Abstract
Objectives:
The aim of the research is to create an experimental data set of coronal section images of a human head.
Methods:
The head of a 49-year-old male cadaver was scanned by computed tomography (CT), then perfused with a green filling material via the bilateral common carotid artery, before being frozen and embedded. The head was sectioned along the coronal plane by a computer-controlled 5520 engraving and milling machine, capable of either 0.03-mm or 0.06-mm interspacing. All images were captured with a Canon 5D-Mk III digital camera.
Results:
A total of 3854 section images were obtained, each with a resolution of 5760 × 3840 pixels. The number of section images at 0.03- and 0.06-mm interspacing were 1437 and 2417, respectively. All the images were stored in JPG and RAW formats. The image size of each RAW format was about 24.5 MB, whereas for JPG format, the equivalent size was about 5.9 MB. All the RAW and JPG images together occupied 117.35 GB of disk space.
Conclusions:
The interspacing of this data set section was thinner than those of any comparable studies, and the image resolution was higher, too. This data set was also the first to take coronal sections of the human head. The data set contains image information from the smallest structures within the human head and can satisfy the needs of future developments and applications, such as the virtual operation training systems for otolaryngology, ophthalmology, stomatology, and neurosurgery, and help develop medical teaching software and maps.
Introduction
In 1989, the United States proposed the creation of the Visible Human Project (VHP), which was intended to present detailed 3-dimensional (3-D) images of the normal male and female human anatomy. 1 In 1994 and 1996, researchers at Colorado University obtained slice image data sets of both a male and a female. The images from both data sets were 2048 × 1216 pixels in size, with each pixel representing 0.33 mm.2,3 Within a few years, researchers in the Republic of Korea proposed a Visible Korean Human (VKH) and produced the first sectioned images of a male cadaver in 2000. This data set consisted of a total of 8590 sections of 0.2-mm slices. Each image was 3040 × 2008 pixels in size, with each pixel representing 0.2 mm. 4 In 2002, researchers completed the collection of data sets of a man and a woman known as the Virtual Chinese Human (Virtual Chinese Human Male No. 1, VCH-M1, Virtual Chinese Human Female No. 1, VCH-F1), with slice thickness of 0.2 mm and images 3024 × 2016 pixels in size.5,6 In 2010, Park et al 7 in South Korea created 2343 independent head image data sets on a horizontal plane with a slice interval of 0.1 mm, the size of each image being 4368 × 2912 pixels.
In terms of the development and application of image data sets for visible humans, South Korea segmented and marked the structures of 937 organs of the VKH as 3-D model resources, to be used for medical simulation systems. 8 China also segmented 869 organ structures of men and 860 of women, using both surface and volume rendering. 9 In the field of otolaryngology—head and neck surgery, an endoscopic surgery simulator with both visual and tactile feedback was developed by Lockheed Martin, using a 3-D nasal model of the VHP data set, and this has been used to help doctors practice sinus surgery and receive tactile feedback when simulating surgical incisions, especially through muscle. 10 Later, Rudman et al 11 successfully developed the functional nasal endoscopic surgical training simulator based on the Visible American Female data set. In building nasal data sets, Vartanian et al 12 obtained 460 digital images by milling a cadaveric head using 0.33-mm intervals and have since performed 3-D reconstructions of the nose for teaching anatomy. Park et al 13 developed an ear surface 3-D model for teaching the virtual otoscope and for training medical students, using an independent head sectional image data set. Tran et al 14 created an electronic anatomical head based on the data set of the Visible American Female, which was used in the study of electronic cochlear implants. Kahrs and Labadie 15 extracted image data of the temporal bone from the data set of the Visible American Female and reconstructed a 3-D model, using volume rendering and real color rendering technology, which was helpful in exploring surgical approaches prior to cadaver tests and clinical practice.
In this study, we aim to create the first coronal section image of the human head and help provide the theoretical basis for anatomical and clinical studies in future literature.
Patients and Methods
Specimen Preparation
The skull exemplar was obtained from the human anatomy laboratory of the medical department at Shenzhen University. The skull belonged to a 49-year-old male whose sudden death was due to cardiac disease and who had wanted his body to be donated to science. The head was free of organic disease and lesions. The donation was received by technicians in the receiving center of remains donation in the department of medicine at Shenzhen University at 12 noon on the day of death. Specimens of the head, showing no obvious abnormality, were obtained from above the cricoid cartilage surface. The use of human remains was approved by the research ethics committee of the Medical College of Shenzhen University.
The perfusion of vessels in the head was performed on the same day at 3
Specimen Embedding
An embedding box 190 mm × 250 mm × 250 mm in size, matching the specimen’s length, width, and height, was manufactured from 6-mm-thick organic glass. The base and sides were removable, and the top surface was put aside to allow for the filling with the embedding liquid. Four locating holes, each 2 mm in diameter, were positioned in the corners of the front and back surfaces, and soft white silica gel rods were inserted and tied to fix them in position. The specimen was embedded with the head upside down and the chin up so that the embedding fluid could drain into the nasal cavity. Methylene blue solution was added to 3% of gelatin for use as the embedding fluid (Figure 1A).
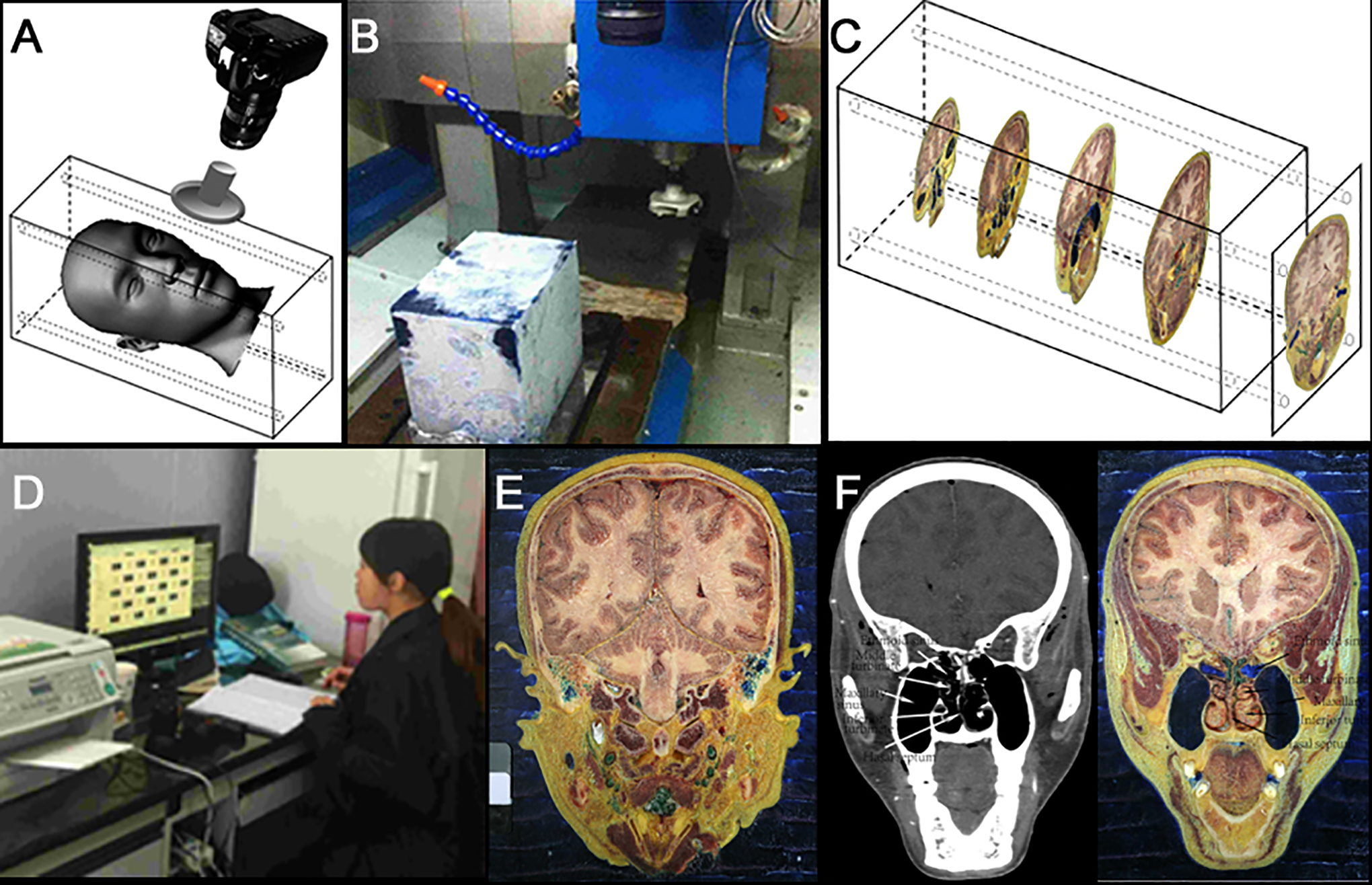
(A) Embedded head for sectioning; (B) milling machine; (C) milling pattern; (D) images collection; (E) a whole section image of the specimen; and (F): computed tomography and the corresponding section image.
Specimen Sectioning and Image Collection
Milling of the specimen was performed by a computer-controlled 5520 engraving and milling machine (produced by the Xuyang machine tool factory, China, and jointly designed by engineers from our team and the factory). This equipment setup was capable of a precision of 0.001 mm/layer. The physical size of the photographic images was 5760 × 3840 pixels, and we used line pairs to determine the resolution. During the milling process, the gelatin solution was used to fill the frontal sinus, maxillary sinus, ethmoidal cellules, sphenoid sinus, middle ear cavity, and mastoid air cells. The frozen state of the embedding box was maintained using a double-temperature icehouse, with a cold laboratory (room temperature below 0 °C), liquid nitrogen, and dry ice. The room temperature was maintained between 0 °C and −20 °C, while the milling chamber was at −5 °C. The whole head was cut layer by layer, from front to rear, in the coronal position. The box was moved through the rotating cutting blade on the cryomacrotome to produce serially sectioned surfaces, with 0.03-mm intervals in the ethmoid area and 0.06-mm intervals elsewhere. Figure 1B shows the specimen prior to milling, and the specimen milling direction is shown in Figure 1C.
When cavity structures such as the sinus and middle ear were to be cut, each structure was filled with more embedding liquid to prevent internal microstructures (such as the sieve interatrial septum and ossicular chain) from collapsing. This also prevented any deep cavity structures distorting the images of these sectioned surfaces. Once frozen by the liquid nitrogen, the cavities could continue being cut by injection filled with warmed embedding liquid. When the cutting was done around the ethmoid sinus, the slice thickness was changed to 0.03 mm.
The whole milling process took more than 270 hours and produced 3854 RAW and JPG images. The images of each sectioned surface were captured with a resolution of 5760 × 3840 pixels by a Canon 5D-Mk III digital camera with a Canon 35 mm f/2.8L II USM lens. Each surface was cleaned with a towel soaked in alcohol. A scale and color card were placed on the margin of the specimen block to ensure consistency between the images. All the images were checked on a computer monitor, and adjustments were made to the camera parameters if the picture quality was poor.
Results
Parameters of the Image Data
A total of 1437 images were made of the ethmoid region, with a thickness of 0.03 mm, and 2417 images of elsewhere were produced, with a thickness of 0.06 mm. With an image resolution of 22.1 megapixels (5760 × 3840), each pixel corresponded to 0.1 mm in the lateral dimension. Each RAW image occupied 24.5 MB of disk space, while the average size of the JPG images was 5.9 MB. The total data size of the RAW and JPG images combined was 117.35 GB.
Contents of Data Sets
The data sets included coronal images of organ structures from the otolaryngology, visual, central nervous, and oral systems.
Image Information
The images we obtained were all clear, the tissues retaining their original color (Figure 1D). Microstructure at a level of 0.2 mm could be identified. The contrast between the arteries and the surrounding tissue was distinct due to the use of green perfusion, and the 0.2-mm diameter arteries were enlarged and could be identified (Figures 1E-F and 2A-J).
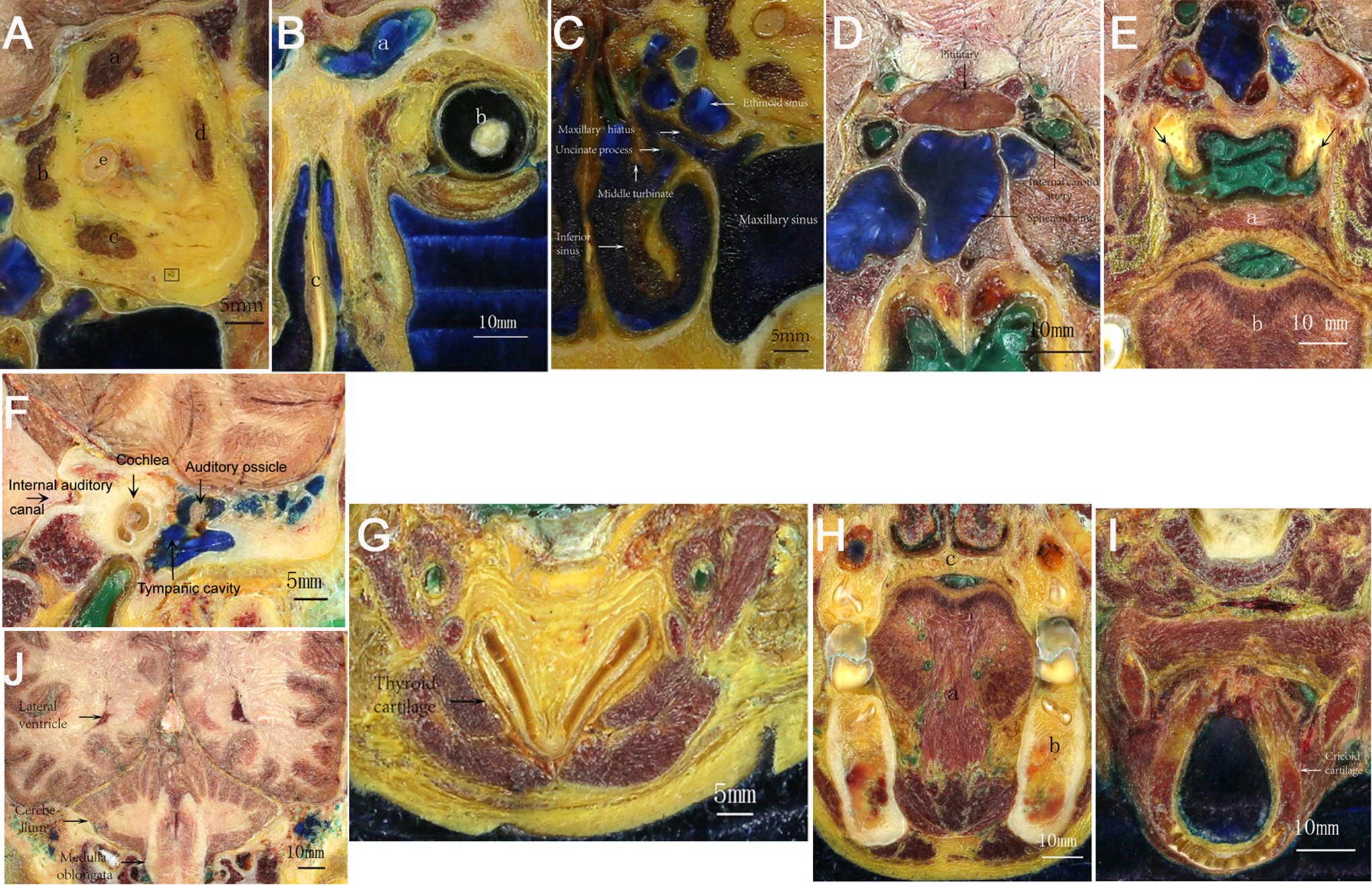
A, Optic nerve (e), superior rectus (a), rectus (b), rectus (c, d), artery. B, Frontal sinus (a), eyeball (b), turbinate (c). C, Uncinate process, ethmoid sinus. D, Carotid artery, pituitary. E, Pharyngeal recess, palatum molle, lingua. F, Ear. G, Thyroid cartilage. H, Oral cavity. I, Cricoid cartilage. J, Brain.
Discussion
The head—the most complex part of the human body—includes soft tissues, bones, and cavity structures. There are many delicate anatomical structures such as the interatrial septum, which is only 0.1 mm in size. Since the milling thickness in this experiment is 0.03 mm, this ensures that there are at least 2 slices that contain image information for any structure. This was essential to prevent the loss of image information of the fine structures, but, unfortunately, it resulted in an extremely large image data set.
At present, in all the visible human data sets created around the world, the slice intervals of male and female specimens are 1 and 0.33 mm, respectively, with a 2.5-megapixel image resolution (2048 × 1216).2,3 In the VKH data set, the slice thickness was 0.2 mm, with a 6.1-megapixel (3040 × 2008) image resolution. 4 In the VCH data set, the thickness and picture size were unchanged, but a slightly different aspect ratio of 3024 × 2016 was used.5,6 The slice interval of the independent head image data set created by Park et al was 0.1 mm, with a 12.7-megapixel resolution (4368 × 2912). 7 In our data set, by comparison, the ethmoid bone area was scanned with a slice interval of 0.03 mm, while the rest of the head used 0.06 mm, and the camera recorded images of 22.1 megapixels (5760 × 3840; Table 1). In addition, the independent head image data sets created by VHP, VKH, VCH, and Park are all horizontal plane images, so our data set is the first to use coronal sections.
Parameters of the Visible Human Data Sets.
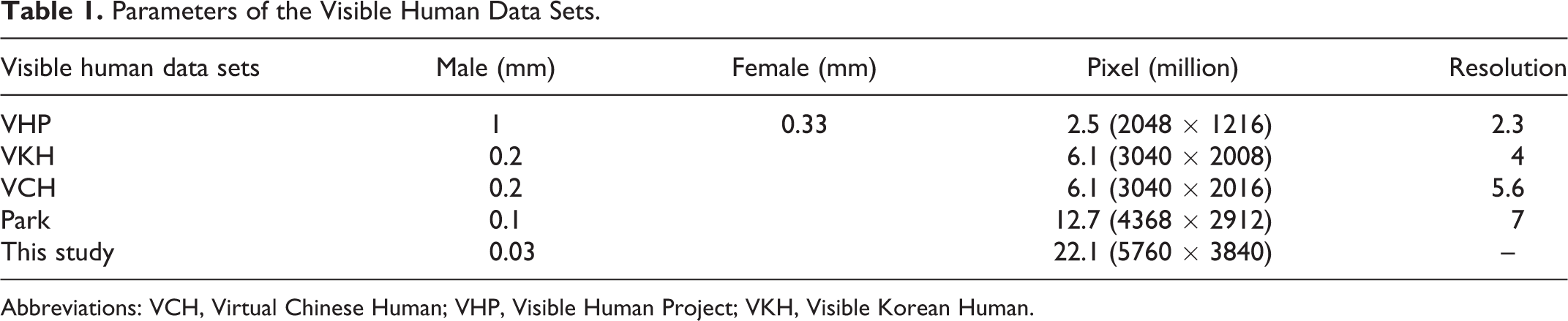

Abbreviations: VCH, Virtual Chinese Human; VHP, Visible Human Project; VKH, Visible Korean Human.
Apart from the application development in the field of otolaryngology—head and neck surgery, the head image data set can also be used for the development of applications in ophthalmology, neurosurgery, dentistry, and other fields. For example, Allen et al 16 have developed an interactive digital app for medical education with 3-D eyes, eye muscles, and nerves controlling eye movement. Park et al 17 identified tracks of the oculomotor nerve (oculomotor nerve, III), the trochlear nerve (trochlear nerve, IV), and the abducent nerve (abducens nerve, VI) in the section images and reconstructed a 3-D surface model based on surface rendering technology. Thus, their image data set has already provided a basis for development in other areas. An advantage of these serial section images for medical education is that it is possible to fully track these 3 nerves, and the 3-D surface model contributes to the understanding of their shape and position as well as their adjacent structures.
The human tongue consists of crisscrossed skeletal muscles, and its structure is associated with obstructive sleep apnea syndrome. In relation to this, Kajee et al 18 extracted the shape and muscle groups of the tongue as well as the direction of muscle fibers from the US visible human data set and built a geometric and structural computer model for studying obstructive sleep apnea. The anatomical structure of the human tongue is quite complex. Sanders and Mu 19 extracted serial slice image data of male and female tongues from the US VHP. The level images produced by computer processing reconstruction were transformed into serial coronary section images. Each image of the tongue’s muscles was segmented by outlining the reconstructed digital 3-D surface model of the tongue so that each muscle could be observed in its original location. This helps clinical and research personnel to recognize each muscle of the tongue. Li et al 20 developed an interactive 3-D digital map, which added anatomical markers to the main structures of the brain in 3 primary planes of section. This digital brain atlas can be used in anatomical studies, user self-testing, student self-assessment, and anatomy teaching. Based on the head image data set, Chung et al 21 reconstructed a 3-D model of the cavernous sinus and its adjacent structure and determined its hexahedral shape. The cross-sectional images have also contributed to the study of the cross-sectional anatomy of the cavernous sinus for radiologists, while the 3-D model has helped neurosurgeons study the sinus’ directional 3-D anatomical structure. These studies have demonstrated that 3-D reconstruction is vital for obtaining accurate head images.
Arora et al 22 show that virtual surgery of the temporal bone and simulated training on endoscopes can be of huge benefit to a doctor training program. Considering the complexity of the anatomical structures of the head organs and the high level of skill needed in surgery, it is necessary to develop high-precision teaching models of the head organs, plus software and virtual surgery training. As the first independent data set showing the coronal section images of the head with both high cutting precision and high image resolution, this current study can satisfy the needs of anatomical researchers, clinical doctors, and medical students. It can also be used to create 3-D anatomy teaching software for the otolaryngology organs, the visual system, the mouth, and the central nervous system. Further, it can be used for developing high-level surgical training systems for otolaryngology, ophthalmology, dentistry, and neurosurgery.
Limitation
Although this data set contains image information of the smallest structures within the human head, it does have certain limitations. The sample size of the study was small (a single adult cadaver head specimen), and a median value of relevant anatomical data could not be obtained. At present, there are no anatomical pictures of corresponding parts of a juvenile, so there is still a need to add to the head axis and coronal image collection. These would be an important basis for subsequent clinical studies and applications, although axial and coronal image reconstruction can still be performed. In addition, further support for image acquisition and software usage needs to be developed. These limitations could impact on future studies and applications, and so a larger patient cohort study is required.
Conclusion
The interspacing of our data set section is the thinnest and the image resolution the highest of any study so far. This data set is also the first to detail the coronal section of the human head and contains imagery of the tiniest structures within the human head. Despite some drawbacks, it should be able to meet the needs of developers of future applications, such as virtual operation training systems for otolaryngology, ophthalmology, stomatology, and neurosurgery, as well as developers of medical teaching software and maps.
Footnotes
Authors’ Note
Xiang-Dong Chen and Jun Wang contributed equally to this study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the Shenzhen Science and Technology Projects (no. JCYJ20120613103507514 and no. JCYJ20170302153015013).
