Abstract
In science and technology studies today, there is a troubling tendency to portray actors in the biosciences as “cultural dopes” and technology as having monolithic qualities with predetermined outcomes. To remedy this analytical impasse, this article introduces the concept styles of valuation to analyze how actors struggle with valuing technology in practice. Empirically, this article examines how actors in a bioscientific laboratory struggle with valuing the properties and qualities of algorithms in a high-throughput setting and identifies the copresence of several different styles. The question that the actors struggle with is what different configurations of algorithms, devices, and humans are “good bioscience,” that is, what do the actors perform as a good distribution of agency between algorithms and humans? A key finding is that algorithms, robots, and humans are valued in multiple ways in the same setting. For the actors, it is not apparent which configuration of agency and devices is more authoritative nor is it obvious which skills and functions should be redistributed to the algorithms. Thus, rather than tying algorithms to one set of values, such as “speed,” “precision,” or “automation,” this article demonstrates the broad utility of attending to the multivalence of algorithms and technology in practice.
What Is a Good Algorithm?
The biosciences are often proclaimed to be going through a data revolution based on high-throughput technologies, online data sharing, and automated tools. Sometimes the biosciences are said to be going from in vivo through in vitro to in silico, a computer-based bioscientific imaginary where research has become “bigger, faster, better” (Davies, Frow, and Leonelli 2013). In response to this enthusiasm, many observers have asked what these technological changes might entail for knowledge production.
In this article, we wish to nuance a troubling tendency in science and technology studies (STS) to treat in silico technologies as having monolithic qualities and pregiven outcomes. Rather than tying high-throughput technologies to one set of values, such as “speed,” “precision,” “size,” or “capitalist automation,” our approach is to examine in detail what Keating and Cambrosio (2012) have referred to as the “hybridization” of the biomedical sciences (p. 37). Thus, rather than tracing how a seemingly nebulous but static set of high-throughput technologies affects the biosciences wholesale, we follow actors’ struggles to value what a good distribution of agency between humans and technology should entail in a high-throughput laboratory.
Consequently, this article approaches the shift toward high-throughput automation in the biosciences through actors’ valuations. Specifically, it examines how different configurations of humans and devices are valued in a high-throughput laboratory. In doing so, we show how actors’ valuations may diverge in ways that are tied to broader historically situated struggles about what should count as “good bioscience.” Concretely, the object of analysis is how laboratory workers value algorithms used for “normalization” and “randomization.”
Today, algorithms are an indispensable part of any high-throughput laboratory that aims to automate a variety of tasks. But what are algorithms? In computer science, an algorithm is defined as a “finite set of rules that gives a sequence of operations for solving a specific type of problem” (Knuth 1997). That is, an algorithm is a computerized instruction sequence to solve certain tasks. In a laboratory, for example, algorithms may be used to clean data or run a sample-handling robot.
However, for those working in the laboratory, the use of algorithms evokes many questions that relate to the valuation of algorithmic automation. What is a good algorithm? What does it achieve? And would the task be better done by an algorithm or a human? The actors’ struggles with such questions ground our research question: what different configurations of humans, devices, and algorithms are performed as “good bioscience?” In short: what configurations of agency between humans and machines do the actors articulate as appropriate and beneficial?
A key finding of this article is that different configurations of algorithms, robots, and humans were valued in multiple ways in the same laboratory: it was not apparent to our informants which configuration of devices should be seen as more authoritative than the other nor was it apparent which skills and functions should be redistributed to algorithms for automation. Instead, the appropriate distribution of agency was a matter of negotiating the value of algorithmic automation in each situation.
To underline the multiple and diverging yardsticks for “good bioscience” that actors articulated in our case, we introduce the concept of styles of valuation (cf. Fujimura and Chou 1994). By doing this, we aim to highlight how different styles of valuing algorithms can coexist within a single laboratory with varying degrees of dispute. 1 In paying attention to valuations in situated practices, we follow a valuographic research program that focuses on the practical enactment of values and stresses how such valuations are an inseparable part of knowledge production (Dussauge et al. 2015).
Approaching the data revolution in the biosciences in this manner enables us to describe and disentangle some of the many yardsticks that coexist in today’s high-throughput biomedical science. Thus, with Knorr Cetina (1999) or Daston and Galison (2007), we are interested in the links between valuations and the machineries of knowledge production. But, rather than studying the disunity of science by comparing different branches of science or tracing the instability of a particular value through history, we examine how different values are attributed to different configurations of human/technology in situated practices.
Technology in the Biosciences
The salience of these issues becomes apparent in relation to STS’s long-standing interest in the automation and digitalization of the biosciences. Recurring questions about the nature of this change have abounded, ranging from questions about how certain representational devices are constructed as more authoritative than others (Fujimura 1999) to questions about how skills and functions are redistributed between human and machines (Keating, Limoges, and Cambrosio 1999).
There has been great interest in how bioscientific research is becoming “big biology” and thus reshaped through “high-throughput” or “data-driven” approaches (cf. Leonelli 2012; Davies, Frow, and Leonelli 2013). Closer to the practices and technologies of knowledge production, questions have been posed about the so-called omics fields (e.g., genomics, metabolomics, proteomics), and how they “produce new genres of difference and variation” (McNally and Mackenzie 2013, 75). Some of these studies have taken an interest in high-throughput technologies and argued that bioinformatics is reshaping the biosciences into a capitalist production machine that emphasizes scale, economy, and surveillance (Stevens 2011). Other studies ask how a new breed of experts without wet-lab experience—such as bioinformaticians and biostatisticians—can lead to new challenges for online data interpretation (Leonelli 2014a, 2014b). One concern is that the data-driven biosciences could lead to an “exaggerated trust in the quality and comparability of the data” and a replacement of “subjective judgment” with a “statistical kind of objectivity” (Strasser 2012, 87).
While these lines of research highlight important developments in the biosciences, their technodeterminist tendencies concern us and we wish to address them here. Our argument is that there is a propensity to produce monolithic accounts of technology in the biosciences. We maintain that these studies sidestep a body of careful research on technology that has shown the variability of technologies in practice (Pinch and Bijker 1984; Mackenzie 1996; Laet and Mol 2000). Consequently, rather than making wholesale assumptions about the properties and effects of technology—for example, that bioinformatics is turning biology into a capitalist production machine—we propose to pay attention to situated valuations of technology in the biosciences and to highlight multiple human-algorithm configurations and concurrent values to which they are linked.
A Valuography of Algorithms
In the social sciences, algorithms have recently received an upsurge of interest (e.g., Kitchin 2014; Dourish 2016). Some researchers are employing the strategy of “going under the hood” of the algorithm, showing, for example, how machine gambling algorithms fabricate “near misses” to entice gamblers to bet again (Schüll 2012) or how algorithms can embody certain values—for example, a utilitarian moral philosophy for allocating organs (Roscoe 2015). In other research, algorithms are analyzed as performing and transforming the objects on which they are brought to bear, for instance, in detecting plagiarism (Introna 2013). We position this article closer to an ethnomethodological perspective that attends to the practices of designing and using algorithms rather than going under the hood to understand what those algorithms really do (cf. Neyland 2015; Ziewitz 2017; Seaver 2017).
To do this, we employ a valuographic research strategy that aims to analyze the multiplicity of values that users and designers attach to algorithms in the biosciences (Dussauge et al. 2015). 2 A particular facet of algorithms, which makes them intriguing as objects of inquiry in this sense, is their multivalence. This implies that their characteristics and their effects and values—just as bikes, missiles, bush pumps, or tomatoes—are difficult to stabilize (Pinch and Bijker 1984; Mackenzie 1996; Laet and Mol 2000). As a consequence, we do not treat algorithms as stabilized nor the values ascribed to them as intrinsic.
Significantly, this approach to analyzing values also entails an impartial stance to which valuations are the correct ones. Just as the symmetry principle in the sociology of scientific knowledge highlighted the need for epistemic impartiality in scientific disputes, this perspective on values means that we do not take sides in arguments about what is valuable in the use of algorithms (cf. Bloor 1976). Concretely, this means that we do not decry the advent of high-throughput algorithms as constituting a “corruption” or an “alienation” of the biosciences. Instead, we focus on the values—both positive and negative—that various actors attach to algorithms in a high-throughput laboratory. 3
Styles of Valuation
Below, we introduce the notion styles of valuation 4 to analyze our informants’ assessment of human–machine configurations. We introduce this concept to explore the problems, values, and yardsticks that are articulated, in this case, around algorithms. We here draw on Fujimura and Chou’s (1994) concept: styles of practice, which is defined as “historically located and collectively produced work processes, methods and rules for verifying theory” (p. 1020). However, while Fujimura and Chou “examine the co-production of facts and the rules for verifying facts over time” (p. 1017), we use styles of valuation to highlight differences in the valuation of scientific work processes, methods, and their relation to different yardsticks for “good science.”
In this view, technologies are intertwined with multiple concurrent sets of values such as tactility, reproducibility, smoothness, neutrality, objectivity, or universality. In the words of Fujimura (1999, 75): Scientific technologies are highly elaborated symbolic systems, not neutral media for “knowing” nature. For example, neutrality, or the idea that one can eliminate “noise” versus “signal” to reach a tabula rasa from which one can then produce “reproducible effects,” is part of a set of “values” historically located in so-called “Western traditions of thought.” These values include realist, objectivist, and empiricist rhetorics, which form the basis for establishing factness and the universality of findings.
An important point is that styles of valuing are situated in historical continuities and discontinuities. For instance, there have been debates about the merits of randomizing samples versus creating balanced comparisons at least since the statistician R. A. Fisher debated the issue of randomness in experiment design in the early twentieth century (Hall 2007). Also, the practice of blinding experiments by employing randomness has a long history, stretching back to early experiments on telepathy and in psychophysics (Dehue 1997; Hacking 1988). Thus, when new algorithms provoke debates on their value, these discussions resonate with historically long-running articulations and valuations of “good science.” Hence, new devices for automation—in this case, algorithms—can sometimes unsettle the status quo and lead to new experimental configurations, but they can also provoke repetitions of well-rehearsed clashes between entrenched styles of valuation.
Consequently, the notion of styles of valuation is meant to draw attention to how actors’ valuations of different configurations of human–machine agencies vary. The notion thus aims to open the possibility to identify situations where multiple styles are concurrent in a situated practice, thereby providing a means for examining frictions between different valuations of the same human–machine configurations.
Fieldwork
This article draws on the first author’s longitudinal and polymorphous engagement with an anonymous research group working with high-throughput biomedical mapping. The empirical work started in 2010 and was concluded in spring 2014, with a return visit to present results in 2016. The data collection consisted of interviews, meeting observations, demonstration sessions, and analyses of published and unpublished laboratory documents (such as articles, protocols, screenshots, and unpublished papers). Interviews were done at the laboratory, audio recorded, and subsequently transcribed. The laboratory demonstrations entailed sessions where different laboratory and software practices were demonstrated and explained, often in conjunction with scheduled interviews. Demonstration sessions were documented in field notes. In total, thirty-four interviews or demonstration sessions were conducted with researchers, lab technicians, doctoral students, and other actors related to the laboratory. Articles from the group and others, unpublished laboratory documents and protocols, screenshots from laboratory management systems, analysis software, and textbooks on research design were analyzed. All interviews and documents were coded and analyzed by the first author using Atlas.ti. To preserve the anonymity of the laboratory, the country, language, publications, and names are not stated.
Entering the Laboratory
The studied lab is situated at a research university in a European country. It consists of about twenty individuals divided into two research groups. The laboratory has been running for roughly ten years. It is focused on biomedical mapping of protein biomarkers or the identification of links between protein “signatures” or protein expression patterns and different diseases. The identified protein signatures, it is hoped, would provide leads for developing novel methods of diagnosis, which perhaps in the longer term would lead to new treatments. One dream being to replicate other laboratories’ successes in finding genomic biomarkers for the diagnosis of breast cancer (the BRCA genes) or prostate cancer (the prostate-specific antigen test).
In this lab, identifying protein signatures entails finding differences between protein expression patterns from “cases”—people understood as having a specific condition—and samples from “controls”—people considered to not have the condition. In order to find these coveted patterns, the lab analyzes samples from patients in the form of various bodily fluids such as blood, urine, and spinal fluid. The samples are supplied by collaborators around the world and have often been gathered in different places, using different techniques, and for different purposes. In the analysis of these samples, the scientists utilize a specific technique to map protein levels in the body fluids. This entails mapping how much of a range of proteins exists in a patient sample. For each sample, the levels of multiple proteins are measured. There are often hundreds of different proteins for each sample.
The high-throughput methods employed are designed to analyze hundreds of samples in one run. In general, the laboratory’s process entails several steps. First, transferring patient samples from different collaborators to the so-called microtiter plates (square plastic “plates” with small indentations or wells for body fluids ordered in a grid formation). Each plate contains a number of wells (up to 384 wells per plate in their fastest machine), and each well contains a sample from a single patient. Second, the plates are fed through the so-called multiplex machines that analyze the protein levels of all the samples on a plate. The third and last step involves comparing the protein levels of different samples to find patterns linked to different conditions. This involves comparing “cases” and “controls” through different types of statistical software, scripts, and algorithms.
The high-throughput methods make automation through algorithms and robotics crucial in the studied lab. The laboratory workflow includes machines to automate the handling of plates and samples, algorithms that control the machines, and algorithms that clean the data. We focus here on two algorithms in use in the lab: randomization and normalization. The randomization algorithm was used in the first step of the process outlined above and generated instructions for sample-handling robots. These robots transferred patient samples from the test tubes they were stored in to the plates that were fed into the multiplex machines. The normalization algorithm was used (in step three above) to separate—as one of our informants expressed it—“biological variation” from “nonbiological variation.” That is, to separate noise from signal when the data from the multiplex machines were analyzed. We will attend to the valuations of these algorithms in detail below. The two algorithms in focus in this article addressed a central matter of concern for our informants—separating signal from noise, pattern from static, and protein signatures from trash.
Randomization: Tinkering versus Ignorance
As described above, randomization algorithms were used to place patient samples on plates. However, achieving a good randomization was not straightforward and included a multitude of devices and practices throughout the lab: it was sometimes introduced manually (literally by hand) by putting samples into cardboard boxes and shaking them, which an informant jokingly called “the box method of randomization.” Sometimes the randomization was brought about using a simple algorithm in the R scripting language that took a “vector” and scrambled it. That is, the algorithm took an orderly list of numbers—1, 2, 3, 4—and randomly reordered the sequence into for example 2, 4, 1, 3. This new sequence of numbers was then utilized to guide the researcher’s hand when manually sorting samples onto a plate. Our informant’s argument was that the vector method ensured random placement without the researchers having to “think about it”—without mental energy being expended on producing a randomization. However, when large numbers of samples had to be randomized—when the lab ran their signature high-throughput analyses—an algorithm produced by the lab’s resident bioinformatician could be used. This bespoke “plate layout algorithm” produced instructions for a sample-handling robot, which automatically randomized the samples on a microtiter plate according to a specific “plate layout.”
What was at stake in these actors’ struggles with randomization is related to well-known issues in experiment design that are brought head-to-head in negotiations of what “good bioscience” should entail. Specifically, the informants struggled with valuing balanced and unblinded ways of organizing experiments versus randomized and blinded ways. Importantly, the valuations of the algorithm related to debates in experimental design that have been running for a century or more (Hall 2007, Marks 2003). Thus, the actors tied the randomization algorithm to different problematizations of good scientific practice and different yardsticks for what a good algorithm should do.
Randomization, Balance, and Human Tinkering
Perhaps paradoxically, the randomization algorithm used in the laboratory we studied was not random—it produced what our informants called “a balanced randomization.” This was by design, since a complete randomization was understood as problematic. Thus, alongside the argument for randomization, our informants contended that the randomization algorithm also needed to produce a balanced distribution of samples. The informants’ reasons for introducing this “balanced randomization” was a concern that the machines and processes used to analyze the samples—the multiplex machines—could introduce nonbiological variation in the resulting data.
The first source of concern was that samples located on different plates could yield different results when they were put through the multiplex machines. At issue was that each run could result in subtle differences that stemmed from slight variations in the machine rather than from measured variances between samples. Hence, an appropriate randomization was considered to be one that located patient samples to be compared on the same plate for analysis in the multiplex machine. By placing samples to be compared on the same plate, they could, our informants argued, neutralize differences in the results from differences in the machine.
A second source of concern for our informants was that samples of a certain kind (e.g., cases/controls or male/female) could become clustered in one location on a microtiter plate. If, for instance, all control samples happened to be grouped in the top left corner, data patterns could be produced that could be picked up in the subsequent analysis. According to this way of thinking, the plate layout algorithm should not (even by chance) place all samples of one type in one cluster on the plates. To counter these two problems, our informants wanted to place samples randomly—but distributed in the right way—on the plates: There are several different methods to do it [the randomization]. But, the important part is to check if all control samples happen to be placed at the start—and all with the condition at the end of the plate. (Excerpt from interview, Biotechnologist 2, 2013)
A common way to algorithmically balance the randomization was to decide which parameters were to be distributed evenly on the plate and feed these parameters to the algorithm. Three parameters frequently seen as being in need of balancing were age, sex, and diagnosis of the patients. One informant told us how their algorithm worked: I can note the information I have about age, gender and diagnosis, and that I want these characteristics to be evenly distributed. Then—say I have four initial plates that are to be combined into a 348 well plate—I want to be sure that the samples are distributed in a balanced way across the four plates. (Excerpt from interview, Biotechnologist 2, 2013)
For our informants, a balanced randomization algorithm that broke up patterns, distributed samples evenly, or balanced samples to be compared was thus seen as the appropriate solution to the problems of noise, clustering, and comparison. This is a style of valuation that foregrounds the benefits of balance and comparability. It emphasizes how the algorithm makes it possible for the experimenters in the laboratory to handle machine noise, thus facilitating comparison.
Randomization to Create Ignorant Researchers and Remove Bias
However, the value of having a balanced randomization was not self-evident to all collaborators. As we outlined initially, the lab depended on partners around the world for the supply of patient samples to be analyzed. These collaborators were an essential part of the high-throughput setup. In a particular misunderstanding between the lab and a collaborator, different ideas about randomization came to a head. Some of them argued for randomness as a tool for creating blindness, thus measuring the value of the algorithm with a different yardstick for “good science”—that of ignorance to remove bias (cf. Dehue 1997; Hacking 1988).
Several of our informants touched on the incident and discussed how their collaborators had a different idea about randomization. The person working in the collaboration recalled: Then they [the partner] randomized the samples and sent them to us, and I didn’t know they were going to randomize them for us. That’s something we usually do ourselves. We weren’t happy with their randomization […] So we had to have a meeting. They wanted us to be blinded. We shouldn’t know which [samples] were [from] healthy and ill [patients]. (Excerpt from interview, Biotechnologist 3, 2013)
Tensions: The Balanced Experiments versus Ignorant Researchers
On the one hand, what was at stake for our informants was the tinkering to produce comparability. These experimental practices are reminiscent of the classic agricultural experiments in the UK in the 1920s, where balanced planning of plots of land were to yield balanced comparisons (Hall 2007). Just as our informants attempted to create a balance on their microwell plates, early agricultural researchers aimed to compare plots that were similar in terms of rainfall, sunshine, soil, and so on. The goal of both the agricultural researchers and our informants was to facilitate comparison by matching the characteristics of agricultural plots—or patient samples.
On the other hand, balanced randomization ran against the practice of randomization favored by a partner supplying samples to the lab. This partner wanted to perform a blinded randomization to render the introduction of human bias impossible. For them, randomization aimed to protect against the specter of human intentionality (see, e.g., Hacking 1988; Kaptchuk 1998). These valuations echo a medical and pharmaceutical understanding of randomization where the ignorance of personnel is a key component of producing unbiased results. In short, it is similar to a key trait in the double-blind randomized controlled trial (RCT). In a textbook on medical research design, the blindness of the clinical team is stressed: If the allocation is predictable, then the investigating physician has knowledge that he or she may subconsciously use to influence their decision to include (or exclude) certain patients from the trial…. […] As a consequence, any prior knowledge by the clinical team or the patient of the allocation can therefore introduce bias into the allocation process, and hence lead to bias in the final estimate of the parameter β1 at the close of the trial. (Machin and Campbell 2005, 70)
Consequently, there are tensions in how actors value different configurations of humans and algorithms, in particular, the value that the actors ascribe to different distributions of agency between humans and algorithms. On the one hand, human knowledge and agency were articulated as crucial for achieving balanced randomization. On the other hand, unblinded humans were articulated as problematic as they could introduce bias. Randomization was thus subjected to two different styles of valuation, each of which favored a particular randomization procedure.
Normalization: The Cooked and the Raw
Another common operation in the lab was algorithmic normalization. This is a frequent operation in different types of signal processing and is used to make data sets more alike. A normalization entails treating data algorithmically, for example, to make similar the volume of different music tracks. A key aspect of algorithmic normalization is that it can automate the process of bringing data into the same dynamic range: you don’t need to constantly fiddle with the stereo volume or sample amplitudes. The algorithm does it for you. Hence, in the studied laboratory, normalization brought protein levels from different patient samples—the data that resulted from the multiplex machines—into the same dynamic range.
Normalization and the Dream of Automation
Early on in the data collection, normalization was introduced by a medical epidemiologist working as a bioinformatician in the lab. Most projects ran through his normalization algorithms at one time or another. In a sense, he functioned as a bioinformatics hub for most of the lab’s projects. In answer to Lee’s questions about normalization, he turned to the practical value of normalization, which, according to him, was to “minimize nonbiological variation” and thus extract the “biological variation:”
What is normalization?
There is an expression: minimizing nonbiological variation. Because we are interested in biological variation, but there’s always some nonbiological variation. Which we should try to remove. But there is no way to remove it, so we try to minimize it.
So, how do you know what nonbiological variation is?
Actually, we don’t know. It’s a really tricky question. […] if we make a plot of the signal and […] plate one generally has a higher signal than plate two, it’s visualized in the plot. Then [we can see that] there’s a big difference between the plates and we try to adjust them, for them to have the same or similar values. That’s normalization. (Excerpt from interview, Bioinformatician 1, 2013)
In the exchange above, the bioinformatician valued normalization for its power to “minimize nonbiological variation.” The ideal was thus to use normalization algorithms to automatically remove noise stemming from variations in samples, machines, and laboratory processes, so that biological variations were highlighted.
Here, we deal with the valuation of one of the normalization algorithms employed in the studied laboratory, probabilistic quotient normalization (PQN), which originated outside the studied lab. In order to preserve anonymity, we deal with an article published by the lab that originated the algorithm rather than one from the studied laboratory. The PQN algorithm’s creators argued that its value lay in its ability “to reduce variances and influences, which might interfere with data analysis” (Dieterle et al. 2006, 4281). The value of the algorithm thus lay in its power to make an automatic separation between biological and nonbiological variation (Figure 1).
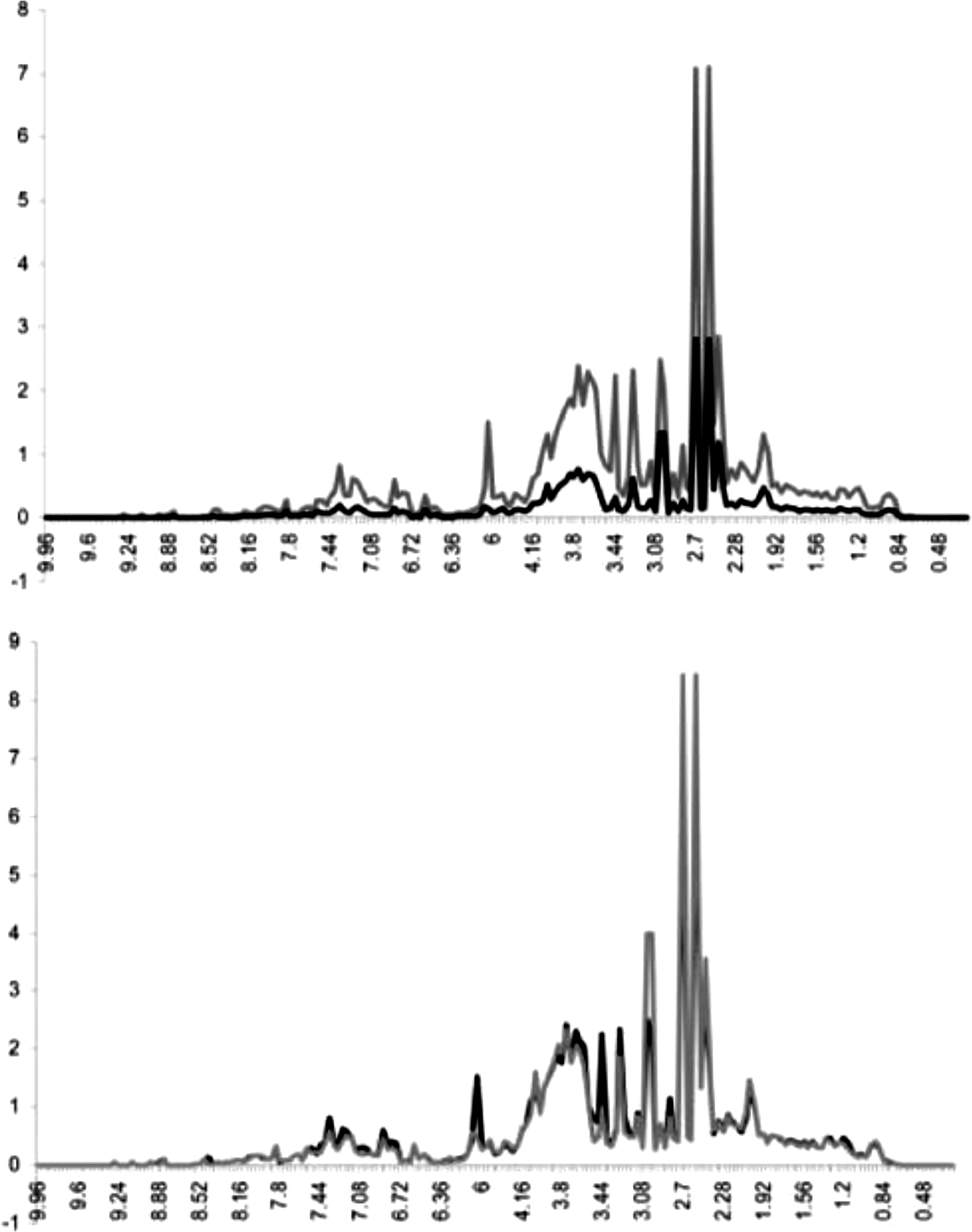
An illustration of the effect of normalization from Dieterle et al. (2006, 4282). The image shows two diagrams. Both diagrams show the same two sets of data (presented in gray and black). The left diagram shows the data sets before normalization, the right diagram after normalization. The gray and black data sets diverge significantly on the left, while they are nearly aligned on the right.
An important part of showcasing this value was the demonstration of its power to automatically produce smoother sets of data. In the above-cited article on PQN, three different normalization methods are compared visually. The argumentation and visualization in the article focused on the power of the algorithm to produce “optimal normalization” results. A diagram shows four different simulated data sets that vary the “nonbiological variation” in different ways. The optimal normalization being represented as a flat line, which only the PQN algorithm achieves in three of the data set visualizations (Figure 2; data set 4 shows a marked “shelf” drop-off for PQN).
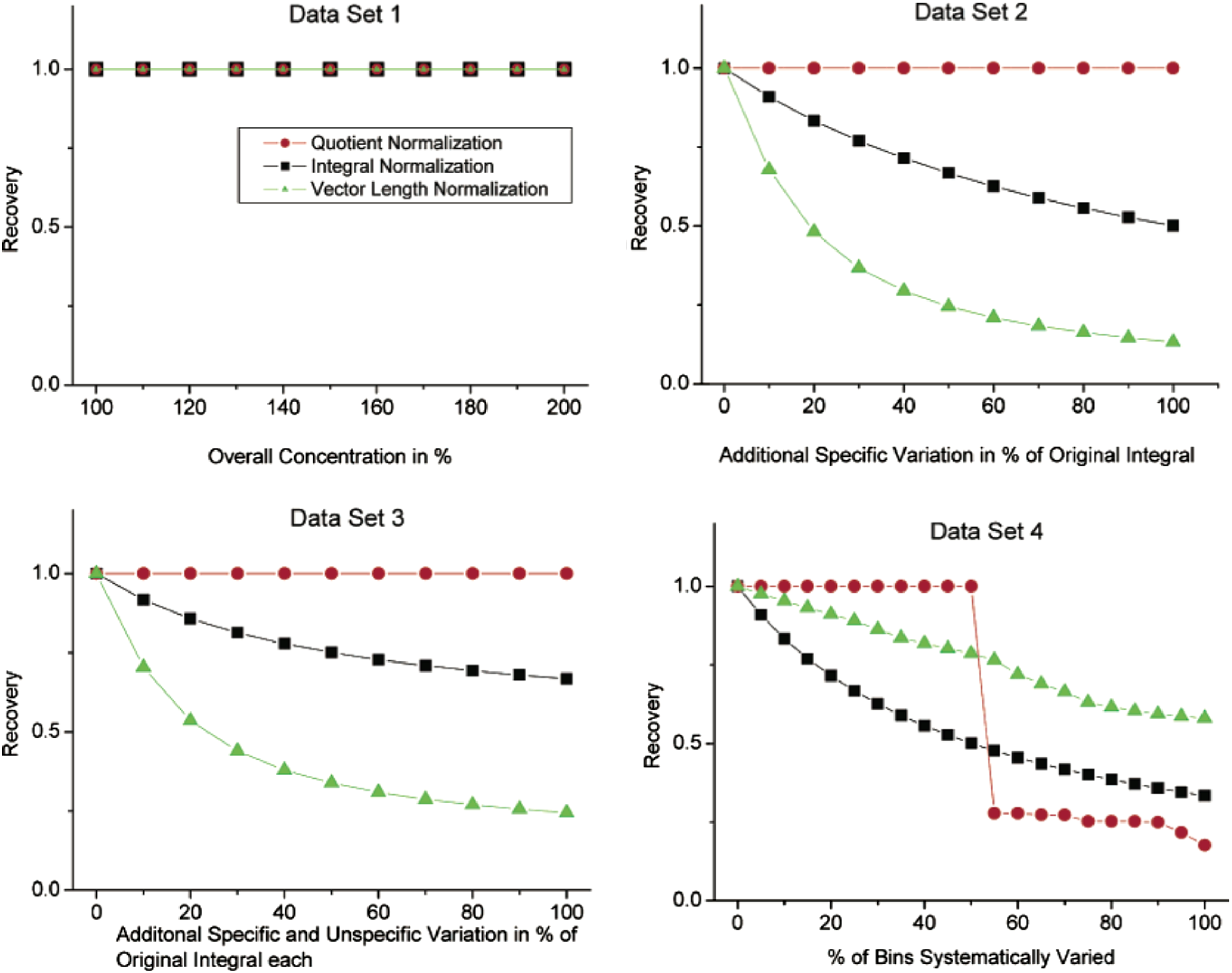
A comparison of different normalization methods, with probabilistic quotient normalization as the clear “winner” (Dieterle et al. 2006, 4286).
The PQN algorithm is thus supposed to automate the smoothing out of nonbiological differences by deciding (based on a set of predetermined calculations) whether a sample varies because of biological or nonbiological differences. The normalization algorithm is thus valued for its ability to produce smooth data sets and for its ability to automate the separation of biology from nonbiology. Hence, the PQN algorithm is assessed in a style of valuation that emphasizes the benefits of automating data processing. This style of valuation thus foregrounds the benefits of automation: high value is attached to the automatic reduction of noise while retaining the signal unchanged.
The Allure of the Raw: The Value of Analyzing by Hand
However, as alluring as the idea of an automatically smoothed biology was for our informants, the algorithmically normalized biology was constantly questioned. The normalized data provoked serious doubts about how biological and nonbiological differences could be told apart. One informant highlighted the difficulty: And then you get stuck in a kind of argument about whether it is age or disease. And if we normalize away the age—which is something you can compensate for in an analysis—then one loses those differences. And are they then biological or are they somehow…(Excerpt from interview, Biotechnologist 2, 2013)
Trust in the algorithmic normalization was at times also contrasted with “destroying the data through normalization.” In these situations, it was argued that nonbiological variation could be introduced by the very algorithmic practices that were used to remove them: I’m not a trained statistician or mathematician. I don’t have the proper understanding or feel for all the effects of throwing a massive number of variables into models. This is also something I hear from others, that you can tweak your data to the point of breaking it. You can destroy the data through normalization. (Excerpt from interview, Biotechnologist 1, 2013)
This skepticism against algorithmic data processing was also expressed by another informant who argued that patterns observed using normalized data should be visible also in “raw data,” protecting against the algorithm introducing nonbiological variations rather than removing them. Here, looking at the unnormalized “raw data”—straight from the multiplex machine—was seen as a safeguard: When you’re working to normalize and trying to get rid of certain things in the data, a helpful rule can be that things should also be visible in the raw data. A difference that you want to point out in a publication shouldn’t be something that has been introduced through normalization. (Excerpt from interview, Biotechnologist 3, 2013)
The algorithmic normalization stuck the researchers between a rock and a hard place. For one informant, getting a feel for how the “raw data” varied was crucial, but the crux of the matter was that this “hands-on disposition” also precluded a high-throughput approach. I’m such a nerd. I like to print out the raw data on paper and have it in a paper table since I like to highlight some things by hand. I’m pretty hands-on. Of course, I do all my analyses in R with the latest statistical software packages and all that jazz, but I think it’s reassuring to be able to go back and actually check if I have done something quick and dirty. It’s way too easy to drop lines and get lost in the normalization. But that’s not possible if you’re working with 10,000 antibodies. (Excerpt from interview, Biotechnologist 4, 2014)
Tensions in Valuing Normalization: Automation versus Tactility
Consequently, algorithmic normalization was subject of two different styles of valuation, one which foregrounded the benefits of high-throughput analyses and one that foregrounded the advantages of tactile human judgment. They each represents a different articulation of the problem of handling signal and noise and what were appropriate means to address these problems.
The drive toward hands-on analysis and tactility is well-documented in research on the cultures of the biosciences. It can be connected to a long series of studies done on the biomedical sciences, emphasizing the hands-on work, the tinkering, or openness of biological experimentation (cf. Jordan and Lynch 1998; Cambrosio and Keating 1992). An early example is Evelyn Fox Keller’s (1983) book A feeling for the organism, which documents, among other things, the geneticist Barbara McClintock’s resistance to the increasing quantification of genetics in the 1930s and 1940s. A classic quote from McClintock shows her passion for knowing the organism: I start with the seedling, and I don’t want to leave it. I don’t feel I really know the story if I don’t watch the plant all the way along. So, I know every plant in the field. I know them intimately, and I find it a great pleasure to know them. (Keller 1983, 198)
The researchers in our laboratory thus ascribed a high value to intimate knowledge of data, which included skepticism toward automated high-throughput approaches and algorithms. The value of human agency and tacit knowledge were central. However, simultaneously, and in contrast, they also foregrounded the benefits of high-throughput automation. It was even deemed impossible to know your data in the traditional manner.
The tension between these two styles of valuation was a constant issue. For example—at a revisit to the lab presenting our results after concluding the fieldwork—we were told by one of our informants that she had discussed normalization and their lab’s criterion to see patterns in nonnormalized data, with an outside biostatistician. The biostatistician’s comment was as follows: “But why normalize at all then?!”
Thus, different styles of valuation coexisted in the same laboratory and surfaced regularly as disagreements about how to define a problem and the proper means to address them. For our informants, there were constant worries about what distribution of agency between algorithms and humans was the right one.
Styles of Valuation: Algorithms and the Distribution of Agency
By paying attention to how actors struggle with the use of algorithms in a high-throughput bioscientific laboratory, we have been able to identify four different styles of valuing algorithms. In this, our emphasis has been on the valuation of different configurations of humans and algorithms and, in effect, different distributions of agency. Thus, we have analyzed how actors struggle with the questions “What is a good algorithm?” and “How should competencies and roles be assigned between humans and machines?”
We have heuristically identified the researchers’ valuations as being done in four different styles. These styles of valuation center on actors’ articulations of problems, solutions, devices, and configurations of agency. Each style is characterized by a particular problematization as well as a particular assessment of what are appropriate solutions to these problems. We argue that styles of valuation is a helpful tool for examining the ambiguous role of algorithms in our examined lab and for analyzing how actors struggle with high-throughput bioscience. Table 1 provides a summary of key characteristics of each style of valuation identified in this case study.
A Summary of Key Characteristics of Each Style of Valuation Identified in This Case Study.
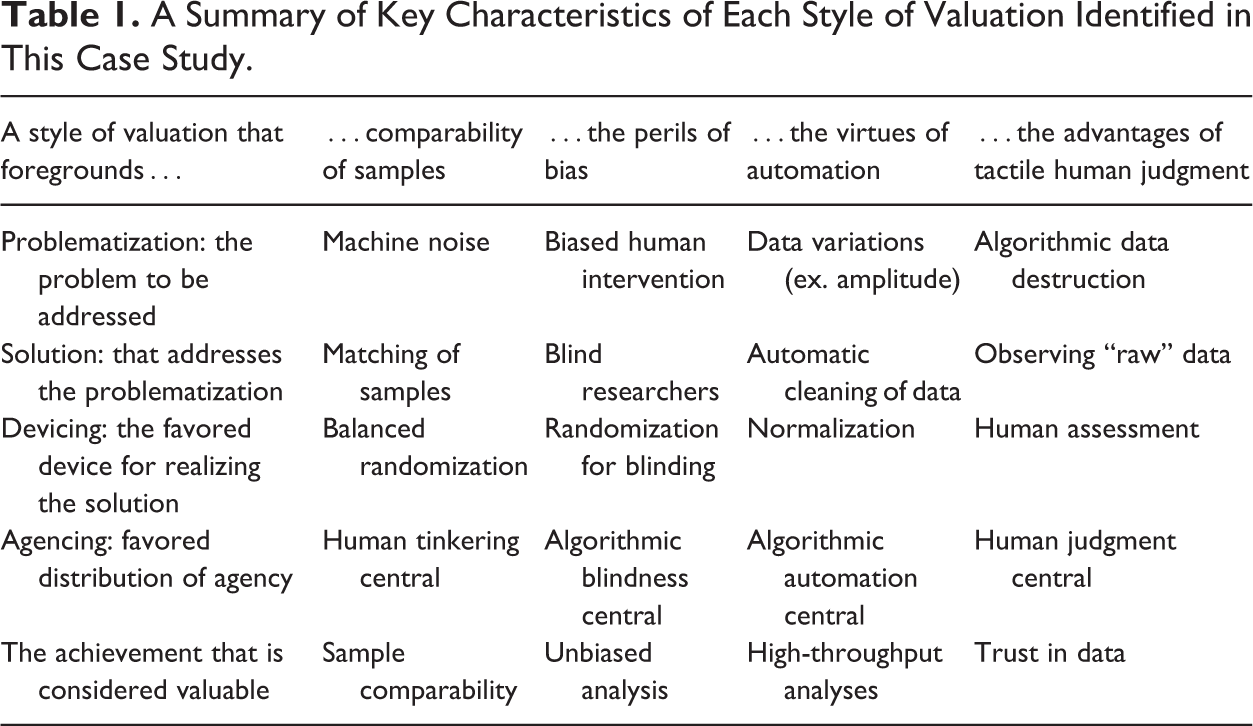
In our analysis, we first attended to algorithms and tensions between the fear of machine noise and the fear of human bias. Tensions between human intervention to achieve balanced comparisons were counterpoised with blinding to achieve unbiased experiments. In one case, noisy machines were articulated as the problem and algorithmic matching of samples as the solution. In the other case, biased human intervention was seen as the central concern and algorithmic blinding as the answer. Second, we examined algorithms and conflicts between automation and human judgment. In this situation, actors valued an algorithm for its capacity to automate the handling of sample differences. This placed the responsibility for separating data and noise in the realm of the algorithm. Clashing with this was the articulation of the algorithm as a destroyer of “raw data.” Here, intimate knowledge of “raw data”—an oxymoron to be sure—was articulated as the remedy against algorithmic destruction (cf. Gitelman 2013).
Our observations should be understood against the backdrop of the ongoing technological shift toward high-throughput bioscience, which has led observers to ask whether subjective judgment will be replaced with statistical objectivity and an exaggerated trust in data (Strasser 2012) or whether singular data points will fade away from high-throughput work (Leonelli 2014b). Our observations, however, both support and contradict these arguments: the tension between subjective judgment and statistical objectivity did not fade away in the laboratory we examined. Rather, tensions were heightened. Individual data points were understood as impossible to know as intimately as desired, but the drive to intimacy and subjective judgment persisted.
In previous studies of high-throughput bioscience, it has been argued that this technological shift might reshape the biosciences in line with a capitalist logic of production (Stevens 2011) or as an enabler of “data journeys” (Leonelli 2016). However, we maintain that tensions and instabilities persist in these practices. Our suggestion has been to analyze these different ways of swimming in the high-throughput tide of data by paying attention to actors’ contradictory valuations of scientific devices and technologies. This approach allows us, as Fujimura (1999) proposes, to understand these scientific devices as part of elaborate symbolic systems, with links to multiple and divergent ideas about how good science should be done.
We see this move to stress the multiplicity of valuations as a remedy to an unfortunate tendency within STS to treat “high-throughput” or “big data” technologies as phenomena that in determinist fashion reshape the biosciences. An unfortunate analytical consequence of this tendency is that “new technology” can be used as a featureless placeholder that allows for deterministic conclusions about how “big data” and “infrastructures” are wholesale reshaping “biomedicine.” Perhaps the quintessence of this simplifying move can be found in Anderson’s (2008) much criticized article “The End of Theory: The Data Deluge Makes the Scientific Method Obsolete”. Some STS analysts may want to argue that high-throughput bioscience undermines the very craft of research. However, we want to refrain from passing such judgments. By sticking instead to a principle of symmetry, we have highlighted the multivalence of these technologies and underlined how bioscientists struggle to work with and value these new sociotechnical configurations.
Today, many STS studies of the biosciences have a troubling tendency to view “algorithms,” “big science,” “big data,” or “data-driven science” as monolithic phenomena that will have pregiven outcomes. It is all too easy to fall into the current “algorithmic drama” (cf. Ziewitz 2015) that highlights either a dystopian future where human agency is curtailed or a utopian dream world of “bigger, faster, better” (Davies, Frow, and Leonelli 2013). We contend that this dualistic take on new technologies in the biosciences—in its seductive simplicity—risks undoing a lot of careful work on the complexities of laboratory practices in STS and elsewhere, reproducing bioscientists as “cultural dopes” who act with preconceived notions of how technology in the biosciences has worked and will work in the future (cf. Garfinkel 1967, 68).
Approaching the data revolution in the biosciences through the concept of styles of valuation enables us to disentangle the multiplicity of valuations that coexist in today’s high-throughput biomedical science. To understand how new technologies become part of the biosciences, we need to be open to actors’ articulations of these challenges and how they value different distributions of agency between humans and algorithms. Our argument is that if we fail to acknowledge the divergent valuations of technology in situated settings, we risk becoming blind to actors’ struggles to work with automation and algorithms. “What is good technology?” will always be an open question, and we want to highlight that tension to bring out the multiplicities, dilemmas, and trade-offs over deterministic accounts of science and technology. As Donna Haraway (2010) phrases it, we wish to provide tools for “sticking with the trouble.”
Footnotes
Acknowledgments
The article has benefited from comments by Steve Woolgar, Nick Seaver, Corinna Kruse, Mikaela Sundberg, Otto Sibum, and Frans Lundgren. We would also like to thank the editors Ed Hackett and Katie Vann as well as our two anonymous reviewers for helpful comments on earlier versions. Presentations at the ValueS research group at Linköping University, the Cultural Matters Group at Uppsala University, and the higher seminar at the Department of History of Science and Ideas at Uppsala University have also been invaluable. The overall project takes an interest in the experimental design of biomedical experiments as a site of valuations.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The article reports a sub-study within the “Trials of Value: On the valuation practices in designing medical experiments” project funded by Riksbankens Jubileumsfond [Grant Number P11-0034:1].
