Abstract
Background:
Chronic rhinosinusitis (CRS) is a prevalent condition frequently evaluated using computed tomography (CT). The application of artificial intelligence (AI) in CRS imaging analysis is expanding; however, comprehensive assessments of its diagnostic performance remain scarce.
Objective:
To systematically review and compare the diagnostic accuracy of AI models for interpreting CT images of CRS, focusing on sensitivity, specificity, accuracy, and area under the curve (AUC).
Methods:
This systematic review followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. Six databases were searched for original studies that assessed AI, machine learning, or deep learning models for CRS diagnosis using CT. Eligible studies reported diagnostic performance metrics for human subjects. The extracted data included the AI model type, validation method, imaging modality, reference standard, and diagnostic outcomes.
Results:
Of the 1502 screened articles, 6 studies involving 2178 patients met the inclusion criteria. Most utilized convolutional neural networks, residual networks (ResNets), or hybrid deep learning models. The sensitivity, specificity, and accuracy ranged from 11.1% to 98.1%, 86.4% to 98.7%, and 63.6% to 98.4%, respectively. AUC values could reach 0.99.
Conclusion:
AI, particularly ResNets-based models, demonstrates promising diagnostic accuracy for CRS CT interpretation. However, methodological heterogeneity limits comparability, underscoring the need for standardized, multicenter validation and integration of clinical data to enhance generalizability.
Plain Language Summary
Chronic rhinosinusitis (CRS) is a long-lasting inflammation of the sinuses that causes nasal congestion, facial pressure, and loss of smell. It is often evaluated using computed tomography (CT), which helps doctors confirm the diagnosis and plan treatment. Artificial intelligence (AI) has recently been applied to CT imaging to improve diagnostic accuracy and reduce interpretation time.
This systematic review analyzed all available studies that assessed the accuracy of AI in diagnosing CRS using CT. Six studies, including 2178 patients, were included. Most deep learning models, particularly convolutional neural networks (CNNs) residual networks (ResNets), were used. The reported diagnostic performance was high with an accuracy of 98% and an area under the curve (AUC) of 0.99.
These results show that AI can reliably assist in the detection of sinus diseases in CT images. However, most studies have been limited to single centers, highlighting the need for larger, standardized, multicenter studies before clinical implementation.
Introduction
Chronic rhinosinusitis (CRS) is a common inflammatory disease of the nasal and paranasal sinuses, characterized by symptoms such as nasal obstruction, facial pain, or pressure, and a reduced sense of smell lasting for at least 12 weeks. CRS is estimated to affect approximately 8% to 11% of adults worldwide, although prevalence rates vary across regions.1 -3 In addition to its high prevalence, CRS poses a substantial clinical and socioeconomic burden, significantly impairing patients’ quality of life and contributing to considerable healthcare expenditures.3,4
Accurate diagnosis and effective management of CRS typically require a combination of clinical evaluation and imaging, particularly computed tomography (CT) of the paranasal sinuses. CT imaging provides essential anatomical details, facilitates surgical planning, and is critical for evaluating disease severity using standardized systems such as the Lund–Mackay score (LMS). 5 However, the interpretation of CT scans may vary among clinicians and can be limited by differences in expertise, time constraints, or access to subspecialty consultation.
In recent years, the integration of artificial intelligence (AI), particularly deep learning algorithms, such as convolutional neural networks (CNNs), has shown promise in improving the diagnostic accuracy of medical imaging across various specialties.6,7 In rhinology, emerging AI applications have primarily focused on automating CT analysis to detect sinus opacification, segment anatomical structures, and predict disease classification. Several studies have reported encouraging performance metrics, including high sensitivity and specificity.8,9
Despite their growing popularity, there is currently no comprehensive evidence to evaluate the diagnostic performance of AI models for CT imaging of CRS. This represents a critical gap, particularly as AI tools are increasingly being proposed for use in clinical decision support.
This study aimed to systematically evaluate and compare the performance of AI models used to diagnose CRS through CT interpretation, with particular attention paid to diagnostic accuracy, sensitivity, specificity, and area under the curve (AUC). By identifying the most effective approaches and highlighting areas for improvement, this review seeks to inform the future development and clinical implementation of AI in rhinology practice.
Materials and Methods
A systematic review of the published English literature was conducted to evaluate the application of AI models in CRS. This review adhered to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines and followed the recommendations outlined in the PRISMA checklist throughout study selection, data extraction, and reporting. 10 The following databases were systematically searched: PubMed (MEDLINE), OpenAIRE, ScienceDirect, Web of Science, Google Scholar, and Springer Nature Journals. A comprehensive electronic search strategy was developed for each database, incorporating both keywords and Medical Subject Heading terms. The search terms included combinations of “artificial intelligence,” “deep learning,” “machine learning,” “computed tomography,” “sinus,” “chronic rhinosinusitis,” and “diagnosis.”
Eligibility Criteria
The articles included in this systematic review were restricted to English-language publications that presented original data. The eligible study designs included prospective and retrospective cohort, cross-sectional, and diagnostic accuracy studies. Randomized controlled trials, although uncommon in this field, were also considered. Two independent reviewers screened all articles for inclusion based on the predefined eligibility criteria. Studies were included if they (1) applied AI, deep learning, or machine learning models; (2) incorporated sinus CT imaging within the diagnostic workflow for CRS; (3) involved human participants; and (4) reported performance metrics such as accuracy, sensitivity, specificity, and AUC. Studies were excluded if they lacked original data; focused on basic science, animal models, or cadaveric studies; used non-CT imaging modalities; or failed to provide sufficient detail on model validation or reference standards. Review articles, expert opinions, abstracts without full text, and studies with unclear methodologies were excluded. Final inclusion criteria were determined following a full-text review after the initial title and abstract screening. Several prior AI-based imaging studies were excluded from the final analysis because they did not report diagnostic accuracy metrics, relied on non-CT modalities, or lacked transparent validation strategies; however, they were referenced for contextual background.
Data Extraction
Data extraction was independently performed by 2 investigators using a standardized data collection form. Any discrepancies related to study inclusion or extracted variables were resolved through discussion and consensus. The extracted data included the publication year, study design, country, sample size, patient demographics, AI model type, imaging modality, and reference standards. Additional methodological details were systematically extracted to improve transparency and comparability across the studies. These included CT data sources (single-center versus multicenter datasets) and model validation strategies. The validation approaches were explicitly categorized as internal (holdout validation or cross-validation within the same dataset) or external (testing on independent datasets). The extracted performance metrics included the sensitivity, specificity, accuracy, and AUC. Variability and incomplete reporting of methodological details across studies were noted and considered during the interpretation of the results.
The methodological quality and risk of bias of the included studies were evaluated using the Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) tool, which assesses bias across 4 domains: patient selection, index test, reference standard, and flow and timing. Given the inclusion of AI models, AI-specific considerations were further evaluated according to the QUADAS-AI extension. 11
Results
Among the 1502 unique research papers initially identified, 63 were selected for primary screening. After a full-text evaluation, 6 studies comprising 2178 patients were included in the final review (Figure 1). All of the included studies provided level III evidence and were published within the past 2 years. Five studies were conducted in China, and one was conducted in the United States. All included studies relied on internal validation approaches, using either holdout validation or cross-validation, and none performed external validation using independent cohort (Table 1).
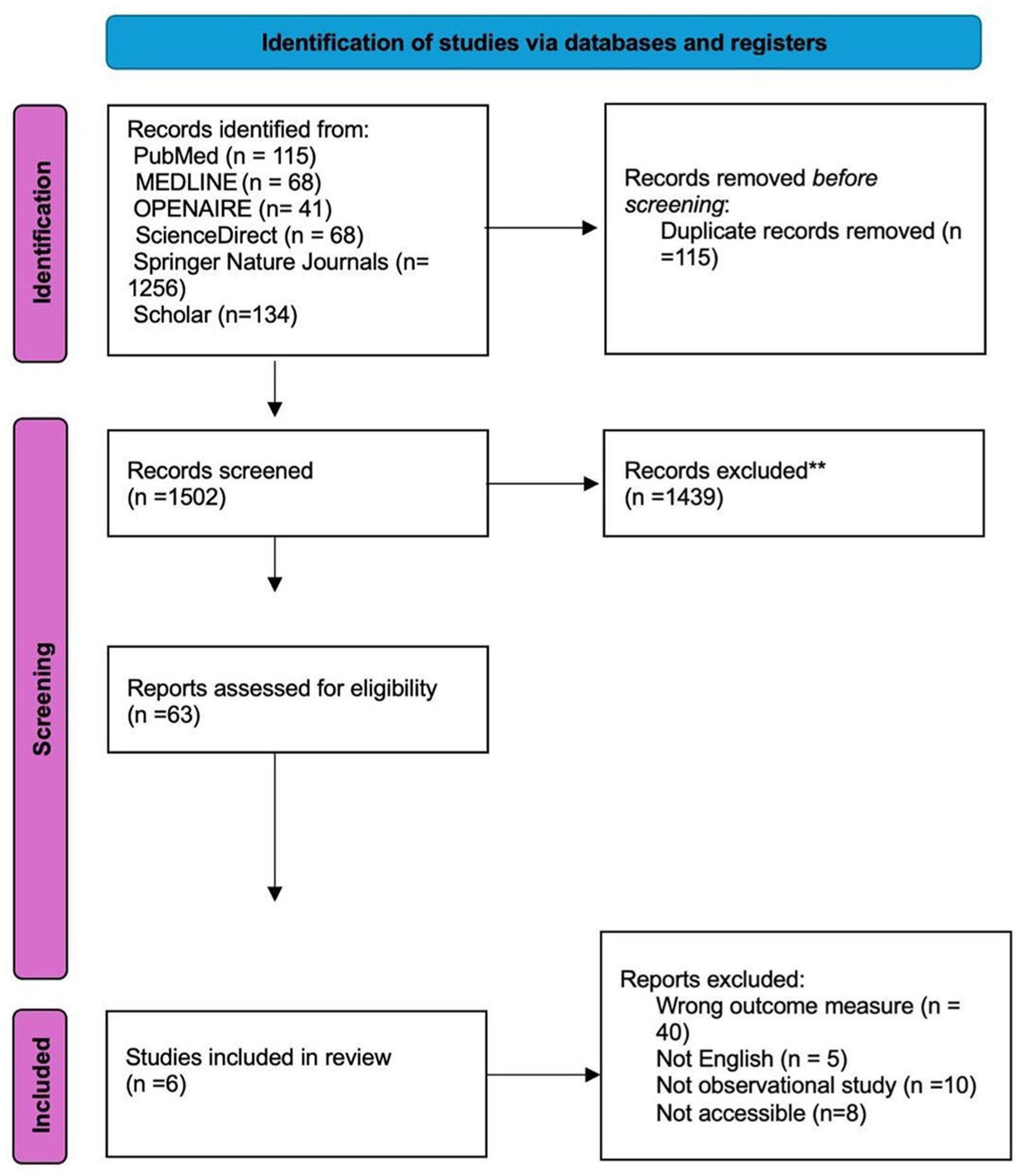
PRISMA flow diagram illustrating the screening process of articles according to the study’s inclusion and exclusion criteria. PRISMA, Preferred Reporting Items for Systematic Reviews and Meta-Analyses.
Characteristics of the Included Studies.
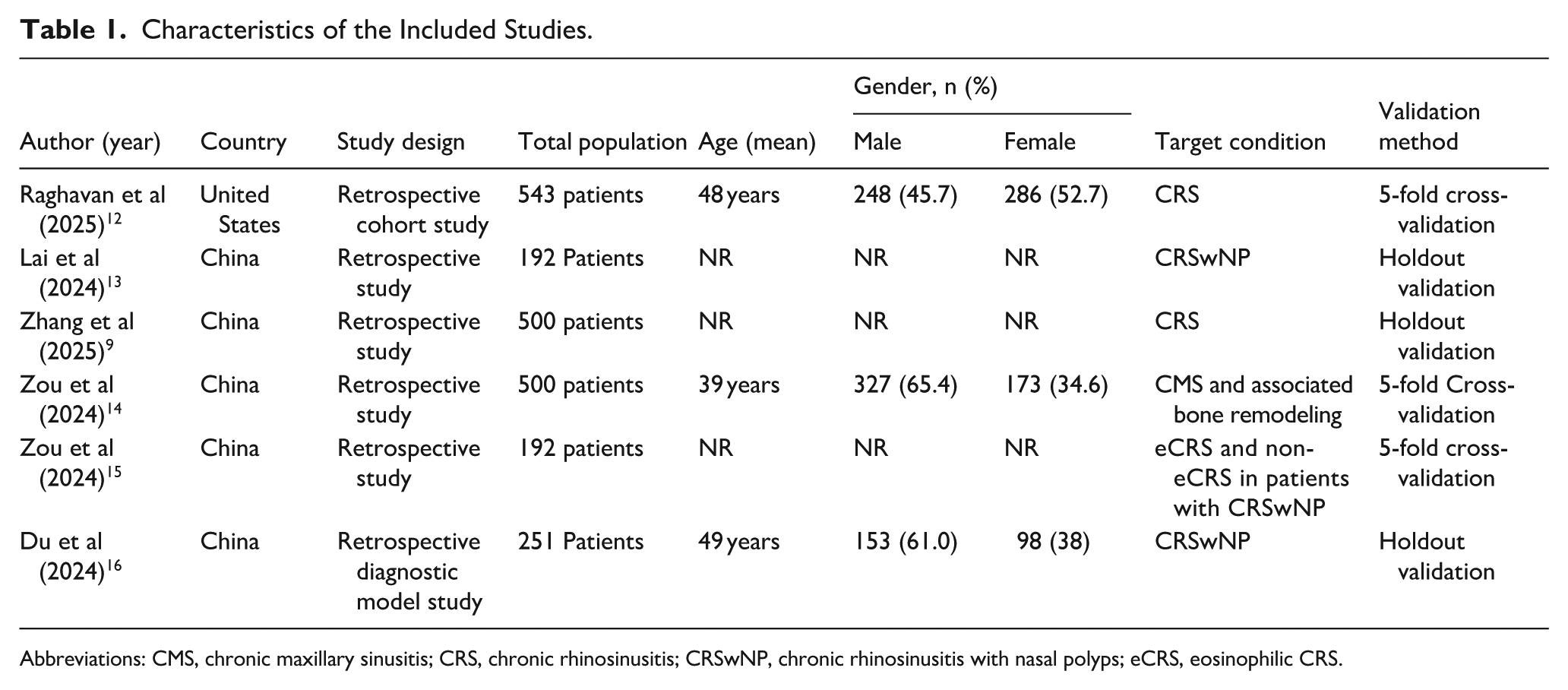
Abbreviations: CMS, chronic maxillary sinusitis; CRS, chronic rhinosinusitis; CRSwNP, chronic rhinosinusitis with nasal polyps; eCRS, eosinophilic CRS.
A range of AI models were used across the included studies. CNNs, residual networks (ResNets), and traditional machine learning models are the most frequently adopted. Deep learning approaches include standard deep neural networks (DNNs) and hybrid models, such as 1-dimensional CNNs combined with long short-term memory networks. Regarding data inputs, all studies primarily relied on CT interpretation, while 1 study additionally incorporated ≥2 cardinal symptoms of CRS. Different reference standards were used to validate the performance of the AI model, primarily based on clinical, radiological, or histopathological criteria. Raghavan et al defined the primary reference standard as radiological evidence of sinonasal inflammation with a LMS ≥5, while the secondary endpoint combined LMS ≥5 with at least 2 cardinal symptoms of CRS. 12 Lai et al applied a clinical diagnosis of CRS based on established guidelines, supplemented by CT-based Lund–Mackay scoring. 13 Zhang et al relied on expert consensus, with diagnoses confirmed by 2 rhinologists using endoscopy and CT imaging. 9 Du et al employed histopathological examination of nasal polyps as the diagnostic reference. 16 These diverse reference standards reflect heterogeneity in clinical practice and emphasize the need for unified criteria when benchmarking AI performance in sinonasal imaging (Table 2).
Diagnostic Performance of Artificial Intelligence (Uploaded Separately).
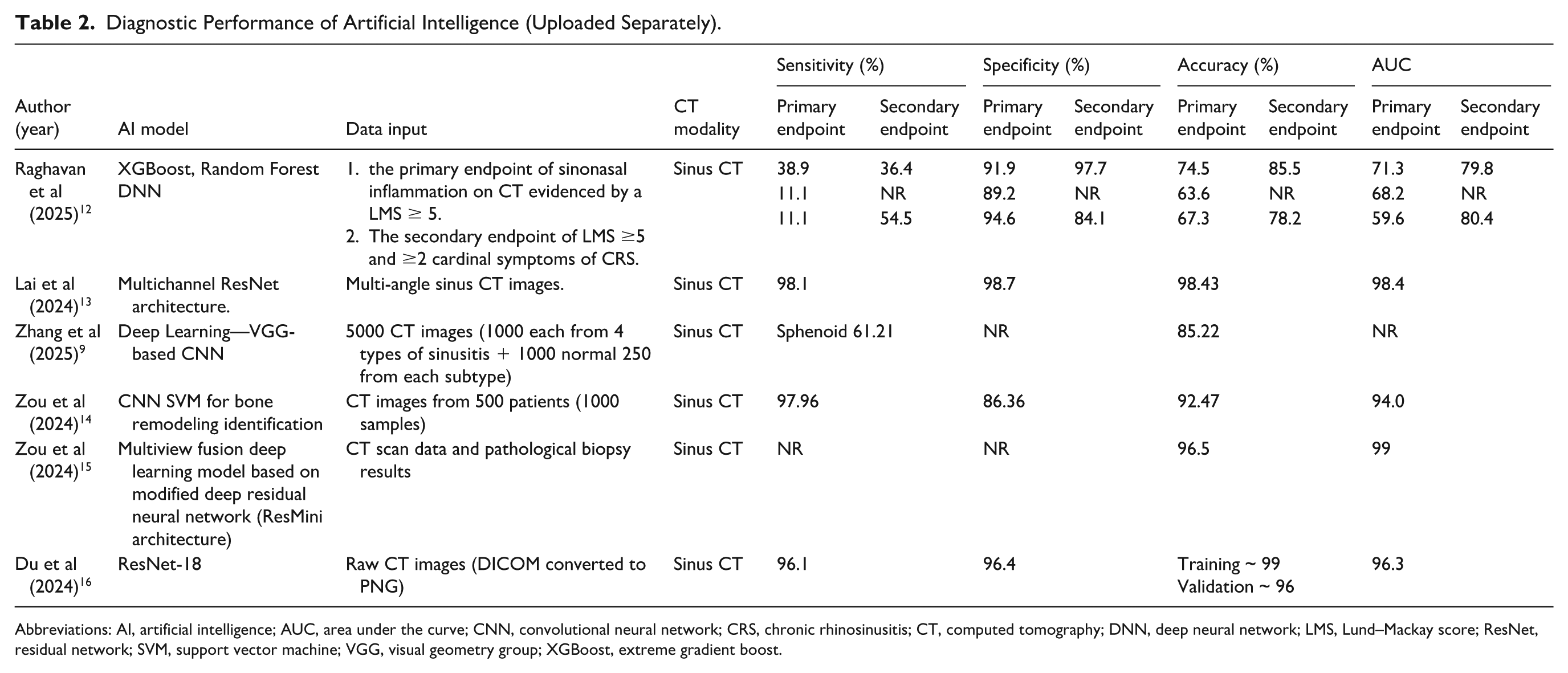
Abbreviations: AI, artificial intelligence; AUC, area under the curve; CNN, convolutional neural network; CRS, chronic rhinosinusitis; CT, computed tomography; DNN, deep neural network; LMS, Lund–Mackay score; ResNet, residual network; SVM, support vector machine; VGG, visual geometry group; XGBoost, extreme gradient boost.
The key metrics used to evaluate the performance of AI models included sensitivity, specificity, accuracy, and AUC. The sensitivity values varied widely among the included studies, ranging from 11.1% to 98.1%. Raghavan et al reported low sensitivity for the primary outcome when using CT alone for diagnosis in both the random forest model and DNN. 12 However, when CT data were combined with the cardinal symptoms of CRS, the sensitivity of the DNN increased by 43.4%. Contrastingly, Lai et al and Zou et al demonstrated substantially higher sensitivities of 98.1% and 97.96%, respectively.13,14 Du et al reported a sensitivity of 96.1%, whereas 1 study did not provide sensitivity values. 16
The specificity was consistently high across most studies, ranging from 86.36% to 98.7%. Lai et al reported the highest specificity at 98.7%, while Zhang et al described specificity as “very high” without specifying the exact value.9,13 Two studies, by Zou et al and Du et al, did not explicitly report specificity values.9,15
The reported accuracy ranged from 63.6% to 98.43%, reflecting variations in model architecture and diagnostic objectives. The lowest accuracy was observed in the study by Raghavan et al (63.6% for the primary endpoint), whereas Lai et al reported the highest accuracy of 98.43%.12,13 Other studies, including those by Zou et al and Du et al, achieved accuracy levels exceeding 85%. Du et al reported training and validation accuracies of approximately 99% and 96%, respectively.14 -16
Overall, the AUC values demonstrated a strong discriminative performance. Zou et al reported the highest AUC of 0.99, whereas Raghavan et al documented the lowest AUC of 0.596 for the primary endpoints (Table 2).12,15
These findings indicate substantial variations in diagnostic performance among AI models applied to sinus imaging, reflecting differences in study design, endpoint definitions, and algorithmic complexity. Although many models, particularly those using deep learning techniques, exhibit high sensitivity, specificity, and AUC values, incomplete or inconsistent reporting across several studies limits the strength of cross-study comparisons.
Importantly, variations in diagnostic performance across studies were closely linked to differences in the model architecture, validation strategies, reference standards, and clinical task definitions. Studies reporting lower sensitivity primarily relied on CT-only inputs for broad CRS diagnosis, whereas models achieving higher AUCs typically address narrower tasks, such as endotype differentiation or bone remodeling assessment, using multiview or attention-enhanced deep learning architectures. These distinctions highlight that the reported performance metrics should be interpreted in the context of the intended diagnostic objective, rather than as a universal measure of model superiority.
Quality Assessment
Most studies demonstrated a low risk of bias across key domains, particularly in patient selection, reference standards, and flow and timing. Raghavan et al, Lai et al, and Du et al showed a consistently low risk across all domains, indicating robust methodological quality.12,13,16 Contrastingly, Zhang et al and Zou et al presented a higher risk in the patient selection and index test domains, with some unclear reporting in flow and timing.9,14,15 Applicability concerns were generally low to moderate across all studies, suggesting reasonable relevance to clinical settings, despite some methodological limitations (Table 3).
QUADAS-AI Risk of Bias and Applicability Assessment for Included Studies.
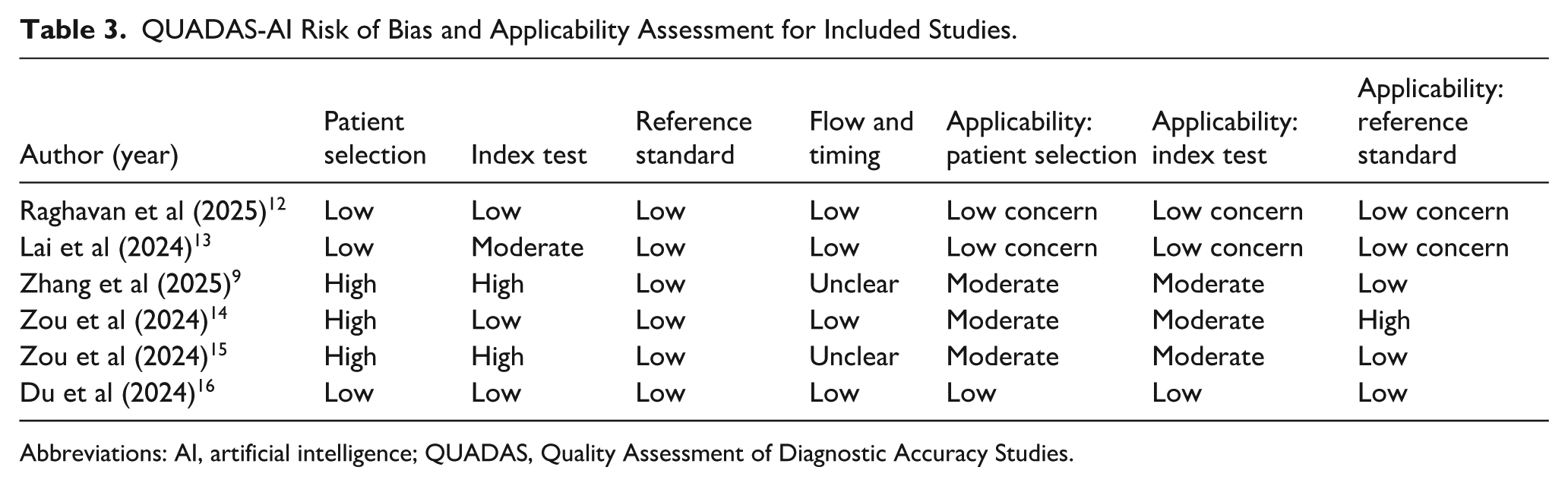
Abbreviations: AI, artificial intelligence; QUADAS, Quality Assessment of Diagnostic Accuracy Studies.
Discussion
AI is rapidly transforming the field of medical diagnostics, offering unprecedented accuracy and efficiency in pattern recognition and clinical decision-making. In rhinology, particularly in the context of CRS, AI has demonstrated promising capabilities in interpreting imaging data, classifying disease subtypes, and potentially guiding patient management. The present review synthesizes the current evidence on the diagnostic performance of AI models applied to sinus CT, emphasizing their potential and limitations in clinical application.
The findings from the 6 included studies indicated that deep learning architectures, particularly CNNs and ResNets, achieved high sensitivity, specificity, and diagnostic accuracy for detecting CRS and differentiating its endotypes. Lai et al achieved a sensitivity of 98.1% and specificity of 98.7% using a multichannel ResNets model trained on multiangle CT views for predicting eosinophilic chronic rhinosinusitis with nasal polyps (CRSwNP), 13 highlighting the importance of architectural depth and multiview input in enhancing diagnostic performance. Similarly, Du et al employed a multiview lightweight attention-enhanced network and reported near-perfect accuracy with an AUC of 0.993, underscoring the scalability of efficient architectures in real-world datasets. 16
Sensitivity values varied considerably among the included studies. While some models demonstrated suboptimal sensitivity, others, particularly those developed by Lai et al and Zou et al, achieved sensitivities of 98.1% and 97.96%, respectively.13,14 This variation suggests that although AI algorithms perform well in detecting pronounced disease patterns, they may be less reliable in identifying subtle or early-stage CRS manifestations such as mild mucosal thickening or partial sinus opacification.
However, the specificity was consistently high across most studies, ranging from 86.36% to 98.7%, indicating that the models were highly effective at correctly identifying patients without CRS. In other words, these AI systems rarely produce false positive results; when a model classifies a CT scan as normal, it is typically accurate in doing so.
These results align with earlier works by Hashimoto et al and He et al, who emphasized that AI models trained on large, well-annotated imaging datasets can outperform human experts in specific radiologic and pathologic recognition tasks.17,18 Furthermore, radiomics-based studies, such as those by He et al, have demonstrated excellent discriminative ability in differentiating sinonasal tumors from inflammatory polyps, suggesting that texture-based features, often imperceptible to the human eye, can significantly enhance diagnostic accuracy. 19
Models employing deeper architectures, such as ResNet–based architectures, particularly multiview and attention-enhanced ResNets models, generally demonstrate superior diagnostic performance than conventional CNNs and traditional machine learning approaches. This performance advantage appears to be driven by enhanced feature extraction, deeper network capacity, and enhanced spatial contextualization across multiple sinonasal regions, which are critical for capturing the heterogeneous and often subtle inflammatory patterns of CRS on CT imaging.
More importantly, the reliability of the reported performance metrics must be interpreted by considering the validation strategies employed. All included studies relied exclusively on internal validation methods, such as holdout validation or cross-validation, within single-center datasets, and none performed true external validation using independent multi-institutional cohorts. While internal validation is appropriate for model development and preliminary performance assessment, the absence of external validation limits evaluation of generalizability and may lead to optimistic performance estimates in real-world clinical settings.
From a clinical perspective, these findings indicate that although AI demonstrates promising diagnostic accuracy, its current role should be regarded as supportive and adjunctive rather than autonomous, particularly for specific diagnostic tasks such as endotype differentiation, rather than for standalone confirmation of CRS. This technology may assist clinicians in streamlining image interpretation and reducing diagnostic variability. However, final diagnostic and management decisions must remain under expert supervision to ensure patient safety and appropriate clinical judgment.
Despite these promising findings, several challenges remain. The studies included in this review exhibited considerable methodological heterogeneity, including variations in the data sources, CT acquisition protocols, AI model architectures, validation techniques, and reference standards. For instance, while Lai et al and Zhang et al used CT and clinical criteria as reference standards, Du et al and Zou et al relied on histopathological analysis.9,13 -15 Such inconsistencies complicate cross-study comparisons and underscore the urgent need for standardized AI validation frameworks in CRS diagnostics.
Furthermore, most of the included studies were retrospective and conducted at single centers, which limited the generalizability of the results (Table 4). Only 1 study was conducted in the United States, whereas the remaining were performed in China, raising concerns about potential geographic and population-specific biases. Prospective multicenter validation is essential before AI models can be widely implemented in clinical practice. Although diagnostic performance metrics, such as AUC and accuracy, have been consistently reported, the issue of interpretability remains a major limitation.
Studies’ Conclusions and Limitations.
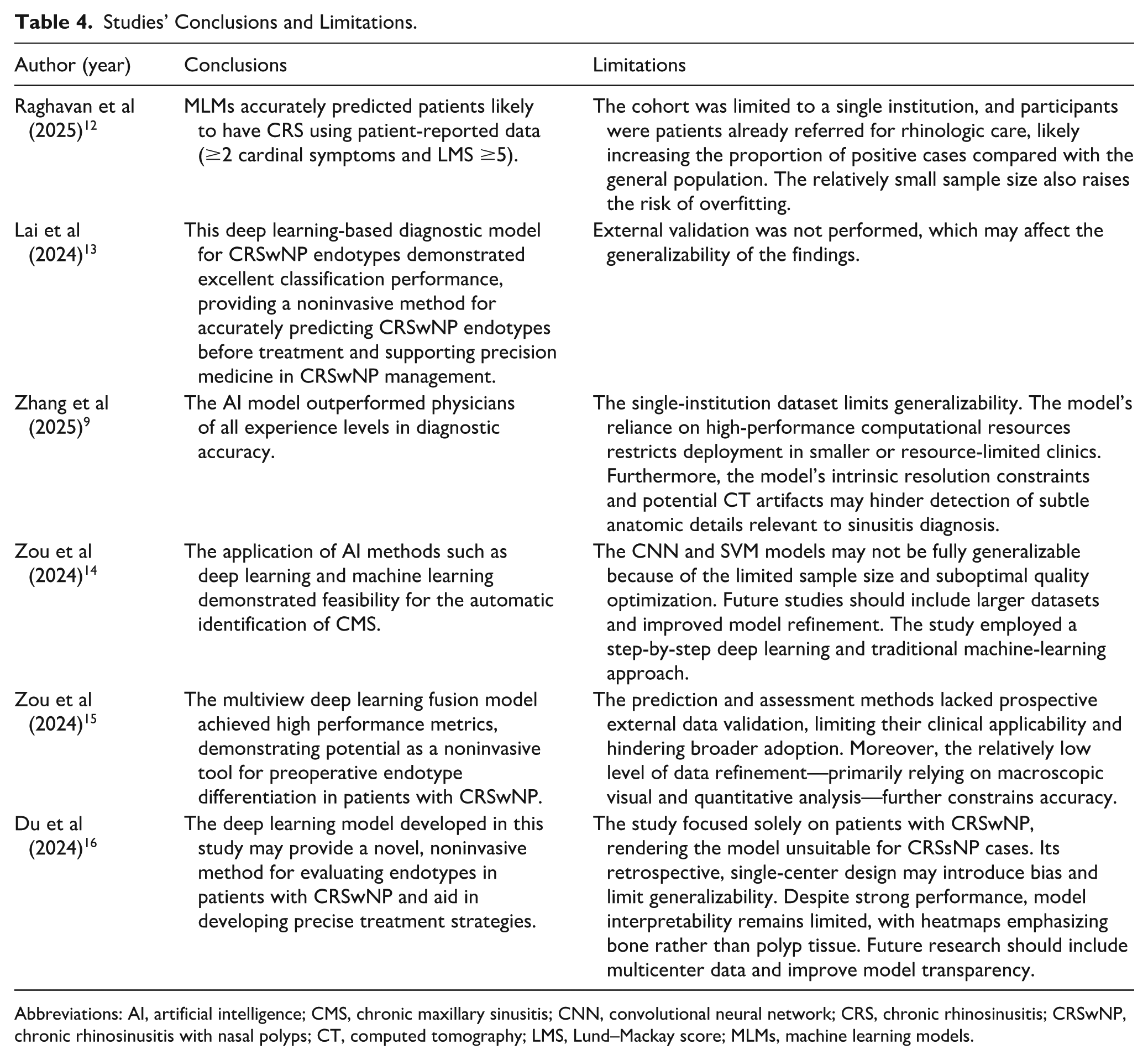
Abbreviations: AI, artificial intelligence; CMS, chronic maxillary sinusitis; CNN, convolutional neural network; CRS, chronic rhinosinusitis; CRSwNP, chronic rhinosinusitis with nasal polyps; CT, computed tomography; LMS, Lund–Mackay score; MLMs, machine learning models.
Future studies should focus on the following key topics. First, the development and validation of AI models should be undertaken through large multicenter studies to ensure generalizability across diverse clinical settings and patient populations. Second, the adoption of standardized methodologies and reporting frameworks is critical for facilitating cross-study comparisons and reproducibility. Finally, integrating CT imaging data with additional patient information, such as symptom profiles, laboratory findings, and clinical history, may further improve diagnostic accuracy and enhance model performance.
In summary, this review reinforces the growing body of evidence that AI, particularly deep learning models, can substantially improve the diagnostic accuracy of CRS when applied to CT imaging. However, to fully realize their clinical potential, concerted efforts are required to standardize methodologies, enhance interpretability, and validate models in diverse prospective clinical environments.
Conclusion
This systematic review demonstrates that AI, particularly deep learning-based models, shows considerable promise for CT-based diagnosis of CRS. Most models achieved high diagnostic performance, with sensitivity, specificity, and accuracy metrics comparable to those of expert interpretations. Nonetheless, the variability in datasets, reference standards, and validation strategies continues to limit the current clinical applicability of these models.
Footnotes
Author Contributions
N.F.A. contributed to the work as the first author; conceptualized the idea; designed the study protocol; assembled the research team; created the tables; and reviewed, edited, and proofread the final manuscript. N.F.A. and T.A. performed the literature review, extracted the data, finalized the tables, drafted the manuscript, and edited and finalized the manuscript. N.A. resolved conflicts and reviewed and proofread the final manuscript. A.A. reviewed and proofread the manuscript. All the authors have read and approved the final version of the manuscript. R.A. supervised, reviewed, and proofread the final manuscript. All the authors have read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data generated or analyzed during this study are included in this published article.
