Abstract
Background:
ARF nucleotide-binding site opener (ARNO) is a guanine nucleotide-exchange factor for ADP-ribosylation factor proteins. ARF nucleotide-binding site opener also binds MyD88, and small-molecule inhibition of ARNO reduces inflammation in animal models of inflammatory arthritis and acute inflammation. However, whether genetic deletion of Arno in mice reduces pathologic inflammation has not yet been reported. Furthermore, its role in the nasal cavity has yet to be investigated.
Objective:
To generate Arno knockout mice and to determine whether genetic loss of ARNO reduces eosinophilic inflammation in the ovalbumin (OVA) murine model of rhinitis.
Methods:
Arno knockout mice were generated and wild type and knockout littermates were subjected to the OVA-induced mouse model of rhinosinutitis. Eosinophilic inflammation was assessed through immunofluorescent quantification of EMBP+ eosinophils in the septal mucosa and cytokine expression was assessed by quantitative polymerase chain reaction.
Results:
Arno knockout mice are viable and fertile without any noted deficits. Arno wild type and knockout mice subjected to the OVA-induced model of rhinitis demonstrated an average of 314.5 and 153.8 EMBP+ cells per mm2 septal tissue, respectively (P < .05). Goblet cells per mm of basal lamina were assessed via Alcian blue and there was no statistically significant difference between Arno wild type and knockout mice. Ovalbumin-induced expression of interleukin-5 (IL-5) was significantly reduced in Arno knockout mice (P < .05). There was no statistically significant reduction in IL-4, IL-13, or eotaxin-1 expression.
Conclusions:
These data demonstrate that deletion of Arno reduces eosinophilic inflammation and IL-5 expression in an OVA-induced model of rhinitis.
Introduction
Allergic rhinitis (AR) is a prevalent immunoglobulin E (IgE)-mediated condition of the nasal cavity.1,2 Characteristic symptoms of AR include itching, sneezing, nasal congestion, and rhinorrhea. Allergic rhinitis is classified based on the overall occurrence of the symptoms (perennial vs seasonal), the frequency (intermittent vs persistent), and the severity.1,2 The estimated direct costs of AR ranges from $1.9 to $3.4 billion of which nearly half is attributed to prescription medications. 3 Commonly used treatments include intranasal steroids, antihistamines, immunotherapy, and others.4-6 Allergic rhinitis in patients and in animal models is also characterized by a release of multiple cytokines and chemokines including interleukin-4 (IL-4), interleukin-5 (IL-5), interleukin-13 (IL-13), eotaxin-1, and others.1,7 Furthermore, there is often an accumulation of eosinophils in the nasal cavity which may further drive the disease pathophysiology. 8
ARF nucleotide-binding site opener (ARNO), also known as cytohesin-2, is a guanine nucleotide-exchange factor for ADP-ribosylation factor GTPases. 9 ARF nucleotide-binding site opener is a member of the cytohesin family, which are characterized by a heptad repeat coiled coil, Sec7, and pleckstrin-homology domains.10-12 The cytohesin family have known functions in cell migration, endocytic trafficking, receptor signaling, and cell adhesion.10,13-16 ARF nucleotide-binding site opener has been shown to bind MyD88 (an adaptor protein important for many innate immune response signaling pathways) and inhibition of ARNO with the small-molecule inhibitor SecinH3 has been shown to reduce pathologic inflammation in the collagen-induced mouse model of arthritis and Carrageenan air-pouch model of acute inflammation.17,18 However, whether genetic deletion of ARNO in mice reduces pathologic inflammation has not yet been reported. Furthermore, its role in the nasal cavity has yet to be investigated.
The ovalbumin (OVA)-induced model of allergic inflammation is a commonly used murine model of rhinitis. 7 In this model, mice are typically first sensitized by OVA via intraperitoneal injection combined with an adjuvant. 19 After sensitization, mice are repeatedly challenged with intranasal instillation of OVA. This technique results in release of inflammatory cytokines and chemokines as well as local nasal eosinophil accumulation. 7 The goal of this study was to generate Arno knockout mice and to determine whether genetic loss of Arno reduces eosinophilic inflammation and inflammatory cytokine release in the OVA murine model of rhinitis. Our rationale also included that should a reduction in eosinophilic inflammation or inflammatory cytokine release be observed, that this study may serve as a springboard to investigation of the role of ARNO in lower airway eosinophilic inflammation, cytokine signaling in the nasal cavity, and potential clinical applicability through small-molecule inhibition.
Methods
Generation of ARNO Knockout Mice
Mice carrying mutations in the Cyth2 (Arno) gene were generated from embryo-derived stem cells purchased from the European Mouse Mutant Cell Repository (EUCOMM). The cell line, designated EPD0140_3_D11, was heterozygous for an allele designated by EUCOMM as Cyth2tm1a(EUCOMM)Wtsi. This allele contained a selection cassette flanked by FRT sites linked to a loxP site inserted into the fourth intron of the Cyth2 gene and an additional loxP in the sixth intron. Embryo-derived stem cells were injected into mouse blastocysts (C57Bl/6) by the University of Utah Transgenic Animal Core Facility to generate chimeric animals that were in turn mated with FLP-expressing mice. Progeny animals lacking the reporter cassette but containing the 2 loxP sites were used as founders for a colony of Cyth2 conditional mice. Crossing of these mice with germ-line expressing Cre mice generated animals carrying the null allele of Cyth2.
Genotyping Protocol
Arno (Cyth2) primers used were oDLKT233: AGAAACAAGCCACATTCACCTCAG, oDLKT235: GTACTCTGATCAGATCTGTAAGCTC, and oDLKT255: CTGCTAGTGTCCCTGCAAGGTG. Primers oDLKT233 and oDLKT235 amplified ARNO+/+ and ARNO FL/FL alleles generating fragments of ∼217 bp (+/+) and ∼374 bp (FL/FL), respectively. The ARNO−/− was detected with primers oDLKT233 and oDLKT255 generating a band length of ∼488 bp. All primers are used within one reaction to detect each band. Polymerase chain reaction samples were composed of 12 µL Promega GoTaq Green Master Mix (Cat# M7123), 6 µL H2O, 5 µL primer total, and 2 µL DNA from tail sample per reaction. Thermocycler settings were stage 1) 94 °C for 3 minutes; 94 °C for 30 seconds; stage 2) 57 °C for 30 seconds; 72 °C or 30 seconds; for 35 cycles; and stage 3) 72 °C for 3 minutes; 4 °C indefinite.
Ovalbumin Sensitization
Studies were approved by the Johns Hopkins Institutional Animal Care and Use Committee. Mice were transferred from the University of Utah to Johns Hopkins (A.P.L.). Arno heterozygous (+/−) mice were bred producing littermate Arno+/+ and Arno−/− mice. Allergic sinonasal inflammation was induced as previously described.7,20 In brief, mice were sensitized with an intraperitoneal injection of 900 µg of OVA (grade V; Sigma-Aldrich) and 77 µL of alum adjuvant (Thermo Scientific). One week later, mice received a second intraperitoneal injection of 450 µg of OVA and 77 µL of alum. Beginning day 14 after the initial OVA sensitization, mice were anesthetized with isoflurane and given a daily nasal instillation of 500 µg of OVA in 20 µL of phosphate-buffered saline (PBS) for 14 days.
Histopathology
Murine tissues were prepared for histologic and immunofluorescence analysis as previously described.20,21 In brief, after cardiac perfusion with 4% paraformaldehyde, mouse heads were fixed, decalcified, equilibrated in sucrose, and cryoembedded in Tissue-Tek OCT (Sakura). Sectioned tissue was used for immunofluorescence analysis of eosinophilic major basic protein (EMBP) to assess eosinophil accumulation or Alcian blue-nuclear fast red staining to assess goblet cell proliferation. For immunofluorescence studies, sections were blocked and permeabilized overnight followed by antigen retrieval with 10 mM sodium citrate and 0.05% Tween-20. Sections were then incubated overnight at 4 °C with EMBP (sc-33938, Santa Cruz Biotechnology) and Krt5 (PRB-160P, Covance). The slides were washed in PBS, incubated with Alexa Fluor conjugated secondary antibodies (Invitrogen) and later counterstained with 4′,6-diamindino-2-phenylindole (Invitrogen). Confocal images were collected with a LSM700 confocal laser microscope (Zeiss). EMBP+ cells in sections of septal mucosa and data were presented as mean with standard deviation.
RNA Extraction and Quantitative Polymerase Chain Reaction
RNA extraction and quantitative polymerase chain reaction were performed as previously described. 20 In brief, septal and turbinate mucosa were removed and placed in RNAlater solution (Ambion). Total RNA was isolated from mucosal homogenates and reverse transcription was performed using the Omniscript RT kit (Qiagen) according to the manufacturer’s protocol. Quantitative polymerase chain reaction was performed using the Applied Biosystems StepOnePlus. Expression levels of 18S, IL-4, IL-5, IL-13, and eotaxin-1 were quantified in each sample in duplicate using the TaqMan gene expression system (Applied Biosystems). Amplicon expression was normalized to 18S. Relative changes in gene expression were calculated as 2−ΔΔCt.
Statistical Analysis
Statistical analyses were performed using analysis of variance followed by an appropriate multiple comparisons test. Tukey’s multiple comparisons test was used when variation was similar among groups, and Dunnett’s T3 multiple comparisons test was used when variation among groups was significant. Data that deviated from a normal distribution were log10 transformed before statistical analysis. Statistical significance was defined as P < .05. Results were analyzed and presented with the use of Prism 8 (GraphPad).
Results
Arno−/− mice were generated and were noted to be viable and born at a Mendelian ratio (Figure 1). Arno−/− mice did not demonstrate any notable deficits. Littermate Arno+/+ and Arno−/− mice were subjected to the OVA-induced murine model of rhinitis and immunofluorescence performed to quantify EMBP+ eosinophils in nasal septal tissue. Wild type animals demonstrated an average of 314.5 EMBP+ cells per mm2 septal tissue (Figure 2). In comparison, Arno knockout mice demonstrated an average of 153.8 EMBP+ cells per mm2 septal tissue (51% reduction, P < .05) (Figure 2). The OVA-induced murine model of rhinitis is also characterized by goblet cell hypertrophy. To assess goblet cell hypertrophy, tissue sections were also subjected to Alcian blue-nuclear fast red staining. Although there was a significant increase in goblet cell hypertrophy in OVA-treated Arno+/+ mice, there was no significant reduction in goblet cell hypertrophy in OVA-treated Arno−/− mice (Figure 3).
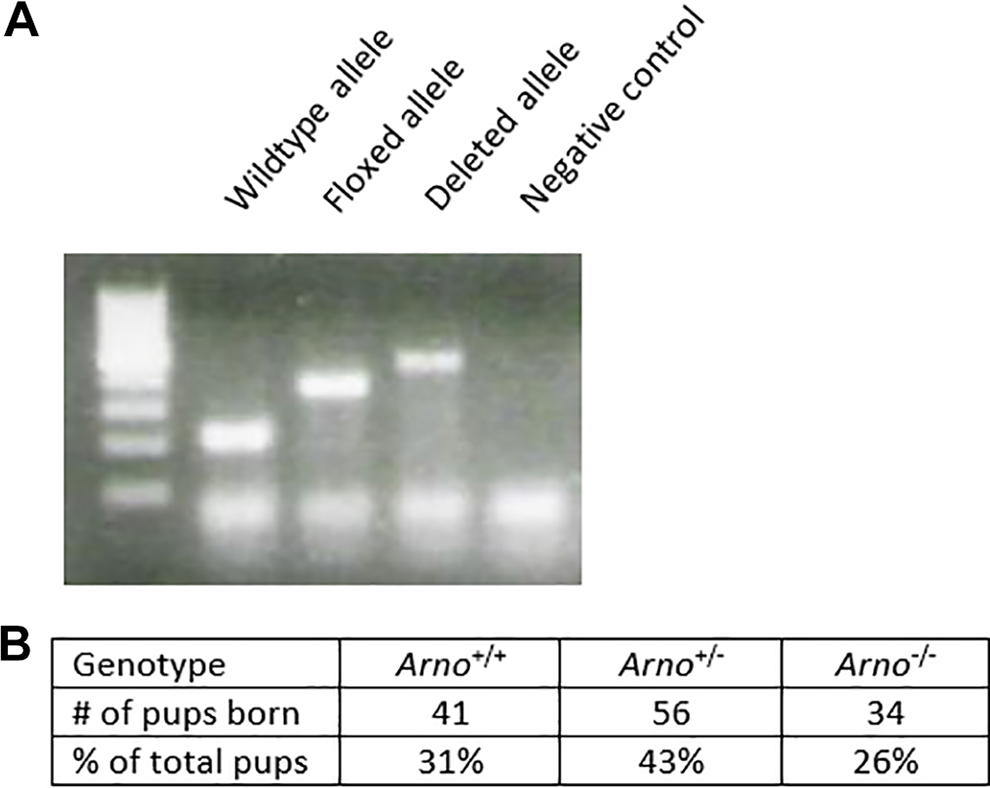
A, Polymerase chain reaction (PCR) genotyping demonstrates Arno wild type, floxed, and deleted alleles. B, Tabulation of 15 litters demonstrates Mendelian inheritance and viability of Arno−/− mice.
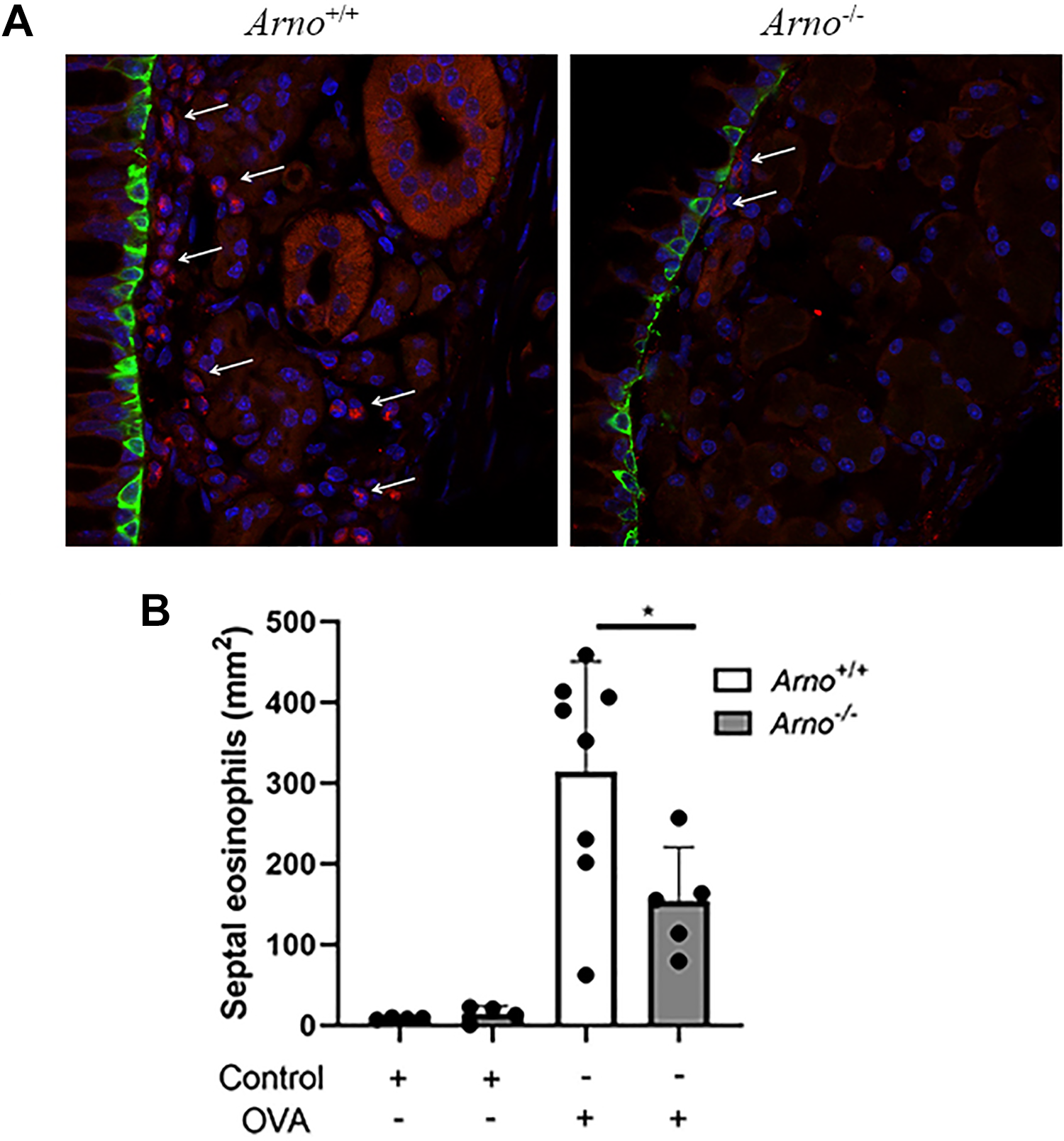
A, Arno+/+ mice (left) and Arno−/− mice (right) were subjected to the OVA-induced model of allergic sinonasal inflammation. A significant decrease in EMBP+ cells in the nasal septum was observed in Arno−/− mice. Representative images are displayed. EMBP (red), KRT-5 (green), and DAPI (blue). B, Quantification demonstrates a significant decrease in EMBP+ cells in Arno−/− mice. For control treated mice N = 4. For OVA-treated mice N ≥ 5. *P < .05. DAPI, 4′,6-diamindino-2-phenylindole; EMBP, eosinophilic major basic protein; OVA, ovalbumin.
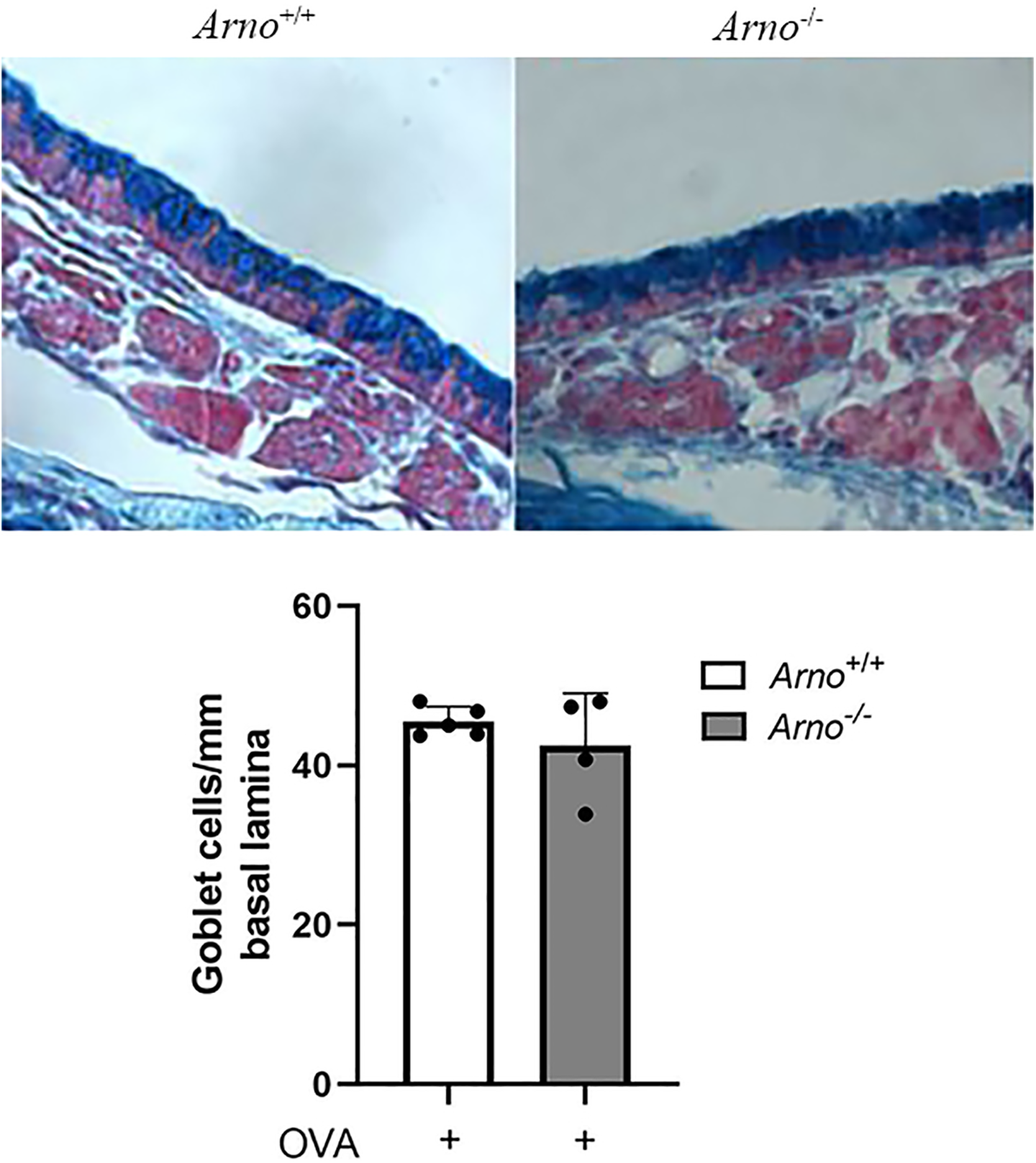
Arno+/+ mice (left) and Arno−/− mice (right) were subjected to the OVA-induced model of allergic sinonasal inflammation. Tissue sections were stained with Alcian blue-nuclear fast red. Representative images are displayed. Quantification demonstrates no statistically significant decrease in goblet cell proliferation in Arno−/− mice. N ≥ 4. OVA, ovalbumin.
Interleukin-5 is a key inflammatory cytokine that regulates eosinophil activation, survival, and tissue accumulation. 22 Thus, we assessed IL-5 expression in sinonasal tissue to determine whether a change in IL-5 could mechanistically explain the decrease in eosinophil accumulation in Arno−/− mice. There was a statistically significant decrease in OVA-induced IL-5 expression in the sinonasal tissue of Arno−/− mice (Figure 4). To determine whether this change in expression applied to additional inflammatory cytokines and chemokines, we also assessed IL-4, IL-13, and eotaxin-1 expression. Interestingly, there was no significant change in expression in these cytokine and chemokines between Arno wild type and knockout mice (Figure 5).
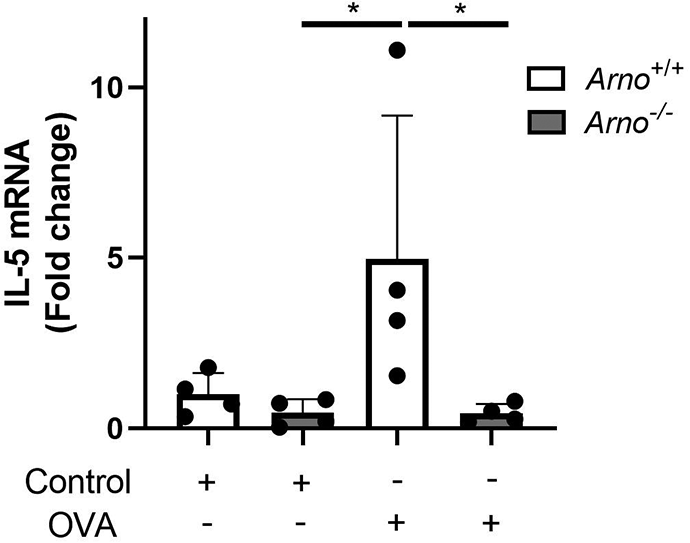
Arno+/+ and Arno−/− mice were subjected to the OVA-induced model of allergic sinonasal inflammation and IL-5 expression assessed demonstrating a statistically significant decrease in Arno−/− mice. N = 4 per group. Error bar represents standard deviation. *P < .05. IL, interleukin; OVA, ovalbumin.
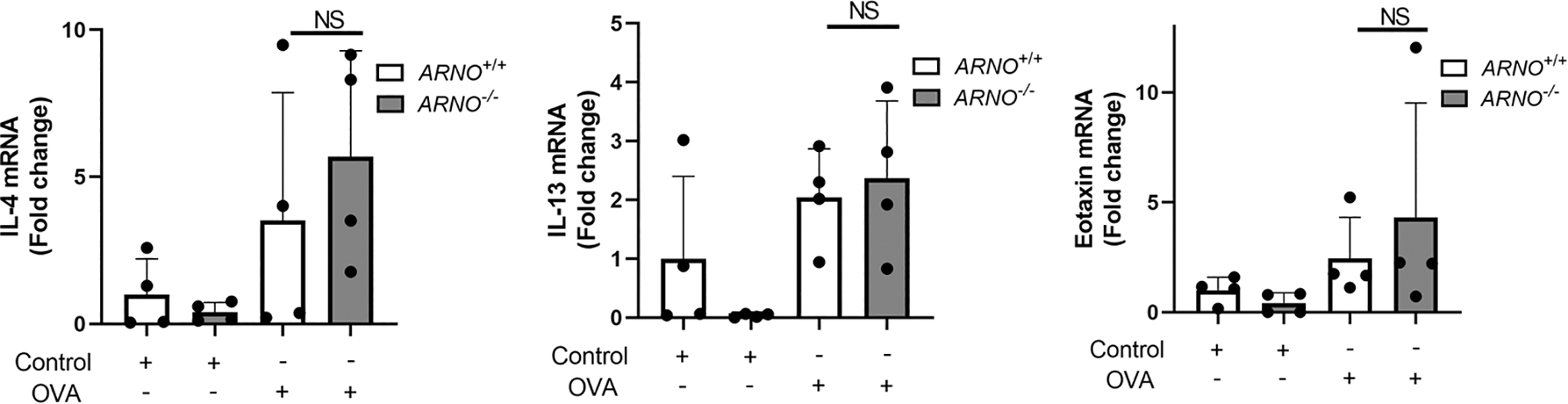
Arno+/+ and Arno−/− mice were subjected to the OVA-induced model of allergic sinonasal inflammation and IL-4, IL-13, and eotaxin-1 expression assessed demonstrating no statistically significant decrease in Arno−/− mice. N = 4 per group. Error bar represents standard deviation. IL indicates interleukin; NS, not significant; OVA, ovalbumin.
Discussion
Allergic rhinitis is a prevalent IgE-mediated condition of the nasal cavity with characteristic symptoms including itching, sneezing, nasal congestion, and rhinorrhea.1,2 Allergic rhinitis in patients and in animal models is also characterized by a release of multiple cytokines and chemokines including IL-4, IL-5, IL-13, eotaxin-1, and the accumulation of eosinophils.1,7 Inhibition of ARNO has been previously shown to reduce pathologic inflammation in mouse models of arthritis and acute inflammation. 17 However, whether genetic deletion of ARNO Arno in mice reduced pathologic inflammation had not yet been previously reported. In this study, we generated a targeted deletion of Arno in mice and demonstrated that Arno−/− mice are less susceptible to OVA-induced IL-5 expression and eosinophil accumulation. Furthermore, this is the first study to demonstrate the effects of ARNO inhibition on nasal inflammation in the nasal cavity. Interestingly, while there was no reduction in IL-4, IL-13, or eotaxin-1 expression, there was a significant reduction in IL-5 in the sinonasal mucosal tissue in the absence of ARNO. Interleukin-5 is a key inflammatory cytokine that regulates eosinophil activation, survival, and tissue accumulation. 22 Thus, the reduction in eosinophil accumulation may be explained by the decreased IL-5 expression observed in this study. Although ARNO has been linked to IL-1β signaling, a direct link between ARNO and IL-5 has not been described and is an area of future study. ARF nucleotide-binding site opener has a well-known function in cell migration and eosinophils are known to produce IL-5. Thus, another potential explanation is that ARNO may be important for eosinophil migration, and the observed decrease in IL-5 may be a by-product of the reduction in accumulated eosinophils.
Sinonasal inflammatory upper airway diseases such as AR and chronic rhinosinusitis affect millions worldwide. However, allergic inflammatory disease of the lower airway such as asthma affects >350 million people worldwide. 23 Asthma may have many severe symptoms including airway hyper-responsiveness, wheezing, and dyspnea. 23 Given the close relationship of the upper and lower airway, it would be interesting to investigate whether a similar reduction in eosinophilic inflammation would be observed in an allergic lower airway inflammation model in Arno−/− mice. 24
There are limitations to this study. First, cytohesin family members are known to regulate cell adhesion and migration. 16 It would be interesting to assess in the future whether loss of ARNO directly leads to decreased eosinophil migration. However, given the low numbers of eosinophils under normal conditions, to directly test a direct effect on eosinophil migration may require IL-5 stimulated expansion of Arno−/− hematopoietic cells, and it is unclear how these cells would respond to IL-5 stimulation. However, this would be an interesting area of further study. It would also be interesting in the future to determine whether the effect of ARNO inhibition is through ARF signaling and whether Arf knockout mice demonstrate similar decreases in allergic eosinophilic inflammation.16,17 Further insights may also be gained by repeating these studies in additional models of AR such as papain, house dust mite, and particulate matter. Lastly, from a clinical applicability standpoint, it would be interesting to test small-molecule inhibitors of the ARNO and ARF signaling pathways in these mouse models of allergic inflammation.17,18
Conclusions
The data presented here demonstrate that deletion of Arno reduces eosinophilic inflammation and IL-5 expression in an OVA-induced model of rhinosinusitis. Future studies will help to determine the applicability of these results to other allergic models of the upper and lower airway. Other areas of future interest include assessing for any connection in the IL-5 signaling pathway and ARNO, determining the impact of ARNO in eosinophil cell migration, and use of small-molecule inhibitors in this and other mouse models to asses for potential translational clinical applicability.
Footnotes
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: N.R.L. holds stock in Navigen Pharmaceuticals, is a patent coinvetor targeting signaling mechanisms related to ARNO and was a consultant for Cooltech Inc.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by an AAO-HNS resident CORE grant from the American Rhinologic Society (N.R.L.) and by NIH 1R21AI119547-01 and NIH R01 (R01AI132590) (A.P.L.). This study was presented in part as a poster presentation at ARS at COSM May 19-20, 2016.
