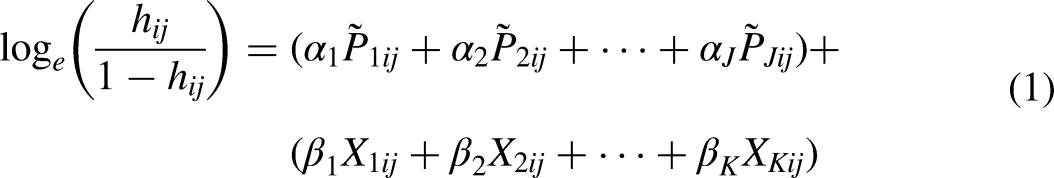Abstract
Discrete-time survival analysis (DTSA) models are a popular way of modeling events in the social sciences. However, the analysis of discrete-time survival data is challenged by missing data in one or more covariates. Negative consequences of missing covariate data include efficiency losses and possible bias. A popular approach to circumventing these consequences is multiple imputation (MI). In MI, it is crucial to include outcome information in the imputation models. As there is little guidance on how to incorporate the observed outcome information into the imputation model of missing covariates in DTSA, we explore different existing approaches using fully conditional specification (FCS) MI and substantive-model compatible (SMC)-FCS MI. We extend SMC-FCS for DTSA and provide an implementation in the
Keywords
Introduction
Many phenomena in the social and medical sciences can be characterized as events—that is, qualitative changes that occur at some point in time. Typical research questions focus on whether, when, and under what circumstances events occur. Examples of sociologically relevant events are divorce or a job offer after a period of unemployment. When analyzing such time-to-event (or survival time) data, one cannot rely on a simple linear regression model, as the event cannot be observed—it is missing (censored)—for parts of the population. For example, some married people never experience divorce, and although everybody dies, data collection will almost certainly not continue until this point for all observations. Different analytical approaches have been developed to deal with the censoring problem. They include, for example, Cox regression for continuous-time survival analysis (Cox, 1972). Cox (1972) also extended the proportional hazard model for discrete-time survival analysis, analyzing the conditional odds of an event occurring at a particular time point, given survival up to that point. For discrete-time survival analysis, data must first be converted from the familiar person-oriented (P) format (one row for each person/observational unit) to a person-period (PP) format (one row for each time period in which a person was observed).
One challenge that arises in the application of these survival models (and in other models) is that often one or more covariates have missing data. Simplistic approaches, such as listwise deletion (LD) and unconditional mean imputation, are still used in the social sciences (e.g. Böttcher, 2006; Cooke, 2006; Arránz Becker and Lois, 2010; Manning, Brown, and Stykes, 2016; Cooper et al., 2018; Stoddard and Veliz, 2019). However, these approaches may be highly inefficient or lead to severely biased variance estimates. Point estimates may also be biased after LD if missingness depends on the outcome (Hughes et al., 2019).
Another approach to handling missing data is multiple imputation (MI) (Rubin, 1987, 1996; Schafer, 1997; van Buuren, Boshuizen, and Knook, 1999). MI leads to unbiased point and variance estimates if certain conditions concerning the missing data mechanism and the imputation model are met (Allison, 2000). However, while there has been researching on how best to impute missing covariate values for Cox regressions (van Buuren, Boshuizen, and Knook, 1999; Clark and Altman, 2003; White and Royston, 2009; Keogh and Morris, 2018), the Cox cure model (Beesley et al., 2016) and the relative survival model (Nur et al., 2009), the imputation of covariates in discrete-time survival analysis is still understudied. For time-varying covariates, Murad et al. (2019) showed that MI approaches using information from the previous and current time points seem sufficient in most situations. However, in discrete-time survival analysis, not only time-varying covariates but also time-invariant covariates are used. This article contributes to the literature by exploring how to specify a suitable imputation model for partially missing time-invariant covariates in discrete-time survival analysis. We present different approximations and an approach using a compatible imputation model now implemented as
This is not an easy or straightforward task. Kenward and Carpenter (2007: 207) shows that including outcome information in the imputation model for partially observed covariates is crucial for unbiased estimates. However, with discrete-time survival models, the time-to-event is not fully observed due to censoring. Nor is it clear in which format the imputation procedure should be carried out: with data in P or PP format? And how should the relationship between the time-to-event variable and the covariates be modeled for imputation?
These are relevant issues, as discrete-time survival analysis is widely used in the social sciences, especially in family sociology (see, e.g., Barber, 2001; Schoen et al., 2002; Cooke, 2006; Nomaguchi, 2006; Arranz Becker and Lois, 2013), and also in the medical sciences (Murad et al., 2019).
The remainder of this article is organized as follows: In the next section, we introduce the formalization of the discrete-time survival analysis model (DTSAM) proposed by Singer and Willett (2003), we address P and PP data set formats, and we briefly discuss the negative consequences of, and ways of dealing with missing covariate data. We then outline the method of MI and present several possible imputation approaches that differ in terms of the data format used and the specification of the imputation model, including a new substantive-model compatible-fully conditional specification (SMC-FCS) approach for DTSAMs implemented in the function
Discrete-Time Survival Analysis Model
The Model
In this section, we introduce the formalization of a DTSAM proposed by Singer and Willett (1993).
The term discrete survival is used when the time-to-event can take only distinct values, for example, one, two, three, or more years/semesters/weeks. Occasionally, discrete-survival data are “truly discrete” (Kleinbaum and Klein, 2012: 325); that is, the event can occur only at distinct values of time (e.g., fertility modeling, particularly the time from puberty to first childbirth). However, in most cases, discrete data are the result of interval-censoring: events might happen in a continuous range of time, but they were observed only in grouped form instead of continuous-time data—for example, the year of divorce is recorded but not the month and day (Kleinbaum and Klein, 2012: 318).
Following Singer and Willett (1993: 163), let
Researchers are usually interested in whether the risk of event occurrence differs systematically between observations. For example, in a study of divorce risks, the risk might depend on age at the beginning of a marriage or on differences in social status between the spouses. For now, we examine time-invariant predictors. We have to distinguish between different individuals
We model the individual hazard
The model to be estimated is that proposed by Singer and Willett (2003: 317):
The Data: Transformations and the Person-Period Format
Having established our model, we take a closer look at our data and the data format needed for a DTSAM. Data are typically available in a P format, with one row (record) for each observational unit (i.e., person). Apart from the covariates,
To estimate a DTSAM (our substantive model), the P data set must be converted into a PP format (Singer and Willett, 1993: 172). In PP format (see Table 1), there is a separate row for each period in which a unit was observed. Apart from the covariates, we usually have observed period indicators into indicator variables
Exemplary data set in person-period (PP) format.
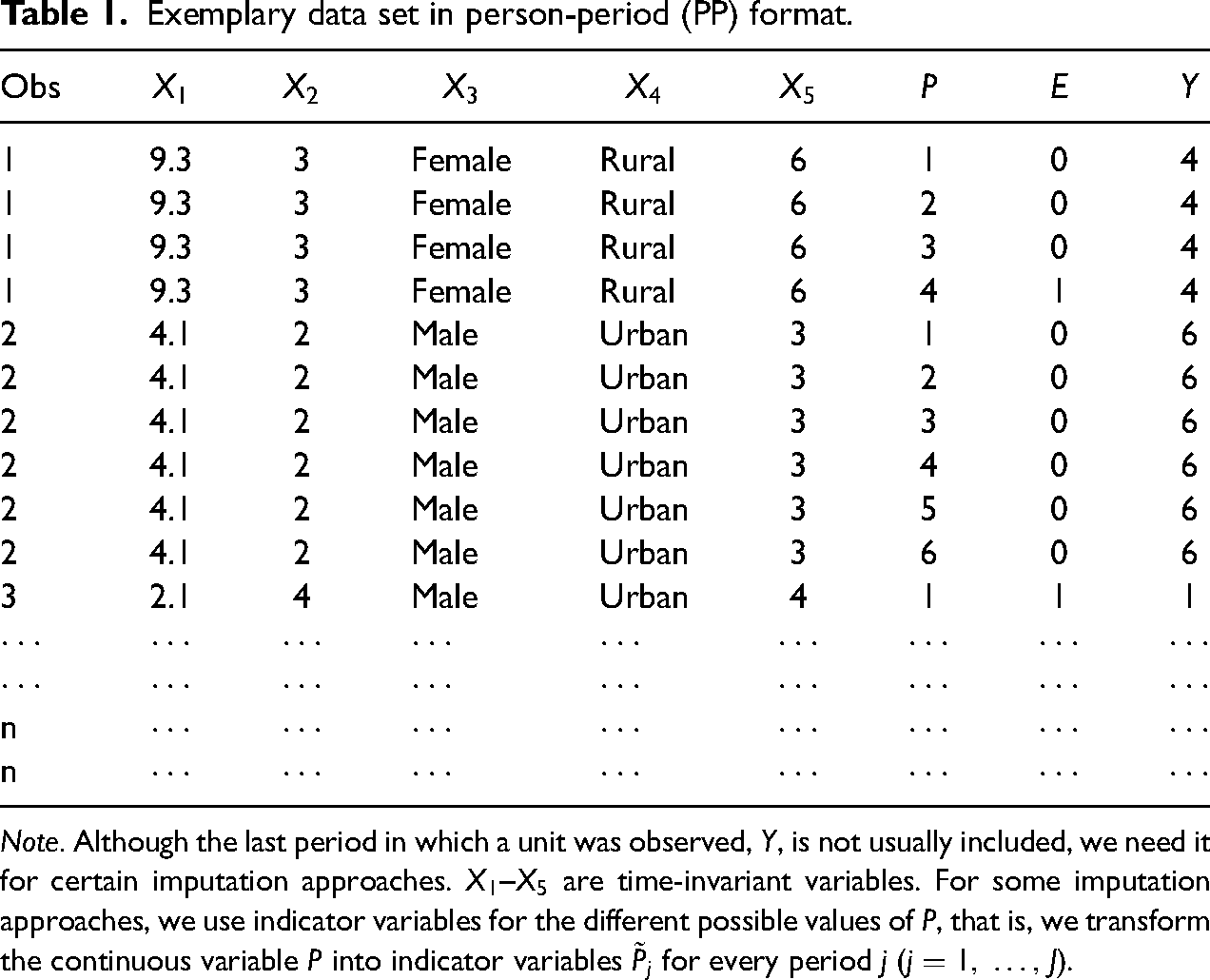
Note. Although the last period in which a unit was observed,
The dichotomous event indicator,
The omission of a critical predictor of the outcome from the model is equivalent to mixing hazard profiles for the different populations defined by the discarded predictor values, that is, the hazards converge. The pooled converged hazard profile does not have to look like any member of the general population. For example, assume that all members of the population have a constant hazard over time, but the height of the risk profile differs between members. Over time, members with a high risk drop out of the population. If we do not include the predictor that shifts the members’ risk, the aggregate profile will show a risk profile that decreases with time (Singer and Willett, 1993: 185).
Building a model that exhausts all sources of individual variation is practically not feasible. However, even in the absence of important predictors, parameters can still be interpreted as an average across population hazard profiles (Xue and Brookmeyer, 1996). For example, if we include obesity as a risk factor for diabetes in our model, and there is unobserved individual heterogeneity, we would estimate the log odds ratio for all people with obesity versus those without. If there were no unobserved heterogeneity, we could interpret the regression coefficient as the log odds ratio for an individual before versus after developing obesity. Apart from complicating the interpretation of factors, Allison (1982: 83) also noted that in the case of frailties, “one would expect this dependance among the observations to lead to inefficient coefficient estimates and standard errors that are biased downward.” As unobserved heterogeneity is not entirely avoidable, we will generate the data sets in our simulations with varying degrees of unobserved heterogeneity. However, before testing the different MI approaches, we take a closer look at the implications of missing data for discrete-time survival analysis.
Implications of Missing Data for Discrete-Time Survival Analysis
Surveys are often subject to missing data, and we have to decide how to treat partially observed covariates in discrete-time survival analysis. Note that, in this article, we are looking only at missing data in the covariates, not in the time-to-event variable.
Before exploring different possible strategies for imputing partly missing covariates in discrete-time survival analysis, we take a more general look at the implications of missing data in regression analysis to demonstrate the need for a suitable MI strategy.
One way to deal with missing data in regression analysis is to exclude incomplete observations. This is known as LD or complete case analysis. One consequence of this approach is a possibly considerable loss of efficiency or information (Carpenter and Kenward, 2013: 9) as only the observations with complete records are used. In other words, even if only one of many covariates is missing, this leads to the complete exclusion of the observation. This becomes especially problematic if the aim is to keep unobserved heterogeneity down and include not only possible confounders but also other important predictors that explain significant parts of the variation.
However, a loss of efficiency, or information, is only one side of the coin; the other is bias. Table 2 gives an overview of the conditions under which logistic regression coefficients will be biased after LD. Here,
Bias in case of logistic regression using complete records.
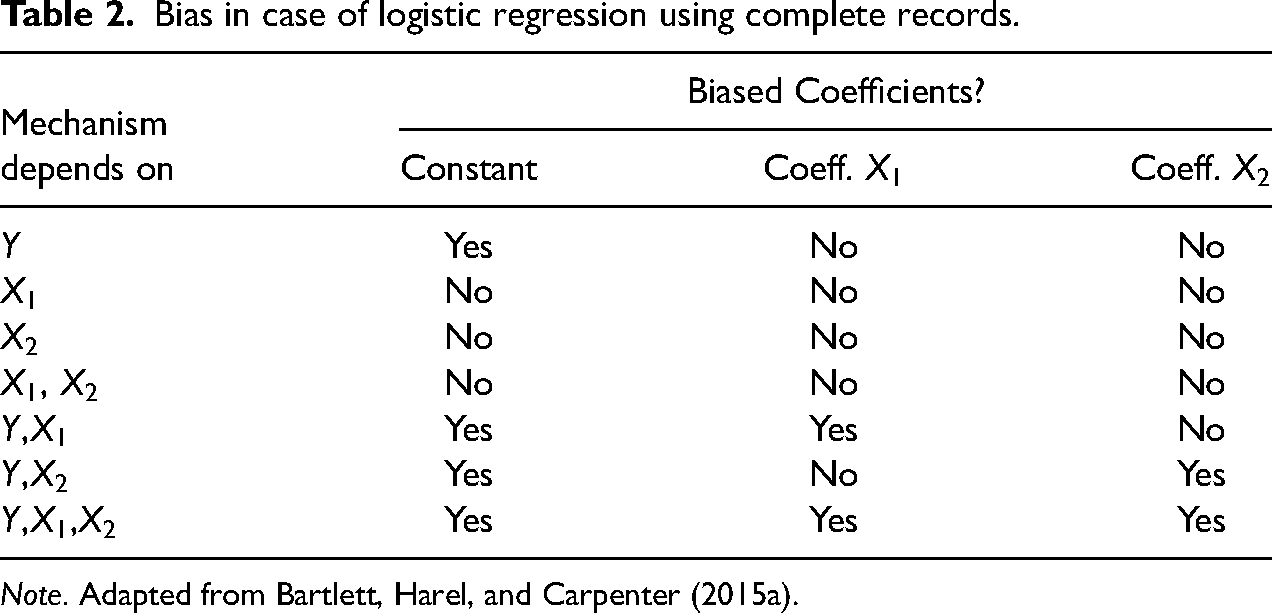

Note. Adapted from Bartlett, Harel, and Carpenter (2015a).
Complete case analysis is valid for linear and logistic regression if missingness depends on the covariates but not the outcome, which in the situation considered here means missingness could depend on
on the binary outcome on the covariate
Therefore, if it must be assumed that the estimated coefficient of our variable
Multiple Imputation
MI in General
One of the most popular approaches to tackle missing data is MI (Little and Rubin, 2002). MI allows for the analysis of incomplete data sets by substituting missing values. Substitution is performed by imputing values of a variable based on other variables, mostly those of the analysis model. In the analyses, these imputed values are not treated the same as observed values, as this would lead to biased variance estimates. Therefore, several values instead of just one are imputed for each missing value in order to avoid treating imputed values as observed. Each data set is then analyzed separately, and estimates and variances are combined across imputations using rules developed by Rubin (1987).
To impute missing values, we need models of their distribution. There are two main approaches, joint modeling (JM) and FCS, also known as multivariate imputation by chained equations or MICE (van Buuren, 2007). 1 Joint modeling MI (Schafer, 1997) draws missing values simultaneously for all incomplete variables using a multivariate distribution. However, specifying such a joint model is often challenging, for example, if there are both continuous and discrete variables with partially missing information in a data set. Newer approaches now allow us to specify the models for each variable one at a time, for example, Erler, Rizopoulos, and Lesaffre (2021). FCS divides the problem into a series of univariate problems (van Buuren, 2007). FCS involves specifying a series of univariate models for the conditional distribution of each partially observed variable, given all the other variables (White, Royston, and Wood, 2011). It is more flexible than the JM approach because adequate regression models can be selected for every variable (e.g., linear regression for continuous partially observed variables, logistic regression for binary partially observed variables). However, for FCS, the joint distribution of variables is only implicitly known and may not actually exist. While this is a serious drawback from a theoretical perspective, it has done little harm to practical application.
When specifying the imputation model, it is crucial to account for the substantive model of interest—which is often also called the analysis model. The associations to be examined in the substantive model must also be represented in the imputation model. Otherwise, bias toward zero will be the likely consequence (Fay, 1992). The imputation and substantive models should be compatible. Compatible means that there exists a joint model with conditionals corresponding to the imputation and substantive models (see Bartlett et al., 2015b and for the related term of congeniality see Meng, 1994). In addition to JM and FCS, a variation of FCS that allows one to more easily specify a compatible imputation model was developed by Bartlett et al. (2015b). It is called SMC-FCS. SMC-FCS is used when it is hard to find a compatible standard FCS imputation model because the substantive model, that is, the model the researcher is interested in, is either non-linear (e.g., a Cox regression) or contains non-linear (e.g., squared or interaction) terms. SMC-FCS can also be used for models without non-linear terms. As with FCS, separate models are specified for the partially observed variables. What differentiates SMC-FCS from regular FCS is that the conditional distribution of a variable (given the other variables) is combined with the specified substantive model to define an imputation model. This combination ensures that the missing covariate data are drawn from models compatible with the specified substantive model.
Handling Missing Covariates Values in Case of a DTSAM as a Substantive Model
Representing the associations in the substantive model of interest is not straightforward in the case of a DTSAM and its complicated outcome structure. Generally, we have two outcome variables: the event indicator,
We will explore several imputation approaches that differ in terms of (1) the data format used, (2) the general imputation approach used, and (3) the imputation model specification. The data format in which we impute has two possible formats: P and PP format. We examine both FCS and SMC-FCS approaches. Our imputation models differ in terms of whether we include variables for different periods or (censored) survival times or whether we also treat the (uncensored) time-to-event,
We want to note that our primary goal here is not to impute (censored) survival times but instead partially missing covariates for the estimation of survival regressions. MI is also helpful if researchers wish to estimate marginal survival distributions, for example. Still, in this article, we are not primarily concerned with imputing the dependent variable of the survival regression. Interested researchers can consult Carpenter and Kenward (2013: 177) for suitable imputation strategies or adapt the strategies proposed for partially classified categorical data in Chapter 13 of Little (2019).
FCS with Data in P Format
We begin with the imputation approaches in a P format, that is, before converting the data to a PP format. We present several approaches, some of which are taken from the literature on MI with continuous survival (cure) data (Beesley et al., 2016). They differ mainly in terms of the implemented conditioning on the (censored) time-to-event. For all imputation models presented, we also condition the other covariates. Imputation is performed using the R (R Core Team, 2019) package
Let
In the following, we will mainly discuss how to incorporate the information included in the event indicator,
The first approach (FCS P
The imputation model is: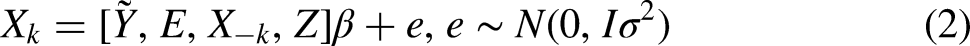
This specification with indicator variables for each possible time point is flexible, but it is possible only if the number of discrete-time points is not too large, the hazard is not expected to be near zero in some periods, and the risk sets are sufficiently large for each time point. Other specifications of the relationship between (censored) time-to-event are possible in these cases. They include, for example, linear, quadratic, cubic, or higher-order polynomials. It is also possible to use the logarithm of time (Klein, Kopp, and Rapp, 2013) or step functions for grouped periods. For our simulation, however, we do not simulate more than 15 possible time points. Thus, we use indicator variables to keep the specification as general as possible.
Another approach in the P format (FCS P
The imputation model is thus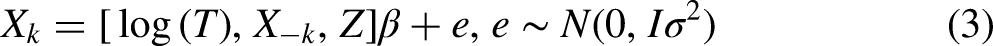
FCS with Data in PP Format
It is also possible to impute after converting the data to a PP format. Note that persons will receive varying imputed values for time-invariant covariates. As in the approaches in the P format, we always impute conditional on all other covariates. Imputation is again done with
For our first approach (FCS PP
The imputation model is thus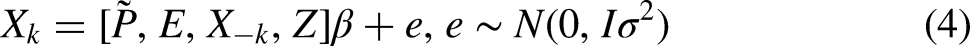
However, we expect that we will lose important information, especially for the rows belonging to the first few periods. If we do not include the last period in which a unit was observed,
Therefore, we also try imputing conditional on the last period in which a unit was observed instead of the current period. We use the set of indicator variables
The imputation model is thus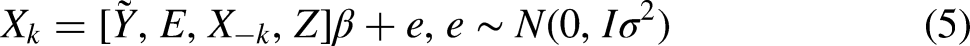
SMC-FCS with Data in PP Format
After presenting several possible FCS approximations, we now examine possible SMC-FCS approaches in this setting. SMC-FCS has shown excellent results in different simulation settings (Bartlett et al., 2015b; Beesley et al., 2016). We explore how this approach fares with a DTSAM as a substantive model. In contrast to the other imputation approaches, we now have to include the substantive model.
When we include the substantive model (see equation (1)) using the R package
A New SMC-FCS Approach: SMC-FCS DTSAM
The approaches presented in the previous sections are all approaches that can easily be implemented in already existing imputation software such as
We therefore propose a new SMC-FCS approach for imputing partially observed covariates from models which are compatible with the DTSAM specified by the user. The new SMC-FCS DTSAM approach leads to imputated covariate values, which are time-invariant for each unit, as they should be. Its derivation, which extends the derivation given by Bartlett et al. (2015b) for continuous survival endpoints, is available in the online Appendix, and it is implemented as
Simulation Study
Data-Generating Mechanisms
We now examine the performance of the imputation approaches in a series of four simulations. We first present the details (data-generating mechanisms, the introduction of missingness, performance measures) of the simulations and then discuss the results.
For all four simulations, we create five continuous covariates,
Concerning the generation of the survival times, we use the DTSAM model (see equation (1)), which will be estimated after the introduction of missingness and the subsequent imputation. This yields (truly) discrete-survival data that fulfill the assumption of proportional odds for the DTSAM (Singer and Willett, 1993: 167), the substantive model of interest. When we do not add a frailty term to the generation of survival times, there is no unobserved heterogeneity when we refit the model that we used for data generation on the simulated data. However, as several authors have noted, it is hard to specify a model without omitting any frailty variables. Ideally, one would diagnose this misspecification and change the model to address this. But this is often not possible and unobserved heterogeneity is therefore present (Allison, 1982: 83). Therefore, no unobserved heterogeneity is simulated only in one of the four simulations. In the other three simulations, we simulate successively larger amounts of unobserved heterogeneity. In all three cases, the frailty term is normally distributed, with a mean of 0 and variance that increases between simulations (0.25, 1, and 4). We add the frailty term after creating the covariates, and before simulating survival times and transforming the data set from a person format to PP format. We use
For each simulation scenario, we generate 1,000 simulated data sets with 2,000 persons each to prevent the Monte Carlo error from masking differences between methods. Data set length in PP format will vary.
Missingness
To introduce missingness in each of the simulations, we set 30% of observations missing in
After introducing different possible imputation strategies and creating data sets with missing observations, we are now able to examine the performance of the different imputation approaches.
Methods Compared and Performance Measures
We perform MI of partially observed covariates in discrete-time survival analysis using the imputation specifications described earlier in this article. For each simulation and method, we create five imputed data sets. We then compute the mean coefficient, relative bias, mean squared error (MSE), confidence interval (CI) length, and coverage for estimated DTSAM parameters across 1,000 Monte Carlo (MC) repetitions for each imputation model specification.
Performance is often evaluated only for regression point estimates and variance estimates. However, logistic regression estimates have come under scrutiny because they do not behave like linear regression estimates. Logistic regression estimates are influenced by unobserved heterogeneity—that is, omitted variables (Mood, 2009: 67). To allow comparisons between models with different covariate specifications, researchers use average marginal effects (AMEs) to interpret substantive results. An AME is the average effect of an independent variable on the predicted probability (Mood, 2009: 75)—in the case of a DTSAM, the hazard. Due to the popularity of AMEs, we also provide a comparison of the AME means for all approaches.
Simulation Results
Figure 1 displays the performance measures for the estimates of
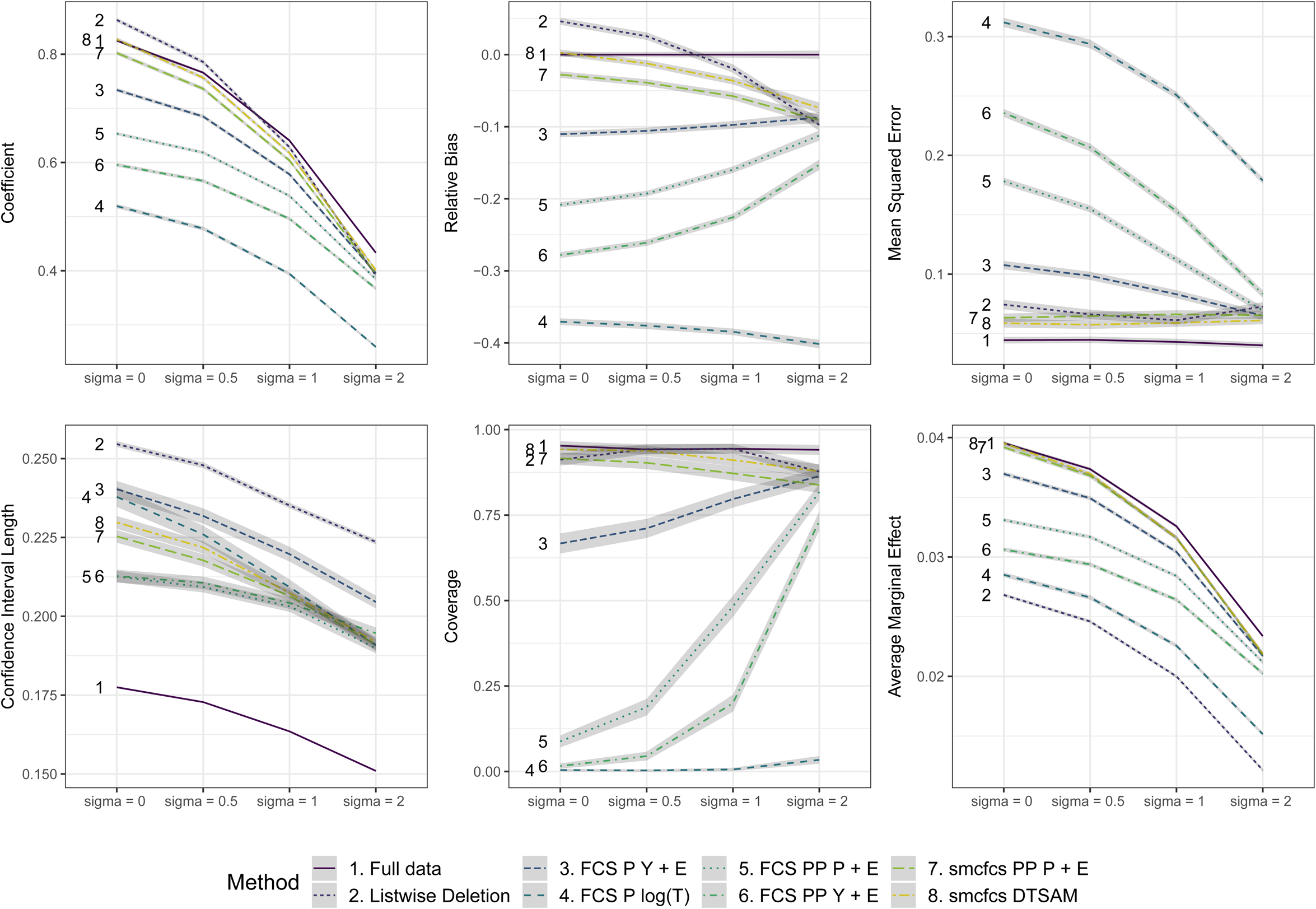
Performance measures for coefficient
We concentrate first on the imputation methods under no unobserved heterogeneity on the left. The assumption of proportional odds is thus fulfilled for the analysis model with full data and the model is correctly specified. As LD overestimates the true coefficient, and the CI length and MSE are about 50% higher than that of the full data, there is room for improvement in terms of bias and efficiency. Turning to the different imputation methods and their performance, we notice profound differences. We register the highest bias for the imputation approach FCS P
The FCS approach in P format using the information from the (censored) survival time and the event indicator (3. FCS P
We now move on to add unobserved heterogeneity to the data-generating process (variance of the frailty term
From all that we have seen so far, the new SMC-FCS DTSAM approach outperforms or performs at least as well as all other imputation models and is therefore recommended. The currently most common approach, LD, leads to high-efficiency losses and possibly high bias, and should therefore be avoided. The amount of heterogeneity that can be modeled in a DTSAM will vary strongly with available variables, but SMC-FCS DTSAM performs comparatively well.
Further Simulation Results
This article focuses on the effect of unobserved heterogeneity on the results of different imputation strategies and presents the new SMC-FCS DTSAM approach. It is also important to check whether the new method is suited to a broader range of circumstances. Therefore, we also conducted additional simulations, varying other factors such as the percentage of observations missing, the data-generating model, the amount of censoring, the correlation of covariates, and the amount of missingness in the online Appendix Figures A3–A7. Briefly summarizing the results of the five additional simulations, we notice that SMC-FCS DTSAM also performs very well under all other simulated scenarios. Several other solutions such as the two FCS imputation approaches in PP are also suited if there is no unobserved heterogeneity. FCS in-person format with
An Applied Example with the German Family Panel Pairfam
To provide an applied example under real-world data conditions, we now conduct an example analysis from the field of relationship and family research using data from the German Family Panel project called pairfam (Brüderl et al., 2017).
The 2008-launched pairfam panel (“Panel Analysis of Intimate Relationships and Family Dynamics”) is a longitudinal study for research on relationships and families in Germany. The data are collected from a nationwide random sample of the three birth cohorts 1971–1973, 1981–1983, 1991–1993, and their partners, parents, and children. For our real-world example, we used data from the data set biopart, which includes prospective and retrospective information on the anchor’s relationships, including relationships, cohabitation, and marriage history. We used the data set 8.0.0 (Brüderl et al., 2017), which includes updated information from the survey waves 1–9 (for more details on the panel, see Huinink et al., 2011).
We use a simple substantive model from the field of relationship stability research. Note that our goal here is not to estimate any causal models but rather to explore how our imputation approaches fare with a real data set. Let us assume that we are interested in the relationship between the probability that a couple
We first include six indicators 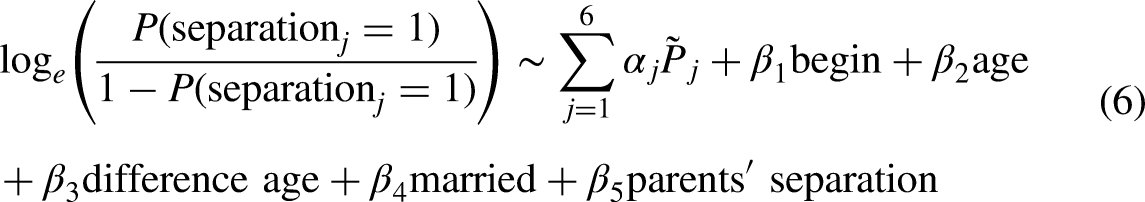
Examining the results in Figure 2, we notice that after LD, the coefficient estimate for begin has a higher variance. We also see that after imputing with our various approaches, the imputation approach FCS P
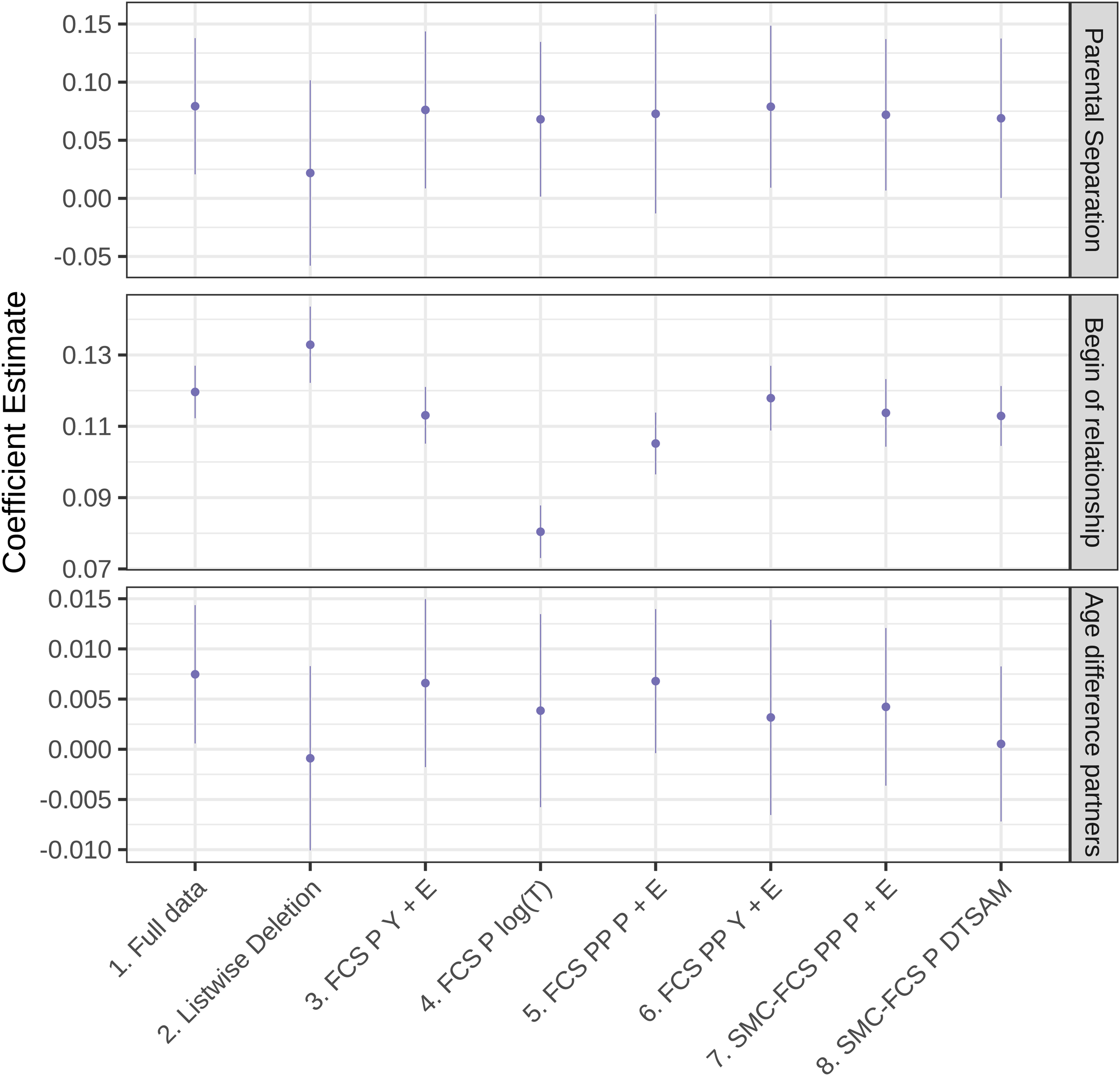
Selected estimated coefficients for a DTSAM of the separation of the first fully observed reported relationships. German Family Panel (pairfam) Data Set 8.0.0 (Brüderl et al., 2017) with MAR data introduced by the authors. For the specification of the DTSAM see equation (6). DTSAM: discrete-time survival analysis model; MAR: missing at random.
In sum, the findings resemble those found in the simulations. They show (1) that some imputation approaches—for example, treating the time-to-event variable as partly unobserved—are not advisable; (2) that imputing in PP format with FCS cannot be recommended; and (3) that SMC-FCS with a compatible imputation model performs adequately.
Discussion
Results
Like many other types of data analysis, the analysis of discrete-time survival data is often challenged by missing data in one or more covariates. Negative consequences of such missing data include efficiency losses and bias. A popular approach to circumventing these consequences is MI. However, in MI, it is crucial to include outcome information in the model for imputing partially observed covariates. Unfortunately, this is not straightforward in the case of discrete-time survival data because (1) we usually have a partially observed (left- or right-censored or both) outcome; (2) we do not have just one outcome variable, but two (i.e., event and time-to-event); and (3) we have to decide whether to impute while the data set is still in the P format or after conversion to PP format, especially if we are looking at time-invariant variables.
In this article, we have tested different approaches for imputing missing covariates in the discrete-time survival analysis model (DTSAM) setting. For this purpose, we performed four simulations that differed in the amount of unobserved heterogeneity. Some of the investigated methods are from the literature on the imputation of time-constant variables for the Cox model (van Buuren, Boshuizen, and Knook, 1999; White and Royston, 2009; Clark and Altman, 2003; Keogh and Morris, 2018; Beesley et al., 2016). We also present a new SMC-FCS approach implemented as
Our findings lead us to agree with Beesley et al. (2016) that treating censored survival times as partially unobserved and imputing other covariates depending on the multiply imputed missing survival times yields unsatisfying results in all cases when one is using existing imputation methods for imputing
Furthermore, the performance of imputation methods in PP format with FCS is disappointing. Apart from the inherent incoherence between the imputed values for different times in the same person, coverage and relative bias are unsatisfying. The new SMC-FCS DTSAM approach using a compatible imputation model performs best in our simulations with and without unobserved heterogeneity and is therefore strongly recommended.
Limitations and Future Research
An open question regarding imputation within the context of DTSAM models is how to best impute missing values for time-varying covariates (for a general overview of joint models of longitudinal and survival data, see Papageorgiou et al. 2019). Imputing with standard FCS or SMC-FCS approaches in PP format is possible but will not reflect correlations over time within individuals of the values of the time-varying variables. Research to confirm the adequate performance of simple approximations or the development of new methods in the case of time-varying covariates has yet to be undertaken.
Another open question is how to deal with the time-varying effects of time-invariant and time-varying covariates. Keogh and Morris (2018) have shown that a variation of the SMC-FCS approach also performs sufficiently well in the case of time-varying covariates. Again, an evaluation for discrete-time survival models has not yet been conducted.
Conclusions
We recommend using the newly developed SMC-FCS DTSAM approach that allows imputations that are time-invariant (do not differ between periods for a unit) and at the same time are imputed from imputation models which are compatible with the DTSAM specified by the user. The general SMC-FCS approach has already shown excellent performance in the case of included quadratic covariates and interaction effects in linear and logistic regression as well as for the imputation of covariates in Cox (cure) regressions (Bartlett et al., 2015b; Bartlett and Taylor, 2016; Beesley et al., 2016; Keogh and Morris, 2018). The new SMC-FCS DTSAM approach performed at least as well as—and usually better than—the other SMC-FCS approaches we explored in this article.
Footnotes
Author’s Note
The code for this article is available at the Open Science Framework: https://osf.io/txvey/?view˙only=04116ea9a8934b5aaf9a41059c254213. It is divided into code for the main part of the article and code for the additional simulations in the online Appendix. The data for the applied example is available as a Scientific Use File on the pairfam website: https://www.pairfam.de/en/data/. The newly developed smcfcs-dtsam method is also documented on CRAN: ![]() .
.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funded partially by the Deutsche Forschungsgemeinschaft (DFG) under project number 316901171.