Abstract
Objectives:
Chronic rhinosinusitis (CRS) is one of the most common inflammations in the upper airway. Despite the wide prevalence of CRS, the pathogenesis of this disease is poorly understood. Several components of the innate immune system may play a significant role in CRS, including Toll-like receptor 4 (TLR4), TLR9, and high-mobility group box 1 protein (HMGB1). This study was conducted to determine the expression of TLR4, TLR9, HMGB1, and pNFκ-B p65 in paraffin-embedded blocks of patients with CRS with nasal polyps compared with those of the control group.
Methods:
Twenty-six formalin-fixed, paraffin-embedded samples from patients with confirmed CRS and 26 patients undergoing septoplasty due to anatomic variations and no other inflammatory nasal diseases as the control group were assessed. Expression patterns of HMGB1, TLR9, TLR4, and pNFκ-B p65 genes were examined using real-time quantitative reverse transcription polymerase chain reaction (Real-Time qRT-PCR). Statistical analyses were performed with SPSS and analyzed using unpaired 2-tailed t tests or 1-way analysis of variance.
Results:
Real-time PCR showed that the expression level of HMGB1 messenger RNA was significantly increased in the tissues of patients with CRS compared with controls (P < .05). The other 3 genes were also upregulated in the patients, but were not significant compared with control. Analysis of the Pearson correlation coefficient (r) revealed a significant positive correlation between HMGB1 and TLR4 (r = 0.79, P < .05) in patients and negative correlation between TLR4 and NfκB in the control group (r = 0.94; P < .05).
Conclusions:
Both HMGB1 and TLR4 are increased in the paranasal sinus mucosa of patients with CRS. These results suggest a possible contribution of HMGB1 and its internal receptor (TLR4) in the pathophysiology of CRS.
Introduction
Chronic rhinosinusitis (CRS) is one of the most common inflammations in the upper airway, which often involves the nasal and sinus mucosa. Different factors are involved in the pathogenesis of CRS, such as histopathology, inflammatory cell and T-cell patterns, tissue remodeling, eicosanoid and immunoglobulin E production, microorganisms, and epithelial barrier malfunction. 1,2 Chronic rhinosinusitis has a significant effect on patients’ physical and mental health status that is parallel to those of arthritis, inflammatory bowel disease, asthma, and cancer. 3 Studies also revealed that CRS diagnosis can be associated with 73 million restricted activity days and 24 million patient visits among American patients, placing it among the top 10 disabling chronic conditions, with a great financial burden. 4 –7 Despite the wide prevalence of CRS and its significant deleterious effect on health-related quality of life, the pathogenesis of this disease is poorly understood. Although it is clear that CRS is a chronic and uncontrolled inflammation, there are still challenges to understand the underlying mechanisms that cause this disease. 8,9 Inflammation is regulated by the immune system and can be conceptually divided into 2 components: adaptive and innate. The innate immune system constitutes the first line of host defense and plays an essential role in the early recognition and subsequent triggering of a pro-inflammatory response to invade pathogens. 10 An attractive theory of CRS represents a dysregulation of normal mucosal immune mechanisms that protect or repair the sinonasal epithelium. These defensive strategies are nonspecific and often constitutively active. 11 Recently, it was discovered that human sinonasal epithelial cells express a series of pattern recognition proteins known as Toll-like receptors (TLRs). 12 Toll-like receptors recognize pathogens and activate the signaling pathway to control the initiation, maintenance, modulation, and termination of innate host defense. On activation of TLRs, epithelial cells may initiate the production of defensive molecules specific to a particular pathogen and perhaps transmit a “danger” signal to warn the elements of the adaptive immune system. 13,14
Recent studies demonstrated an interplay among TLRs, pathogen-associated molecular patterns (PAMPs), and high-mobility group box 1 protein (HMGB1), an ancient evolutionary protein belonging to the damage-associated molecular patterns (DAMPs) family. Secretion of DAMPs in the epithelium such as HMGB1 may induce the selective recruitment of immune cells and lead to airway inflammation and thus play an important role in the progression of the inflammation response. Under homeostatic conditions, HMGB1 acts as a nonhistonic DNA-binding protein; however, when released extracellularly, it plays the role of a potent pro-inflammatory cytokine. 15,16 Extracellular HMGB1 binds to different membrane receptors (TLR4, TLR9, and RAGE), and through activation of nuclear factor κB (NF-κB), it releases pro-inflammatory mediators, cytokines, and chemokines; induces endothelial activation; and increases survival of inflammatory cells. 17 –19
Through TLR4 signaling, bacterial products (eg, lipopolysaccharides [LPS]) may subsequently trigger the intracellular signaling pathways, including MAPK-p38 and NF-κB. Many inflammatory responses to LPS are mediated by enhancing the transcription of HMGB1 products of the activated macrophage. Once HMGB1 is released into the extracellular medium, it exerts its effects by binding to its receptors (RAGE, TLR2, TLR4, and TLR9) and activating the NF-κB signaling pathway to cause cytokine production. Consequently, HMGB1 indirectly causes epithelial cell barrier dysfunction. 20 Understanding the mechanisms of inflammatory responses directly or indirectly through HMGB1 may provide significant insight into the pathogenesis associated with rhinosinusitis. Even with therapeutic intervention, treatment response is often only partial and frequently ineffective. Glucocorticosteroids are currently considered as the first-line therapy in the chronic inflammatory diseases of the nose and paranasal sinuses. They directly interact with NF-κB and block its binding to DNA and inhibit the transcriptional activity of the inflammatory mediators. Several cases are resistant to the usual forms of therapy. 21 Chronic rhinosinusitis can be divided into 2 major subgroups: (1) chronic rhinosinusitis with nasal polyps (CRSwNP) and (2) chronic rhinosinusitis without nasal polyps. Chronic rhinosinusitis with nasal polyps appears to be a more refractory and aggressive form of CRS. 22 The current study aimed at investigating the innate immunity mediators in patients with CRSwNP and those of the ones without CRS as a control group. The current study was conducted to determine the expression of TLR4, TLR9, HMGB1, and pNFκ-B p65 in paraffin-embedded blocks of patients with CRSwNP compared with those of the control group.
Methods and Materials
The current study was conducted on 26 formalin-fixed, paraffin-embedded (FFPE) sinus biopsies from patients with CRSwNP, 1 to 2 years old, obtained from the archive of Pathology Lab at the 5th-Azar Hospital affiliated to Golestan University of Medical Sciences, Gorgan, Iran. A total of 26 patients undergoing septoplasty due to anatomic variations and no other inflammatory nasal diseases such as allergic rhinitis or sinusitis from 2016 to 2018 were enrolled as the control group. The groups were matched by gender and age. Patients with HIV infection, hepatitis virus infection, immunodeficiency disease, or other lung diseases were excluded from the study. Demographic characteristics of the patients are reported in Table 1. Freshly obtained tissue as the control group was stored at −80°C until further assessment by Real-Time qRT-PCR. All participants signed informed consent, and the study protocol was approved by the ethics committee of Golestan University of Medical Sciences (code number: IR.GOUMS.REC.1395.24).
Demographic Data of the Patient.
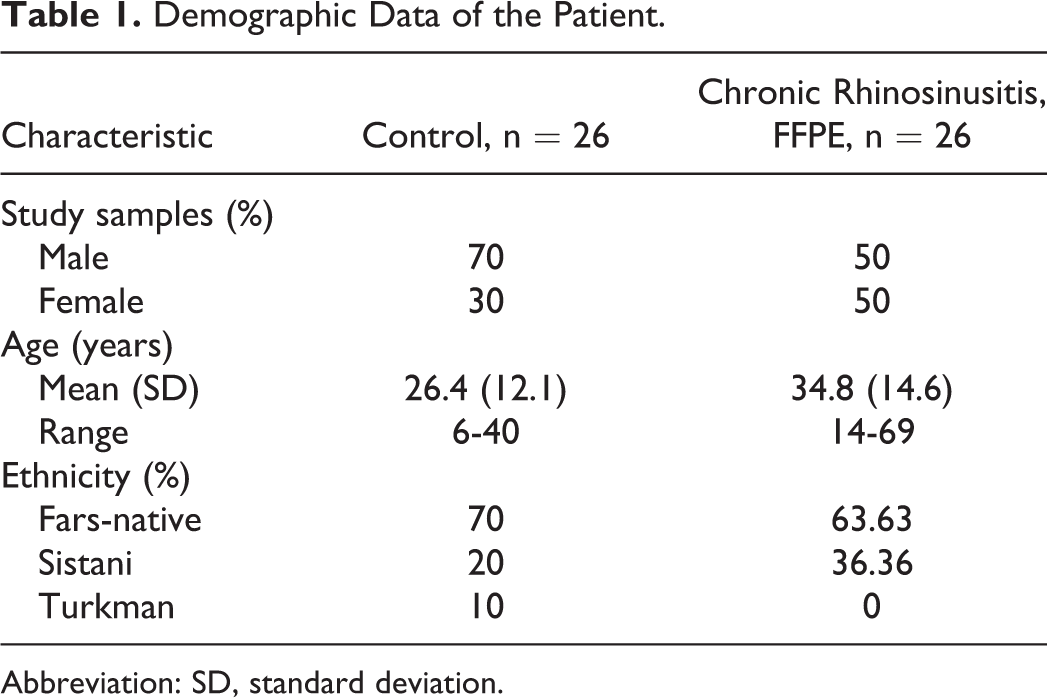
Abbreviation: SD, standard deviation.
Deparaffinization of the FFPE Tissue
Twenty-six, 8-µm-thickness, FFPE tissue sections (1-2-year-old) were deparaffinized by 3 prewarmed changes of xylene incubated for 15 minutes at 65°C and washed with descending concentrations of ethanol as 100%,70%, and 30%. Then, tissue samples were washed with phosphate-buffered saline (PBS) once and dried at room temperature for RNA extraction. 23 To achieve better yield and concentration of RNA, deparaffinization method and time of proteinase K treatment for tissue digestion were optimized. To completely remove paraffin residues from the tissue, the tissue sections were placed in 2-mL tubes containing xylene and incubated for 15 minutes at 65°C. Then, all tissue sections were washed with descending concentrations of ethanol as 100%, 70%, and 30% and once with PBS. To examine the effects of incubation time in the proteinase K digestion buffer, 2 durations were tested (15 minutes according to the manufacturer’s protocol and overnight). All samples were incubated at 56°C in a heating block. Then, proteinase K was inactivated the next day by incubation at 80°C for 15 minutes.
RNA extraction
Total RNA was extracted from fresh-frozen sinonasal samples using CinnaPure RNA extraction kit (SinaClon BioScience, Tehran, Iran), according to the manufacturer’s instructions. In summary, 25 to 50 mg fresh tissue samples were grinded by mortar and pestle in liquid nitrogen and then added 400 μL lysis solution immediately. Then, it was completely homogenized, 300 μL precipitation solution was added and centrifuged at 12 000g, 13 000 rpm, for 1 minute. The supernatant was then discarded and 400 μL wash buffer was added to the precipitate and centrifuged at 12 000 g, 13 000 rpm, for 1 minute. The precipitate was washed with wash buffer II and then centrifuged at 12 000g, 13 000 rpm, for 1 minute. Finally, 100 μL preheated RNase-free water was added to the precipitate and incubated for 3 to 5 minutes at 55°C. Thereafter, the solution was centrifuged at 12 000g, 13 000 rpm, for 1 minute to elute the RNA and digested with DNase I to prevent DNA contamination. Then, all the extracted RNA samples were stored at −80°C. The quality of the isolated RNAs was verified by Nano-Spectrophotometer (Denovix, USA) and electrophoresis before reverse transcription. For FFPE samples, Qiagen RNeasy FFPE kit (Qiagen, Hilden, Germany) was used, according to the manufacturer’s instructions; all pellets were digested with 10 μL proteinase K and 240 μL proteinase K digestion (PKD) buffer (provided by the kit) for overnight. Then, all extracted RNA samples were stored at −80°C. The quality of isolated RNAs was verified by Nano-Spectrophotometer (Denovix, USA) and electrophoresis on a 1.5% agarose gel.
Alternatively, RNA extraction was performed on fresh-frozen and FFPE tissue using TRIzol (Invitrogen Life Technologies, USA) following the manufacturer’s protocol. Following the isolation of RNA, the concentration of each RNA sample was measured and DNase (Thermo Scientific, USA) treatment was performed to remove genomic DNA carryover (Table 2).
RNA Extraction Protocols.
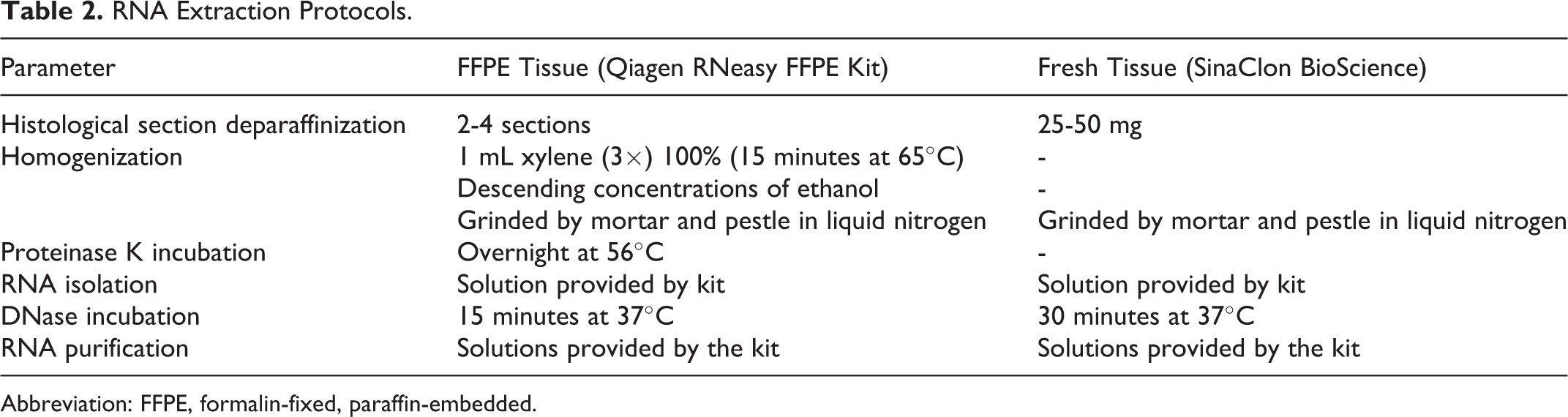

Abbreviation: FFPE, formalin-fixed, paraffin-embedded.
Complementary DNA synthesis and PCR
RNA (1 μg) was reverse transcribed (RT) using a first-strand complementary DNA (cDNA) synthesis kit (Thermo Scientific, USA), according to the manufacturer’s instruction. Briefly, RT reactions were conducted in a 20 μL reaction solution containing 1 μL random hexamer, 1 μL oligo-dt, 4 μL 5× reaction buffer, 1 μL RNase inhibitor buffer, 2 μL dNTP 10 mM, and 1 μL RevertAid M-MuLV RT and RNase-/DNase-free water. Detailed procedures were followed according to the manufacturer’s instructions. The cDNA samples were stored at −80°C until further analysis. In addition, cDNA of the peripheral blood mononuclear cell RNA was also synthesized as the positive control.
The expression of HMGB1, TLR9, TLR4, and pNFκ-B p65 genes was assessed by Real-Time qRT-PCR using SYBR Select Master Mix (ABI, USA) reagent in ABI Real-Time qRT-PCR system (ABI, USA). Polymerase chain reaction primers were designed using online website Primer-BLAST of NCBI (National Center for Biotechnology Information) and purchased from Metabion (Planegg, Germany). The primers are indicated in Table 3. The PCR reactions were amplified at 95°C for initial denaturation followed by 40 cycles at 94°C for 30 seconds, 60°C for 30 seconds, and 72°C for 30 seconds. Each run was completed with a melting curve analysis to confirm the specificity of the amplification and the absence of primer dimers. Relative expression level of messenger RNA (mRNA) was determined with 2−ΔCt value using β-actin as an internal housekeeping gene for normalization. The Ct method uses the difference in Ct value obtained from normalizing housekeeping gene and target gene to calculate relative quantification (ΔCT; the difference in the threshold cycles for target and housekeeping genes).
Primers Used for RT-PCR Quantification.
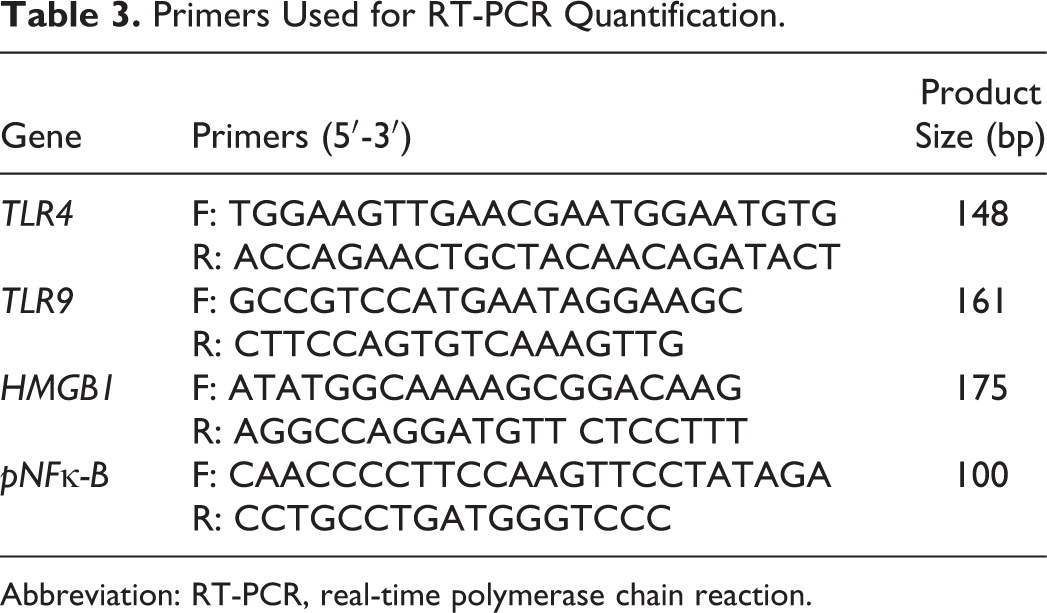

Abbreviation: RT-PCR, real-time polymerase chain reaction.
Statistical Analysis
All statistical analyses were performed with SPSS version 21. The normality of data distribution was assessed by the Kolmogorov-Smirnov test. Normal data were analyzed using unpaired 2-tailed t tests, 1-way analysis of variance (ANOVA), or 2-way ANOVA, as appropriate. The Spearman and Pearson correlation tests were appropriately applied to analyze the correlation of mRNA expression of target genes. Data are expressed as mean ± standard error of the mean, with 95% confidence interval and statistical significance for a P value <.05.
Results
A total of 26 samples of the control group and 26 patients were examined. The demographic characteristics of the samples are shown in Table 1. The gene expression was detected in only 60% of the control and 50% of patient groups. Therefore, the extraction of mRNA was not successful in a number of samples. All stages of deparaffinization method, extraction, production of cDNA, and PCR were optimized. Statistical analyses of the data indicated that deparaffinization of the tissue at 65°C and treating the tissue with proteinase K for overnight duration conferred beneficial effects with regard to total yield of the extracted RNA in comparison with those of the main protocol (P < .05). Average concentration of the extracted RNA and purity was 200 µg/μL with OD260/280 = 1.7. The results of the PCR showed that the β-actin reference gene was expressed in all specimens, but the studied genes were not detectable in some specimens. All 4 examined genes were expressed in samples, and gel electrophoresis was carried out to show the expression of all genes derived from patients and controls samples (Figures 1 –3).

Representative agarose gel electrophoresis image of HMGB1 gene. Lines 1 to 7 from FFPE tissue patients, line 8 the negative control, and line 9 the molecular weight marker. FFPE indicates formalin-fixed, paraffin-embedded; HMGB1, high-mobility group box 1 protein.
Representative agarose gel electrophoresis image of NfκB gene. Lines 2 to 5 from FFPE tissue patients, line 2 the negative control, and line 1 the molecular weight marker. FFPE indicates formalin-fixed, paraffin-embedded; NfκB, nuclear factor κB.

Representative agarose gel electrophoresis image of TLR4 and TLR9 genes. Line 1 TLR9 and line 3 TLR4 both from FFPE tissue patients, line 2 the negative control, and line 5 the molecular weight marker. FFPE indicates formalin-fixed, paraffin-embedded; TLR, Toll-like receptor.
Considering Real-Time qRT-PCR data, distribution of data was normal based on the Kolmogorov-Smirnov test results. Normal data were analyzed using t tests.
According to data analysis, HMGB1 was significantly upregulated in the patient group compared with the controls (P = .03). The other 3 genes were also upregulated in the patients, but were not significant compared with control (Figure 4).
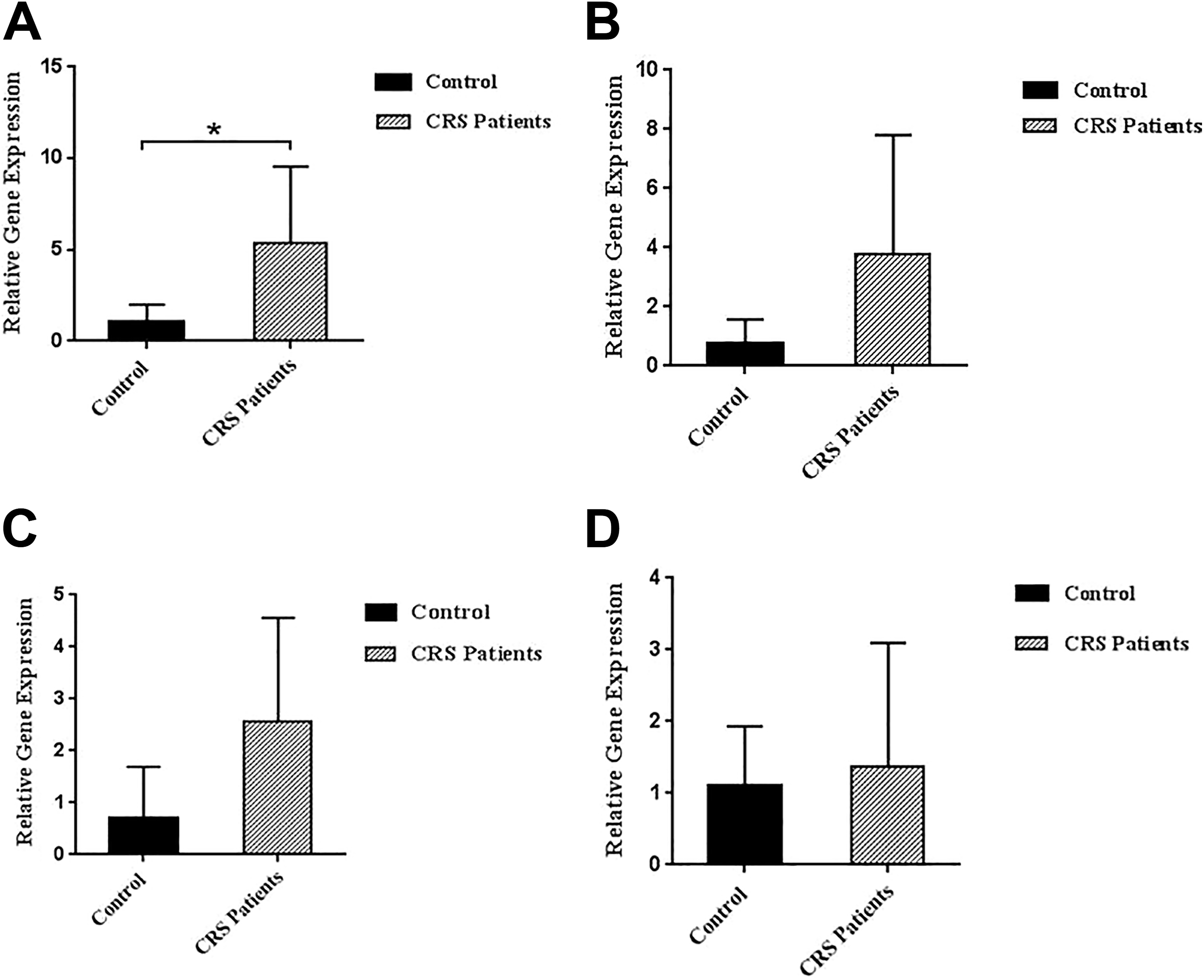
Expression profile of HMGB1, NfκB, TLR4, and TLR9 mRNA. According to Real-Time qRT-PCR results, comparison of the expression of HMGB1 (A), NfκB (B), TLR4 (C), TLR9 (D), and values in the control (dark bars) and patients with CRS (light bars) are normalized to the housekeeping gene β-actin. HMGB1 is significantly increased in comparison with controls (P = .03). HMGB1 indicates high-mobility group box 1 protein; mRNA, messenger RNA; NfκB, nuclear factor κB; Real-Time qRT-PCR, real-time quantitative reverse transcriptase polymerase chain reaction; TLR, Toll-like receptor.
Analysis of the Pearson correlation coefficient (r) revealed a significant and positive correlation between HMGB1 and TLR4 (r = 0.79, P > .05) in patients and negative correlation between TLR4 and NfκB in the control group (r = 0.94; P > .05).
Discussion and Conclusion
Sinusitis is one of the most common diseases of the ear, nose, and throat. The average prevalence of sinusitis is relatively high worldwide, and in some regions of Iran, the prevalence of the disease is reported to be 50% in the population older than 30 years. 24 Although chronic sinusitis is not a severe life-threatening condition, it can lead to serious complications and can cause serious problems due to the proximity of sinusitis to the eye sockets and brain; for example, about 15% of all eye socket infections are directly related to chronic sinusitis, about 3.7% to 10% of the brain abscess cases are due to chronic sinusitis, and 14% of the meningitis cases caused by acute frontal sinusitis are lethal. 25 Unfortunately, despite the high prevalence of this disease in various populations and the economic burden imposed on individuals and work environments, the exact pathogenesis of the disease is not clearly understood yet, and it is even reported that the administration of corticosteroids was useless, despite the side effects. 26 One of the clearest horizons in responding to the diagnostic and therapeutic defects of the disease, and possibly its prevention and control, is to study the immune system functioning in this disease, which could be the cornerstone for immunotherapy in the disease. Knowing the molecules involved in the activation pathway of the immune system in sinusitis and examining the changes that probably occur in their release may play a key role in clarifying this pathway. Therefore, in the current study, 2 receptor molecules that are important in innate immunity can play roles in bacterial and viral sinusitis, TLR9 and TLR4, respectively, as well as 2 important molecules in inflammation, HMGB1 and pNFκ-B p65, were studied.
The Real-Time qRT-PCR technique was employed in the current study to detect and evaluate the expression level of the abovementioned molecules in sinus mucosal tissue samples of the patient and control groups at the RNA level. Patient samples were stored in paraffin blocks and the fresh samples were used in the control group. Since most of the efforts to extract and detect mRNA from paraffin-embedded tissue were useless, unfortunately and usually RNA was not obtained with sufficient concentration and purity. 23 In the current study, with the optimization of various deparaffinization methods, the extraction, and all stages of gene expression detection, RNA was extracted with appropriate purity and concentration for RT-PCR, and all 4 molecules were evaluated and detected in samples. In a study by Tengroth et al, TLR3, TLR7, and TLR9 were expressed on nasal mucosal cells at the RNA level. 27 Also, in the study by Lane et al on chronic sinusitis, expression of TLR2, as well as the inflammatory genes macrophage inflammatory protein α, RANTES, and granulocyte-macrophage colony-stimulating factor in the patient group, was significantly higher than those of the control group. Also, the expression of TLR2 and TLR9 reduced in patients with recurrence of polyps after surgery. 28
In the current study, important molecules in innate immunity, including TLR4 and TLR9, were found in the sinus mucosa of the control and patient groups. In the patient group, both molecules had higher expression compared with the control group, but the expression level of TLR4 was higher than that of TLR9, although the difference was not statistically significant, which was probably due to the small sample size. Since TLR9 and TLR4 are more involved in viral and bacterial infections, respectively, higher TLR4 expression in chronic sinusitis can be justified scientifically.
In the current study, the expression of inflammatory molecules, HMGB1 and pNFκ-B p65, were also investigated and HMGB1 expression was significantly higher in the patient group than the control. In a study by Iosif et al on the delayed expression of HMGB1 in patients with rhinosinusitis, in the control group, the expression of this protein was mostly observed in the nucleus and it was less secreted or released in the cytoplasm, compared with those of the patient group. In the patient group, the expression of this protein was higher in the cytoplasm and outside the cell. 16 Recently, HMGB1, as the main protein in the pathogenesis of several inflammatory diseases, provided a new research context in chronic nasal inflammation. HMGB is a signaling protein opposed to membrane-specific receptors (TLR4, TLR2, and RAGE) on the surface of macrophages and monocytes. The function of this ligand and receptors is to produce pNFκ-B p65, which plays a pivotal role in the phenomenon of inflammation, and is important to produce cytokines, chemokines, and adhesion molecules. 29 In the current study, the expression of pNFκ-B p65 in the patient group was higher than that of the control group, although the increase was not significant. The mechanism of HMGB1 function is through the TLR4, HMGB1 acts as an endogenous ligand for the TLR4, meaning that the expression of these molecules is correlated with the disease or health conditions. Recent studies indicated that chronic sinusitis results from irregular immune responses to microorganisms or their products. In CRS, the abnormal mucosal immune response disrupts the function of microorganism removal and causes inflammation in sinus cavities. Toll-like receptors play an important role to detect pathogens and induce and regulate natural and specific immune responses. Recent studies showed that CRS is associated with increased activity of such receptors. 15,16,18,19 The current study results also indicated the increased expression of receptor molecules as well as inflammatory factors, particularly HMGB1, indicating the significant role of immune responses as well as inflammatory molecules in CRS. Another important point was the relationship between the expression of these molecules; statistical analyses showed a significant direct correlation between TLR4 and HMGB1 expression in patients. This finding indicated the role of inflammation and important molecules in the immune system in CRS, and the scientific justification for this association is that, as already mentioned, HMGB1 is the internal ligand of TLR4, and by the increase of one, the other also increases. As a general conclusion, HMGB1 and its receptor, TLR4, can be considered as important target molecules in immunotherapy and the correction of the immune responses pathways in CRS. Further studies are suggested on the function of such molecules in CRS and their mechanism of alteration in their expression in the immunotherapy pathway of CRS.
Footnotes
Acknowledgments
The authors gratefully thank the Research and Technology Deputy, Golestan University of medical Science, for financial support.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Research and Technology Deputy, Golestan University of Medical Sciences with grant number IR.GOUMS.REC.1395.24.
