see Fatumo et al. 2022; Lee et al. 2022), how might the shift to deep phenotyping and subclinical traits exacerbate long-standing inequalities? As deep phenotyping pushes researchers to see carriers as “manifesting heterozygotes” and recast incomplete penetrance as variable expressivity, how many more people will find themselves newly “affected” by a genetic condition? Do we really need to salvage genomics in psychiatry and other fields, and at what cost?
The answer to that final question is partly one of ideology. As Hans-Jörg Rheinberger argued, the stark division between phenotype and genotype that generations of classical geneticists relied on was not “natural” or inevitable but “ideological” (in Canguilhem's sense). It represented a turn-of-the-20th-century epistemic moment in which the sciences broadly (including social sciences) were interested in reductionist models that excavated the hidden structures underlying complex phenomena. This invocation of scientific ideology is not meant to evoke “bad science,” but simply frameworks that draw upon the dominant scientific notions of their times. Nevertheless, such systems produce what Rheinberger calls “hyperbolic objects” which can outlive their ideological origin and become stabilized within new epistemic formations. It is an open question if they are, or were ever, truly fit for purpose. As Rheinberger (2013, 489) puts it, “in a coming age of postgenomics the distinction [between genotype and phenotype] may well be pushed to the verge of its dissolution as a research-driving categorial partition.” The endophenotype concept is a response to that same tension—and one that may allow for reformulation rather than dissolution. At the same time, there is also the need to maintain all the hope, human capital, and “biocapital” that has been invested in human genetics and genomics (Nelkin and Lindee 2004; Sunder Rajan 2006), even as proven environmental and public health initiatives struggle for funding (see Tabery 2023 for the latest in a long line of forceful critiques). Ilana Löwy (2022), building on Rheinberger, has identified the promise of drug development as a driving ideological force in postgenomic research, justifying continued investment despite decades of disappointing results. Endophenotypes and deep phenotyping allow these and other research programs to continue apace, repurposing a genotype-centered ideology across a range of academic and commercial fields.
In postgenomic research, incommensurability and failure have loomed large. The endophenotype concept was rehabilitated and deployed by experts across a wide range of postgenomic research programs to quilt together two otherwise-irreconcilable modes for making sense of illness and disease—a batting hidden beneath the surface of high-profile science. But their relationality does not render them trivial: by standing in between genotypes and established phenotypes, endophenotypes and deep phenotyping are enabling the creation of radical new visions of human difference.
Footnotes
Acknowledgments
We are very grateful for invaluable feedback from Hannah Landecker, Jenny Bangham, Emily Merchant, Gil Eyal, Cathy Gere, and Ilana Löwy; audiences at the Social Science History Association 2022 annual meeting, the Sciences Po médialab, the Max Planck Institute for the History of Science, and École Normale Supérieure; and the anonymous reviewers and ST&HV editors.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
