Abstract
Olfactory neuroblastoma (ON; Esthesioneuroblastoma) is a malignant tumor that arises from the olfactory neuroepithelium. Very rarely, ON can histologically display a biphenotypic pattern, with only 7 cases reported in the literature to date. We describe a case of this poorly understood entity and review the patient’s histology, pathology, and treatment. An 85-year-old man presented with endoscopy and imaging findings of a sinonasal mass arising from the olfactory cleft. Biopsy and further pathological review established Hyams grade 3 and Kadish stage C tumor. The patient underwent combined endoscopic and open-approach resection due to the involvement of the nasal bone and subsequent adjuvant radiation therapy. Specimen contained Flexner-Wintersteiner rosettes and stained positive for the neuroendocrine marker synaptophysin consistent with ON. A second, epithelial component pattern of goblet cells and mucin was observed suggesting divergent differentiation. We are the first group to report next-generation sequencing of this tumor, which revealed a pathogenic mutation in PIK3CA and a likely pathogenic variant in RUNX1 (AML1). ON with divergent differentiation is very rare, and more robust studies characterizing molecular drivers and pathology may aid in clinical management.
Keywords
Introduction
Esthesioneuroblastoma, also known as olfactory neuroblastoma (ON) accounts for approximately 3% to 5% of nasal cavity cancers. 1 These malignant tumors arise from the olfactory neuroepithelium and typically occur in a bimodal age distribution in the second or fourth to sixth decade of life in patients. 2 Patients often present with epistaxis, nasal obstruction, or with symptoms of tumor spread to the sinuses, brain, and orbit. The 5-year overall survival rate is up to 80% with multimodal therapy, including surgical resection and adjuvant radiation, with or without chemotherapy. 3 Histologically, ON is evaluated by the Hyams grading system. A score of 1 to 4 is given based on adverse features such as rosette formation, mitotic activity, pleomorphism, necrosis, and disorganized architecture. 4 Clinical staging of the tumor is conducted with the Kadish system that classifies lesions into low grade (A, B) or high grade (C, D), with higher grade correlating with greater tumor extension into surrounding structures. 5
An exceedingly rare subset of these tumors can display a biphenotypic pattern on histology. They are often described as having divergent or mixed differentiation. Existing literature on this topic includes case presentations and histological findings, with only 7 cases published in the literature to date (Table 1).6-12 However, there is a lack of consensus on how to classify this tumor type due to the limited number of reported cases. Important clinical variables for this patient group including prognosis have yet to be elucidated. Here, we present a rare case of ON with divergent differentiation, review its histopathology, and discuss the role of next-generation sequencing (NGS) in guiding treatment decisions and future research.
Summary of Literature for Olfactory Neuroblastoma With Divergent Differentiation.
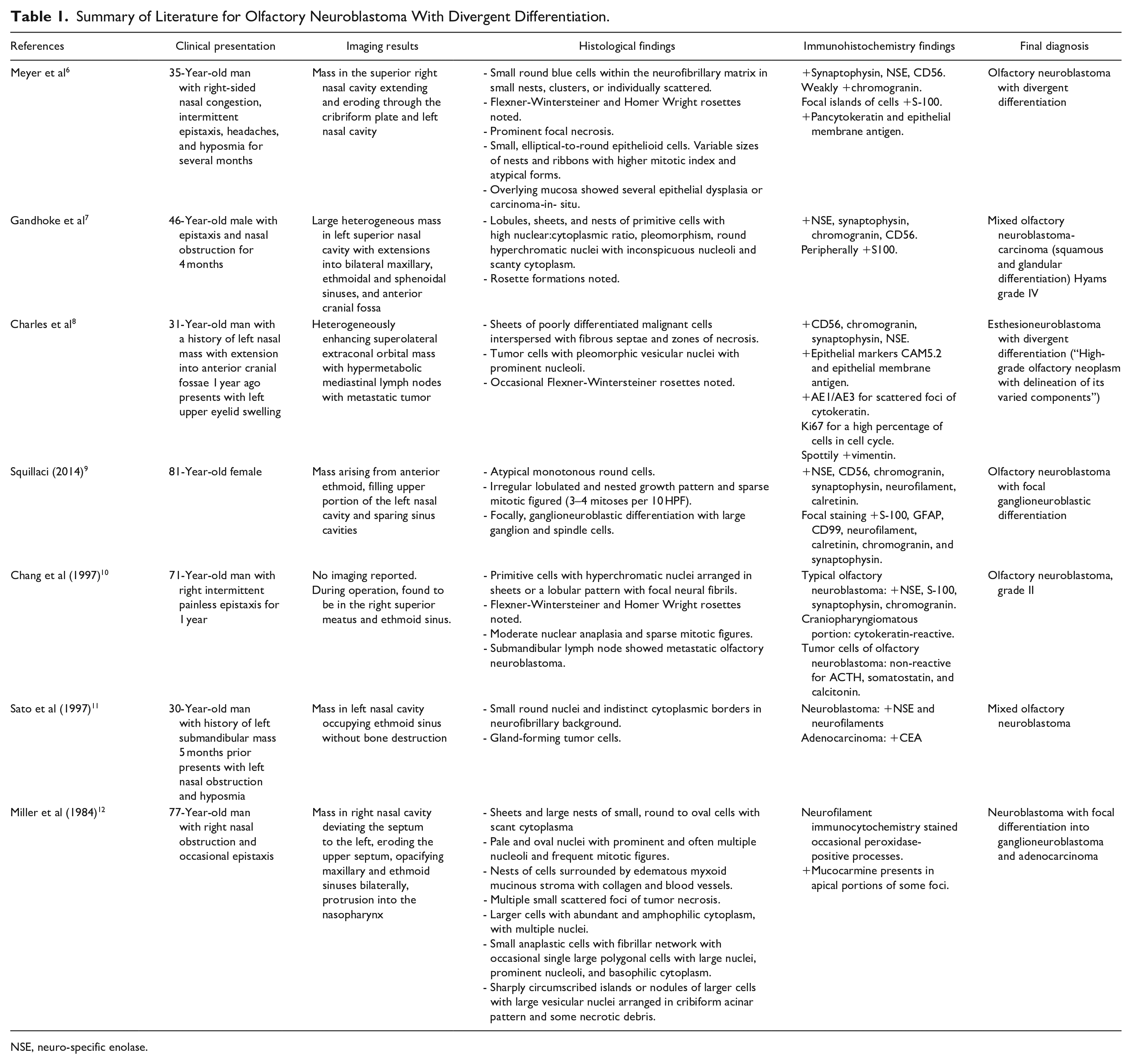
NSE, neuro-specific enolase.
Case Presentation
An 85-year-old man with an unremarkable medical history presented to the otolaryngology clinic with increasing epistaxis, which upon workup revealed a right-sided sinonasal mass. Non-contrast sinus computed tomography and contrasted magnetic resonance imaging demonstrated a right nasal mass, arising from the olfactory cleft and anterior skull base involving the nasal bones with no significant intracranial or orbital extension (Figure 1A-D). An initial endoscopic biopsy of the mass was reported as an infiltrative malignant tumor with neuroendocrine differentiation. The patient underwent resection of the tumor via an extended endoscopic approach including an open lateral rhinotomy to remove the involved nasal bone and undergo reconstruction (Figure 2A). Reconstruction of the nasal bone was accomplished with a titanium plate (Figure 2B) and a pedicled nasoseptal flap (not shown).
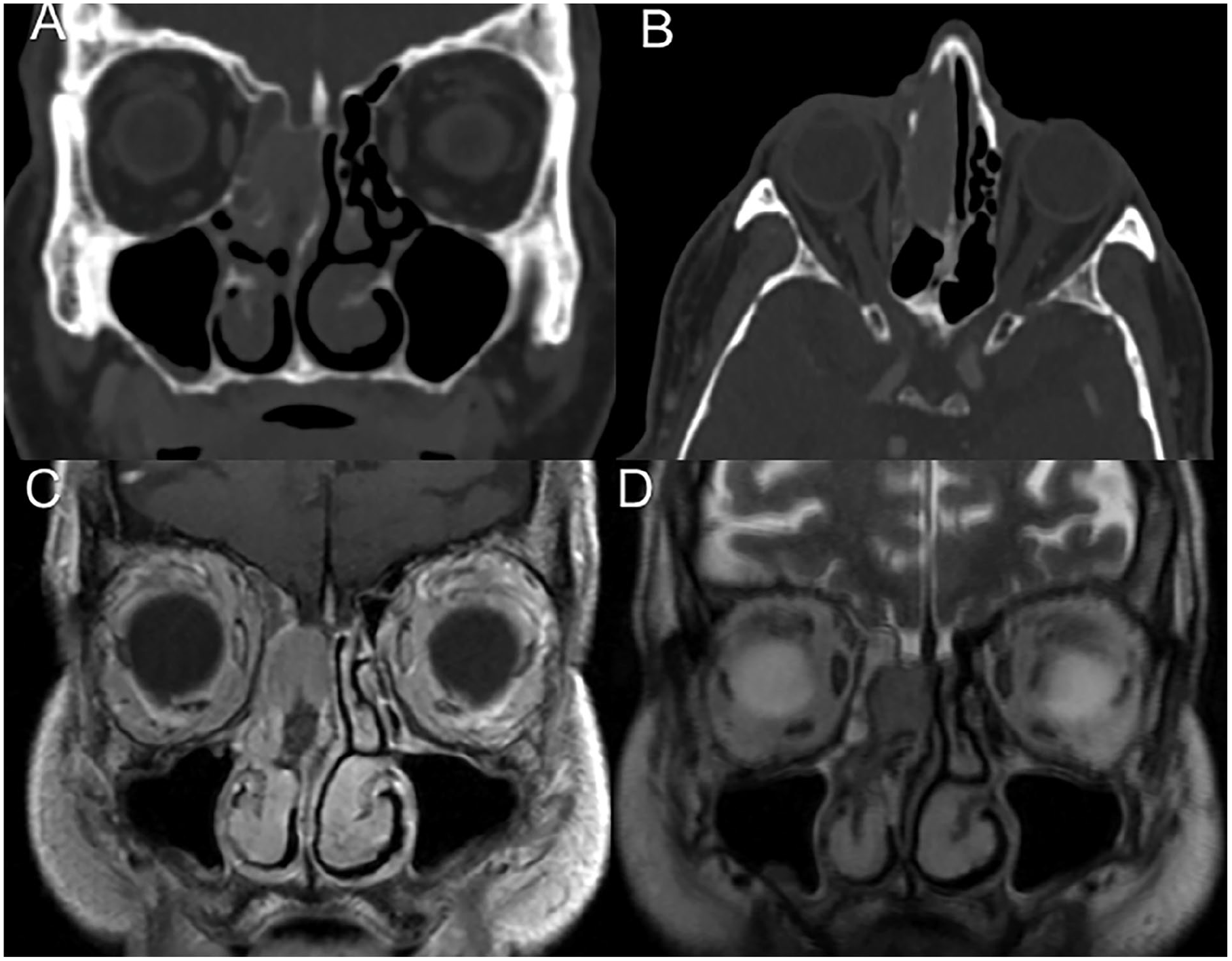
CT scan (A, B) showing right sinonasal mass with erosion of nasal bone and septum. MRI with contrast (C) and T2 image (D) showing a mass arising from the olfactory cleft and anterior skull base, but no significant orbital or intracranial extension.
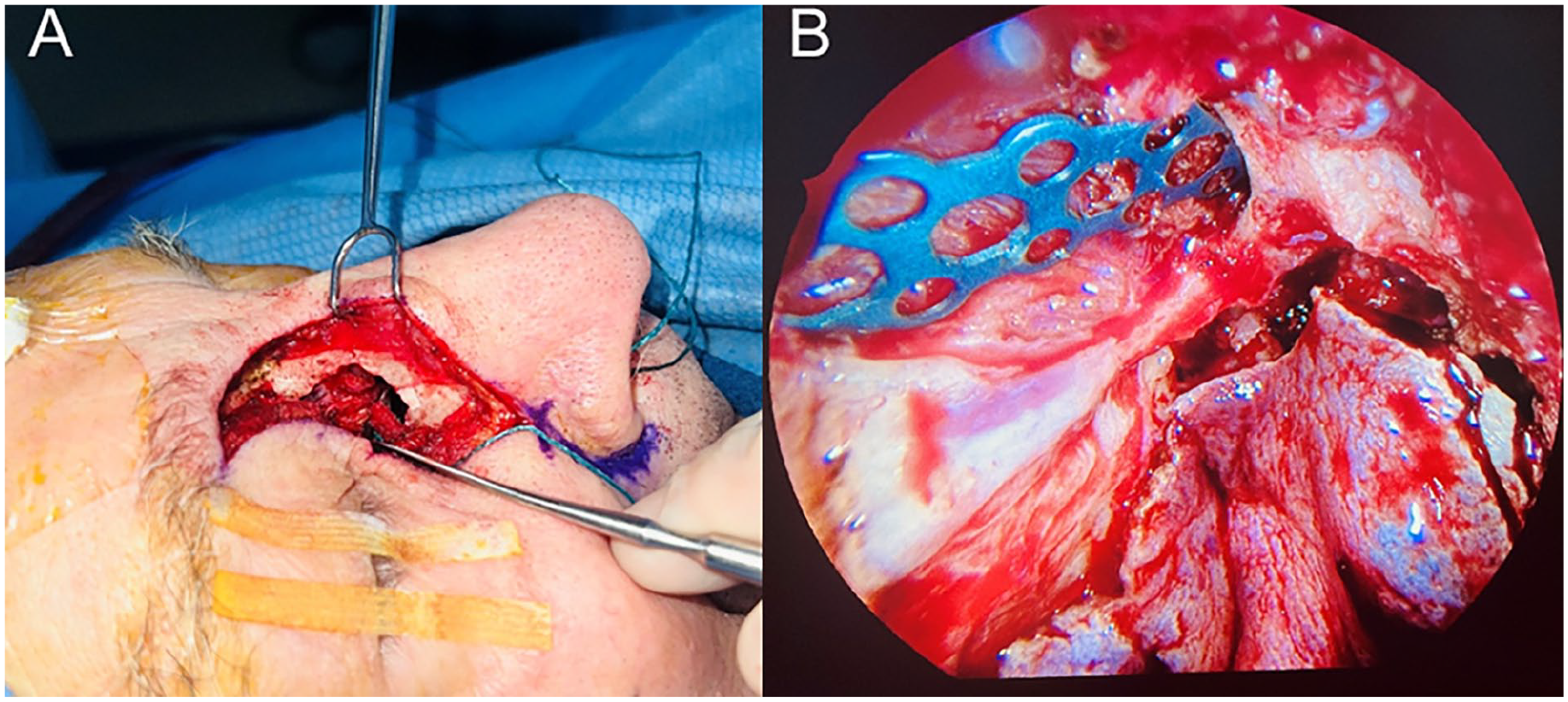
(A) The lateral rhinotomy approach to resect the tumor involving the nasal bone, and (B) endoscopic titanium plate reconstruction for the nasal bone after tumor resection.
Histologic and Molecular Profile
Pathology assessment revealed a biphenotypic tumor. The majority of the tumor demonstrated histologic findings consistent with ON, including small round or oval blue cells arranged in nests and ribbons with Flexner-Wintersteiner rosettes (Figure 3A). The tumor cells in this region had a high nuclear:cytoplasmic ratio, with coarsely stippled chromatin, abundant apoptoses, and mitoses. Immunohistochemical staining in this region showed positivity for chromogranin, synaptophysin, and CD56. The minority region was composed of moderately to well-differentiated epithelial cells with mucocytes and gland formation, nonreactive to neuroendocrine markers, indicating divergent differentiation (Figure 3B-D). Immunohistochemistry did not show loss of INI1 expression and in situ hybridization for Epstein Barr virus was negative. Patchy reactivity was seen for p53, and background structures were stained for CK5/6, p63, and p40 as appropriate. A Hyams grade of 3 was determined from histological findings, and Kadish stage of C was chosen because of tumor extension into the paranasal sinuses.
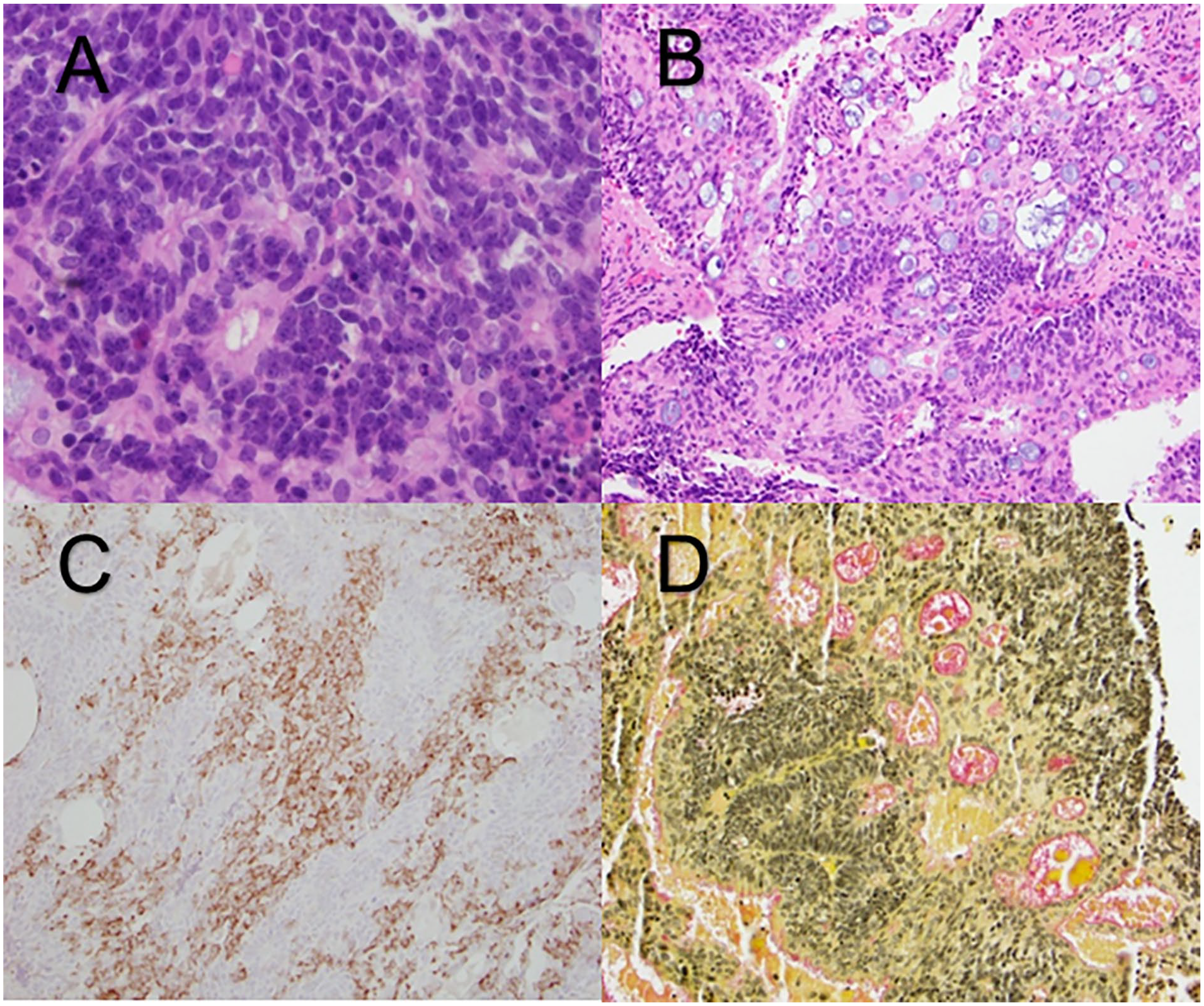
(A) Small blue cells with abundant apoptosis and Flexner-Wintersteiner rosettes. (B) Gland-like formations with goblet cells and mucin. (C) Synaptophysin highlighting a subset of cells. (D) Mucicarmine stain highlighting mucin pink.
Tumor tissue was sent for NGS at the University of Florida Health Pathology Laboratories. Genomic DNA extracted from the tissue was amplified using the GatorSeq NGS Panel and sequenced on the Illumina NextSeq. The sequence was processed using the genomic analysis application DRAGEN enrichment version 3.9.5 (GenomOncology LLC, Cleveland Ohio, USA), designed to accurately detect base substitutions, insertions, deletions, and copy number variations. NGS revealed a pathogenic mutation in PIK3CA (3140A>T) and a pathogenic variant in RUNX1 (AML1). Other detected genetic variants of unknown significance included CD276, ATM, NCOR1, PIK3R2, DOT1L, AXL, POLD1, SEC23B, LRP1B, CUL3, and MAGI2.
The patient underwent an uneventful surgical recovery and adjuvant treatment involved radiation therapy. Chemotherapy was considered but not recommended due to the patient’s advanced age.
Discussion
ON with multiple differentiation patterns is a very rare entity within primary sinonasal tumors with only 7 cases reported in the literature to date (Table 1). Treatment guidelines for ON have remained largely unchanged with surgical resection as the standard of care. There is an opportunity to update our understanding as recent studies suggest that postoperative chemotherapy improves survival, especially for high-grade tumors. 13 Often, this is decided on a case-by-case basis; consequently, clinicians and patients would benefit from an updated protocol regarding postoperative chemotherapy. 8
To advance therapy and understand the prognosis for this patient population, we suggest scrutinizing the current tools used to characterize the features of ON, such as the Hyams grading system, the Kadish staging system, and immunohistochemistry markers. There is limited available evidence of the utility of the Hyams and Kadish systems. Some studies demonstrate that they provide some benefit in understanding prognosis and disease progression.14,15 However, these studies do not include ON with divergent differentiation, which limits their generalizability. Immunohistochemistry markers such as chromogranin, S100, and synaptophysin are used by convention but have not consistently predicted clinical outcomes for patients with ON. 4 There is a need for tools that can account for additional tumor characteristics, including those with divergent differentiation. A molecular methodology like NGS has the ability to fill this gap.
Here, we present the use of NGS in ON with divergent differentiation for the first time. Our histological findings showed alteration in TP53, PIK3CA, CDKN2A, and CDK2NC, which have been previously described in the literature on genomic alterations in ON without divergent differentiation. 16 Each of these proteins has targeted chemotherapeutics available, although none have been specifically indicated for ON. We additionally found mutations in RUNX1 (AML1), which is a transcription factor that acts as a tumor suppressor in myeloid and breast cancers. This demonstrates one of the key advantages of NGS—its ability to provide insight at the nucleotide level, which facilitates a more detailed interrogation of the tumor genome. We encourage clinicians to incorporate NGS into the routine work-up for patients with this rare tumor because it may reveal more detailed information about the tumor genotype and contribute to the research community’s effort to better characterize the ON genetic landscape. 17 As genetic evidence grows, we can move closer to potential future therapeutics and unveil the molecular mechanisms driving divergent differentiation. 18
Conclusion
ON with divergent differentiation is a rare tumor with varying pathological features. Leveraging NGS can expand our understanding of this rare tumor and potentially deliver more personalized treatment to patients with ON with divergent differentiation.
Footnotes
Author Contributions
Anil Patel: Conceptualization, data gathering, writing—original draft. Eunice Im: Writing—review & editing, data gathering. Jesse Kresak: Writing—review & editing, data gathering. Erica Olgaard: Writing—review & editing, data gathering. Jason E. Blatt: Writing—review & editing, data gathering. Brian C. Lobo: Writing—review & editing. Nikita Chapurin: Conceptualization, data gathering, writing—original draft.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics Statement
Our institution does not require ethical approval for reporting individual cases or case series.
Consent Statement
Appropriate written consent was obtained to publish necessary patient information.
Data Availability Statement
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
