Abstract
In view of the end of the golden years of the Norwegian oil economy, ocean genetic resources are being advertised in policy environments as holding great potential for the future of the country. Microbes have increased in popularity as promising agents of the Norwegian new bioeconomy, as advances in gene sequencing technologies and genomics have made them more accessible. This paper examines the turn toward digital sequence data in bioprospecting to inquire about its political implications. It draws on a combination of empirical materials to describe the infrastructural work that goes into extracting microbes from their in situ locations in arctic and subarctic waters to in silico collections and databases. I argue that in that infrastructural work, microbes may become both more promising and more elusive public and political matters. As biodiversity is turned into data, bioprospecting appears as less dependent on material samples, which may ultimately render policy frameworks for biodiversity governance obsolete. The shift toward big sequence data in bioprospecting entails shifts in how promise is attributed to biodiversity, which life forms appear to be more promising, and how such life forms come to appear as public good.
An Ocean of Opportunities
“Data is the new oil” has become a global metaphor, one that perhaps works in a more literal fashion in Norway. Since the 1970s, Norwegian public investments have heavily supported ocean exploration and the development of an infrastructure for oil extraction (Ryggvik 2018). State-driven oil extractivism has turned Norway into one of the wealthiest nations in the world. Today, the Norwegian state turns to the ocean again, hoping to move away from the oil economy and toward a new bioeconomy. The ocean is again home to expedition vessels, divers, and underwater prospecting technologies, this time seeking not oil but genetic materials. Marine bioprospecting appears as a form of infrastructural work, bringing together stories of the past and present with the promise of a still wealthy future (Appel 2018).
Icebreaker “Kronprins Haakon” (Crown Prince Haakon) in bioprospecting expedition. Picture published in Svalbardposten August 27, 2019—Picture by Ole Magnus Rapp.
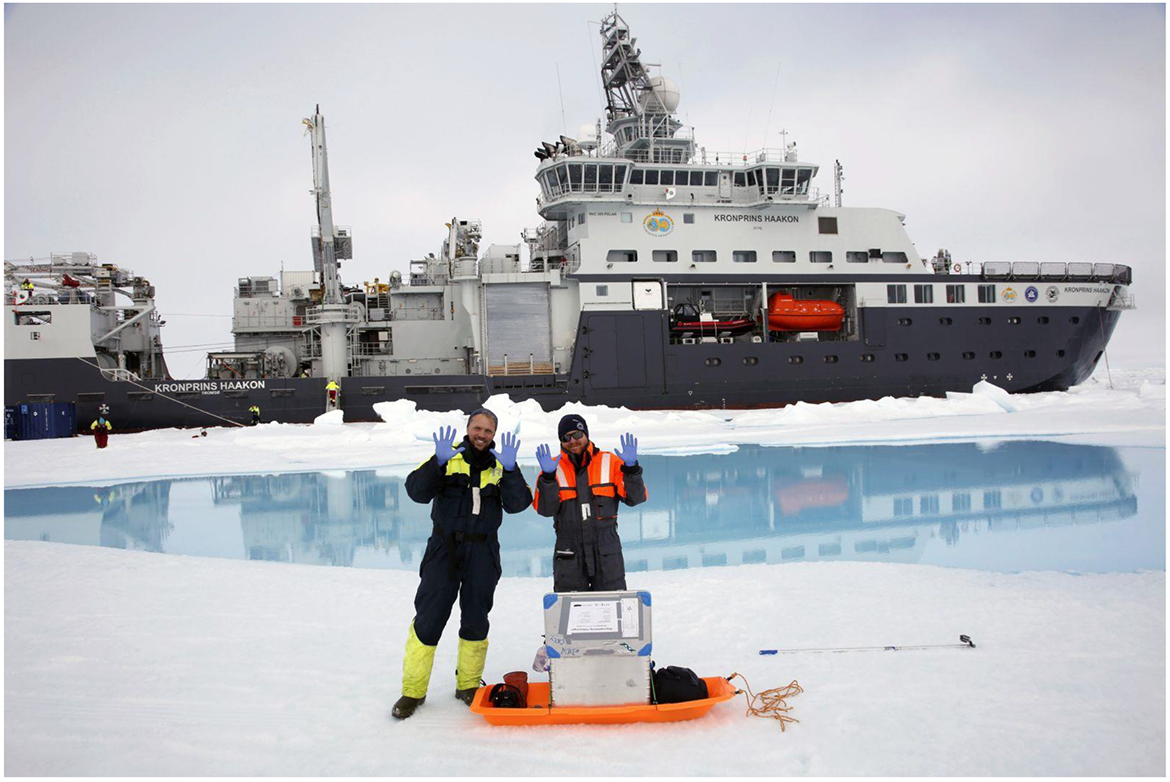

In Norwegian policy and research environments, the technical capacity to access and exploit genetic resources from the sea is advertised as opening a whole new ocean of opportunities, right at the end of the golden years of the oil economy (Norwegian Ministry of Education and Research 2011; Partnerskapet Bioverdi 2014). In policy discourse, big data—including sequence data—is suggested as holding great innovation potential (Research Council of Norway [RCN] 2018). This way of talking falls between metaphor, hope, and speculation, serving to justify significant public investment in biotechnology infrastructures and capacity building. Scholarly work in Science and Technology Studies (STS) has accounted for the speculative and promissory character of the bioeconomy in its various versions (Cooper 2008; Fortun 2008; Sunder Rajan 2006). What is particular about the Norwegian case may be the extent to which the state takes on the roles of both an investor and a representative of the public good and how, in practice, this generates access to national genetic resources. The efforts to build national health biobanks in Norway and their legal intricacies have been the focus of recent scholarly analysis (Befring 2020). In this paper, I turn attention to the extraction of marine genetic resources, particularly to recent efforts directed at bioprospecting arctic and subarctic waters. I inquire into how promises are performed through such infrastructural work and how public matters are configured in the process. In so doing, I address future-making as practice (Delgado 2013; 2016), attending to how scientists work to identify the prospects of marine microbial biodiversity. Previous work in STS has shown how expectations of the future shape social action in the present, particularly political decision-making (Borup et al. 2006). Inspired by this work, I look specifically at how infrastructural work is performed to realize promise (Mackenzie 2013). I draw on recent scholarly work on promise and infrastructures (Anand, Gupta, and Appel 2018) and how publics and public issues are performed through infrastructuring (Anand 2017; Barry 2016; Kelty 2009) to explore how bioprospecting is configuring Norwegian marine microbial biodiversity as a promising public good and a public issue.
By attending to the turn toward the use of sequence data in bioprospecting that has taken place since the early 2000s, I provide insights into how oceanic genetic materials are coming to be seen as valuable national resources. As genetic materials are reconsidered as data, marine life forms such as microbes and algae have taken on new potential. I focus on the case of the arctic and subarctic marine microbial biodiversity in Norway to ask: How have these microbes become reconfigured as political and public matters in the turn toward sequence-based bioprospecting? In addressing this question, this paper gives an account of extractivist practices as performed by scientists in bioprospecting facilities in three major cities along the Norwegian coast: Bergen, Trondheim, and Tromsø. 1 This paper describes how scientists work to gain access to deep-sea sites, archaic microbial metabolisms, difficult-to-cultivate life forms, and novel genetic sequences. In the infrastructural work that goes from extracting material samples to digital mining, microbes become both more promising and more elusive public and political matters. As this paper describes, turning microbes into sequence data is generally seen as a process of dematerialization that produces an augmented sense of promise among scientists. In parallel, the process of extraction—from on site to in silico collections—produces an effect of deterritorialization that is difficult for a state to handle. The infrastructural work that the state is supporting and that scientists are developing to realize the promise of microbial life ironically also serves to set that promise in suspension. By looking at the case of Norway, I argue that the shift toward big sequence data in bioprospecting entails shifts in how promise is attributed to biodiversity, which life forms appear to be more promising, and how such life forms come to appear as public good.
Making Nature Public Again: Digital Sequence Information (DSI) and How the Nagoya Protocol Has Not (Yet) Been Enforced in Norway
Entering into force in the early 1990s, the United Nations Convention on Biological Diversity (CBD) is the main global policy framework for biodiversity governance. It acknowledges states’ sovereignty over the biodiversity found within their jurisdictions. It also places responsibility on states to ensure a regulated and fair extraction and exploitation of genetic resources. The CBD established guidelines for the fair and equitable access to and sharing of benefits from the utilization of genetic resources (referred to as Access and Benefit Sharing [ABS]). In 2010 the treaty body passed the Nagoya Protocol, establishing specific and legally binding standards for national ABS regulations. In the Southern latitudes, the so-called megadiverse countries have typically contained the most biodiversity hotspots and thus also have appeared as the most common sites of bioextractivism and biopiracy (Greene 2004; Laird 2002). In CBD debates, these countries typically appear as biodiversity “providers,” while Northern countries with developed biotechnology industries appear as “users.”
In When Nature Goes Public, Hayden (2003) provides a comprehensive ethnographic account of how the extraction of genetic materials was regulated in Mexico to comply with the CBD. In Mexico, as in most Southern biodiversity hot spots, plant biodiversity has been prioritized as holding great potential for pharmaceutical purposes. Indigenous knowledge and practices of plant use appeared to be a gateway for private capital to capitalize on these promising molecules and chemical reactions. Science—particularly ethnobotany—played a crucial role as a “translator.” As recounted by Hayden, researchers in public universities acted as intermediaries to render visible potential bioprospecting targets for industry partners. In this paper, I explore the actions that bring that kind of visibility to the promises of biodiversity today. I am particularly interested in how that visibility is produced in extractivist practices and the kind of access enabled by particular forms of infrastructural work. ABS contracts, Hayden (2003) argues, came inscribed with “a promise of redistributed value” (p. 3). What happens when the means for producing “access” to biodiversity change, and when that change is promoted by the state itself?
In the last two decades, bioprospecting practices have changed significantly, with rapid advances in gene sequencing, software for data storage and mining, and gene editing and assembling technologies (CBD Ad Hoc Technical Expert Group on DSI on Genetic Resources [CBD AHTEG] 2018; Scott and Berry 2018). A change in approaches to life is occurring; organisms and phenotypes as research objects have increasingly given way to genetic, genomic, and metagenomic information. The policy term “Digital Sequence Information (DSI)” has been proposed to capture this shift in use of sequence data. The CBD and its Nagoya Protocol have been grappling for several years now with whether DSI should count as “genetic material” and thus be covered by these legal agreements. DSI lacks in definition (Aubry 2019; Laird et al. 2020; CBD AHTEG 2020b), and it is a highly disputed term. Debates are playing out between provider and user countries. What is at stake is the very definition of “biodiversity,” with competing interests on how much the term should cover.
It is difficult to situate Norway within provider/user categories. It is a Northern country but lacks a strong biotechnology industry. Yet, in contrast to countries such as Mexico, Norway is not “megadiverse,” the issue of indigenous knowledge has not been given much consideration, and plant biodiversity has never really attracted much attention as an issue of public concern. Perhaps more than in other places, the state has remained a dominant actor in bioprospecting activities—both as an investor and as a representative of the public good. Norwegian scholars specializing in ABS have suggested a “cost-sharing” principle that would oblige potential beneficiaries to compensate for infrastructural costs (Rosendal, Myhr, and Tvedt 2016). However, although Norway ratified the Nagoya Protocol in 2014, no concrete administrative practice for the effective and fair regulation of the extraction of genetic materials is in place yet, despite two attempts by the government (in 2012 and 2017) to pass an administrative order for the “extraction and utilization of genetic materials” (Norwegian Ministry of Trade, Industry and Fisheries 2012; NMTIF 2017). Today, debates on DSI within the CBD and Nagoya Protocol are pushing Norway to decide whether to consider sequence data as part of its biodiversity.
The following sections describe in more detail how the turn to sequence-based bioprospecting is challenging the view of biodiversity in Norway as not much of a public issue. With enhanced access to marine life forms, policy actors are seeing them as strategic national resources in moving Norway toward a postpetroleum economy. Among this marine life, microorganisms have received special attention. In key scientific conferences, 2 scientists present microbes as encoding a large range of functions with potential industrial applications. Some of these potential uses are newer and greener, such as plastic degradation and the production of biomaterials, while others are more classic, such as the production of novel antibiotic compounds. Sequence-based bioprospecting is supported by the state as it is expected to make the screening and identification of such functions more efficient through the infrastructural work of data storage, access, and mining. The next section provides insight on how microbes have been infrastructured as a public issue in Norway.
Building an Infrastructure, Realizing Promise
Norwegian public efforts to develop infrastructures for bioprospecting must be situated within the state’s larger strategy for developing biotechnology research and innovation. Public investments are often justified in terms of a future public good, with the aim to move toward a new bioeconomy (RCN 2018). As I describe below, in this context, infrastructural work is fueled with promise and targeted to realize that promise.
Map of Norway. The cities where fieldwork was conducted are highlighted. Accessed August 31, 2021. Source: http://mapsopensource.com/norway-map-black-and-white.html.
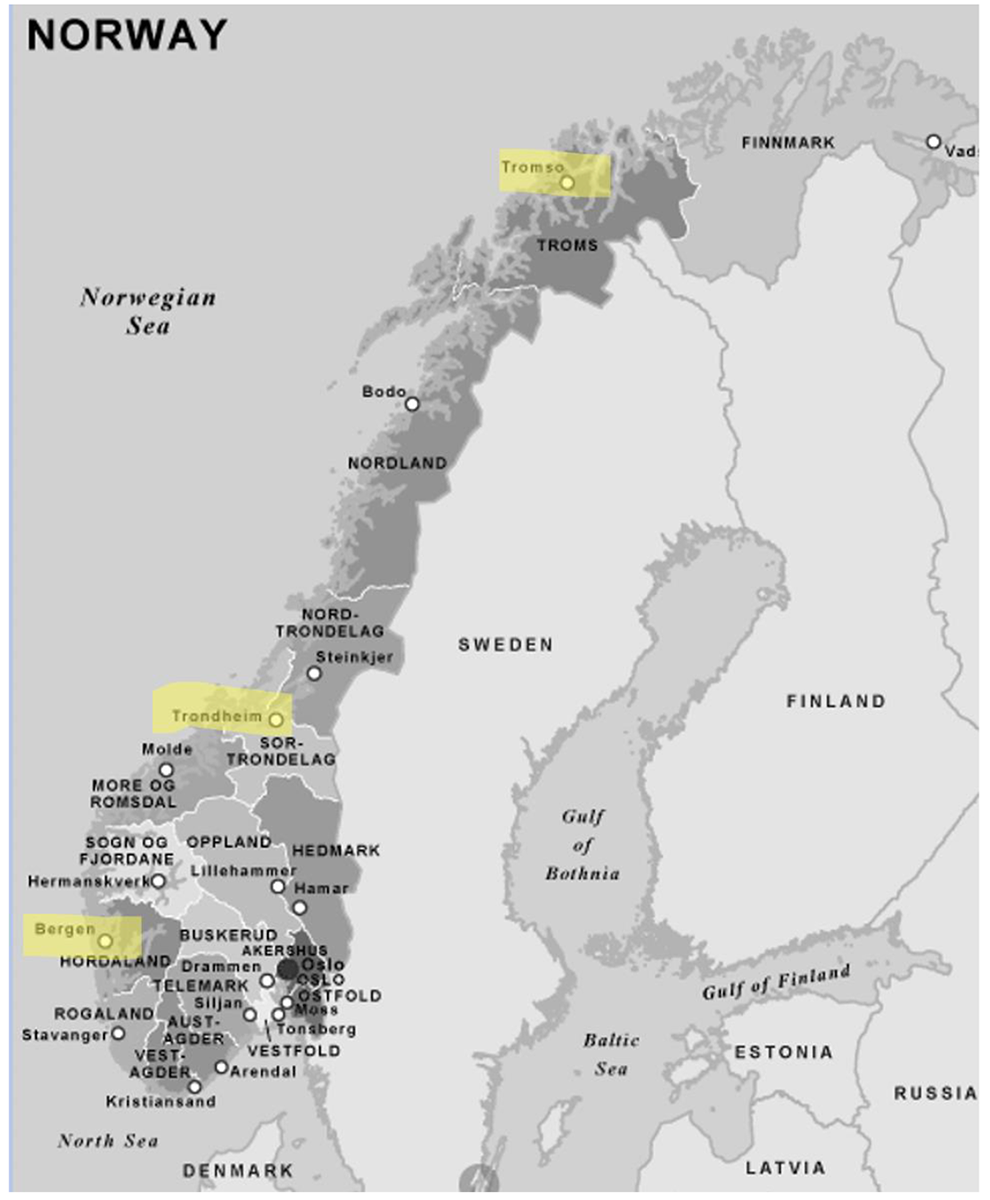
To meet the objectives of the CBD, in 1998, the RCN launched the program Biodiversity: Dynamics, Threats and Management. Since 2001, bioprospecting has been given high priority (Rosendal, Myhr, and Tvedt 2016); the government has supported a number of platforms for bioprospecting, promoting a redistribution of these activities across the country. The arctic city of Tromsø was chosen to host Marbank, the national marine biobank, and Marbio, an associated screening platform for the identification of commercially interesting compounds. The Arctic Bioanalytical Platform was also established at the University of Tromsø. From 2007 to 2015, the RCN financed the Centre for Arctic Marine Bioprospecting (MabCent) in the same city. In Trondheim, the Norwegian University of Science and Technology became involved early on in bioprospecting activities, collaborating on projects with SINTEF, a research organization with a strong industrial orientation that depends to a large extent on public funding. Today, SINTEF runs a unit specifically dedicated to industrial microbiology. At the University of Bergen, the RCN financed the Centre for Geobiology, a national center of excellence. This center has worked in collaboration with the KG Centre for Deep Sea Research (2017-2021), financed on philanthropist donation. These are not the only ones, but they are the main institutions for marine bioprospecting in Norway.
The first sight of a visitor to one of these facilities would likely be wet lab tools and computers, sitting alongside each other. In the last two decades, both microbiology and bioinformatics have expanded in Norway. Two of the facilities I visited were advertising new professorship and research positions for people with backgrounds in both fields. Researchers with these specialties as well as genomics, metagenomics, and systems and synthetic biology have been recruited nationally and from abroad to build national capacity. Increased attention to these fields has occurred at the same time as shifts in the RCN’s research programs, from the first biodiversity program’s emphasis on organisms and ecosystems, to the Functional Genomics program’s emphasis on DNA (RCN 2003), to the BIOTEK 2021 program’s emphasis on big data biology (RCN 2018). In biodiversity research, the turn to big data implies a reconsideration of ways of seeing (Daston and Galison 2007), classifying (Sommerlund 2006; Waterton, Ellis, and Wynne 2013), and even collecting biodiversity (Bowker 2000). In bioprospecting practices, this turn comes with a move from microscopy and taxonomic and chemical analysis to a focus on genetic sequence structures, such as gene networks, gene clusters, and genome maps. This information is stored in and retrieved from gene sequence libraries and databases. This entails a different way to see potentials in life as well as a different idea of how those potentials can be realized (Mackenzie 2003), often in redistributed infrastructural work.
A need to map, decode, and explore ocean biodiversity is expressed in key policy documents such as the Norwegian National Strategy for Bioprospecting for 2009-2021, Marine Bioprospecting—A Source of New and Sustainable Wealth Growth (Norwegian Ministries 2009). Norwegian marine biodiversity is presented here as holding great potential because it remains largely unknown and is “unique.” The government’s second draft for an administrative order to regulate the extraction and utilization of genetic materials in Norway follows the same rhetoric. It postulates that “peculiar genetic codes typically occur as part of adaptation to different living conditions” (NMTIF 2017, 5). The uniqueness of marine oil reservoirs and arctic waters is emphasized as they are qualified as “extreme environments” (ibid.), where most commercially interesting genetic functions, metabolisms, and forms of biochemistry are likely to develop. Beyond Norwegian policy documents, the Arctic is typically presented as not just unknown and unique but also subject to rapid environmental changes and therefore at a higher risk of biodiversity loss (Leary 2008; Dempsey 2015). This may almost turn its exploration into a public duty for both conservation and exploitation. The need for “reading” the ocean and decoding its potential is in such rhetoric endorsed with a sense of urgency. Biodiversity prospects are projected through a rhetoric that aligns remote and rare environments, in situ adaptability, economic potential, and the need to gain access: Genetic material in Norwegian nature has not been fully mapped, but it is clear that in general, biodiversity in Norway is not as extensive as in countries with warmer climates. On the other hand, Norway has habitat types and natural conditions which in some areas have led to the development, on an international scale, of peculiar diversity with a high degree of specialization. This increases the possibility of finding genetic material with very special properties…. There is good reason to assume that several of these marine organisms and microorganisms have properties which can be exploited and form the basis for various products and processes in a number of industrial areas. (NMTIF 2017, 5; author’s translation and emphasis added) Technological developments allow for ever easier access to and deeper insight into genetic material from publications which are open to all, so-called access in silico. Developments in DNA sequencing have increased interest in this type of access. (NMTIF 2017, 4)
Extractions: From Local Sites to Digital Databases
Extraction 1: Mining the Deep Ocean
In 2018, I attended the annual conference of the Centre for Digital Life Norway in the city of Bergen. This center is an initiative of the RCN to promote big data biology as part of a transition to a new Norwegian bioeconomy. A principal investigator at the Centre for Geobiology and leader at the KG Centre for Deep Sea Research at the University of Bergen gave a keynote entitled “Biodiscovery and Bioprospecting of Norwegian Deep-Sea Hydrothermal Vent Systems” (Steen 2018). 3 In her talk, she described her group’s research on bacteria and viruses as frontier research that will provide access to the still largely “unknown” arctic marine biodiversity. She showed a video featuring the highly advanced and expensive technology her team uses to collect samples from the deep-sea floor, often in remote and extreme environments such as deep-sea hydrothermal vent systems (Figure 1). The video showed a tank developed by her team to collect up to fifty liters of seawater and microorganisms at 2,400 feet. After microbial samples are sequenced, the resulting genomic and metagenomic information can be accessed through PubMed and in common formats for large databases such as GenBank.

Video shot showing sampling technology. Source: Steen (2018, 00.06:33).
In this conference keynote, the unexplored and promising conditions of the arctic marine environments were repeatedly emphasized. The Arctic hydrothermal vents were particularly presented as rare and remote environments to which archaic and very special forms of microbial life (archaea) had adapted since ancient geological times. Discovered in 2005 by scientists at the Centre for Geobiology, the Norwegian hydrothermal vent systems are considered unique, as highlighted by the keynote address: “A lot of this biodiversity has not been seen before, it is really novel biodiversity, and it is also really novel biochemistry…. Since many of these (micro)organisms are very close to the roots of the tree of life, it could be very interesting to look into how different enzyme families have evolved…. This is a really nice database to look for new enzymes for new value chains” (Steen 2018, 00:21:23).
In the keynote, the commercial potential of this type of archaea was argued in terms of their temporal and spatial remoteness. These microorganisms are close to the origins of life, with archaic metabolisms that have adapted to the special geological conditions of the vent system, specifically to very high temperatures, low pressure, and lack of light. Their capacity to function in such extreme pressure conditions has led scientists in Bergen to try using enzymes from these archaea in bioreactors for industrial processes. Determining how such ancient metabolisms work is a matter of decoding microbial secrets (Ferrer et al. 2019) to understand how certain gene structures code for certain functions (i.e., the capacity to resist high pressure or to produce a certain chemical compound).
These Arctic archaea are categorized as extremophiles, a type of microorganism with the capacity to adapt to extreme environments. First recorded in the 1980s, microbial extremophiles are often classified for what “they can provide humans” (Helmreich 2009, 99). Their rare metabolisms and biochemistry are seen as opening a whole new range of applications, and so extremophiles have from early on been presented as promising agents of a new bioeconomy (Cooper 2008). 4 Interest in extremophiles has grown in parallel with the development of gene sequencing. They are often described as being in close interdependency with their environment, including genetic interactions with other microorganisms. 5 Extremophiles could thus be better accessed and studied once sequencing technology enabled simultaneous reading of large amounts of genetic information beyond single genomes. Today, metagenome screening and analysis are common practices in bioprospecting. But what is the final product of this work?
As I was told in an informal talk during my visit to Bergen, these scientists consider their job done when they openly publish the sequence data for their samples of archaea and other microorganisms. Helmreich (2009) has argued that, “with the application of gene sequencing to consortia of marine microbes, the ocean is becoming something to be textually scanned, deciphered, read, at ever higher resolutions” (p. 51). Further, this infrastructural way of seeing the ocean often ends up with data being deposited in databases. Data are then largely available to bioprospectors for further screenings and multiple “rereadings.” As one of the researchers in Bergen told me, they expect that by publishing their data openly, others “who know how to look” and know how to see bioprospecting potential will be able to further use this data. Published sequence data often include partial and selected information about the environment and site of collection. In the context of the CBD and Nagoya Protocol, this is called “subsidiary information” (CBD AHTEG 2020b), and it is still of unclear importance for gene expression (i.e., for microbes to express functions). The scientists I engaged with acknowledged the technical difficulties of annotating environmental and ecological information. 6 What is interesting here is how the in situ origins of these microbes—their Arctic origin—endow them with specific prospects as genetic resources. Yet simultaneously, sequence-based bioprospecting practices seem aimed at removing, “washing away,” and replacing that environment. The next section further elaborates on this point.
Extraction 2: From Less DNA to More Information
Sequence data in bioprospecting enhance the scope of what can be achieved, as explained in the keynote at the Digital Life Norway Conference: “When we started in 2007, the sequencing technology didn’t allow us to make metagenomics from all the samples that we collected from these systems…because we really had to have high biomass…. With the new sequencing technologies, we really need to feed much less DNA. We can now start doing metagenomics on samples that we couldn’t do before” (Steen 2018, 00:10:25).
Across different bioprospecting facilities, one topic that repeatedly came up in conversation was the key challenge in traditional bioprospecting of ensuring the availability of large, good quality material samples. In the Arctic city of Tromsø, the mission of Marbank, the national marine biobank, has been to ensure access to high-quality samples. 7 Through Material Transfer Agreements, Marbank provides its users with good quality genetic material. Information on the specific location where the sample was taken is important; if they run out of specific samples in the collection, they have to go back to the site of extraction and collect a new sample to replace the old one.
In the Marbank facilities, the samples collected in vessel expeditions are first cleaned up to remove sand and other waste and then torn apart, their parts separated, selected, and prepared into extracts (Figure 2). In traditional chemical bioprospecting, those extracts undergo chemical analyses through isolation of chemical compounds. Isolates are then screened for interesting forms of bioactivity. For instance, one could insert cancer cells with an extracted compound into a multiwell plate (Figure 2) to see whether a reaction might indicate possible anticancer properties in the isolate.
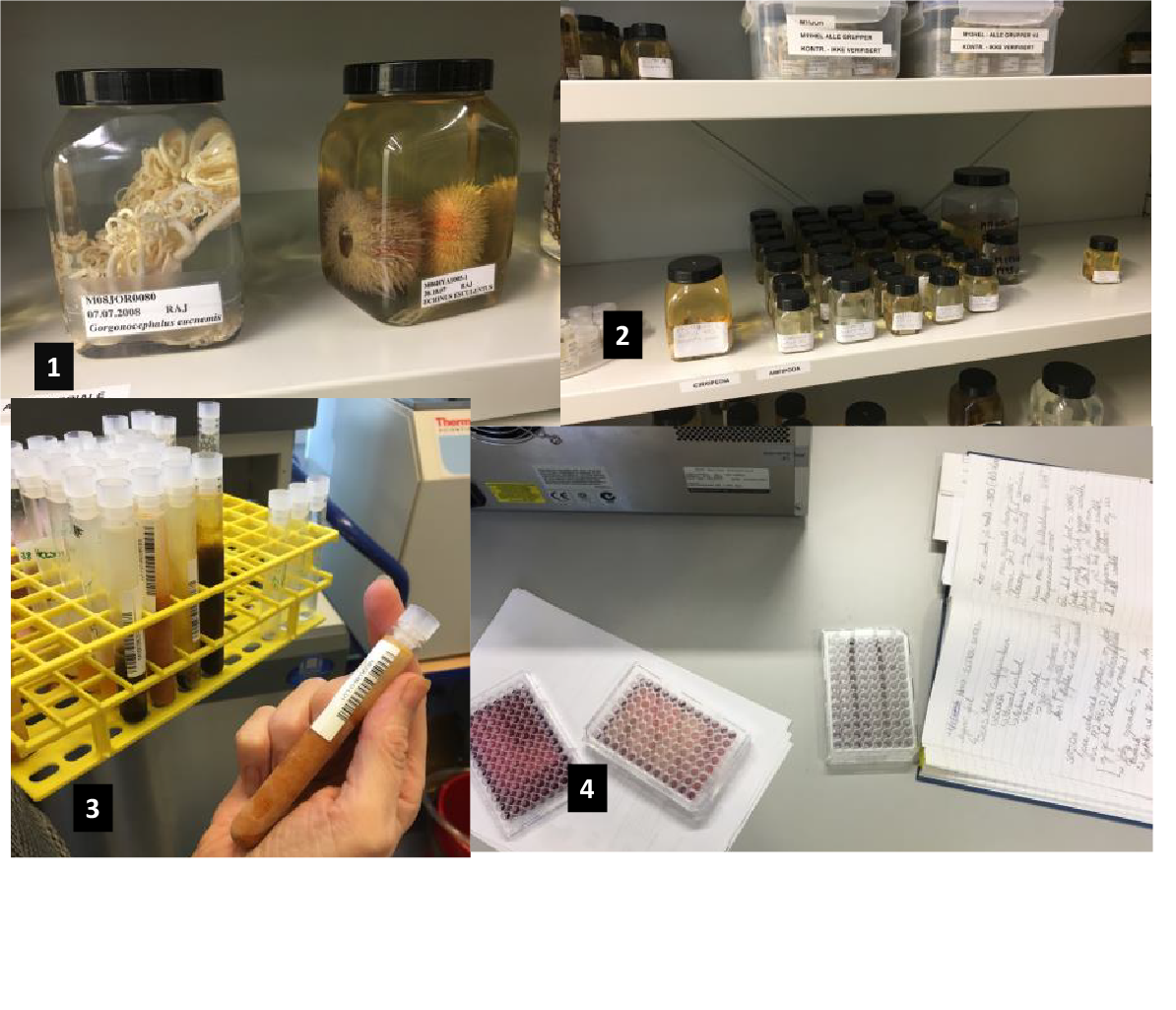

Marbank collection and facilities. Photos taken by the author.
In contrast, sequence-based bioprospecting enables a different way of seeing by looking directly into genetic structures and sequences. Advances in gene sequencing technology, as the quote introducing this section indicates, enable the extraction of more information from smaller samples. Metagenome analysis amplifies the scope of what can be seen, as information of all the DNA contained in a small sample—say, a drop of marine water—can be extracted at once. This kind of analysis reduces not only dependency on large samples but also the need to preserve the samples, as sequencing can be done quickly. During my fieldwork, I was told with enthusiasm how the latest advancements in gene sequencing enable in situ sequencing. That would make the “scanning” and “reading” of the ocean cheaper and also lower dependence on material samples. An important advantage in this regard, I was told, is that there is less need to return to the original site of sample extraction for new samples once the sequence information has been extracted and stored in databases. As sequence data, the collection remains easily available, no matter how many times it is accessed. The next section further elaborates how microbes are recontextualized from in situ to in silico infrastructures and how, in this translation, they are invested with augmented promise.
Extraction 3: From Difficult Microbes to Easy Hosts
Microbes have not been easy laboratory creatures. To characterize them, scientists have traditionally depended on microscopy. In chemical-based bioprospecting, scientists need to make microbes produce the bioactive compounds in question in order to run screenings, which is not necessarily easy to do. A main problem in traditional bioprospecting of microbes has been that they tend to resist cultivation in agar laboratory media. Marine microbes in particular are difficult to cultivate out of marine water. For example, since 2003, the Trondheim facility has been curating a collection of marine actinomycetes bacteria from the Trondheim fjord. The researchers explained to me that actinomycetes are a very common kind of bacteria and long been researched for commercial purposes; 70% of known antibiotics are naturally produced by actinomycetes, mainly extracted from soil. However, marine actinomycetes remain largely unknown, less than 5% of these microorganisms are culturable with currently existing methods. 8 Advances in DNA extraction and sequencing technologies have shifted this situation, enabling cultivation-free practices. 9 Sequence reading technology produces a sort of visibility different from microscopic optics. Visualized as gene clusters, genome maps, or phylogenomic trees, sequence information appears as positional—one can “see,” for example, where particular sequences are placed in whole genome structures.
Such informational display (Figure 3) makes it possible for scientists to identify sequences that could code for interesting functions but are not expressed in natural environmental conditions. Researchers in Trondheim described such nonexpressed sequence information as “silent genes,” which they work to express. They are developing a bioprospecting screening platform that enables the identification of sequences encoding promising functions (even if silent), to then trigger their expression. Both in Trondheim and Tromsø, scientists in bioprospecting facilities told me that triggering this type of gene expression is one of their main challenges.
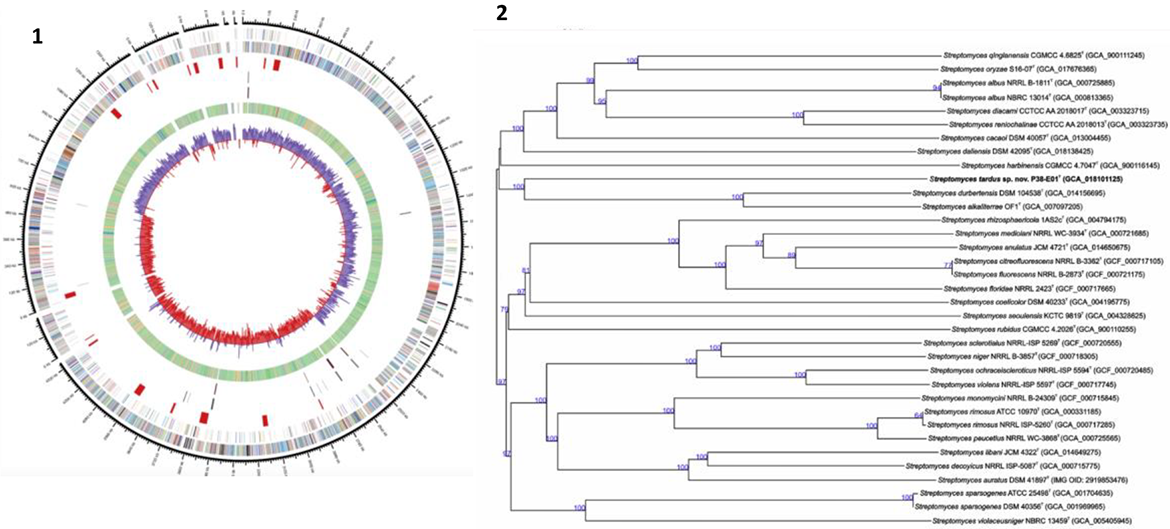
(1) Bacterial genome map (Schouw et al. 2018). (2) Phylogenomic tree from actinomycetes (streptomyces) collected at the Trondheim fjord (Králová, et al. 2021).
Advancements in gene editing and synthetic biology make it possible for scientists to trigger gene expression by transferring DNA into a different host. Often a well-known and domesticated lab microorganism host is chosen, such as E. coli. The Trondheim facility is working with an archaea collection mined from the subsurface of an oil reservoir, which is also characterized by these scientists as an “extreme habitat” (Lewin et al. 2017, 183). 10 A lead researcher from the team explained to me that they use E. coli as a host to facilitate the expression of potentially interesting sequences but frequently experience problems of incompatibility (the desired genes are not expressed by the host). They use predictive software to calculate rates of incompatibility, combined with gene assembling methods to transfer the most promising gene candidates to the new host environment. If cultivation media is used to control the time of life in the laboratory (Landecker 2007), here the digital media is used to activate and optimize archaic metabolisms. Combining remote and archaic microbes with domesticated lab bacteria was presented as a way to trigger ancient and otherwise silent properties seen to hold bioprospecting potential because of their novelty.
Attending to scientists’ work across various facilities in Norway, one can easily argue that the promise attributed to microorganisms relies on a series of recontextualizations (Thacker 2004) from local environments to informational media and domesticated hosts such as E. coli. Sequence-based bioprospecting appears as an extractivist and future-oriented practice that draws on radical recontextualization. In Latour’s ethnography on the Amazonian forest, the environment was the “context” to be removed; soil samples from the forest were removed from those environments, isolated, and classified toward the production of static biological facts (Latour 1999). As a future-oriented practice, bioprospecting configures the local environment as both a source of potential and as a limitation to the realization of potential. We have seen how the adaptive properties of microbes made them appear as a source of novelty and so a source of promise. At the same time, culture-free and DNA extraction technologies have enabled scientists to work with marine microorganisms that previously resisted removal from marine waters, amplifying their promise. The next section provides a description of how databases turn into preferred sites of extraction to further amplify such promise.
Extraction 4: From Mining the Ocean to Mining Databases
Once sequences are available, data-mining software is often used to further screen for bioactivity. It produces a sense of enhanced vision as it enables rapid retrieval and screening of large data sets. In the facilities I visited, the mining software tools antiSmash and BLAST were often used to screen microbial collections. The bioinformatician in charge at the Trondheim facility explained in an interview how they use antiSmash to make predictions on promising “candidates,” that is, gene clusters that are likely to code for interesting compounds. The software extracts information from existing databases and runs comparisons and automated analysis (Medema et al. 2011). Drawing on sequence information comparison, the program makes rapid predictions about the expected novelty and interest of the chemistry associated with the sequence(s) under investigation.
Data-mining technology runs on access to large data sets. Large amounts of data appear as both a product and a condition of possibility in “data-centric biology” (Leonelli 2016; Strasser 2019), including in sequence-based bioprospecting (Neimark and Wilson 2015). In turn, the production of large pools of data sets relies on large research collaborations and sharing (Edwards 2010; Kohler 1994). Increasing demands for open science and enhanced data flow in bioprospecting practices are a general trend (Laird et al. 2020). Sequence data exchange is seen as opening up new opportunities in bioprospecting, as a research leader in the Tromsø platform put it: I could send the genome information to another group somewhere else who do not have the physical sample and they can look at the genome and try to get information and try to express it in another host. So if they have the genome data, they do not need the original sample. They are able to express this chemistry without the sample. It could be a new chemistry that was originally encoded by a Norwegian bacterium…which then could be expressed somewhere else, just based on the digital information that you are providing if you are sharing it openly. (Researcher at the Marbio platform; emphasis added) In the future, what people can do is, they probably can reconstruct the whole gene sequence, the whole gene clusters to produce a novel compound that is independent of the original physical one…. Let’s say you need fifty genes and then you can just select from either from different type of strains, or different type of public knowledge, you’re reassembling it into the very novel ones that nobody has seen before, it’s not in nature. So then it could, at that time it could be you don’t need to depend on the physical strain that you need to sequence any more. So I think when we reach that point, probably (it) will be interesting to see how, let’s say, how can you prove that the component of the clusters come from these microorganisms or not. (Bioinformatician at Trondheim)
Infrastructural Displacements: Promising Microbes, Elusive Politics
Difficulty in “proving” the origin of a sequence has implications for scientific publishing and crediting as well as for determining sequence ownership. During my fieldwork, I frequently heard statements along the lines of “the same sequences can be found in different sites.” This observation coexists with strong claims of functional specificity, particularly for microbes adapted to extreme environments. 11 O’Malley (2008) describes how the field of microbiology has built on this tension between the belief that “everything is everywhere” (so all microbes can be found everywhere) and yet, at the same time, “the environment selects.” This ambivalence makes microbial bioprospecting promising. As microbial bioprospecting moves toward big data genomics and metagenomics, the question of whether the same microorganisms can be found in different places is reformulated into whether the same microbial sequences can be found in multiple locations. And of all locations, databases seem to be gaining in prominence as sites of biodiversity extraction, possibly eventually displacing the need for natural environments, physical samples, and particular jurisdictions.
An increasing replacement of in situ sampling with data mining could complicate the Norwegian state’s role as an administrator of biodiversity as a public good. The CBD acknowledges states’ sovereignty over the biodiversity within their jurisdictions, meaning biodiversity is primarily managed under national law and policy. In Norway, policymakers have promoted investment in infrastructures that will make the marine biodiversity in Norwegian waters more accessible. This is presented as strategic investment that will attract private investment and foster scientific collaboration. It is also in the interest of local scientists, who see data flow and collaboration as the ultimate conditions for the realization of bioprospecting’s prospects. The possibility of extracting large amounts of data from databases is seen as opening unprecedented opportunities in bioprospecting. Genomic and metagenomic analysis allows researchers to find more information from smaller samples, and it lessens the need to preserve and replace material samples. Ultimately, scientists’ vision is that, with enough public data, one day they could be able to work entirely independently from material samples. As scientists work to realize this promise of dematerialization, forms of life that are difficult to work with in the lab, such as marine microorganisms, see their prospects enlarged.
But while biodiversity as data becomes more portable and easier to exchange, it also becomes more difficult to track and capture under existing governing and administrative tools (Parry 2004). Arguably, the turn to sequence-based bioprospecting augments the prospects of some life forms while turning them into evasive political matters in the same move. It is thus not surprising that the Norwegian state has not determined how it will regulate the extraction and utilization of genetic resources in Norway. No matter what the state does, it will lose something: either international interest in Norwegian marine biodiversity or control over it. In a combination of presumed dematerialization and unwanted deterritorialization, microbes as data become a kind of outlaw form of public goods, promising but elusive as they are relocated across local, national, and global sites.
Not only scientific imaginaries hold this vision of an ultimately dematerialized bioprospecting in which databases are the main extraction sites. Policy debates on “DSI” at the CBD and Nagoya Protocol are often formulated in similar terms. The debates are permeated with fear that the Nagoya Protocol could become obsolete and frustration with state authorities’ difficulties in capturing the flow of data. Discussions have tended to focus on whether DSI counts as “genetic material”—the Nagoya Protocol’s original object of regulation and core to the definition of biodiversity itself. Countries with developed biotechnology industries, such as those in the European Union negotiating block, have tended to argue against considering DSI as genetic material (Karger 2018). Following this argument, sequence data would simply fall out of the scope of the CBD and Nagoya Protocol, and Parties would not have to consider DSI in their ABS regulations. Against this, countries that have traditionally been sites for material sample extraction argue that genetic materials and their digital counterparts must be understood as equivalent or else ABS will become obsolete (Bond and Scott 2020).
In these debates, as in my fieldwork in Norway, accumulation of data in databases appears as a condition for a dematerialized bioprospecting. As we saw, scientists (and also industry) consider this as opening up unprecedented opportunities. Effectively, databases have tended toward integration and upscaling. For example, nucleotide sequence data (the core of what is considered DSI under the CBD) is accessible through the International Nucleotide Sequence Data Collaboration infrastructure, which brings together the European Nucleotide Archive, GenBank (United States), and the DNA Data Bank of Japan. Against this tendency toward data accumulation, social movements across various countries have problematized the new “digital commons” as creating illicit pathways to accessing and exploiting genetic resources and so enabling an emerging digital biopiracy (Ambler et al. 2020; Bond and Scott 2020). Monitoring and tracing systems for both genetic materials and digital counterparts have been suggested as a technical solution to such problems (Ambler et al. 2020). Yet, as we saw, traceability is technically difficult because published sequence data can be copied or assembled from a variety of sources. Questions about how to improve data annotation to enable data tracing are on the debate table (CBD AHTEG 2020a). A key question is which actors have the material and infrastructural capacity to exploit data and thus “capture” biodiversity as a digital common good. Norway is a country difficult to place as either purely a biodiversity provider or a user, and its position within CBD debates remains unclear. 12 This is perhaps because the same infrastructural work that augments the promise of its marine biodiversity turns it into a volatile policy object, difficult for a state to capture.
Coda: Infrastructuring as Promise Making
Extensive scholarly work on infrastructure and data at the crossroads of media studies, STS, and anthropology challenges scientific ideals of dematerialization (Anand, Gupta, and Appel 2018; Balbi, Delfanti, and Magaudda 2016; Dourish 2014; Harvey, Jensen, and Morita 2016; Nadim 2021). Contributing to this literature, the empirical materials of this paper show how infrastructural work is carried out to realize the promise of dematerialized bioprospecting, augmenting the prospects of some life forms while simultaneously turning them into elusive political matters. Arguably, in Norway, it is the possibility of being “infrastructured” that is turning microbial marine biodiversity into a public issue and a political matter. Enhanced access to marine life forms augments the promise attributed to them that they will help move Norway toward a postpetroleum economy. However, that promise may remain suspended as microbes become elusive forms of public goods.
The kind of digital extractivism described in this paper runs on a series of displacements of life forms, political agency, and promises. Ultimately, infrastructuring biodiversity into sequence data affects the very definition of what counts as biodiversity and the kind of promises it encodes and entails. Concern for organisms, phenotypes, and taxonomic practice increasingly has been replaced with an attention to displays of life in the form of gene clusters, genomic maps or phylogenomic trees, and perhaps someday lab-assembled genetic constructs. Could lab-assembled life forms come to be claimed as biodiversity in the future? Could infrastructuring and datafication come to be seen as means for biological diversification, beyond adaptability to local environments? Such questions underpin the vision of a dematerialized bioprospecting. Further, as bioprospecting turns to genomic and metagenomic data, some life forms see their prospects particularly augmented. As we saw, as the focus of bioprospecting is scaled down to sequences, the prospects of microbial life are scaled up. And as some versions of what counts as promising biodiversity gain in prominence, others may become displaced. Colder Nordic waters have increasingly gained visibility as biodiversity hot spots as biodiversity is more easily accessed as data. This could displace the traditional “megadiverse” countries and their corresponding focus on plants and indigenous knowledge from the center of global biodiversity politics. Further, as databases displace in situ biodiversity and ex situ collections as extraction sites, the ways in which biodiversity is made public are radically reconfigured. Moving from nationally based public good to a global commons may broaden access to biodiversity, but not necessarily the spectrum of publics who could effectively benefit from its use. How data are extracted from particular sites, what information is taken into account and what information is excluded in data formations, the kind of infrastructural work that is institutionally supported, and the prospects attributed to infrastructural building—all these are stories worth reflecting upon in reconsidering biodiversity as promising political matters. A situation in which biodiversity tends to be datafied and exists as a global common may require renewed political agencies across local, national, and global sites for future (less elusive) biodiversity politics.
Footnotes
Acknowledgments
I would like to thank the Res Publica project team for their useful comments on earlier versions of this paper. Many thanks also to my colleagues at the TIK-STS research group for some really valuable last-minute comments. Special thanks to Heidrun Åm and to Deborah Scott for very constructive observations and suggestions on structure and editing. I am very grateful to all the researchers who took the time to talk with me and to walk me around in their lab facilities and to those who attended the Digital Life Norway Bioprospecting Workshop right before Norway went into lockdown. Many thanks to the anonymous reviewers and to Ed Hackett and Katie Vann for their helpful feedback.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Norges Forskningsråd (Res Publica 190743).
