Abstract
Objective:
To evaluate an embedded artificial intelligence (AI) system for otoscopic image-based screening and prediagnostic triage of pediatric otitis media, with an emphasis on point-of-care deployment on a microcontroller.
Methods:
We retrospectively analyzed 19 522 tympanic membrane images labeled by otolaryngologists as acute otitis media, otitis media with effusion, or normal. A lightweight convolutional neural network derived from AlexNet was trained using a patient-level split. Two posttraining INT8 variants (per-channel and per-tensor quantization) were deployed on an STM32H7S78-DK board using STM32Cube.AI. We evaluated accuracy on a held-out test set (n=600) and measured activation memory, total random access memory (RAM) usage, and average inference latency on the target device.
Results:
The full-precision model achieved 97.67% test accuracy. Per-channel INT8 preserved accuracy (97.67%). Per-tensor INT8 showed a small decrease (97.50%) but reduced activation memory by 16%, total RAM by 8%, and latency by 13%.
Conclusion:
On-device otoscopic image analysis is feasible on an STM32H7-class microcontroller and may support screening and triage workflows (eg, repeat examination, tympanometry, monitoring, or referral). The tool is not intended to provide a final diagnosis or to direct treatment decisions; clinical diagnosis and management remain clinician-led.
Introduction
Otitis media is among the most common pediatric conditions evaluated in routine practice and remains a major driver of antibiotic use and follow-up visits.1 -5 Differentiating acute otitis media (AOM), otitis media with effusion (OME), and normal tympanic membranes can be difficult, particularly in high-volume outpatient clinics and telemedicine workflows where image quality and operator technique vary.6 -10 In settings with limited access to otolaryngology expertise, decision support that improves consistency of image interpretation may help prioritize appropriate next steps, while preserving clinician oversight.11,12
Deep-learning models for otoscopic image classification have shown promising results, including evidence from systematic reviews and meta-analysis.12 -19 However, many implementations rely on cloud or workstation resources, which can introduce latency, connectivity requirements, and deployment constraints.20 -22 Tiny machine learning enables on-device inference under strict memory and compute budgets and is well suited to handheld or portable workflows.21,22
In this study, we present an embedded, point-of-care otoscopic image screening system designed to support prediagnostic triage for pediatric otitis media. Using a specialist-labeled dataset of 19 522 tympanic membrane images, we trained a lightweight AlexNet-derived CNN 23 to classify AOM, OME, and normal findings and then deployed INT8-quantized variants on an STM32H7-class microcontroller. Our main contribution is a workflow-oriented evaluation that links model outputs to practical next steps (eg, repeat examination, tympanometry, monitoring, or referral) and quantifies the on-device trade-offs that determine feasibility in real clinics, including random access memory (RAM) footprint and inference latency. The system is intended as decision support for screening and triage rather than a standalone diagnostic device; final diagnosis and management remain clinician-led and should incorporate symptoms, history, and objective testing when indicated.
Materials and Methods
Dataset and Labeling
Otoendoscopic tympanic membrane images were retrospectively collected at Shenzhen Children’s Hospital (January 2016 to December 2019) from pediatric patients younger than 18 years, including infants as young as 2 months. The study was approved by the Ethics Committee of Shenzhen Children’s Hospital, and all images were de-identified prior to analysis.
Images were acquired using multiple clinical systems, including Otocope 0° φ2.7 × 105 (Zhejiang Tiansong Medical Instrument Co., China), EndoSTROBD (XION GmbH, Germany), and EPK-i5000 VNL-1070STK (PENTAX, RICOH Imaging Company, Japan). Board-certified otolaryngologists labeled each image as AOM, OME, or normal tympanic membrane. The dataset contained 19 522 images: 6210 AOM, 7548 OME, and 5764 normal. Figure 1 shows representative pediatric otoendoscopic images for each category.
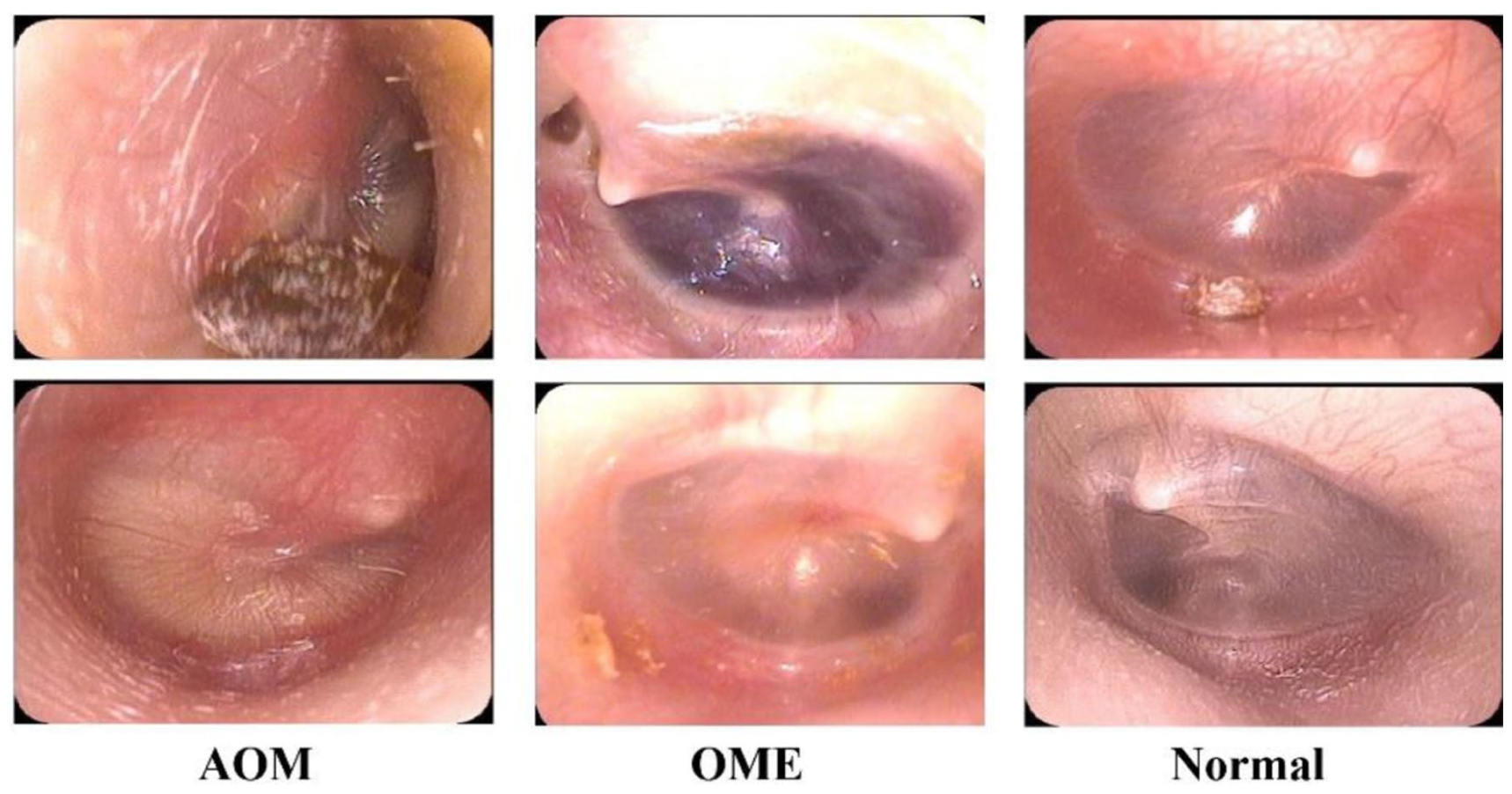
Representative otoendoscopic tympanic membrane images for AOM, OME, and normal. AOM, Acute otitis media; OME, Otitis media with effusion.
We prespecified a held-out test set of 600 images (approximately 3%) to provide stable estimates of overall and per-class performance while preserving a large development set under a patient-level split. With an expected overall accuracy of 0.97 to 0.98, a test set of 600 yields a 95% confidence interval with an approximate half-width of 1% for overall accuracy, and—given class-balanced sampling—supports per-class estimates with acceptable precision for a screening/triage tool evaluation. The remaining 18 922 images were used for model development. The held-out test set was constructed with 200 images per class (AOM, normal, and OME) to support balanced per-class evaluation, while preserving patient-level separation from the development set.
Model Architecture and Training
We trained a lightweight convolutional neural network derived from AlexNet for 3-class classification. 23 The classification model was based on a standard AlexNet architecture, consisting of 5 convolutional layers followed by 3 fully connected layers. The numbers of output channels in Conv1-Conv5 were 96, 256, 384, 384, and 256, respectively. ReLU activation was applied after each convolutional layer and after the first 2 fully connected layers, and max-pooling was used after Conv1, Conv2, and Conv5. Dropout (P = .5) was applied to the first 2 fully connected layers. For the present study, the final classification layer was adapted to output 3 classes (AOM, OME, and normal). No pruning was applied. The full layer-by-layer specification is provided in Table S1.
Training ran for 800 epochs with a batch size of 64. We used the Adam optimizer 24 and applied standard data augmentation (random rotation, scaling, and horizontal flipping) to improve robustness. 25 Early stopping was enabled based on validation loss, and the checkpoint with the best validation performance was selected for final testing. The training and validation learning curves are shown in Figure 2.
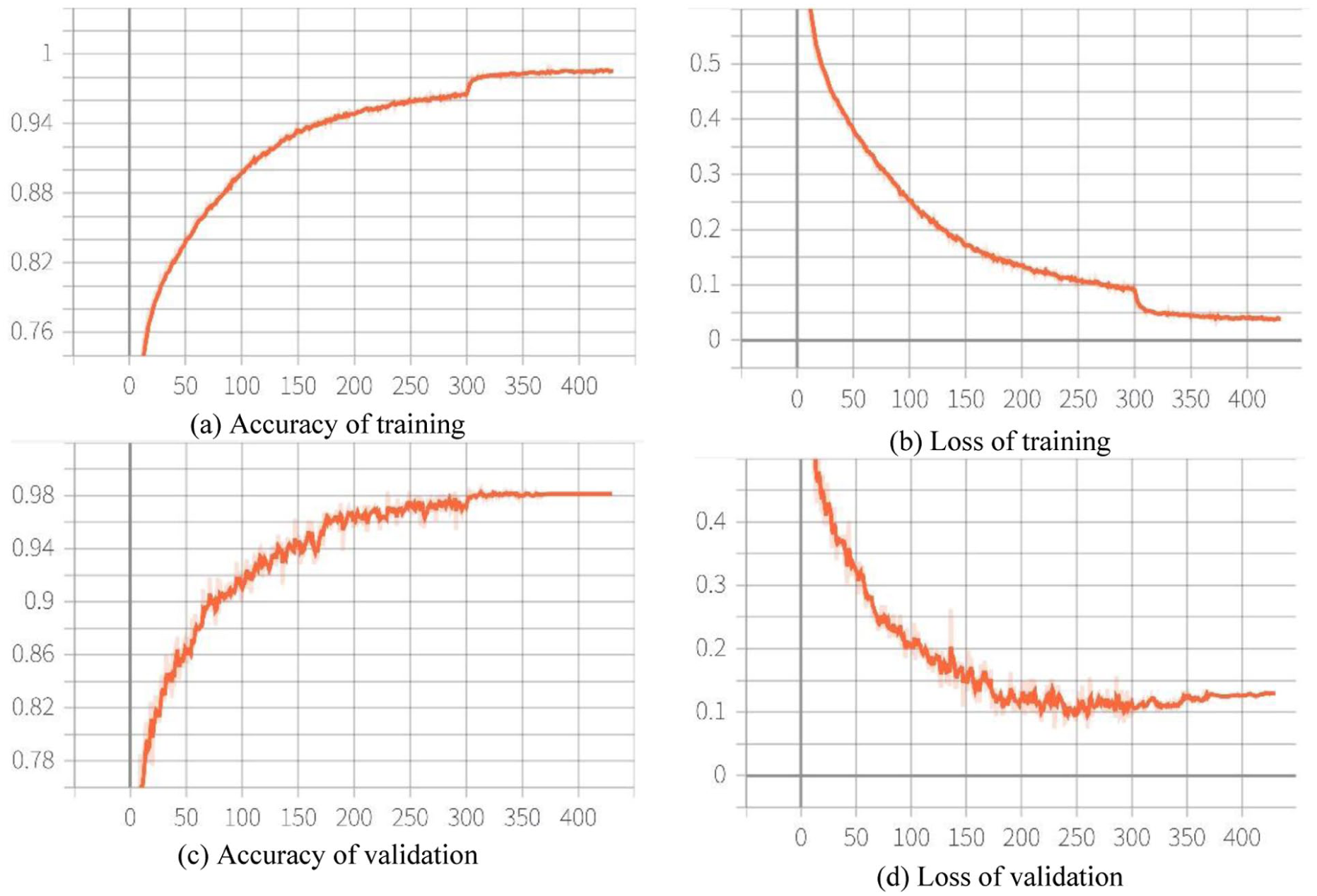
Training and validation accuracy and loss over 800 epochs. (a) Accuracy of training (b) Loss of training (c) Accuracy of validation (d) Loss of validation.
Quantization and Embedded Deployment
For microcontroller deployment, we implemented post-training INT8 quantization using 2 granularities: per-channel and per-tensor.26,27 Models were deployed to an STM32H7S78-DK board (ARM Cortex-M7) using the STM32Cube.AI toolchain (X-CUBE-AI).28,29 Model weights were stored in external Octo-SPI flash on the STM32H7S78-DK, while intermediate activations were allocated in on-chip static random access memory under a fixed firmware configuration. The key steps of deployment are shown in Figure 3. We measured activation memory, total RAM usage, compute-node count in the generated inference graph, and average inference latency under a consistent firmware configuration.
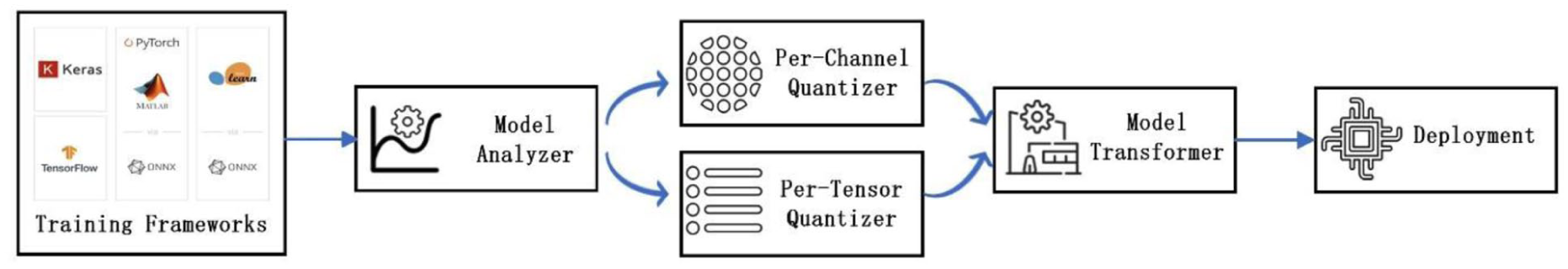
Embedded deployment pipeline and software stack.
Results
After deployment, classification performance was assessed on the held-out test set using overall accuracy and confusion matrixes. Embedded benchmarks were performed on the same test set to ensure comparability across quantization schemes.
On the held-out test set (n = 600), the full-precision model achieved 97.67% overall accuracy (Figure 4). Most errors occurred between AOM and OME, reflecting overlapping otoscopic appearances in borderline presentations.
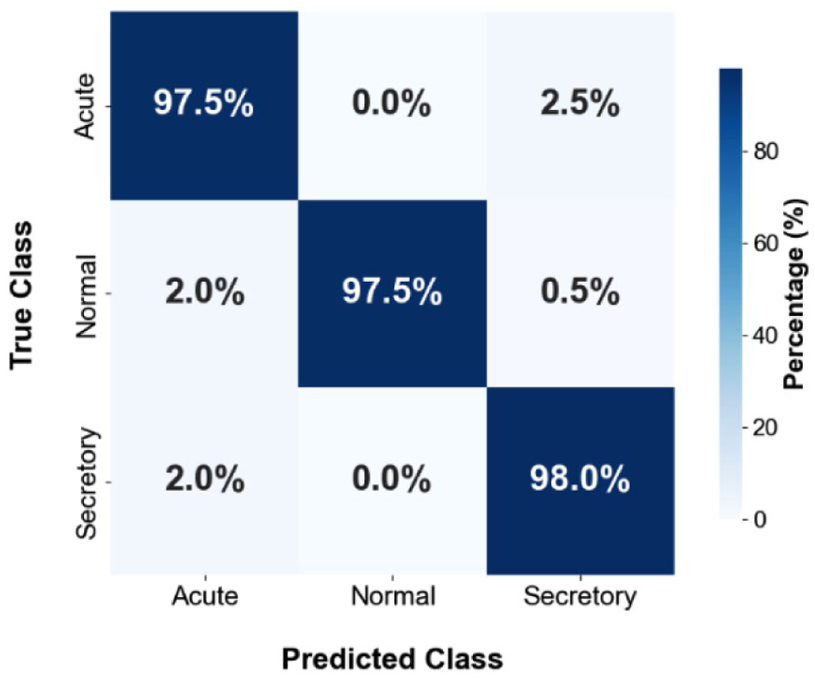
Confusion matrix of the original unquantized model on the held-out test set.
We benchmarked per-channel and per-tensor INT8 AlexNet deployments on the STM32H7S78-DK to quantify accuracy–efficiency trade-offs. As summarized in Table 1, per-tensor quantization reduced activation memory by 16% and total RAM by 8%, simplified the execution graph (9 vs 13 compute nodes), and lowered average latency by 13%. These gains come at the cost of reduced quantization granularity, whereas per-channel quantization better preserves numerical fidelity but with higher resource demand.
Resource Utilization Metrics for Per-Channel and Per-Tensor Models.
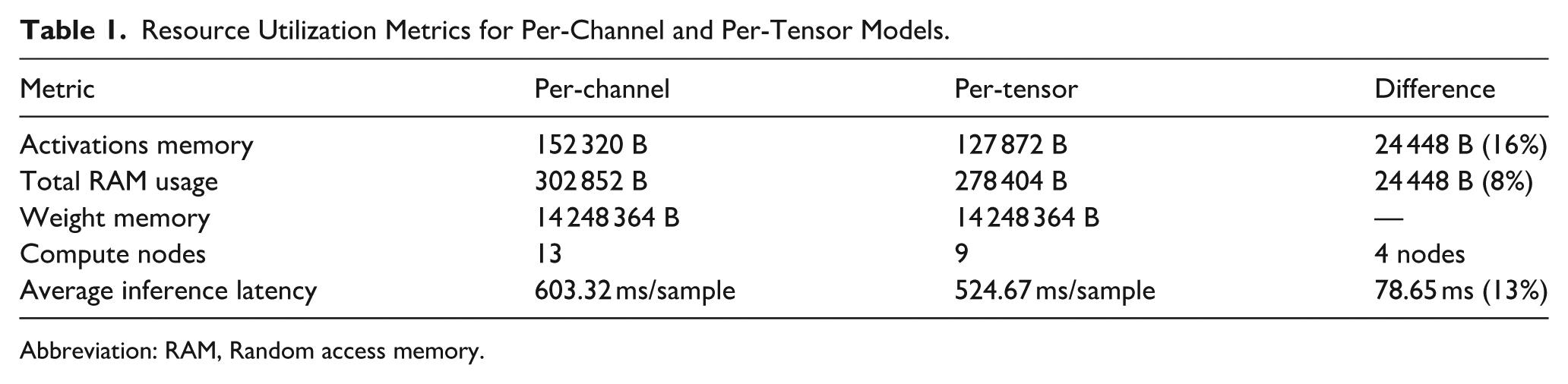
Abbreviation: RAM, Random access memory.
Confusion matrixes for both models (Figure 5) show subtle differences. Per-channel INT8 preserved test accuracy (97.67%, Table 2). Per-tensor INT8 produced a small decrease (97.50%, Table 2) while improving RAM and latency. The per-channel and per-tensor INT8 models showed similar classification performance on the held-out test set (97.67% vs 97.50%). Given the small absolute difference, the results are presented descriptively and are best interpreted in the context of deployment trade-offs rather than as evidence of superiority of 1 quantization strategy over the other.
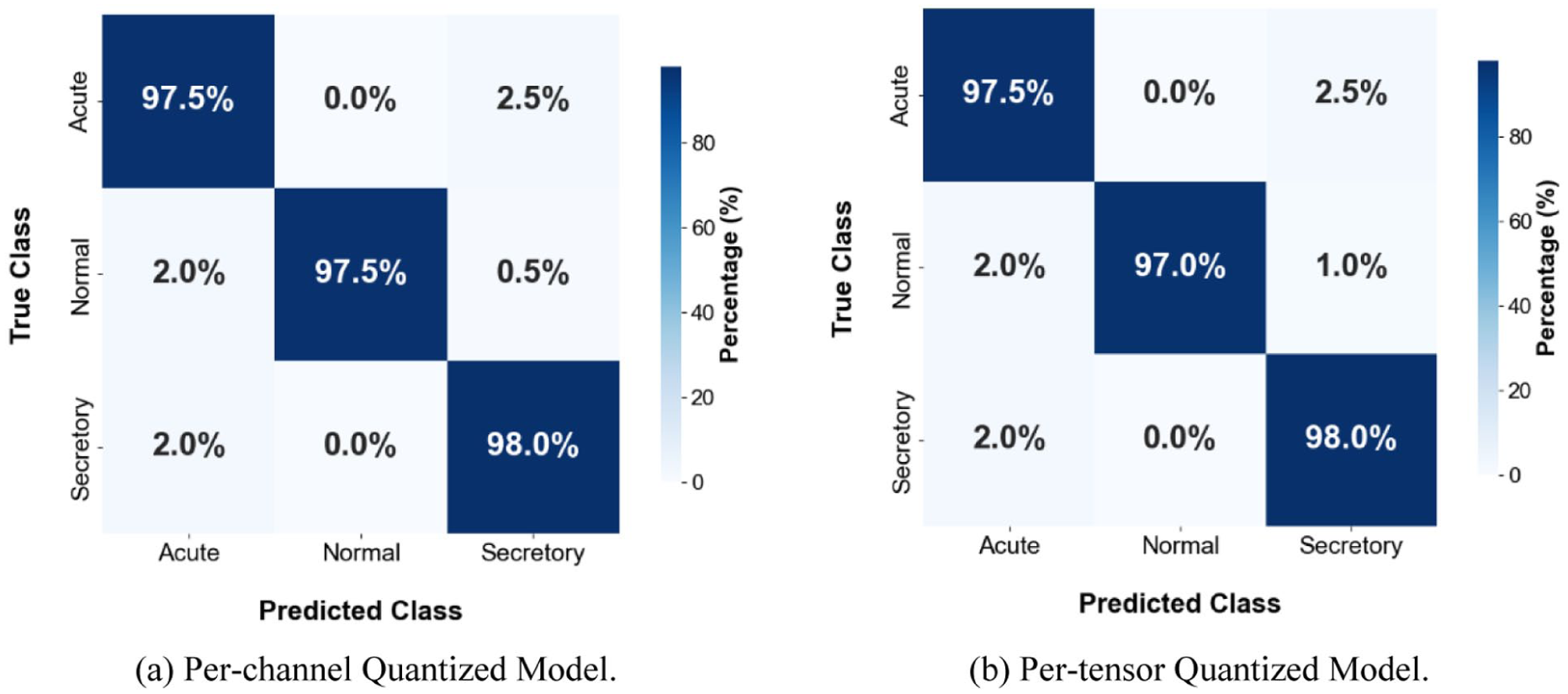
Confusion matrixes for INT8 per-channel and per-tensor models on the held-out test set (a) Per-channel quantized Model (b) Per-tensor quantized model.
Accuracy Comparison Between Per-Channel and Per-Tensor Models.
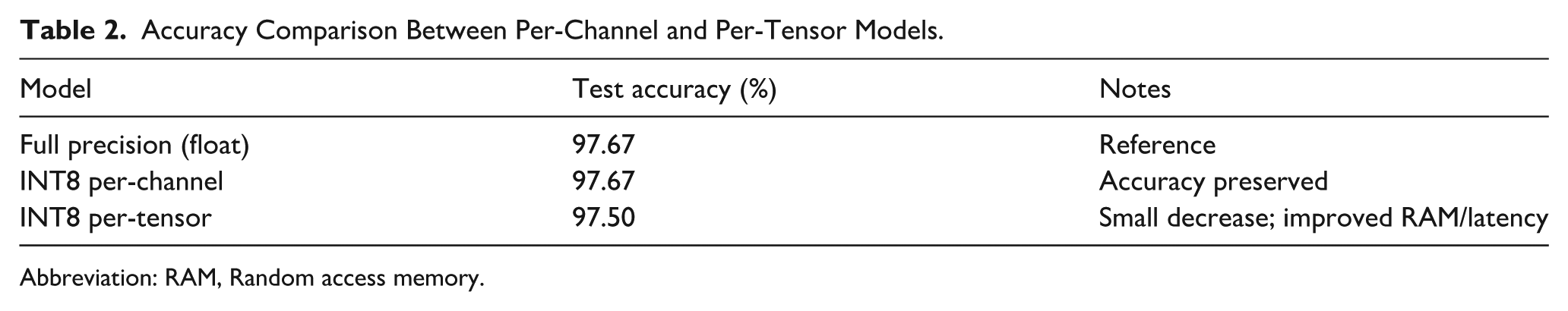

Abbreviation: RAM, Random access memory.
To provide a clinically relevant evaluation for this 3-class clinical classification task, we further computed per-class sensitivity, specificity, positive predictive value, and negative predictive value using a one-vs-rest formulation, with 95% confidence intervals estimated by the Wilson score method, as is shown in Table 3.
Per-Class Clinical Performance Metrics for the Per-Channel and Per-Tensor Quantized Models, Computed Using a One-vs-Rest Formulation.
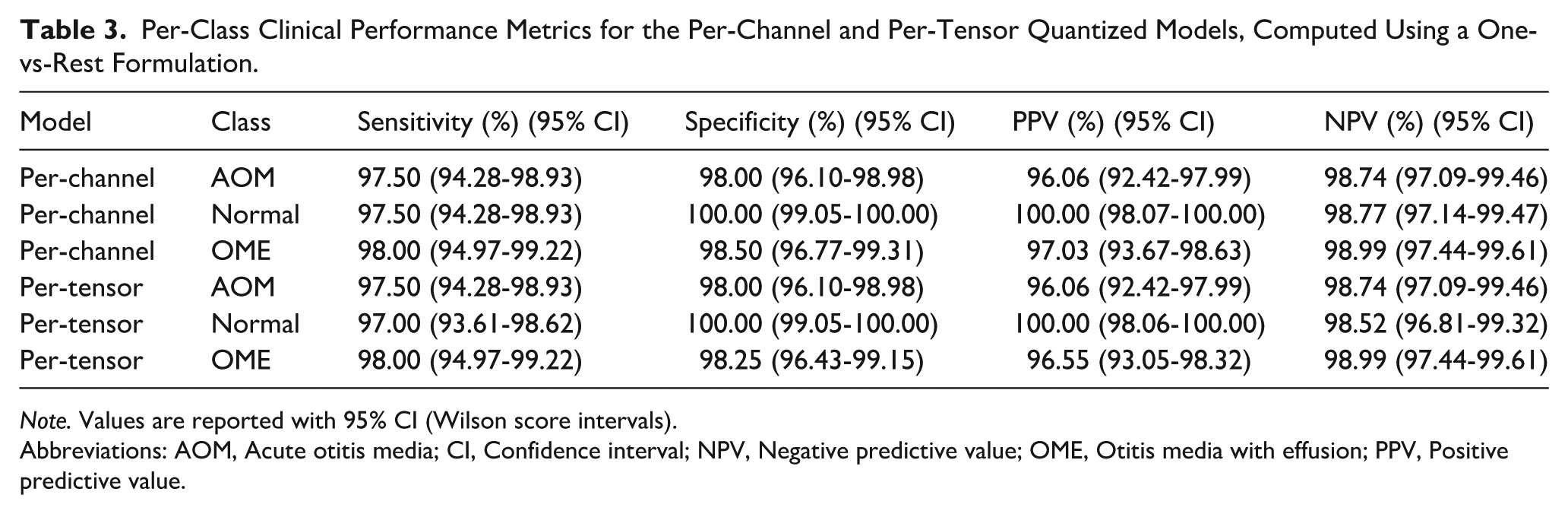
Note. Values are reported with 95% CI (Wilson score intervals).
Abbreviations: AOM, Acute otitis media; CI, Confidence interval; NPV, Negative predictive value; OME, Otitis media with effusion; PPV, Positive predictive value.
Discussion
We evaluated an embedded, on-device otoscopic image screening system for pediatric otitis media and quantified practical trade-offs between 2 INT8 quantization strategies.26,27 Per-channel quantization preserved accuracy relative to the float reference, whereas per-tensor quantization reduced activation memory and latency at the cost of a small decrease in accuracy. Although the per-channel model showed a numerically higher accuracy, the difference was small. Therefore, the practical choice between the 2 INT8 variants is better interpreted as a trade-off between fidelity to the reference model and deployment efficiency, with per-tensor quantization offering lower RAM usage and faster inference on the target hardware.
Prior work has reported high performance for deep learning-based analysis of otoscopic images, but most systems assume smartphone, cloud, or workstation inference.12 -19 Embedded inference offers a complementary path that can reduce reliance on connectivity and simplify privacy-preserving deployment. In our implementation, quantization reduced the model size substantially (approximately 163 MiB to 13.6 MiB. For clarity, the 163 MiB refers to the end-to-end deployment footprint measured in our toolchain, including network parameters, intermediate activation buffers, and runtime/graph metadata required by the embedded inference engine), enabling fully on-device inference under tight memory budgets.
This work is positioned as screening and prediagnostic triage support, not autonomous diagnosis. Otitis media diagnosis depends on integration of symptoms, history, and objective assessments when indicated (eg, tympanometry and audiology).2,3,8 A realistic role for embedded AI is to flag images that may warrant repeat examination, tympanometry, closer follow-up, or referral—especially in primary care and telemedicine contexts where specialist review is not always available.9 -11
Limitations include use of a single-center retrospective dataset and evaluation of a single model family. External validation across institutions and devices, including low-cost digital otoscopes, is needed. Future work will focus on generalizability, robustness under realistic degradations (blur, low illumination, specular reflection, cerumen occlusion), and prospective workflow evaluation to determine clinical impact. We did not perform a formal inferential statistical comparison between the 2 INT8 variants; future studies using larger and externally validated test sets should include such analyses in addition to descriptive deployment benchmarks.
Finally, we propose a possible clinical workflow for image-based screening and prediagnostic triage as shown in Figure 6. The system first performs an image quality check to determine whether the input is suitable for automated analysis. If acceptable, the AI model provides a screening/triage output to support recommended next steps (eg, repeat examination, follow-up, or referral). The clinician integrates symptoms/history and objective assessments to make the final diagnosis.

Proposed clinical workflow for image-based screening and prediagnostic triage.
Conclusions
A lightweight AlexNet-derived CNN can be quantized to INT8 and deployed on an STM32H7 microcontroller for on-device screening of pediatric otitis media. Per-channel INT8 preserved accuracy, whereas per-tensor INT8 improved latency and reduced activation memory with a minimal accuracy decrease. The system is intended to support screening and triage; final diagnosis and management remain clinician-led.
Supplemental Material
sj-docx-1-ear-10.1177_01455613261441511 – Supplemental material for Embedded AI-Assisted Otoscopic Image Screening for Pediatric Otitis Media
Supplemental material, sj-docx-1-ear-10.1177_01455613261441511 for Embedded AI-Assisted Otoscopic Image Screening for Pediatric Otitis Media by Changwei Lv, Desheng Jia, Zebin Wu, Bo Gao, Linzhong Xia and Xuansheng Wang in Ear, Nose & Throat Journal
Footnotes
Ethical Considerations
The study was conducted in accordance with the Declaration of Helsinki, and approved by the Institutional Review Board of Shenzhen Children’s Hospital for studies involving humans.
Consent to Participate
Patient consent was waived due to the retrospective nature of the study and the use of anonymized clinical data.
Author Contributions
Conceptualization, Changwei Lv. and Xuansheng Wang.; methodology, Changwei Lv, Desheng Jia and Zebin Wu.; software, Bo Gao and Changwei Lv.; investigation, Changwei Lv, Desheng Jia and Xuansheng Wang.; resources, Linzhong Xia and Xuansheng Wang.; data curation, Desheng Jia, Zebin Wu and Changwei Lv.; writing—original draft preparation, Changwei Lv.; writing—review and editing, Xuansheng Wang.; funding acquisition, Changwei Lv, Xuansheng Wang. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by National Natural Science Foundation of China, grant number 92467204.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The original otoendoscope tympanic membrane images analyzed in this study are not publicly available due to ethical approval constraints and patient privacy protections. Access to de-identified data may be considered upon reasonable request to the corresponding author, subject to institutional approval and execution of an appropriate data use agreement. To facilitate reproducibility without disclosing protected clinical images, the authors will provide trained model weights, inference code, scripts used for robustness/degradation experiments, and anonymized aggregate statistics (eg, per-class performance metrics and confusion matrixes) via a controlled repository or upon request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
