Abstract
Objective
Training and validating vision transformer-based endoscopic assisted detection models for chronic atrophic gastritis (CAG) to assist endoscopists in detecting and localizing atrophic lesions.
Methods
In this retrospective study, gastroscopy images stored in the endoscopy center were collected between June 2019 and March 2023. On the basis of pathological findings, the images were manually classified into CAG and chronic nonatrophic gastritis (CNAG) using Labelme software, and the atrophic areas were further manually annotated in the CAG images. Furthermore, the anatomical structures were meticulously documented on the CNAG images. The labeled images were subsequently employed to train the Swin transformer and SSFormer for the task of detecting the anatomical structures of the stomach, CAG and atrophic lesion regions.
Results
The test results revealed that the trained Swin transformer model had an accuracy of 0.98 in recognizing the anatomical structures of the stomach (gastric sinus, stomach angle, lesser curvature, cardia fundus, and greater curvature). Moreover, the accuracy, specificity, and sensitivity of the model in recognizing the CAG and CNAG images were 0.91, 0.95, and 0.86, respectively, which were significantly superior to those of the junior endoscopists who participated in the test (p < .05). In addition, the test results suggested that the trained SSFormer model had a similar ability to segment lesions as the senior endoscopist did, with the overlap of atrophic lesion regions indicated by both exceeding 0.90.
Conclusions
In this study, a set of vision models was trained to identify not only CAG and intragastric structures but also the extent of atrophy. The application of these models to the diagnosis of CAG is expected to increase the accuracy of this process.
Keywords
Introduction
Gastric cancer is a malignant tumor originating from the epithelium of the gastric mucosa, and it accounts for more than 95% of all malignant tumors of the stomach. 1 Gastric cancer is a prevalent and deadly condition worldwide. While the incidence of tumors varies geographically, China, Japan and Korea have particularly high rates, and China bears the greatest burden of this disease. According to the latest data, China accounts for 43% of the global new cases and 48% of the global deaths caused by gastric cancer each year, making it the leading country in terms of both incidence and mortality. 2 Most gastric cancers follow a long and progressive process (Correa's cascade), in which atrophic inflammation and intestinal epithelial hyperplasia are the early stages of tissue damage, and timely discovery of these lesions and treatment of the associated risk factors (e.g. eradicating Helicobacter pylori) are effective in preventing further damage to the mucosa and reducing the risk of gastric cancer.3–5 Currently, the main clinical method for detecting chronic atrophic gastritis (CAG) is still gastroscopy, and atrophic lesions often present features such as whitish color and exposed blood vessels. However, the accuracy of conventional gastroscopy in diagnosing CAG, when pathological findings are used as a reference, is only approximately 50%.6–8
Numerous studies have recently explored the application of artificial intelligence (AI) technology to assist in clinical decision-making. A significant aspect of this involves training AI models to interpret clinical imaging data, including X-ray images, computed tomography (CT) scans, and magnetic resonance imaging (MRI) examinations.9–11 Furthermore, the continuous innovation of models has resulted in encouraging accuracy in AI recognition of complex clinical images, such as tongue images and skin lesions.12,13 Subtle color variations in these images often indicate distinct diseases, posing interpretation challenges even for clinicians. In the field of digestive endoscopy, previous studies have revealed that trained AI models can achieve high levels of accuracy in detecting diseases such as gastrointestinal polyps, tumors and ulcers; moreover, these models can assist in the quality control of endoscopes.14–17 These studies often utilize models based on the structure of convolutional neural networks (CNNs), such as DenseNet and ResNet. Notably, these CNN models fail to model long-range spatial topological relationships among gastrointestinal anatomical structures (e.g. cardia–pylorus associations) because of constrained local receptive fields and hierarchical downsampling operations (stride convolutions and pooling).18,19 The progressive degradation of feature map spatial resolution across layers compromises their ability to detect minute lesions. Furthermore, their adaptability to image rotations and scale variations relies on data augmentation strategies. In contrast, vision transformer (ViT)-based architectures use global self-attention mechanisms to establish semantic dependencies across entire endoscopic images.19,20 Their sequential processing paradigm prevents spatial information loss, increasing the detection sensitivity for subtle lesions.18,21 Critically, the inherent permutation equivariance of self-attention synergizes with the absolute spatial perception from learnable positional encodings to achieve rotation robustness, reducing the dependence on data augmentation after large-scale pretraining. 22
Although previous research has involved training AI models to assist in the detection of CAG, for example, Guimarães et al. employed a VGG16 model trained on 100 images of atrophic gastritis and 100 images of nonatrophic gastritis, achieving an accuracy of 0.929 on a test set of 70 images; their scope has been limited to the effectiveness of detecting the CAG image in its entirety, with little to no consideration of the boundary information of the atrophic lesions, and the trained models have been incapable of detecting the extent of the atrophy.23–33
Because of the potential adverse effects of CAG and the current challenges associated with endoscopic diagnosis of CAG with low accuracy and poor consistency, in this study, ViT models were trained for the classification and segmentation of CAG images.
Methods
Ethical approval
This study complied with the requirements of the Declaration of Helsinki (1975) and its 2024 revision and was approved by the Ethics Committee of Xuzhou Municipal Hospital affiliated with Xuzhou Medical University (xyyll2023059). In accordance with national legislation and institutional requirements, signed consent and preregistration were waived by the Ethics Committee of Xuzhou Municipal Hospital, which is affiliated with Xuzhou Medical University, because of the retrospective nature of this study. The reporting of this study conforms to CLAIM guidelines. 34
Gastroscopic image data acquisition and processing
The gastroscopy images utilized in this retrospective study were obtained from the Digestive Endoscopy Center of Xuzhou Municipal Hospital, which is affiliated with Xuzhou Medical University. The gastroscopy images stored in the endoscopy center between June 2019 and March 2023 were selected for this study, and the equipment models used to capture these images included the Olympus GIF-H290, GIF-H260, GIF-HQ290, GIF-XP290, GIF-TJF240, and GIF-TJF260. The specific steps of image processing are as follows: First, logging into the endoscopy system and inputting the keyword “atrophic/atrophic gastritis” were required for the initial screening of the previous examination reports. Second, the qualified images were exported, and the examination order number was recorded to search for the corresponding pathological report. Relying on the pathological results, the exported images were subjected to secondary screening, and the inclusion criteria were as follows: (1) A clear pathological examination result confirming a CAG was obtained, and the exclusion criteria were as follows: (1) images with hemorrhage, ulcers, polyps, tumors and other lesions; (2) images with large amounts of mucus, foam, bile reflux or food residue; (3) images that were too dark or too bright; (4) lens-shake or other causes of blurred images; (5) images too close to the gastric mucosa; and (6) images with special imaging or staining. The “labelme” software was used to label the images and delineate the atrophic areas in the CAG images. A graduate student was assigned to perform the labeling task. Following the labeling of each image, two senior doctors with more than 15 years of clinical experience and more than 10,000 cases of independent endoscopic operations were invited to review the images. The images could progress only to the subsequent stage of the training model when both reviewers gave their approval; images that did not receive approval from either reviewer were discarded. If the opinions of the two reviewers differed, the images were reviewed by two other doctors with the same degree of seniority, and the images were discarded when the review opinions were disapproved or when there was disagreement. The complete processing of the images is shown in Figure 1. Images of chronic nonatrophic gastritis (CNAG) were collected with the same procedure and labeled as the gastric sinus, stomach angle, lesser curvature, cardia fundus and greater curvature. All the images used in this study had any potentially identifiable patient information removed.
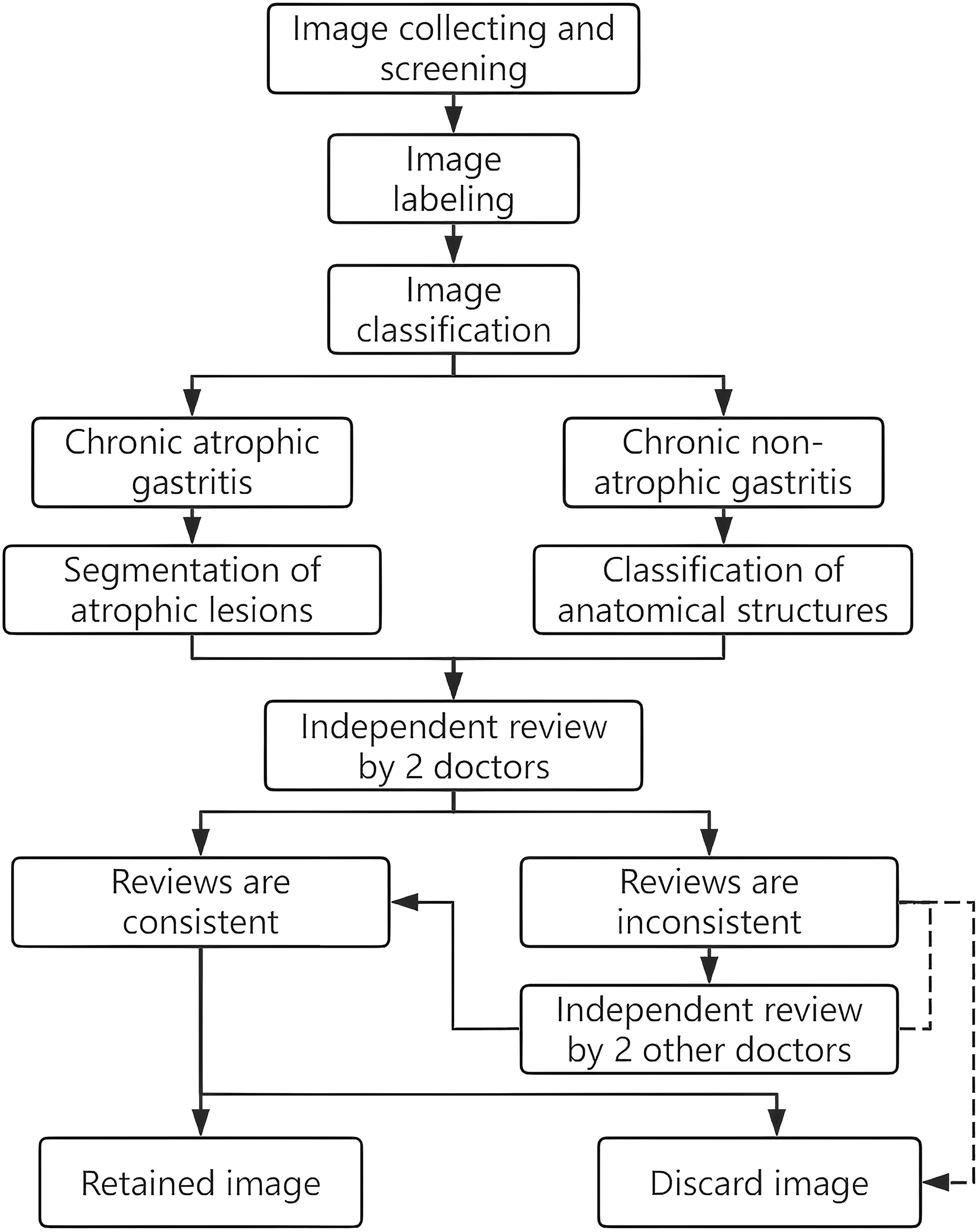
Images collection and processing flow.
Gastroscopic image grouping
In this study, three datasets were prepared; Dataset 1 consisted of 1300 gastroscopic images, including 650 CAG images confirmed by pathology (Supplemental Table S1 provides deidentified patient information) and 650 CNAG images. Dataset 2 comprised 1250 CNAG images, including 250 images each of the gastric sinus, stomach angle, lesser curvature, cardia fundus and greater curvature. Both datasets were then divided into training, validation, and testing sets at a ratio of 8:1:1, with the caveat that no images were to be shared between these data subsets. Furthermore, an additional 500 images with endoscopic diagnosis of CAG (without pathological diagnosis) were prepared for the supervised pretraining of the model (Dataset 3). The characteristics of each dataset are summarized in Supplemental Table S2.
AI model training
In this study, the Swin transformer model was trained to recognize the CAG and the anatomical structures of the stomach, whereas the SSFormer model was trained to segment the atrophic lesions.35,36 The Swin transformer is a novel ViT-based model. The model relies on a hierarchical downsampling structure analogous to CNNs to generate multiscale feature maps, enabling it to adapt well to tasks such as object detection. Through local window self-attention and shifted window mechanisms, the model significantly reduces computational costs while equipping the model with global perceptual capabilities. After its release, the model achieved outstanding performance across multiple visual tasks. SSFormer was initially designed for segmenting intestinal polyps. After it was trained on a limited dataset of intestinal polyp images, the model can accurately segment polyp lesions within complex intestinal environments and has generalizability across different data sources. This study similarly requires segmenting specific atrophic lesions within a complex gastric environment. Considering that the progressive local decoder of SSFormer improves local feature learning and suppresses background interference—capabilities crucial for addressing challenges such as gastric mucosal background interference, variable lesion morphology, and blurred boundaries—the SSFormer model was selected for training. The source codes of both models are freely available at https://github.com/microsoft/Swin-Transformer and https://github.com/Qiming-Huang/ssformer, with the models subsequently configured in the PyTorch environment after the codes are obtained. The training of the models is accelerated by an NVIDIA RTX2080Ti GPU. Prior to the commencement of the training, the input images are resized to 352 × 352 pixels. The specific settings are as follows: The models were trained for 250 epochs with a batch size of 8. An early stopping strategy (patience of 20 epochs) was employed. The models were selected on the basis of the epochs that achieved the best validation performance. The data augmentation strategies included random flipping (probability of 50%), rotation (±15°), brightness adjustment (±10%), and random cropping (ratio of 80%). The adaptive moment estimation (Adam) algorithm is utilized during the training process, with an initial learning rate of 1e-4.
Evaluating the model performance
The models were tested after complete training, and six endoscopists were invited to participate in the trial as controls. The participants were divided into three groups on the basis of their work experience: junior endoscopists (those who performed gastroscopy < 1000 times independently), middle-level endoscopists (those who performed gastroscopy between 1000 and 2000 times independently), and senior endoscopists (those who performed gastroscopy >2000 times independently). Each group contained two members (Figure 2). The performance of the trained Swin transformer model in recognizing CAG and stomach anatomy was evaluated by calculating the accuracy, sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), and F1 score. The differences between the model test results and the endoscopist test results were assessed via McNemar's test, and the differences were considered statistically significant when p < .05 (two-tailed). Moreover, the consistency between the test results was assessed by the kappa test, with p < .05 indicating consistency across both tests, and the strength of consistency was evaluated on the basis of the magnitude of the kappa value. The performance of the trained SSFormer model in segmenting atrophic lesions was evaluated by calculating the mean dice coefficient (mDice), the mean intersection over union (mIoU), and the mean absolute error. The dice coefficient and intersection over union are frequently utilized in the domain of image segmentation to evaluate the extent of overlap between the region delineated by the model and the ground truth region. In this study, the ground-truth region refers to the atrophic region that was manually delineated and reviewed, and the relevant calculation formula is as follows:

Prior endoscopy experience of six endoscopists.
Results
Ability of the model to identify CAG
The gastroscopy images contained within datasets 1 and 3 were utilized to train the Swin transformer to recognize CAG, and the model was subsequently tested after the training process (comprising 65 CAG and 65 CNAG images) was concluded. The test results were then compared with those of the clinical endoscopists (Table 1, Supplemental Table S3). The results revealed that the model achieved a higher level of accuracy in recognizing CAG images than the five endoscopists did. McNemar's test revealed no statistically significant difference between the test outcomes of the model and those of the senior endoscopists (p > .05), but it outperformed the junior endoscopists (p < .05). Kappa's test indicated high concordance between the model test results and those of the senior endoscopists (0.71 and 0.80, respectively).
Test results of endoscopists and AI in identifying CAG images.

Note: PPV: positive predictive value; NPV: negative predictive value; AI: artificial intelligence; CAG: chronic atrophic gastritis.
*Kappa test p values were all <.01.
95% confidence intervals are presented.
Because the prevalence of CAG in the general population is approximately 15%, 6 this study additionally collected gastroscopy images retained in the center during April 2023 and readjusted the proportion of image composition in the test set (including 30 CAG and 170 CNAG images) to repeat the test of the model. The results revealed that the accuracy, sensitivity, specificity, PPV, NPV, and F1 score of the model were 0.96, 0.80, 0.98, 0.89, 0.97, and 0.84, respectively (Supplemental Table S4). In addition, heatmaps were constructed utilizing gradient-weighted class activation mapping (Grad-CAM) to examine the regions that the model prioritized when the images were being processed, as shown in Figure 3. The warm regions in Figure 3 represent the regions on which the model focuses and where the atrophic lesion is most pronounced.

Model performance for recognizing chronic atrophic gastritis and corresponding heat maps.
Ability of the model to identify the gastric anatomy
The images from dataset 2 were used to train Swin transformer to recognize the anatomical structures of the stomach (gastric sinus, stomach angle, lesser curvature, cardia fundus and greater curvature), and the model was tested after the training (involving 25 images of each part). The results revealed that the model achieved an accuracy of 98.4% (123/125) in identifying the gastric anatomical structures, and the six endoscopists achieved 100% (125/125) accuracy. To assess the learning effect of the model, heatmaps were likewise used (Figure 4). The heatmaps showed that the model accurately captured the endoscopic features of each anatomical site: gastric sinus-pylorus, stomach angle-protrusion, lesser curvature-smoother planes, greater curvature-mucosal folds, cardia fundus-cardia, and fiberoptic endoscopy.

Model performance for recognizing gastric anatomical structures and corresponding heat maps.
Ability of the model to segment atrophic lesions
The images labeled with atrophic regions in dataset 1 are used to train SSFormer, and the test results are shown in Figure 5. Column A in Figure 5 represents the input images, and column B represents the initial segmentation results of the model, where white indicates the region of interest (atrophic region) and black indicates the region of noninterest (nonatrophic region). To present the segmentation results more intuitively, the initial segmentation result map is color-converted and fused with the input image via the Blend algorithm, with the original pixels being retained in the nonatrophic regions and 50% of the pixels of each of the original and color-converted maps being retained in the atrophic regions (Supplemental Figure S1). In the third row of Figure 5, the input images are characterized by blurriness and numerous reflections; however, the model exhibits satisfactory segmentation performance. The capacity of the model to segment atrophic regions was evaluated by utilizing manually annotated and reviewed images as a criterion, and the results revealed that the atrophic regions delineated by the model were highly congruent with those delineated by the human (Table 2).
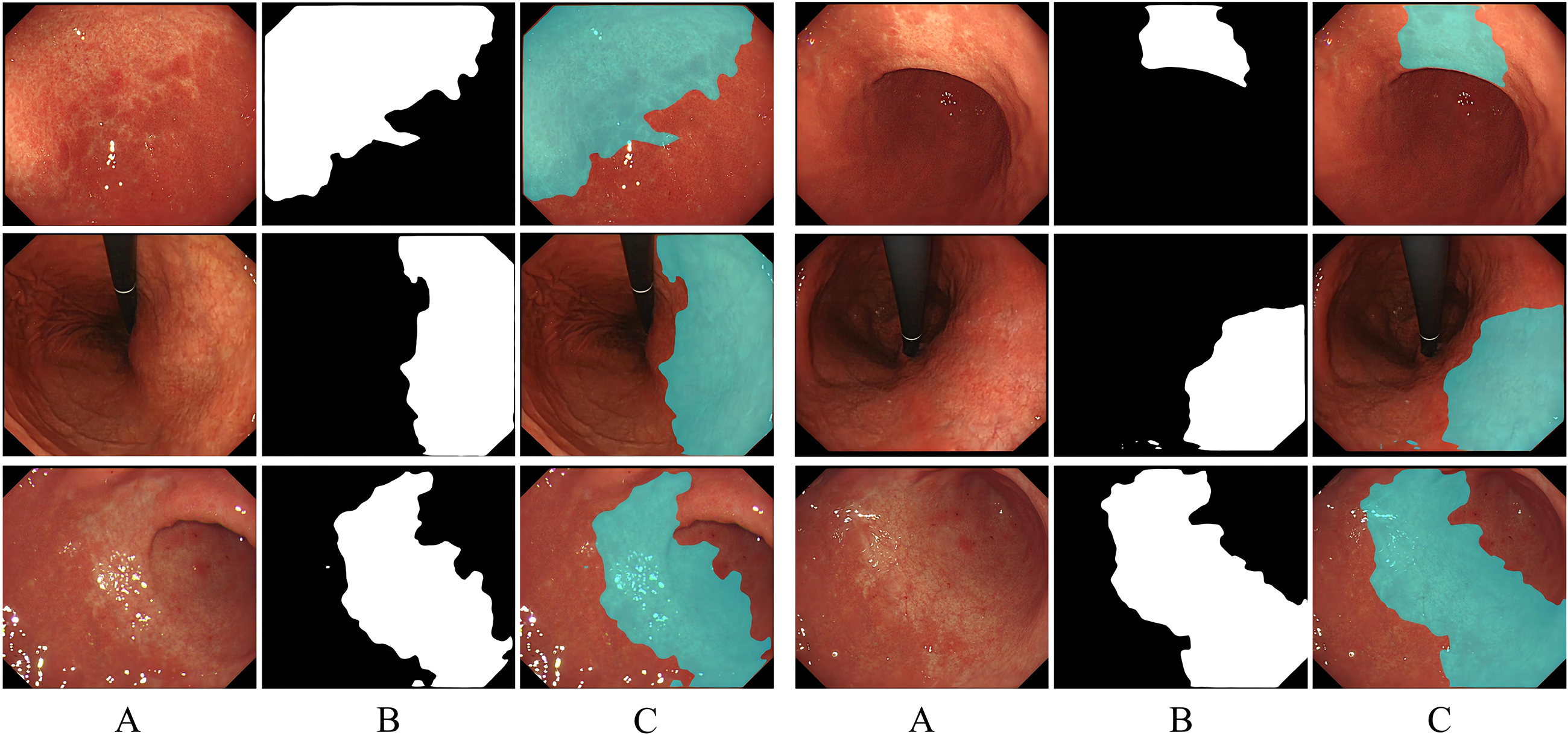
The performance of the model in segmenting the atrophic lesions.
Evaluation of the ability of the model to segment atrophic lesions with reviewed images as a criterion.

*mDice: mean dice coefficient: mIoU: mean intersection over union; MAE: mean absolute error.
Discussion
This study represents the first application of trained ViT models for both identifying CAG images and segmenting atrophic lesions. Furthermore, the trained model demonstrates the ability to recognize specific gastric anatomical structures. The test results revealed that the accuracy of the model in recognizing CAG images was greater than that of the control group of five endoscopists. Statistical analysis revealed high consistency between the model test results and those of the senior endoscopists, which indicates that the differences in diagnostic ability between endoscopists due to differing clinical experiences can be rapidly reduced with the model, enabling detection of the disease and improving the diagnosis and treatment level of young doctors. The application of the model in primary healthcare organizations can improve the quality of gastroscopy without increasing the investment of resources. Furthermore, in this study, the image segmentation model was trained to detect specific atrophic regions in the images, and the test results revealed that the segmented atrophic regions strongly overlapped with the manually delineated atrophic regions, which can help endoscopists determine the atrophic boundaries and subsequently evaluate the disease.
CAG is pivotal for the pathogenesis of intestinal-type gastric adenocarcinoma, and the Correa cascade demonstrates that CAG and intestinal metaplasia constitute the foundation for the potential development of dysplasia and gastric cancer and increase the eventual risk of gastric cancer occurrence. 37 In light of the potential risks of CAG and the challenges posed by the low diagnostic accuracy of gastroscopy, relevant guidelines suggest that the clinical diagnosis of CAG should be substantiated by pathological findings.6,7,38 However, the process of pathological sampling is associated with the risk of bleeding, particularly in patients who have recently taken antiplatelet/anticoagulant drugs. Furthermore, the imbalanced distribution of healthcare resources in some countries or regions has resulted in some endoscopy centers being required to conduct many endoscopies per day. 39 This necessitates that endoscopists perform as many examinations as possible within a constrained timeframe, reducing the time allocated for each case and increasing the risk of missed diagnoses. In some primary health care centers, the pressure from the number of examinations is usually low, but these centers often suffer from a lack of highly qualified endoscopists, which can result in missed or misdiagnosed cases. Another noteworthy aspect is that, influenced by the theoretical basis and clinical experience, the tolerance level of different endoscopists for mucosal changes varies greatly, and CAG is in the early stage of mucosal injury. Therefore, when lesion characteristics are not obvious, it is easy for different endoscopists to make opposite interpretations of the same lesion, which indicates that the consistency of diagnosis among doctors is poor. Combining the above factors may result in the clinical oversight of some CAG patients, which could account for the high incidence of gastric cancer.
Our trained CAG recognition model achieved high accuracy (0.91) in the balanced test dataset, with a sensitivity and specificity of 0.86 and 0.95, respectively. In the additional imbalanced test dataset, the model achieved higher accuracy and specificity but slightly lower sensitivity. Heatmap analysis revealed that the model focused primarily on areas of pale mucosa and exposed vasculature, which align with the characteristic manifestations of mucosal atrophy, indicating successful learning of atrophy features. When mucosal atrophy coexists with hyperemia and edema, partial atrophy features may be obscured, interfering with model recognition. Furthermore, scarring from prior biopsy sampling or endoscopic procedures (e.g. endoscopic mucosal resection and endoscopic submucosal dissection) adversely affects model recognition and segmentation. In a previous study, Zhang et al. trained a DenseNet121 model on 5470 gastric antrum images, with test results revealing an accuracy of 0.942, a sensitivity of 0.946, and a specificity of 0.940 in identifying antral atrophy. 26 Luo et al. similarly trained a ResNet50 model on more than 5000 gastroscopy images for CAG detection, with external validation revealing an accuracy of 0.854, a sensitivity of 0.870, and a specificity of 0.850. 30 Although the latter model had relatively low accuracy, its detection capability extended beyond the gastric antrum. The majority of these studies have focused exclusively on the identification of the atrophic image in its entirety; that is, whenever the model identifies the presence of atrophic lesions in the image, it determines that the image is an AG. This results in the inability of the model to detect specific atrophic areas in the image and to determine the extent of their extension. The consequence of this approach is that the model may output the same results when an image with only minor atrophic lesions and a full-map atrophic image are detected. Full-image atrophy frequently signifies severe mucosal damage. During endoscopy, both atrophic and nonatrophic mucosa are present within the same image, and when the energy of the endoscopist decreases or visual fatigue occurs, he or she is often prone to erroneous judgment of the boundaries of atrophy, which affects the assessment of the condition of the patient. To address this issue, in this study, an additional model that focuses on segmenting atrophic lesions in addition to training the model to recognize AG lesions is trained, which assists endoscopists in focusing on the boundary information of atrophic lesions. Image segmentation techniques have historically been employed more frequently in the interpretation of ultrasound, CT, and MR images, which is related to the nature of the images.40–43 In general, the color composition of ultrasound, CT, and MR images consists of white and black, whereas digestive endoscopy images are more colorful and richer in image pixel information, which renders model training more challenging. Several studies have trained AL models to segment gastric cancer lesions, and the results show that the Dice coefficients of the models are between 0.87 and 0.91.44,45 Another study trained a model to segment varicose veins in the esophagus and stomach, and the test results revealed that the mIoU of the model in segmenting varicose veins exceeded 0.80. 46 The atrophy lesion segmentation model developed in this study achieved an mDice of 0.94 on the test set, achieving high consistency with regions segmented by clinical experts. Further analysis revealed superior model performance in images exhibiting more severe atrophy. As noted previously, the characteristic features of advanced mucosal atrophy tend to be more pronounced and exhibit greater apparent differences from adjacent normal mucosa, facilitating accurate identification and segmentation.
In the Kimura–Takemoto classification, the categories C-3 and above are designated as moderate-to-severe atrophy, and studies have demonstrated that the risk of developing gastric cancer increases with increasing degrees of atrophy.47–49 In this study, the model was trained to recognize the anatomical structures of the stomach to assist in classification, and the test results revealed that the model was highly accurate at recognizing five areas, namely, the gastric sinus, stomach angle, lesser curvature, cardia fundus, and greater curvature. Previous research has demonstrated that the trained DL model can accurately identify 26 types of gastroscopy images, thus paving the way for its utilization in real-time supervision of the blind spot during gastroscopy, with the model test results revealing an accuracy of up to 90.40% in monitoring the blind spot. 50
From the analysis of images misidentified by the model, we determined that the folds of the gastric mucosa become less visible in the inflated state, which can result in the model misidentifying the greater curvature as the lesser curvature.
This study has several shortcomings. First, although the images used to train the model segmentation were subjected to strict expert review, there is no guarantee that the labeled atrophic region completely overlaps with the actual lesion area. The strategy of excising large pieces of tissue for pathological examination is needed to obtain accurate atrophic boundaries. However, this is not aligned with ethical principles. Furthermore, the appearance of the excised isolated tissues clearly differs from that of the in vivo tissues, which hinders their use for training models. Second, the models have been tested only on the image dataset. Owing to the safety considerations for gastroscopy, as well as limitations in terms of model size and hardware equipment, evaluating the performance of the models in clinical practice is not yet possible. Hence, this study focuses on how to seamlessly integrate the models into the current operating system of endoscopy centers in the future. Last, because the images collected in this study were obtained from a single endoscopy center using a specific gastroscopic model, the dataset has a limited scale and potential selection bias in terms of image inclusion. These disadvantages may compromise the detection performance of the model on images obtained by different endoscope models across multiple centers. Future efforts will focus not only on expanding the dataset within our center but also on collaborating with other endoscopy institutions to collect additional gastric endoscopic images for model optimization. After optimization, multicenter external validation will be conducted. Furthermore, we will systematically integrate images from various manufacturers and models of gastroscopy into both training and testing.
Integrating models into clinical workflows presents several critical challenges. First, as previously discussed, the model performance in dynamic video analysis and its real-time processing capability are paramount. Clinical diagnosis and treatment constitute complex and continuous activities where any oversight can directly affect patient health and safety. Consequently, both model misdiagnosis and processing delays are clinically unacceptable. Second, although the primary intent of model deployment is to assist clinicians in decision-making, the process of utilizing these models may inadvertently divert the attention of the physician. More significantly, excessive reliance on these models risks weakening the diagnostic skills and clinical judgment of clinicians over time. Last, the ethical implications of employing models in clinical settings warrant careful consideration.
Conclusions
This study developed a set of ViT-based models for computer-aided detection of CAG, whose clinical application is expected to increase the diagnostic accuracy of CAG under gastroscopic conditions.
Supplemental Material
sj-jpg-1-sci-10.1177_00368504251381972 - Supplemental material for Constructing and validating vision transformer-based assisted detection models for atrophic gastritis: A retrospective study
Supplemental material, sj-jpg-1-sci-10.1177_00368504251381972 for Constructing and validating vision transformer-based assisted detection models for atrophic gastritis: A retrospective study by Hu Chen, Shiyu Liu, Yanzi Miao, Xin Yang, Tao Li, Chuannan Wu, ZhenTao Li, Yahui Guo, Sijin Yu and Guangxia Chen in Science Progress
Supplemental Material
sj-docx-2-sci-10.1177_00368504251381972 - Supplemental material for Constructing and validating vision transformer-based assisted detection models for atrophic gastritis: A retrospective study
Supplemental material, sj-docx-2-sci-10.1177_00368504251381972 for Constructing and validating vision transformer-based assisted detection models for atrophic gastritis: A retrospective study by Hu Chen, Shiyu Liu, Yanzi Miao, Xin Yang, Tao Li, Chuannan Wu, ZhenTao Li, Yahui Guo, Sijin Yu and Guangxia Chen in Science Progress
Footnotes
Acknowledgments
We sincerely thank the Digestive Endoscopy Center of Xuzhou Municipal Hospital affiliated with Xuzhou Medical University for providing support for this study and thank the endoscopists who participated in this study but were not included in the list of authors.
Author contributions
Conceptualization: HC, SL, and GC; methodology: HC and YM; software: YM and HC; formal analysis: HC, XY, and SL; resources: GC and SL; data curation: HC, TL, CW, ZL, YG, and SY; writing—original draft preparation: HC; writing—review and editing: SL and GC; supervision: GC; and funding acquisition: GC and SL. All the authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Xuzhou Medical Leading Talent Training Program (Grant No. XWRCHT20210025), Xuzhou Key Research and Development Program (Grant No. KC22095), and Xuzhou Municipal Health Commission Youth Innovation and Technology Program (Grant No. XWKYHT20220073).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All the data are accessible and available from the corresponding author upon request.
Supplemental material
Supplemental material for this article is available online.

