Abstract
A thorough review of the literature is the basis of all research and evidence-based practice. A gold-standard efficient and exhaustive search strategy is needed to ensure all relevant citations have been captured and that the search performed is reproducible. The PubMed database comprises both the MEDLINE and non-MEDLINE databases. MEDLINE-based search strategies are robust but capture only 89% of the total available citations in PubMed. The remaining 11% include the most recent and possibly relevant citations but are only searchable through less efficient techniques. An effective search strategy must employ both the MEDLINE and the non-MEDLINE portion of PubMed to ensure all studies have been identified. The robust MEDLINE search strategies are used for the MEDLINE portion of the search. Usage of the less robust strategies is then efficiently confined to search only the remaining 11% of PubMed citations that have not been indexed for MEDLINE. The current article offers step-by-step instructions for building such a search exploring methods for the discovery of medical subject heading (MeSH) terms to search MEDLINE, text-based methods for exploring the non-MEDLINE database, information on the limitations of convenience algorithms such as the “related citations feature,” the strengths and pitfalls associated with commonly used filters, the proper usage of Boolean operators to organize a master search strategy, and instructions for automating that search through “MyNCBI” to receive search query updates by email as new citations become available.
Keywords
The ability to perform an effective search strategy comprises the foundation of any evidence-based discipline. Whether one is functioning in a clinical or research capacity, the assumption is that the expert suggestions offered arise from a complete review of the relevant literature. In systematic reviews and meta-analyses, this is taken a step further, necessitating that the methods of these exhaustive literature reviews be reproducible. The conveniences afforded by the basic PubMed search engine and algorithms, such as the “Related Citations” feature, provide powerful search tools but take much of the control out of the hands of the user. Blind usage of these hands-off methods weakens the user’s ability to ensure a thorough, systematic, and reproducible search. A common solution to this problem is to limit the search strategy to include only the MEDLINE portion of the PubMed database, allowing for the usage of more robust, user-controlled search methods. Unfortunately, depending on the journal indexed, MEDLINE is often as much as a year behind the current research, 1 leaving many of the most recent and relevant research citations out of the search results. The strategies presented in this article offer a solution to this problem.
The general landscape of PubMed has been previously described in a recent article by Lindsey and Olin. 2 The objective of the current article is to augment that landscape by proposing a detailed method through which its many components may be employed and integrated into one highly efficient, reproducible, and exhaustive search strategy. True comprehension of this system provides the ability to create a master search for any topic. Furthermore, the many free tools provided by the National Center for Biotechnology Information (NCBI) and the National Library of Medicine (NLM) through PubMed make it possible to receive timely notifications of citations, directly relevant to any area of one’s field as they become available. Part I of this article discusses theories, methods, and key concepts for using medical subject headings (MeSH) to search PubMed’s MEDLINE database, the less robust strategies necessary for searching the remaining non-MEDLINE database, and methods for restricting the less robust search tools to search only the non-MEDLINE database. Part II provides a step-by-step tutorial for implementing part I as well as methods for automating a search strategy so users may receive daily, weekly, or monthly ongoing updated results to their search query by email.
Part I: Theory, Methods, and Key Concepts for Performing a Master Search
PubMed, MEDLINE, and MeSH: The Filing System for Evidence
PubMed is a free citation database from the NLM. It currently houses >23 million citations dating back to 1809 with new citations added daily. The largest portion of PubMed citations comes from the MEDLINE database, which is an NLM database specific to the fields of biomedicine. The database covers citations from 1946 to present 3 and currently presents 88.9% of all citations in PubMed. Of the remaining 11.1%, 23.3% are in the process of becoming MEDLINE indexed, 15.3% have been submitted by the publisher and may become MEDLINE indexed in the future, 51.7% are available on PubMed but will not be indexed by MEDLINE, and 9.7% belong to an older version of MEDLINE that was indexed under less rigorous standards. 4 The distinctive feature of a MEDLINE-indexed citation is the manual rigor under which it is cataloged and categorized. Highly skilled information technicians read each article to determine the key points as well as its smaller discussion points. These points are then manually catalogued under a series of searchable Medical Subject Headings (MeSH terms) and subheadings. 5
The current staff that manages citations and MeSH terms at the NLM generally have a master’s degree or higher in some form of biomedical field. 6 Their collective expertise embraces a diverse range of biomedical topics. Each attends rigorous training in document analysis and cataloguing for MeSH (visible on the NLM website). 7 They then complete an internship/apprenticeship with an indexing mentor who checks every indexed article for quality and consistency with MEDLINE cataloguing methodology.
The MeSH system is essentially a digital filing cabinet with 16 biomedical topic drawers that match to the broadest MeSH terms. These drawers are filled with a 12-tier hierarchy of folders within folders, which systematically narrow into increasingly refined topic areas; each topic area is accessible through its MeSH term. Each MeSH term comes with a selection of up to 83 possible subheadings to further refine the topic area to a specific focus of interest, such as the “adverse effects,” “economics,” “ethics,” or “methods,” associated with a specific term. 7 This system is intricate, but all the tools necessary to simplify its navigation are available for use through PubMed.
A general, unqualified PubMed search will retrieve citations through several different pathways. Wherever possible, PubMed will translate the user’s search terms into MeSH terminology, a process known as Automatic Term Mapping. Frequently, however, this is not possible, and PubMed must rely on the less sophisticated indexing system of word searching. 8 When a search word does not map to a MeSH term, PubMed looks for the search phrase in other fields of the document. It first attempts to match the phrase to a journal name, then to an author name, then to a name of a research investigator, and finally it searches for the phrase in [all fields]. 5 A search in [all fields] includes areas such as the title field, the author field, the publication date field, and an abstract field when available. 9 It then breaks the search phrase into its component words and repeats the above process for each individual word. While this may retrieve many relevant articles, these articles will be interspersed with an abundance of irrelevant articles. Many users circumnavigate this inconvenience by employing the “PubMed Related Citations” feature available on the right side of any PubMed search results page (Figure 1).
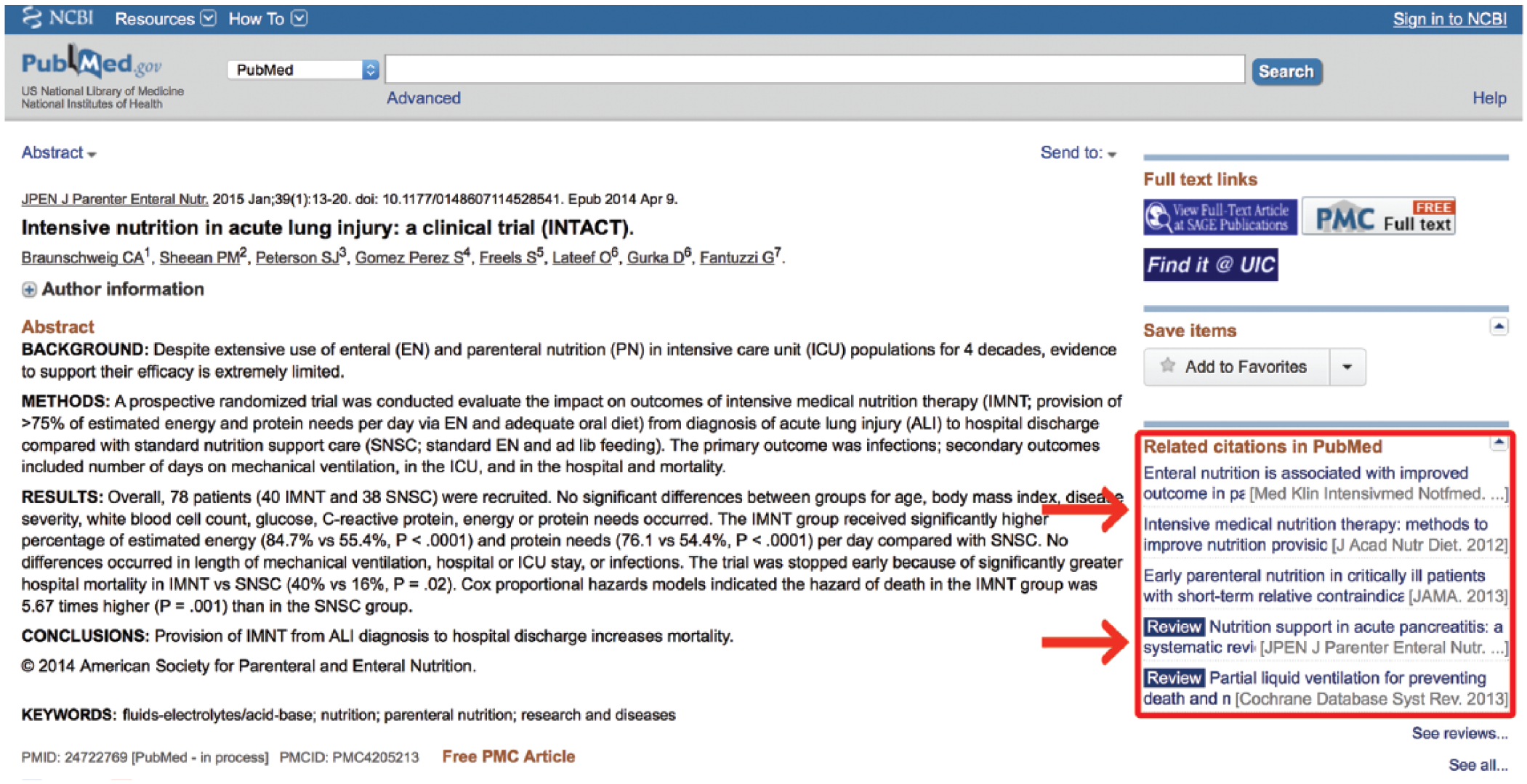
Related citations.
Pros and Cons of “Related Citations”
When a user clicks on a citation, the “PubMed Related Citations” feature finds the 5 citations that contain the most verbiage in common with the selected citation. This algorithm-driven search utility is essentially an intricate word counter, which tracks the words in a citation and weighs them according to their frequency and location. 10 It counts the number of times a word is repeated in the title, abstract, and list of MeSH terms in a document. It then applies different weights to the words according to where the word is found in the citation (ie, title, abstract, etc), the number of times it occurs in the document, and the frequency with which the word is found throughout all titles and abstracts in the database. A small mathematical adjustment is made for the length of the document, and a word score is assigned to the citation, which is used to determine similarity with other citations. This feature requires very little skill on the part of the user and is exceptionally good at quickly providing relevant articles. For more scientific pursuits, however, this feature may be inappropriate. While it is a useful feature, it is not a systematic search approach. Most research is built on the idea that an exhaustive literature review has been performed. With the related citations feature, an investigator does not know for sure which articles they have missed. Furthermore, science is built on the principle that someone’s methods of discovery must be reproducible. Every time a promising citation is selected, the “Related Citations” algorithm resets to that new citation and produces 5 more citations based on that new input, making the pathway of the search impossible to track. While this automated approach is a valuable feature, the ability to formulate an effective manual search strategy will enable investigators to know exactly which areas they are including and which they are ignoring, a vital component for any formal literature review.
Key Concepts of the Master Search Strategy for PubMed
As previously stated, 88.9% of the PubMed database comes from the MeSH-indexed MEDLINE database. While MEDLINE is a more robust search system, it can take weeks to months to years after release for an article to become MEDLINE indexed. 1 The other 11.1% of non-MEDLINE citations are available in PubMed but without MeSH indexing. While this non-MEDLINE search system is less robust, its contents contain the most recent and temporally relevant research often many months before it would be found in a MEDLINE search. For this reason, a full review of the literature must have both a MEDLINE component and a non-MEDLINE component. The MEDLINE component may employ both MeSH and text searching (non-MeSH) strategies when necessary, but its results will always be limited to the MEDLINE database. By design, the non-MEDLINE component will use less robust non-MeSH search strategies, but its results will be limited to only include the 11.1% of citations that are not indexed in MEDLINE (Figure 2).

Overview of the Master Search Strategy. Capital letters A, B, C, and D represent different or synonymous search terms in combination or separate. An “*” represents a truncated search term. MeSH, medical subject headings.
MeSH Strategies
The discovery of MeSH terms relevant to the chosen topic area is the foundation of an effective search strategy, as this prequalified collection of citations will comprise approximately 89% of the final search results. There are 2 main tools to determine which MeSH terms are most appropriate for a given search topic: (1) the MeSH database and (2) the “MeSH Terms” listed underneath the cataloged abstract of a relevant MEDLINE-indexed article.
Key concept 1: The MeSH database
The MeSH database uses a Unified Medical Language System to automatically map a search term to its relevant MeSH terms and may be accessed from any PubMed page. A drop-down box to the left of the main PubMed search box may be activated and the “MeSH” option selected (Figure 3). Then, any term typed in the search box, providing it translates to MeSH terminology, will lead to a page of possible MeSH terms. Figure 3 provides an example using the search word “fasting.” This word yields 3 MeSH results (fasting, hypoglycemia and Angptl4 protein, mouse). The user may then click on any MeSH term to discover its definition, its possible subheadings, and its place in the MeSH hierarchy, allowing the user to view which broader term the MeSH term came from and which subterms that term includes (Figure 4; the MeSH term “fasting” was selected). In this way, the user may be very clear on exactly which article topics the term will cull if used in a search. This allows the user to restrict the search to one or more smaller MeSH terms or to conflate terms by choosing a single broader MeSH term to encompass them all. For example, “feeding behavior” is the next broadest term after “fasting.” This MeSH term will include the articles from the “fasting” MeSH folder, as well as other behavioral topics surrounding feeding. The user may also restrict a search to yield only citations wherein the MeSH term is the key topic of the article or restrict the search so that it does not contain any of the subterms within the main MeSH term.
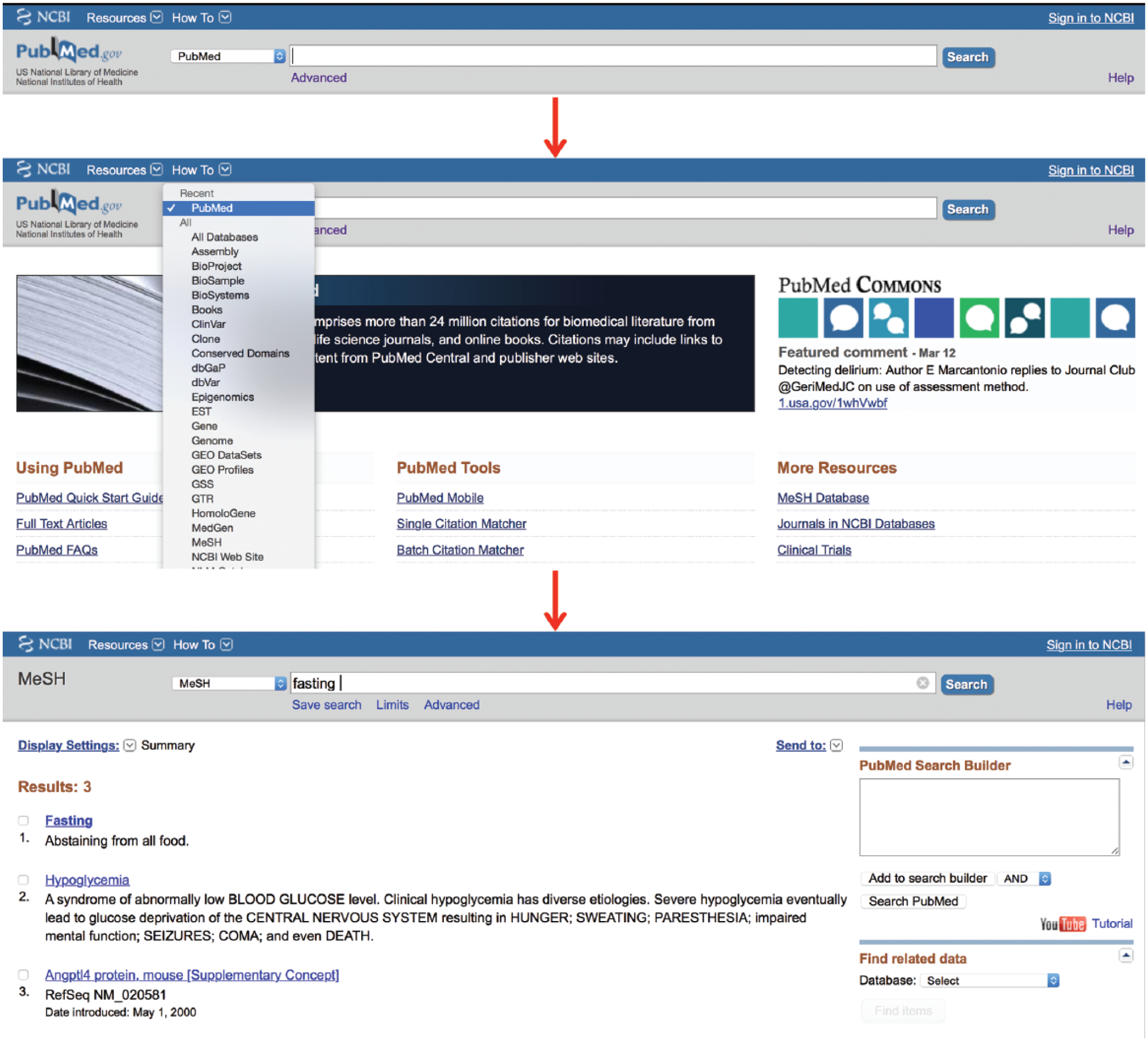
The medical subject headings (MeSH) database.
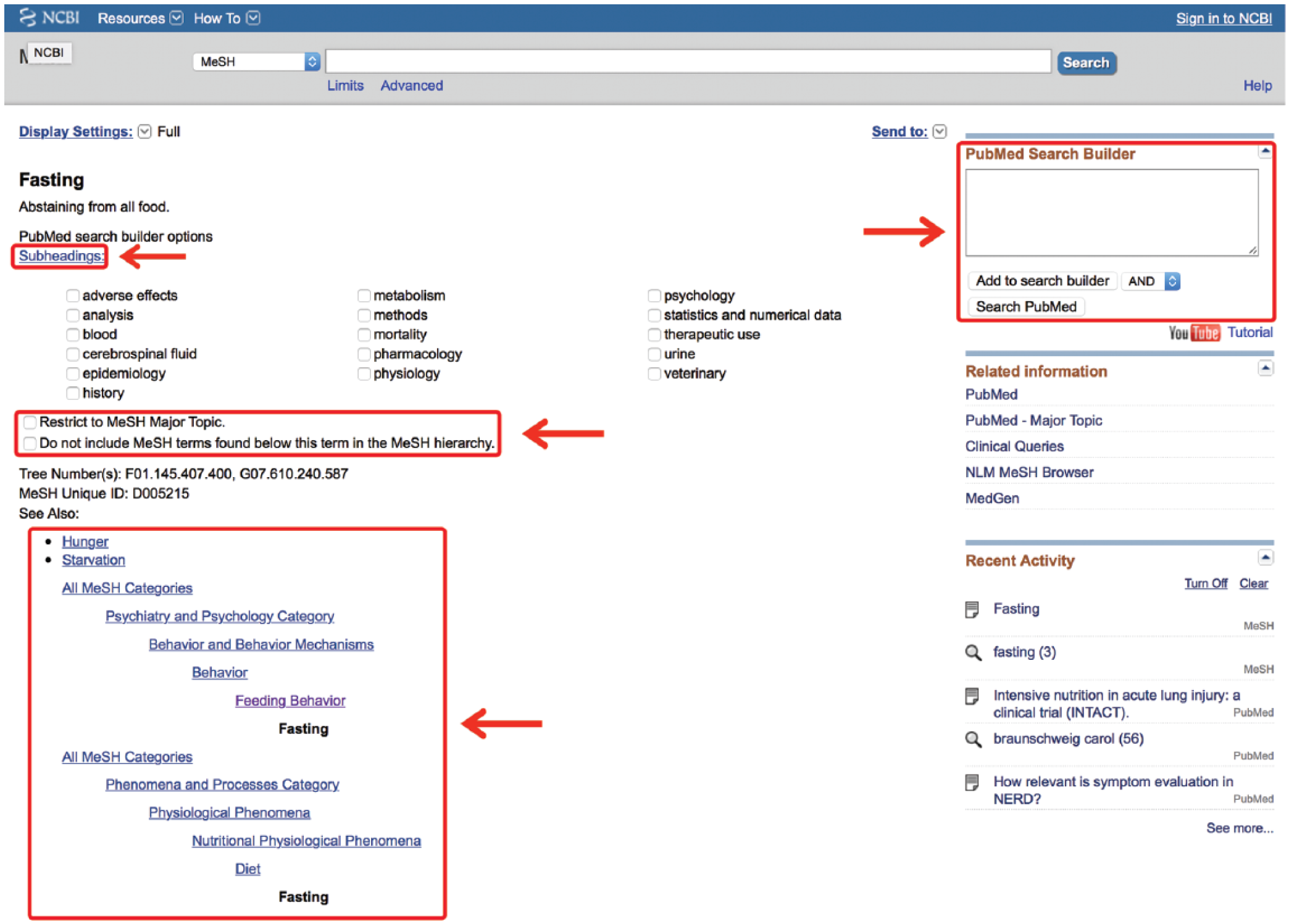
Inside the medical subject headings (MeSH) term.
Key concept 2: Articles of known relevance
Often, it is helpful to search documents of known relevance to the search topic to assess which MeSH terms have been ascribed to that article. This may be done simply to make sure the users did not miss any terms, which may be helpful to their search, or because the non-MeSH term they searched did not successfully translate into a MeSH term. This is accomplished by clicking on the link located below the article’s abstract. This link typically contains a string of names, which vary from article to article. Provided the article has been MEDLINE indexed, one of those names will be “MeSH Terms” (Figure 5). Clicking on this link provides the user with all the possible MeSH terms under which this article may be found. This list may then be searched for other synonymous or relevant MeSH terms that match the concepts being queried. It should be noted that some terms have an asterisk (*) after their names. This is used to signify a term that represents a main topic of the article. All other MeSH terms listed will have been discussion points in the article but were not the main thrust of the article.
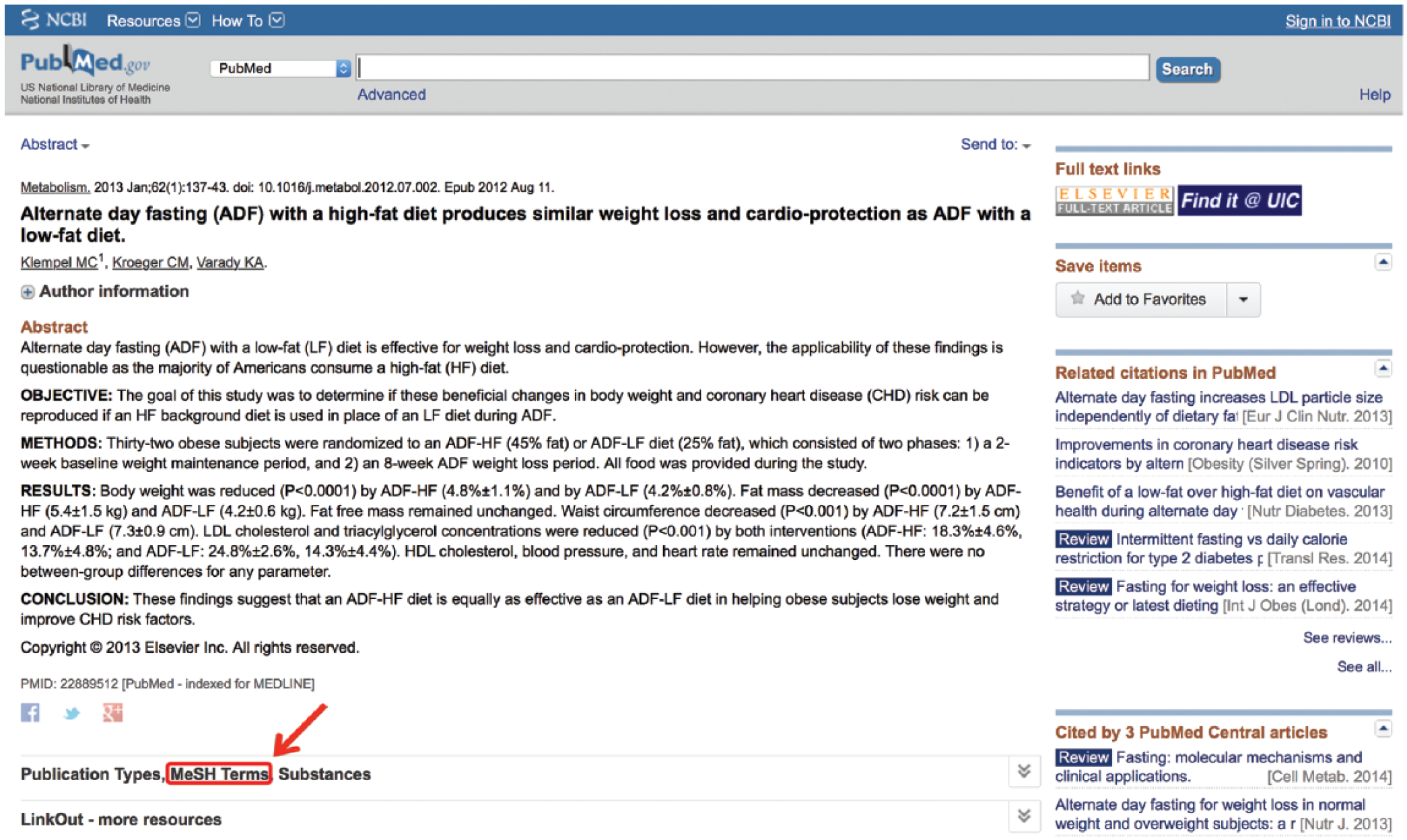
Search medical subject headings (MeSH) terms of relevant articles.
Key concept 3: Working with Boolean operators
Three highly useful commands may be used in both PubMed and MEDLINE searching systems. They are
Perhaps the most appropriate use of

Click the “Advanced” link.
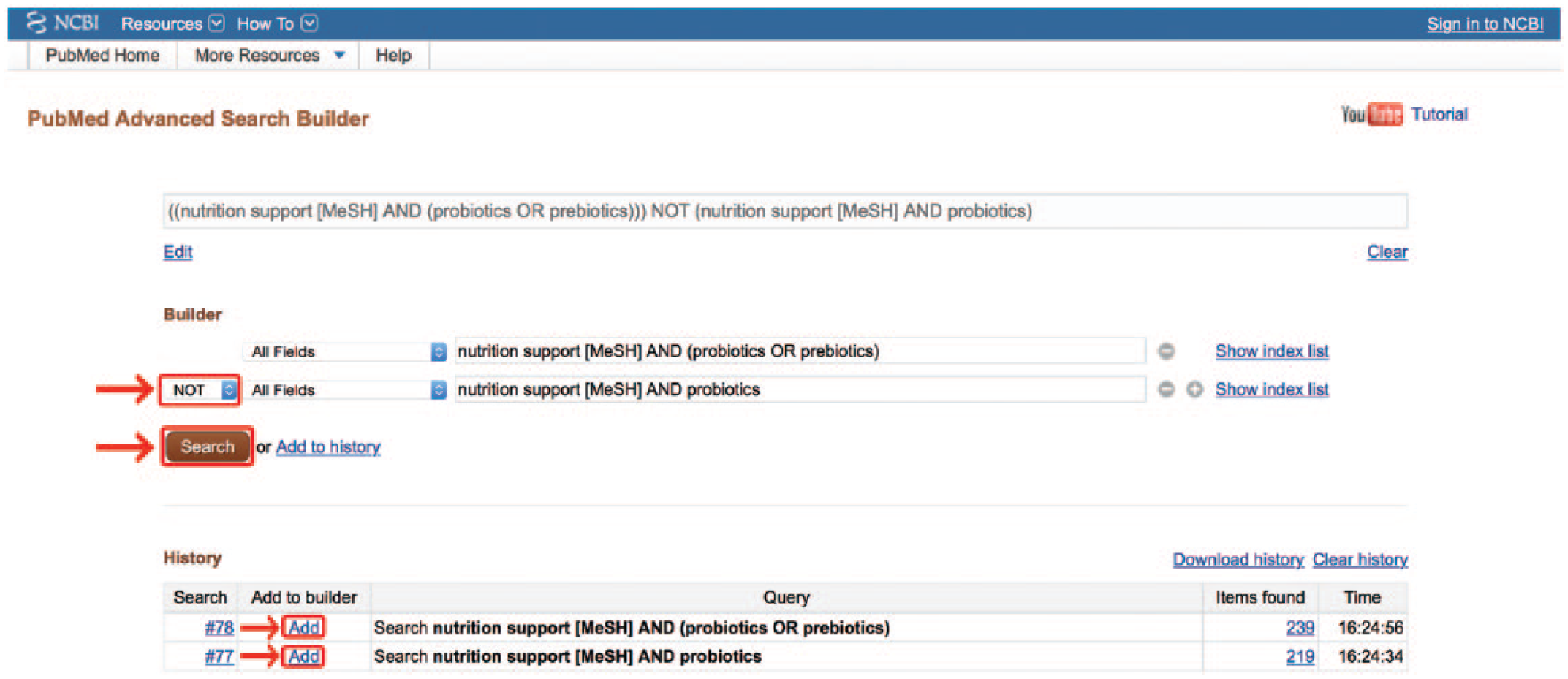
Boolean manipulation of entire searches. MeSH, medical subject headings.
Often, a search strategy contains several terms connected by many Boolean operators. When this occurs, it becomes important that these terms are grouped properly. Similar to algebraic equations, parentheses are used to delineate whether the operator is meant to act upon a single term, a grouping of terms, or even upon 2 or more full searches.
1
For example, (a
Key concept 4: The MeSH Search Builder
The PubMed Search Builder box, located on any MeSH Term page, allows the user to directly add the page’s MeSH terms to a search strategy and modify them with Boolean operators. 11 From the web page of the chosen MeSH term, one simply clicks “Add to Search Builder” and chooses the appropriate Boolean operator from the drop-down menu (Figure 4). The MeSH term will then remain in the box as other terms are searched, allowing the user to build on that term. The Search Builder tool will attempt to use parentheses to group the items properly, but often the user will need to alter the parenthetical grouping of the terms. Fortunately, the search box is directly editable, allowing users to perfect their MeSH search before submission. Once the search is built, clicking “Search PubMed” will yield a robust list of article citations.
Non-MeSH Search Strategies
Depending on the availability of MeSH terminology for a particular search topic, non-MeSH strategies for both the MEDLINE and non-MEDLINE portion of the search may be necessary. Unlike the MeSH database, which uses the intelligence of trained humans to catalog the
Key concept 5: Non-MeSH search tips
The following tips1,8,11 may be helpful when constructing a non-MeSH search:
Use “quotations” to search a specific phrase or exact spelling of a word.
Be sure to include alternate spellings (eg, (color OR colour)).
Use a thesaurus to look up common synonyms or related words.
Include plural versions of appropriate words.
Include any acronyms with and without periods.
Include different endings for the same word (eg, microbiology, microbiologist, microbiologia, etc).
Using an asterisk (*) to truncate a word (eg, microbiolog*) will automate the culling of plurals and alternate suffixes as well as include all subheadings associated with the word in MeSH.
Key concept 6: The Search Details box
In a nonqualified default search, PubMed attempts to interpret the search terms into MeSH terms or non-MeSH variations. Unfortunately, its efforts are often incomplete or incorrect. If a search phrase is entered into PubMed nonqualified, it will first attempt to find MeSH terms to match the words. Often, the MeSH terms chosen will match the words but not the meaning of the phrase. For example, a search for
This valuable information regarding how PubMed interpreted the search may be found in the “Search Details” box on the middle right side of any PubMed results screen (Figure 8). 11 The Search Details box is an invaluable tool for diagnosing problems in a search strategy. It instantly tells the user whether MeSH terms were found, lists what liberties PubMed may have taken to expand the search, and provides a snapshot of the cause-and-effect relationship between a search strategy and its outcome.
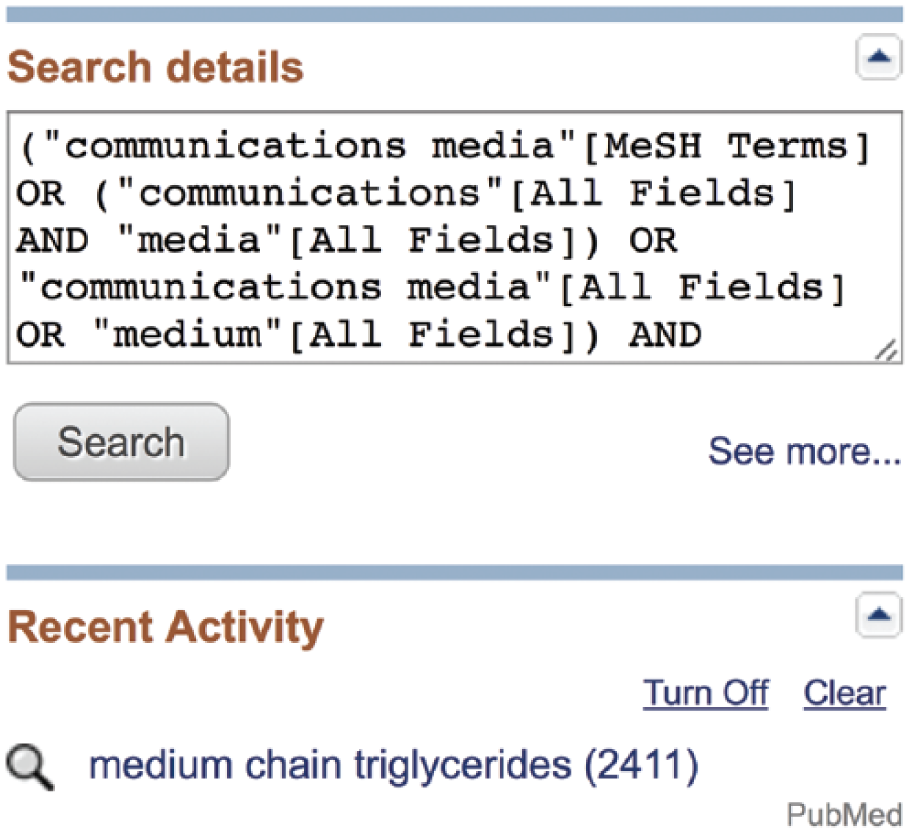
Search details box.
Key concept 7: Filters
As previously stated, any exhaustive search strategy will have a MEDLINE component and a non-MEDLINE component. The MEDLINE component should use MeSH-based strategies wherever possible and non-MeSH strategies only where necessary. The non-MEDLINE component must use only non-MeSH strategies, because only MEDLINE articles are cataloged with MeSH terms. It is then necessary to restrict the non-MEDLINE portion of the search to only the small fraction of citations that are not indexed in MEDLINE. Several filters, known as “subsets,” select for these non-MEDLINE citations. Subset filters may be recognized by their tag, [sb]. Examples of non-MEDLINE subset filters include publisher[sb], inprocess[sb], oldmedline[sb], and pubmednotmedline[sb].
12
The simplest filter that will collect citations from all 4 of these categories is to add a Boolean
It is important to acknowledge that any articles culled from the non-MEDLINE portion of the search may become MEDLINE articles by the time another researcher attempts to reproduce the search. If one is performing a systematic review, it is imperative for methods reproducibility that they provide a list of which articles were chosen from the non-MEDLINE search to account for their repositioning from the PubMed to MEDLINE indexing system.
Some Further Filter Issues
For the convenience of the user, PubMed makes several filters available on the left-hand side of any search results page with checkboxes.
14
Many of these automatic filters on the left of the main search page are MeSH-based filters. They will only pull articles that have been indexed with MeSH terms. When these MeSH filters are selected, they are automatically attached to the entire current search with an
To enter these filters by hand, one must know how they are coded. Discovering this involves entering any term into the search box and clicking “Search.” Then, deleting that term and checking the box of the desired filter will cause the “Search Details” box to reveal how that particular MeSH term is catalogued (Figure 9). An alternative option is to find the term in the MeSH database and add it with the PubMed Search Builder discussed earlier. In either case, that exact term, including its [MeSH] tag, may be inserted into the MeSH section of the search with an
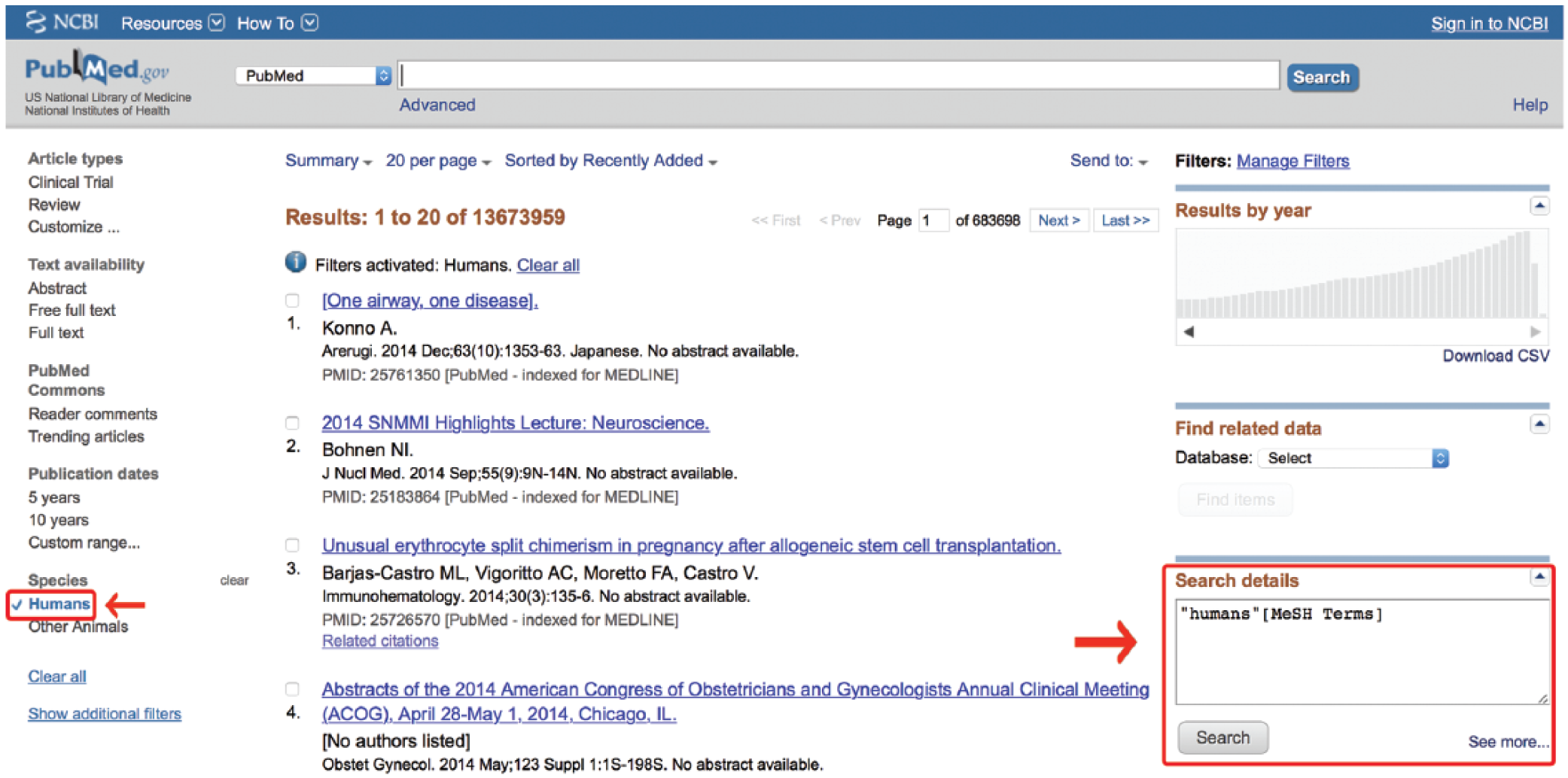
Obtaining a filter tag from the Search Details box. MeSH, medical subject headings.
Key concept 8: The Master Search Strategy
With all the components of an exhaustive but efficient search in place, it is possible to assemble the search strategy. Below is a simplified way to state the basic Master Search Formula for PubMed from Figure 2:

This formula allows the users to use their knowledge of MeSH terms to focus their search on human-cataloged MEDLINE articles while limiting their default PubMed search to only those articles that have no MEDLINE equivalents. In this way, their search will be exhaustive but efficient.
Part II: A Step-by-Step Tutorial
The previous sections of this article have outlined the components of the Master Search Strategy. This next section demonstrates these components in action as one performs a Master Search. A screen-capture tutorial of this process may be found at www.ResearchRDN.com. Building a Master Search occurs in phases. Some phases are simple and linear, while others require more reflection and exploration of terms. Key concepts (KCs) from the previous sections of this article will be referred to throughout the search to provide clarity. While learning, it is helpful to open a document where one may copy and paste the components of the search strategy as they are created. These components may then be assembled within that document and pasted back into the PubMed search box once the search strategy is complete.
As an example, consider the nonqualified search terms alternate[All Fields] AND day[All Fields] AND (“fasting”[MeSH Terms] OR “fasting”[All Fields])
Notice that the only term that mapped to a MeSH term was “Fasting”[MesH]. The current search is indeed 2 separate searches. One is searching within the “Fasting”[MesH] term for citations that also include the words “alternate”
The MEDLINE Portion
We will begin by constructing the MEDLINE portion of the search.
1. From the PubMed.gov homepage, make sure the “MeSH” option is selected from the drop-down menu to the left of the PubMed search box.
2. Enter the word “Fasting” and click “Search.”
This will display a page with the term’s description, possible subheadings, and a visual of the term’s place within the hierarchy of MeSH terms (Figure 4) (
3. Perform a similar search for “Alternate” and “Day.”
When “Alternate” and “Day” are searched in the MeSH database, the Search Detail box reveals they do not directly translate to MeSH terms. It may be useful to search an article in PubMed of known relevance to alternate-day fasting (
4. Click on the drop-down box to the left of the PubMed search box and change the selection from “MeSH” to “PubMed.”
5. Type the name of the above article into the PubMed search box and click “Search.”
Below the abstract of this article in PubMed is a string of links entitled
6. Click on “Publications Types, MeSH Terms, Substances.”
It appears from this list that some additional relevant MeSH terms may include “obesity/diet therapy” and “weight loss/physiology” (Figure 10). The terms after the “/” signify subheadings. Record these terms. They will be added to the search shortly. At this point, it may be beneficial to further restrict the search to human subjects.
7. Check the humans box on the left-hand side of any PubMed search results page and look in the Search Details box to discover its code (Figure 9) (
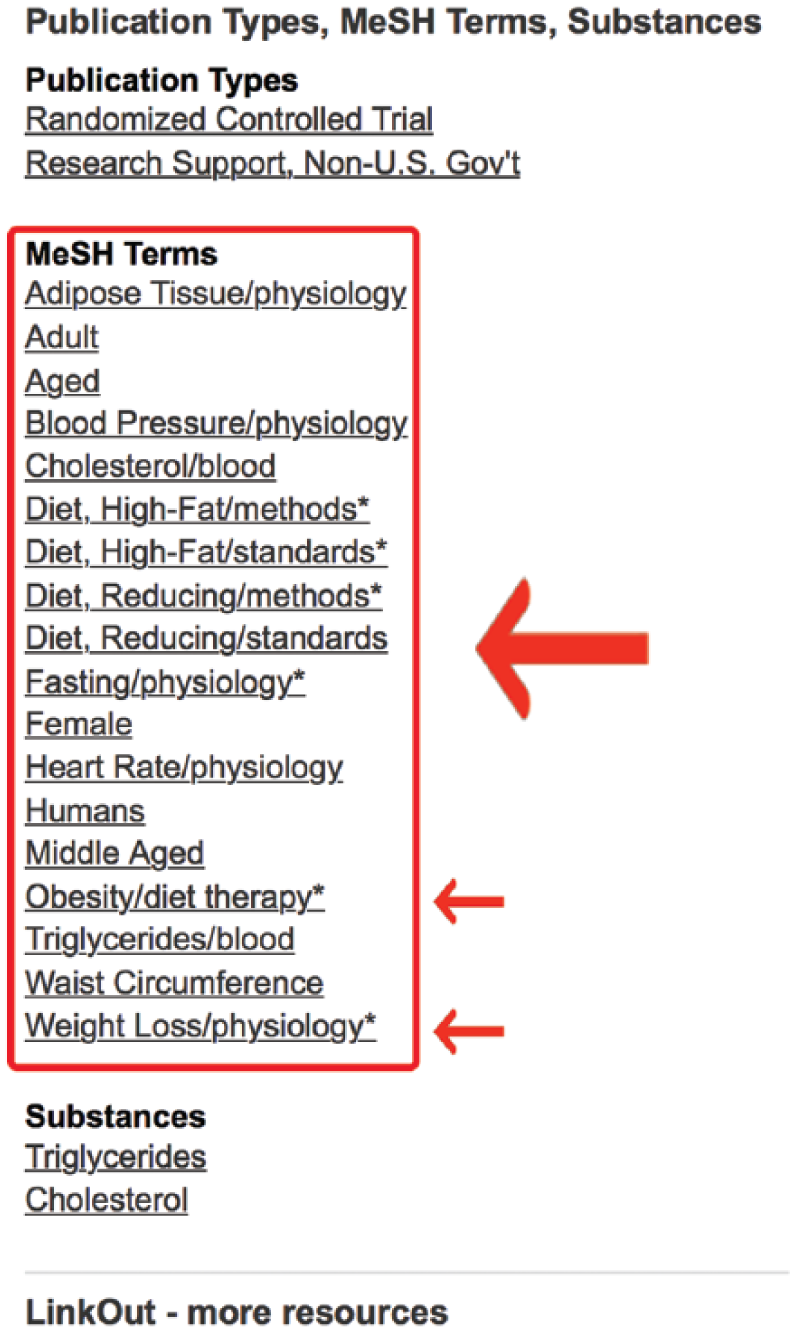
Medical subject headings (MeSH) terms for article of known relevance.
In this case, “Humans” is a MeSH term (“Humans”[MeSH]), so it would go on the MEDLINE side of the search strategy, connected with a Boolean
Now that all the relevant MeSH terms are known, it is time to build the MeSH portion of the search. Begin by systematically building the search in the Search Builder utility in the MeSH database (
8. Click the drop-down menu next to the search box and select “MeSH.”
9. Type in the MeSH term “Fasting” and click “Search.”
10. Select the “Fasting” MeSH term from the resulting search output page. (This will load the dedicated “Fasting” MeSH term page.)
11. Click “Add to Search Builder.”
12. Return to the main “MeSH” search box at the middle of the screen and type “Obesity” and click “Search.”
13. Select “Obesity” from the result MeSH Search results page.
14. From the list of checkboxes, check the subterm “diet therapy.” (Notice the term “Fasting” has been retained in your Search Builder box so that you may simply add to it.)
15. Set the drop-down Boolean operator in the Search Builder box to “
16. Return to the MeSH search box at the top middle of the screen; type in the known MeSH term, “weight loss”; and click “Search.”
17. Select “Weight Loss” from the output screen and check the subheading “physiology” on the resulting MeSH term page.
18. Set the Boolean operator to
19. Return to the MeSH search box in the top middle of the screen, type “Humans,” and click “Search.”
20. Click on “Humans” in the resulting output.
21. Set the Boolean operator to
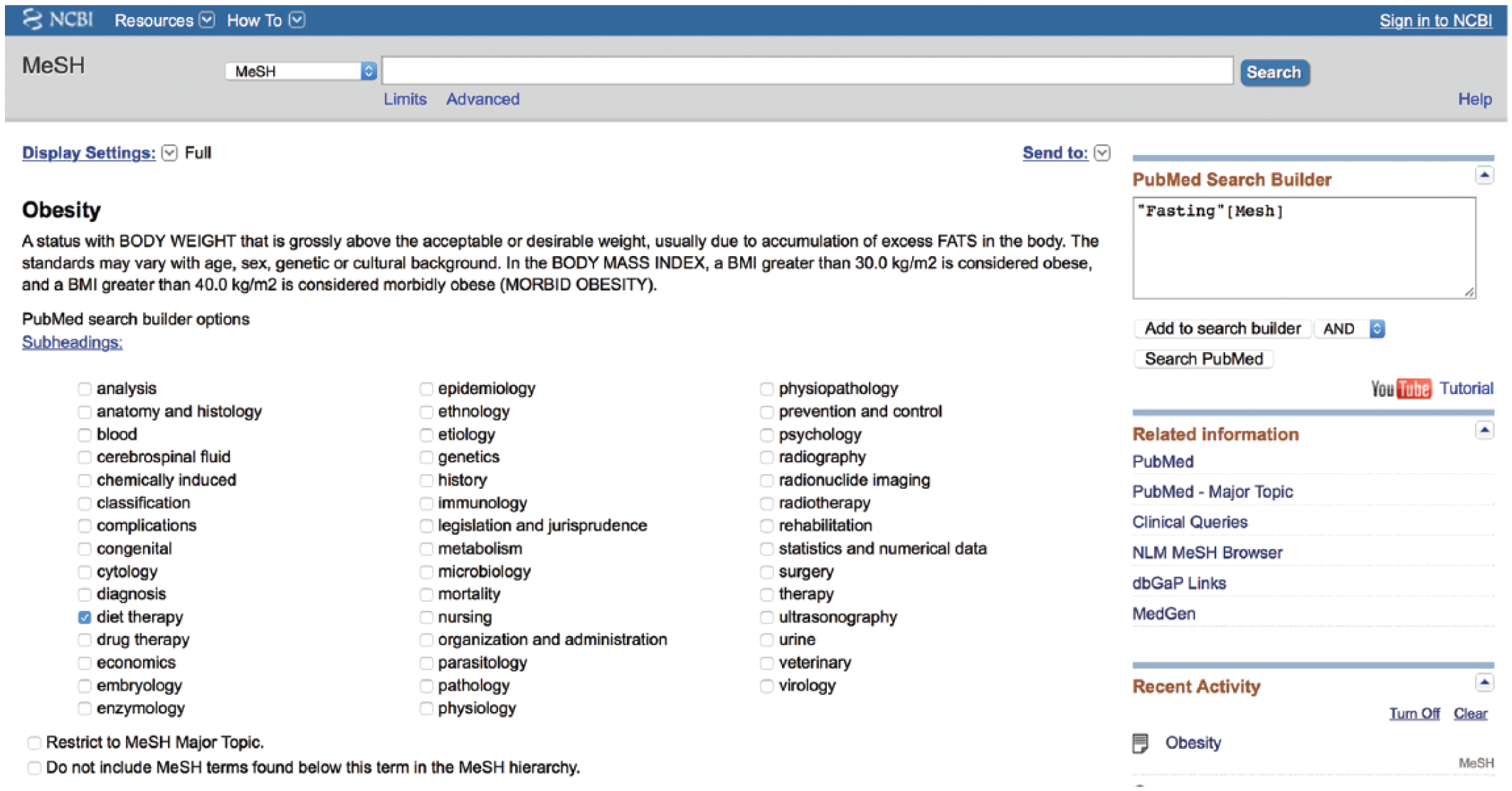
Building the medical subject headings (MeSH) portion of the search (example: obesity/diet therapy [MeSH]).
This will create a grouping of properly worded MeSH terms all connected with an
((“Fasting”[MeSH] OR “Obesity/diet therapy”[MeSH] OR “Weight Loss/physiology”[Mesh]) AND “Humans”[MeSH])
Recall that the words “Alternate” and “Day” did not map to MeSH terms. These may still be added as text terms to the MEDLINE search. The rules associated with non-MeSH strategies may be used, such as creating suitable synonyms and related words (
22. Create a string of synonyms for any words that did not map into MeSH terms.
23. Connect them to each other with Boolean
24. Connect this grouping of synonyms to the rest of the MEDLINE search with a Boolean
25. Contain the entire MEDLINE strategy in its own set of parentheses
This completes the MEDLINE portion of the search strategy (see below).
The (((“Fasting”[MeSH] OR “Obesity/diet therapy”[MeSH] OR “Weight Loss/physiology”[Mesh]) AND “Humans”[MeSH]) AND (“alternate day fasting” OR “fasting on alternating days” OR “alternate-day fasting”))
This alone yields 17 relevant citations but may be missing newer articles that have not yet been indexed in MEDLINE.
The Non-MEDLINE Portion
The non-MEDLINE portion requires the creation of synonyms, alternate suffixes, and any other combination of necessary terms (
26. Create a string of synonyms for the expression “alternate day fasting” that you might expect to find in the title or abstract of a paper relevant to your topic.
27. Connect them to each other with Boolean
28. Outside of these parentheses, connect the MEDLINE filter with the Boolean operator
The non-MEDLINE portion of the search strategy could be assembled as follows:
((“Alternate Day Fasting” OR “fasting on alternating days” OR “alternate-day fasting”) NOT medline[sb])
This yields 12 results.
29. Place the entire non-MEDLINE search in its own set of parentheses and connect it to the MEDLINE search with a Boolean
30. Switch the search drop-down box to PubMed if needed.
31. Copy and paste the completed search strategy into the main PubMed search box.
32. Click Search.
The combined master search is
(((“Fasting”[MeSH] OR “Obesity/diet therapy”[MeSH] OR “Weight Loss/physiology”[Mesh]) AND “Humans”[MeSH]) AND (“alternate day fasting” OR “fasting on alternating days” OR “alternate-day fasting”)) OR ((“Alternate Day Fasting” OR “fasting on alternating days” OR “alternate-day fasting”) NOT medline[sb])
This yields a manageable 29 exhaustive yet highly efficient results covering 13 years of research on the topic of alternate-day fasting in humans.
Automated Citation Updates: Saved Searches in MyNCBI
Once a search strategy has been created, PubMed has a powerful tool to help any researcher or clinician stay as current in his or her field as possible. Once the past literature has been reviewed through an exhaustive search strategy, that strategy may be saved into a MyNCBI account. MyNCBI is a free service offered by the National Center for Biotechnology Information. By clicking on the MyNCBI link in the upper right corner of the PubMed search page, the user is led through a series of steps to set up a free account (Figure 12). Once completed, MyNCBI may be used to save a current search strategy by clicking “Create Alert” under the search box. Here, the user is given the option to set up a plan whereby he or she may receive an email monthly, weekly, or daily with any new citations indexed in PubMed that meet his or her search criteria (Figure 13). In this way, a well-created search strategy allows the user to be the first to read any new articles related to his or her search.

Signing into MyNCBI.
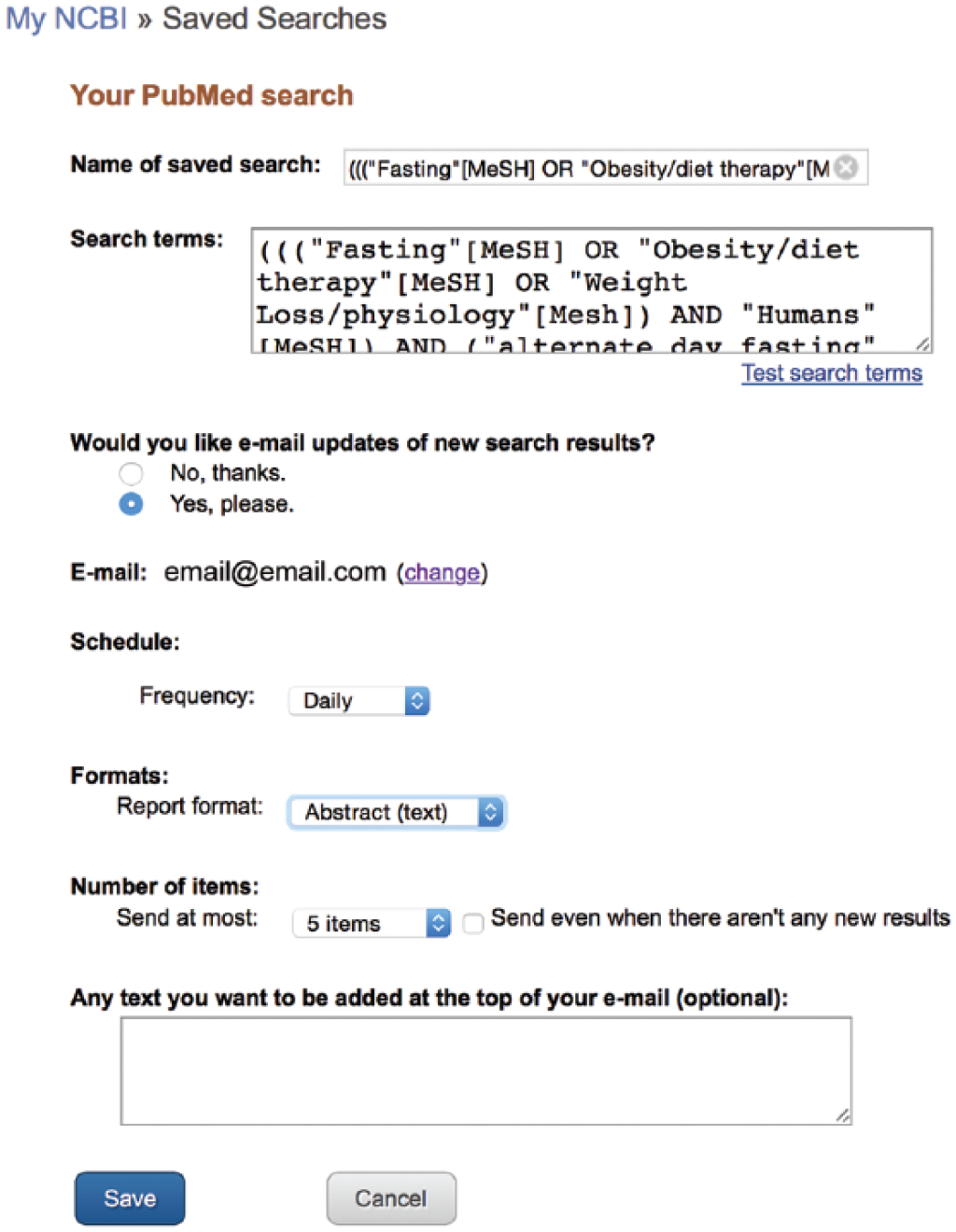
Setting up automated citation delivery.
Steps to Assembling a Master Search Strategy (Summary)
The steps to assembling a master search strategy may be summarized as follows:
Determine which terms best describe the search concept.
Create the MEDLINE portion of the search using the MeSH database to see if the search terms map to acceptable MeSH terms and articles of known relevance to capture any terms you may have missed.
For terms that map well, use the PubMed Search Builder to construct a MeSH search strategy.
If a portion of the terms does not map well to MeSH terms, create a grouping of words using the non-MeSH default search techniques. Connect the words with
When the entire MEDLINE portion of the search is complete, click “Search.”
To construct the non-MEDLINE portion of the search, use non-MeSH techniques to generate appropriate search terms, connecting the synonyms with
Next, filter for only non-MEDLINE citations by connecting the filter “medline[sb]” to the non-MEDLINE search with a Boolean
Connect the MEDLINE search to the non-MEDLINE search with a Boolean
Log into MyNCBI and click on “Create Alert,” selecting the option of receiving daily updates for any new citations that meet the search parameters.
A tip for those who do not have access to a medical library: There is a simple filter that will allow users to search only articles that offer free access to their publications. These results may be filtered by connecting
Example:
For articles that are not automatically free in PubMed, it may be worthwhile to visit the journal’s homepage. Often, simply registering with the journal will be enough to allow access to free articles. If no free method is available, the main service for ordering articles is Loansome Doc. Loansome Doc connects PubMed users with medical libraries for the delivery of paid articles. To learn more and set up a Loansome Doc account, visit their webpage. 16
Conclusion
The ability to competently search a citation database is the foundation of all evidence-based practice and research. Timely access to the right information is the currency of an expert in any field. The MEDLINE system continues to be a powerful resource for prequalified information but only if intelligently used and combined with select subsets of non-MEDLINE articles. Armed with these skills, researchers and clinicians may be assured that their recommendations are well informed and that they have performed a truly exhaustive review of the literature.
Footnotes
Statement of Authorship
L. McKeever, V. Nguyen, S. J. Peterson, S. Gomez-Perez, and C. Braunschweig contributed to the conception or design of the research; L. McKeever contributed to the acquisition, analysis, and interpretation of the data; L. McKeever drafted the manuscript; L. McKeever, V. Nguyen, S. J. Peterson, S. Gomez-Perez, and C. Braunschweig critically revised the manuscript; and L. McKeever, V. Nguyen, S. J. Peterson, S. Gomez-Perez, and C. Braunschweig agree to be fully accountable for ensuring the integrity and accuracy of the work. All authors read and approved the final manuscript.
Financial disclosure:
None declared.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
