Abstract
Objective:
Illicium verum is an endemic plant of southern China, which is the primary country for its production. Trans-anethole (t-anethole) is the key component of the volatile aromatic essence in I. verum, and it has therapeutic effects such as anti-cancer and anti-diabetes. However, its biosynthetic pathway in I. verum is rarely reported.
Methods:
In the present study, we cloned and expressed the cDNA encoding t-anol/isoeugenol synthase (IvAIS1) in Escherichia coli. The characteristics of the IvAIS1 were determined and its gene expression in different tissues was measured by real-time polymerase chain reaction.
Results:
The IvAIS1 protein is 76% identical to Schisandra chinensis isoeugenol synthase, and the two proteins were clustered closely together in the clade of IGS and EGS. IvAIS1 exhibits NADPH-dependent enzyme activity and dual product specificity, and it converts coumaryl acetate and coniferyl acetate to t-anol (the precursor of t-anethole) and isoeugenol, respectively. The Km values for coniferyl acetate and coumaryl acetate were 438.4 ± 44.3 μM and 480.30 ± 86.61 μM, respectively. The catalytic efficiency of IvAIS1 for coniferyl acetate was found to be higher than that for coumaryl acetate. The gene expression profiles showed that IvAIS1 accumulated in the roots, leaves, and fruits, but the levels were relatively low in the stems and flowers of I. verum.
Conclusions:
This study showed a putative t-anol/isoeugenol synthase responsible for converting coumaryl acetate to t-anol in I. verum. It expands our current knowledge of the enzymes involved in t-anethole biosynthesis.
Introduction
Illicium verum, an ancient member of the most basal angiosperm lineage, is a characteristic endemic plant of Guangxi, China, which supplies almost half of the total world consumption of this plant. 1 The fruit of I. verum has a strong fragrance, which is not only used as a spice, but it is also a traditional Chinese medicine. It possesses pharmacological effects and has antioxidant and anti-inflammatory properties, and it can be used to treat vomiting, abdominal pains, and lumbago. It is also used to dispel colds and stimulate nerves.2–4 The volatile oil from the fruits of I. verum contributes to its aromatic smell and pharmacological activities. 5 The Chinese Pharmacopoeia (2020 edition) stipulates that the volatile oil in the I. verum fruit should comprise a minimum of 4.0% (mL·g−1) of the total oil content. The Chinese national standard GB/T 7652-2016 stipulates that the total amount of volatile oil in I. verum fruit must exceed 7.5%. Trans-anethole (t-anethole) is the most important bioactive compound in I. verum volatile oil, accounting for 50–80% of the total volatile matter. 6 Its concentration determines the difference in flavor and the quality of this medicinal herb. It has emerged as a potential anti-cancer agent and has demonstrated effectiveness against breast, prostate, lung, and colorectal cancers by its ability to modulate critical signaling pathways including NF-κB, PI3 K/Akt/mTOR, and caspases. 7 Additionally, t-anethole can protect the gonads from testicular ischemia reperfusion injury, potentially through the transcription factors, STAT3 and NF-κB. 8
Structurally, t-anethole belongs to the phenylpropene group of compounds, and they are derived from phenylalanine by modification of the benzene ring (C6) and reduction of the propyl side chain (C3). In most plants, the phenylpropene biosynthesis pathway starts with the upstream products of the glycolytic and shikimate pathways. 9 Following the sequential catalytic action of phenylalanine ammonia lyase, cinnamic acid 4-hydroxylase, and 4-coumarate-CoA ligase, phenylalanine is converted successively into cinnamic acid, p-hydroxycinnamic acid, and p-coumaryl-CoA. 10 Subsequently, 4-coumaric-CoA is converted to p-coumaryl acetate by the catalysis of downstream enzymes cinnamyl-CoA reductase, cinnamyl alcohol dehydrogenase, and coniferyl alcohol acyltransferase, respectively. 11 This monolignol acetate is reduced by the NADPH-dependent reductases, to form an allyl-phenylpropene (with a double bond between C8 and C9) or propenyl-phenylpropene (with a double bond between C7 and C8). 12 Estragole and t-anethole are then produced after further methylations of the benzene ring, respectively. 13
The NADPH-dependent aromatic alcohol reductases in the above catalysis reactions belong to the PIP reductase family (named after the three initially identified enzymes, pinoresinol-lariciresinol reductase, isoflavone reductase, and phenylcoumaran benzylic ether reductase). 14 Proteins belonging to the PIP reductase family possess a NADPH-binding domain, GXXGXXG, in their N-termini. In addition, they contain a highly conserved lysine residue indispensable for enzymatic activity and an active site motif, which serves as a crucial catalytic residue. 15 The reaction mechanism of these NADPH-dependent reductases involves the reductive cleavage of acetate from the propenyl side chain of the monolignol acetate substrate by the formation of a quinone methide intermediate. 16 Its substrate specificity analysis revealed that a hydroxyl group of benzene ring in the substrates was required in order to form the quinone methide intermediate, regardless of whether the propenyl side chain was changed to a propionate, butyrate, or benzoate group. 17 In Clarkia breweri flowers, the NADPH-dependent reductases, eugenol synthase (CbEGS1) and isoeugenol synthase (CbIGS1) were found to exhibit high sequence similarity, with only two amino acid differences found. 18 They are Class I members in the EGS/IGS subfamily of the PIP family, together with basil (Ocimum basilicum) EGS 19 and petunia (Petunia hybrida) IGS. 18 However, the CbEGS2 belongs to the Class II subfamily, and it exhibits a reduced substrate specificity when compared to those in Class I. FaEGS2 from garden strawberry (Fragaria × ananassa) catalyzes the formation of eugenol and isoeugenol, but the latter has a lower catalytic efficiency. 20 The anise t-anol/isoeugenol synthase 1 (AIS1) can biosynthesize t-anol and isoeugenol from coumaryl acetate and coniferyl acetate, respectively. 21
The biosynthetic pathway of phenylpropene volatiles has been studied in several herbs and fruits, and many biosynthetic genes have been isolated and characterized.22–24 Although the fruit of I. verum is considered the richest source of t-anethole, the biological and molecular mechanisms involved in its biosynthesis, accumulation, and regulation have not been extensively investigated. To understand the potential key catalytic steps of t-anethole biosynthesis, we generated transcriptomic data from a mixed tissue sample of I. verum. Candidate unigenes homologous to PIP reductase were identified, cloned, and expressed. The catalysis characteristics of the putative PIP reductase were determined. The results lay the foundation for further elucidating the formation mechanism of the characteristic flavors and the biosynthetic pathway of t-anethole in I. verum.
Material and methods
RNA extraction, cDNA synthesis, and transcriptomic sequencing
The roots, stems, leaves, and flowers of I. verum that were collected and stored in the Guangxi Botanical Garden of Medicinal Plants (22°85′73″N, 108°36′72″E, Nanning, China). Total RNA was isolated using the RNAprep pure plant plus kit by following the manufacturer's instructions (Code No. DP441, Tiangen, Beijing, China). The integrity of RNA obtained was assessed by electrophoresis and quantified using a spectrophotometer. High-quality RNA was used to prepare cDNA with the oligo dT primer by following the manufacturer's protocol (Code No. 6210A, Takara, Dalian, China). Then, a sequencing library was constructed and this was sequenced by the PacBio Nanopore platform (Novo Zhiyuan, Beijing, China).
Gene cloning and expression plasmid construction
One set of primer for the CDS of IvAIS1 was designed on the annotation sequence of the transcriptomic analysis (Supplementary Table S1). The cDNA was used as a template for in vitro amplification by polymerase chain reaction (PCR). The purified products were cloned into pET22b(+) and pMAL-p5x vectors by following the manufacturer's protocol (Code No. BL1046B, Biosharp, Hefei, China). The recombinant plasmids were transformed in the Escherichia coli strain, DH5α. The positive clones were selected by using LB plates, and these were confirmed by sequence analysis (Sangon Biotech, Shanghai, China).
Sequence analysis of IvAIS1 protein
ExPASy (https://web.expasy.org/protparam/) was used to calculate the number of amino acids, molecular weights (MWs), theoretical isoelectric points (pIs), instability indexes, aliphatic indexes, and grand average of hydropathicity (GRAVY) of the proteins. Wolf PSORT (https://wolfpsort.hgc.jp/) and Plant-mPLoc (http://www.csbio.sjtu.edu.cn/cgi-bin/PlantmPLoc.cgi) were employed to predict their subcellular localization, and SOPMA (https://npsa-pbil.ibcp.fr/cgi-bin/secpred_sopma.pl) was applied to predict their secondary structures. The conserved motif composition was assessed by using the MEME 5.5.5 online program (http://memesuite. org/tools/meme).
Homology modeling and molecular docking
The X-ray crystalline structure of EGS from C. breweri (CbEGS1, SMTL ID: 3c1o.1) was used as a template, 18 and the structure of IvAIS1 protein was homologously modeled using the SWISS-MODEL server (https://swiss model.expasy.org/). The ligand structures of coumaryl acetate, 25 coniferyl acetate, 26 and NADPH 27 were obtained from PubChem (https://pubchem.ncbi.nlm.nih.gov/). AutoDock v4.2 (Scripp Research Institute, CA, USA) was used to explore the molecular docking of IvAIS1 with substrates and NADPH as previously report. 28 The IvAIS1 and ligands were modified by adding hydrogen atoms, incorporating nonpolar hydrogen and calculating Gasteiger charges. Subsequently, molecular docking of the IvAIS1 proteins and ligands was carried out by using AutoDock with the exhaustiveness setting at 10. The best conformation was selected based on the minimal binding energies. The ligand-protein interactions (hydrogen bonds and hydrophobic) were analyzed and visualized by using PyMol software packages (Version 4.60, DeLano Scientific LLC, CA, USA).
Sequence alignment and phylogenetic analysis
The protein sequences of different species were obtained by performing BLAST comparisons in the NCBI database (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The PIP family amino acid sequences of basal angiosperms (Amborellales, Nymphaeales, and Austrobaileyales) and some model plants (including Arabidopsis thaliana, Zea mays, and Populus trichocarpa) were compared by using the DNAMAN software package (V6 version, Lynnon Biosoft, USA). The neighbor-joining method was used to construct the phylogenetic tree by using MEGA7.0 with the bootstrap value set at 1000 replicates. 29
Protein expression and purification
Recombinant plasmids of pMAL-p5x and pET22b harboring IvAIS1 were transformed into the E. coli strain BL21 (DE3) competent cells, respectively. Expression of the combined proteins was performed as described previously. 30 The culture was induced with isopropyl 0.1 mM β-D-1-thiogalactopyranoside (IPTG) for 12 h at 20 °C. E. coli was harvested by centrifugation at 8000 rpm for 20 min at 4 °C. The cell pellets obtained were suspended in Tris-HCl buffer (20 mM, pH7.4, containing 200 mM NaCl, 1 mM ethylene diamine tetraacetic acid, 1 mM DL-dithiothreitol, and 1 mM phenyl methane sulfonyl fluoride. Then, the cells were sonicated, and the lysates were centrifuged at 12,000 rpm for 10 min at 4 °C. IvAIS protein from cytoplasmic extract was purified using a dextrin affinity column by following the manufacturer's protocol (Code No. SA77025, Smart-Lifesciences, Changzhou, China). The eluted proteins were analyzed using 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and western blotting. MBP-tag mouse monoclonal antibody (Code No. M1001, Lablead, Beijing, China), peroxidase AffiniPure goat anti-mouse IgG (Code No. 115-035-003, Jackson ImmunoResearch, West Grove, PA, USA), and hypersensitive ECL western blotting substrate (Code No. 17046, Zenbio, Chengdu, China) were used for western blotting. The protein concentration was determined with a Bradford protein assay kit (Code No. BL521A, Biosharp, Hefei, China).
Measurement of enzyme activity
The catalytic activity of IvAIS1 was determined as previously described. 31 Enzyme activity assays were performed in 200 μL MES-KOH buffer (50 mM, pH 6.5) with 20 μg purified IvAIS1 protein, 1 mM NADPH, and 1 mM substrate. Reactions lacking co-substrates NADPH and those containing heat-denatured IvAIS1 served as controls. The assay mixture was incubated at 30 °C with shaking for 30 min. Then, it was extracted three times with ethyl acetate (200 μL for each time). The organic phase was filtered through a 0.22 μm microporous filter, and it was supplemented with 0.1 mM linalool as internal standard. The samples were then measured by gas chromatography–mass spectrometry (GC-MS). GC-MS analysis was performed with an Agilent 8090 GC & 5977B GC/MSD System (Agilent Technologies, CA, USA) equipped with a 30 m × 0.25 mm i.d. capillary column coated with a 0.25 μm film of HP-5MS (Agilent Technologies, CA, USA). Samples (2 μL) were injected at 270 °C in split mode with a split ratio of 4:1 and flow rate of 4.8 mL·min−1. The carrier gas was ultrapure helium (99.999%) used in a constant-flow mode (1.2 mL·min−1). The oven temperature was held at 70 °C for 2 min and then programmed at 10 °C·min−1 to 270 °C, which was held isothermally for 2 min. The detector operated in the electron-impact mode at 70 eV using single ion monitoring at 3.1 scan·s−1 (dwell time 323.43 ms for each ion). The electron multiplier potential was set to 1224 V. The transfer line, source, and quadrupole temperatures used were 280, 230, and 150 °C, respectively.
Biochemical characterization of recombinant IvAIS1
The specificity toward the reaction substrate was analyzed using coumaryl acetate and coniferyl acetate. The optimum temperature was determined by analyzing the IvAIS1 activity at different incubation temperatures, ranging from 25 to 60 °C. The optimal pH was determined by measuring the enzyme activity at pH values ranging from 4 to 9. The kinetic parameters of Km and Vmax of the enzyme were determined using different substrate concentrations and analyzed by using GraphPad Prism 9 (GraphPad Software Inc., San Diego, CA, USA).
IvIGS expression analysis in different tissues
Expression levels of IvAIS1 in the tissues of I. verum were determined by quantitative real-time PCR. It was performed by using the Roche real-time PCR detection system (Roche, lightcycler96, Basel, Switzerland) and Takara TB GREEN Premix Ex Taq™ (Code No. RR420Q, Takara, Dalian, China). The RNA samples from the roots, stems, leaves, and flowers were extracted, and first-strand cDNAs were synthesized as described above. Specific primers (Supplementary Table S1) were designed accordingly based on the IvAIS1 sequence by using Primer Premier 5 software (Premier, Canada). Beta-actin was employed as reference gene in this study. Each qPCR reaction was conducted in a volume of 10 μL. The thermal cycling conditions consisted of an initial step at 95 °C for 30 s. This was followed by 40 cycles of two-step amplification at 95 °C for 5 s and 60 °C for 30 s, and then, a denaturation step at 95 °C for 5 s, 65 °C for 60 s, and 97 °C for 1 s. Finally, there was a cooling step at 50 °C for 30 s. The relative gene expression levels were analyzed using the 2−ΔΔCt method. 32
GC was used to quantify the concentrations of t-anol and t-anethole in the various tissues of I. verum plant. Fresh samples (0.1 g) were grinded homogeneously with 5.0 mL of ethyl acetate. The mixture was extracted by ultrasonic at room temperature for 0.5 h, followed by centrifugation at 12,000 r·min−1 for 10 min. The supernatants obtained were filtered through a 0.22 μm microporous filter, and these were passed through an Agilent 8090 GC system (Agilent Technologies, CA, USA). Separation of the analytes was performed on an HP-5 capillary column (30 m × 0.32 mm i.d., with a 0.25 µm film thickness, Agilent Technologies, CA, USA). Helium (99.999%) was employed as the carrier gas at a constant linear velocity of 1 mL min–1. The injection temperature was kept constant at 250 °C. The injection volume was 2 µL. The oven temperature was regulated as follows: an initial temperature of 50 °C (held for 2 min), and then, it was elevated to 270 °C at a rate of 6.5 °C min−1 and held for 1 min. The detector temperature was fixed at 250 °C.
Statistical analysis
Enzyme activities, biochemical characterization, gene expression data, and content determinations of t-anol and t-anethole were performed in triplicates. Excel 2023 (Microsoft, USA) and SPSS 25 statistical software (IBM, USA) were used to assess the obtained. The means were compared using one-way ANOVA, and P ≤ 0.05 was considered to be statistically significant.
Results
Cloning and expression of IvAIS1
Using I. verum leaf cDNA as the template, a PCR fragment of 957 bp was amplified by using the specific IvAIS-F and IvAIS-R primers (Supplementary Table S1). For protein expression and function validation, the sequence was constructed in pET22b(+) and pMAL-p5x vectors (Figure 1(a)), subsequently named pET22b-IvAIS1 and pMAL-p5x-IvAIS1, respectively. SDS-PAGE and western blotting analysis showed that the target protein was induced, and then, it was expressed by IPTG, and it could then be purified by using the MBP tag (Figure 1(b) to (d)). The purified protein expressed by E. coli contained the pMAL-p5x-IvAIS1, and it was chosen for its better solubility in the following studies.
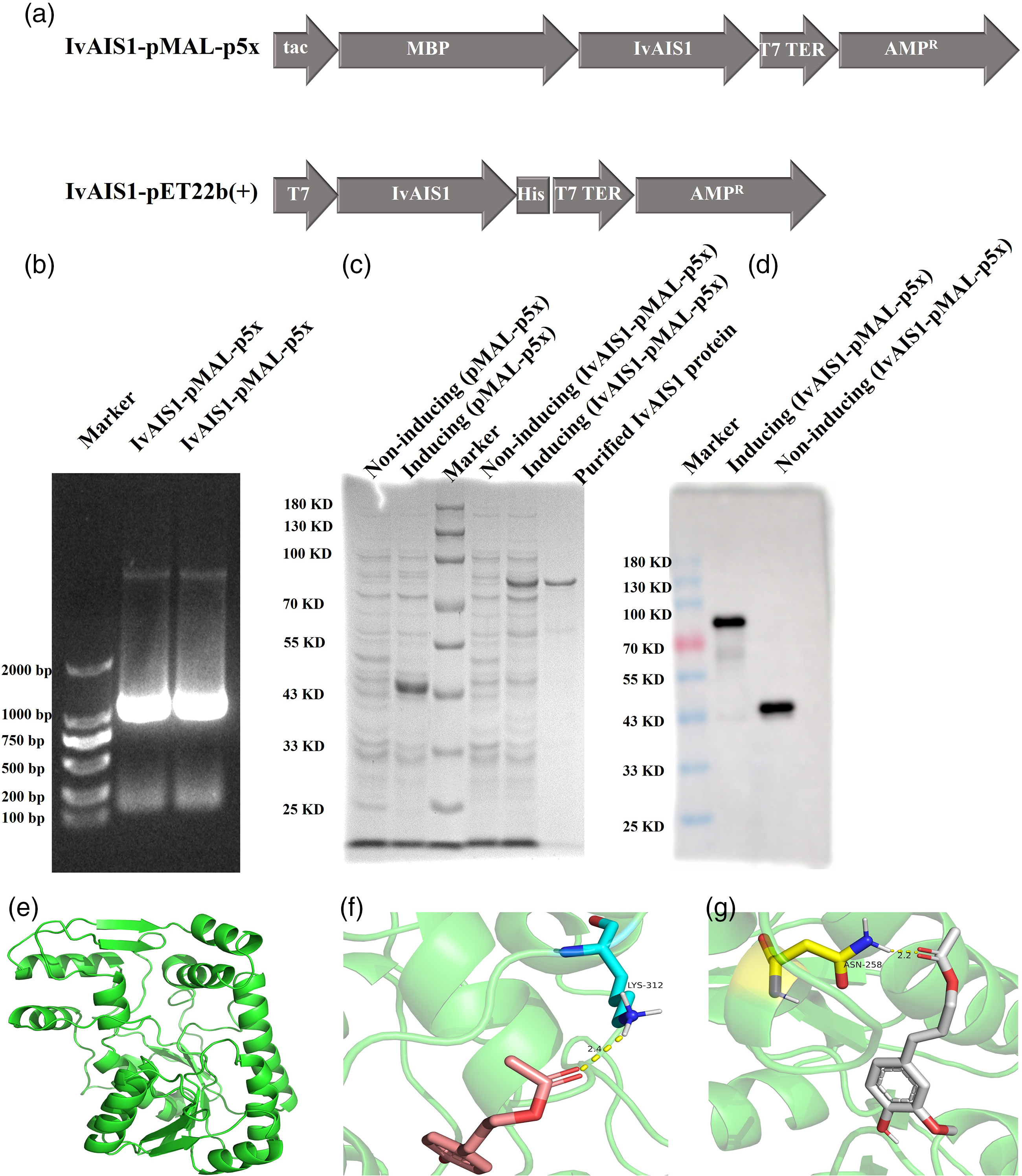
The recombinant plasmid construction, protein expression, and enzyme affinity analysis of IvAIS1. (a) The IvAIS1 was constructed with the pMAL-p5x and pET22b(+) vectors, respectively. (b) The IvAIS1 in plasmid, pMAL-p5x, was verified by PCR and gel electrophoresis. SDS-PAGE (c) and western blotting (d) were used to identify the recombinant IvAIS1 protein. The IvAIS1 protein was subjected to homolog modeling (e) and docking with coumaryl acetate (f) and coniferyl acetate (g).
Sequence analysis of the deduced IvAIS1 protein
The IvAIS1 protein consists of 317 amino acids and a MW of 35.39 kDa. The theoretical pI was found to be acidic at 5.59. According to the instability index of 35.26, the IvAIS1 is a stable protein. The aliphatic index and grand average of hydropathicity of IvAIS1 was determined to be 100.25 and −0.018, respectively. Furthermore, as the subcellular localization predicted, the IvAIS1 protein was most likely to be located in the cytoplasm. The alpha helix, extended stand, beta turn, and random coil made up 34.38, 17.35, 6.9, and 41.32% of the 317 amino acids sequence, respectively. Based on the protein sequence identity, an X-ray crystalline structure of EGS from C. breweri (CbEGS1, SMTL ID: 3c1o.1) was applied as template for homology modeling of IvAIS1. The protein structure of IvAIS1 was then generated using the SWISS-MODEL (Figure 1(e)). The stereo-chemical quality of the 3D structure of IvAIS1 was validated using Ramachandran plot analysis. The proportions of amino acid residues within both the favored and permitted regions were found to be 99.6% and ≥90% is considered to be a good quality model (Supplementary Figure S1). The QMQE score of the predicted model was 0.87, and the QMEAND value was 0.83 ± 0.05. Further, the validated model of IvAIS1 was capable of docking with coumaryl acetate + NADPH and coniferyl acetate + NADPH, respectively (Figure 1(f) and (g)). The docking scores was estimated to be −3.15 and −4.3 kcal·mol−1 for coumaryl acetate and coniferyl acetate, respectively. The coumaryl acetate + NADPH was shown to dock in the same binding pocket as that of coniferyl acetate + NADPH. Asn-258 formed hydrogen bonds with all of the small molecules. And the distances between Asn-258 and coumaryl acetate + NADPH were 2.68 Å and 2.10 Å, while those between Asn-258 and coniferyl acetate + NADPH were 2.96 Å and 2.10 Å. The interactions between IvAIS1 and coumaryl acetate were found to be weaker than that with coniferyl acetate.
Sequence alignment and phylogenetic analysis
To characterize the features of IvAIS1, a multiple alignment analysis was performed based on the amino acid sequences of the putative IvAIS1 and other plant PIP family proteins, including PCBER, LAR, IFR, IGS, EGS, and PLR proteins (Supplementary Table S2). The alignment analysis results revealed that IvAIS1 shared relatively high identities to these IGS proteins, ranging from 52% for C. breweri CbIGS1 to 76% for Schisandra chinensis ScIGS1 (Figure 2(a)). All these IGS proteins shared the NADPH-binding domain, GXXGXXG, in their N-termini. There are conserved lysine residues that determine specificity. There is also substrate binding at the C-terminal region and an upstream that contributes to the active site. In IvAIS1, these were GATGYIG, Lys-134, and Val-87, respectively. The phylogenetic tree with various plant PIP family proteins is shown in which indicates that IvAIS1 exist in the clade of IGS and EGS (Figure 2(b)). The position of these genes within the phylogenetic tree is compatible with where they evolved. Therefore, the IvAIS1 and IGS of S. chinensis clustered closely together in the same branch due to their close evolutionary relationship.

Sequence alignment of IvAIS1 with EGS and IGS obtained from other species (a) and the phylogenetic tree of the PIP family (b). The NADPH-binding sites and substrate specificity for amino acids are indicated. The phylogenetic tree was constructed by using the maximum likelihood method and applying 1000 bootstrap replicates. The GenBank accession numbers of the PIP family proteins in this tree are listed in Supplementary Table S2.
Identification of IvAIS1
To investigate the catalytic function of IvAIS1 protein, coumaryl acetate and coniferyl acetate were used as substrates in reactions. Coumaryl acetate could be catalyzed by purified IvAIS1 protein to form t-anol, while coniferyl acetate was converted to isoeugenol (Figure 3(a) and (b)). Both the two reaction systems used the cofactor, NADPH. Instead of using purified IvAIS1, the spent mediums of E. coli bearing empty or recombinant vectors were used in this study. Chavicol could be detected as a byproduct of coumaryl acetate in the reaction systems, which used either the spent mediums with expressed IvAIS1 or not (Supplementary Figure S2). Isoeugenol and eugenol were detected simultaneously in as products catalyzed by the medium containing expressed IvAIS1. In addition, the eugenol was biosynthesized by the medium without IvAIS1 (Supplementary Figure S2). These complicated products were produced not just by catalysis with IvAIS1 but also by the inherent protein activity of E. coli. However, the results confirmed that the IvAIS1 possessed t-anol/isoeugenol synthase enzymatic activity, which was identified from the phylogenetic tree.

GC-MS analysis of the reaction products of recombinant IvAIS1 protein with coumaryl acetate (a) and coniferyl acetate (b), respectively. The substrates and products were compared to reference standards including t-anol, coumaryl acetate, isoeugenol, and coniferyl acetate. Reaction systems with and without heat-denatured samples of IvAIS1 were included as negative controls.
Enzyme activity measurements
To determine the enzyme kinetic parameters of IvAIS1, the optimal pH and temperature for the catalysis reactions of coumaryl acetate and coniferyl acetate were determined (Supplementary Table S3). Both the substrates were most suitable for the reaction to occur at pH 7.0 (Figure 4(a) and (b)). The optimal temperature for the catalytic of coumaryl acetate was 40 °C, while it was 45 °C for coniferyl acetate (Figure 4(c) and (d)). With the optimal reaction systems, the Km value for coumaryl acetate was determined to be 480.30 ± 86.61 µM, Kcat was 0.33 ± 0.02 s−1, and the maximum reaction rate Vmax was 4.05 ± 0.23 nmol·s−1·mg protein−1. The Km, Kcat, and Vmax values for coniferyl acetate were 438.40 ± 44.29 µM, 0.67 ± 0.03 s−1, and 8.18 ± 0.30 nmol·s−1·mg protein−1, respectively (Figure 4(e) and (f)). These were 1.10-fold, 0.49-fold, and 0.50-fold as the values for coniferyl acetate, respectively. Similarly, the Km, Kcat, and Vmax values of NADPH (coumaryl acetate) were 0.80-fold, 0.44-fold, and 0.44-fold of NADPH (coniferyl acetate), respectively (Figure 4(g) and (h); Supplementary Table S3).

The t-anol/isoeugenol synthase activity of the recombinant IvAIS1 protein. Determination of the optimal pH and temperature for IvAIS1 activity with coumaryl acetate (a and c) and coniferyl acetate (b and d), respectively. The steady-state kinetic measurements of IvAIS1 using coumaryl acetate (e), coniferyl acetate (f), and NADPH (g and h) with varying substrate concentrations.
Gene expression profiles of IvAIS1
The relative expression levels of IvAIS1 in various tissues of I. verum were quantified by RT-PCR analysis (Figure 5(a)) using the primer sequences shown in Supplementary Table S1. IvAIS1 was expressed in the roots, leaves, flowers, stems, and fruits. The highest relative expression level of IvAIS1 presented in the roots, which was significantly greater than the other tissues. The levels from high to low were leaves, fruits, flowers, and stems. In addition, GC was used to quantify the concentrations of t-anol and t-anethole in various tissues. The overall concentration of t-anethole was about 35.7 times higher than that of t-anol. The concentration of t-anol was highest in the stems, followed by the fruits and leaves, with minimal levels in the roots. t-anol and t-anethole were rarely detected in the flowers (Figure 5(b)). t-anethole exhibited significant levels in both the fruits and stems, followed by the leaves, while being present only minimally in the roots (Figure 5(c)).

Relative expression levels of the IvAIS1 (a), t-anol (b), and t-anethole (c) in various tissues of I. verum. The results represent the average of triplicate determinations. The letters above the error bars indicate significant differences (P < 0.05) as assessed by one-way ANOVA.
Discussion
Phenylpropenes contribute to the flavor and aroma of many aromatic spices and herbs. t-anethole, a phenylpropene that is extensively utilized in the flavoring, fragrance and pharmaceutical industries, is abundant in the essential oils obtained from I. verum. However, the enzyme involved in the biosynthetic pathways of t-anethole has not yet been identified in I. verum. The NADPH-dependent PIP reductases catalyze the ester forms of monolignol coniferyl and coumaryl alcolols to produce eugenol and t-anol, respectively. Therefore, in this study, we cloned a gene consisting of 957 bp from the I. verum, and it was named IvAIS1. The deduced protein possesses the typical sequence characteristics of the PIP family. It has a NADPH-binding domain, GATGYIG, a conserved lysine residue Lys-134 and an active site at the Val-87 position. It appears that the three sites are critical for the IvAIS1 and their substrates (coumaryl acetate and coniferyl acetate) to interact, based on the in silico docking analysis. Despite the valine residue being positioned at different positions in the amino acid sequences among the different IGS/EGS proteins (Val-84 of CbIGS1, 18 Val-87 of PhIGS, 30 Val-86 of AsIGS 12 ), they play a crucial role in maintaining and influencing enzymatic activities. The enzyme activity of the prokaryotic-expressed IvAIS1 protein was determined, and it was capable of catalyzing both coumaryl acetate and coniferyl acetate reduction reactions, yielding the products of t-anol and isoeugenol, respectively. The protein encoded by the gene sequence was found to be a t-anol/isoeugenol synthase. As the previous reports, the biosynthetic precursors, t-anol and isoeugenol, undergo an O-methylation reaction in order to produce t-anethole and estragole, respectively. 21
The optimal reaction systems exhibited Km of 480.30 ± 86.61 and 438.40 ± 44.29 µM, Kcat of 0.33 ± 291 0.018 and 0.67 ± 0.025 s−1, and Vmax values of 4.05 ± 0.23 and 8.18 ± 0.30 nmol·s−1·mg protein−1 for coumaryl acetate and coniferyl acetate, respectively. These three parameters consistently showed that IvAIS1 is more active in catalyzing coniferyl acetate to produce isoeugenol in vitro. In reality, t-anethole accounts for 70–90% of the volatile substances, while isoeugenol and methylisoeugenol are hardly detected.33,34 There is a difference in substrate preference between in vivo and in vitro for IvAIS1. Similarly, anise AIS1 has demonstrated dual catalytic potential with greater activity toward coniferyl acetate rather than coumaryl acetate. 21 However, there was no isoeugenol found in the anise plant, and it might only catalyze coumaryl acetate in vivo. The AsIGS converted coniferyl acetate into both isoeugenol and eugenol in vitro, but isoeugenol has not been detected in A. sieboldii. 12 Site-directed mutagenesis was used to explore the influence of amino acid residues on the substrate specificity of PIP reductase. The substitution of residues at positions 110 and 113 impacted the product selectivity of ScIGS1, and more eugenol was discovered in the final product. The mutation of L157A eliminated catalytic activity. 15 Further, the enzyme activities of anise AIS1, AsIGS, and IvAIS1 were all studied with the prokaryotic-expressed recombinant proteins. As is well known, prokaryotic-expressed proteins lack post-translational modifications. However, enzyme function is significantly impacted by phosphorylation,35,36 acetylation (typically to lysine residues),37,38 and other processes, which can dramatically alter an enzyme's stability or interactions with substrate molecules. The activity differences of the PIP reductases between in vivo and in vitro conditions would suggest that further studies of the in vivo action mechanism of IvAIS1 needs to be conducted.
The optimal pH and temperature for the catalysis were determined to be pH 7.0 and 40 °C and pH 7.0, 45 °C for coumaryl acetate and coniferyl acetate, respectively. Compared with the optimal temperature for sweet basil eugenol synthase (25 °C), 14 anise AIS1 (30 °C), 21 carrot DcE(I)GS1 (30 °C), 22 and petunia IGS1 (25 °C), 17 a higher temperature implies that IvAIS1 may be a high-temperature responsive protein that exhibit isoeugenol synthase activity during heat stress. However, it mainly catalyzes the reduction reaction of coumaryl acetate at subtropical temperatures in I. verum.
We analyzed the relative expression of IvAIS1 in various tissues in order to validate the function of IvAIS1 in I. verum. The concentrations of both t-anol and t-anethole were determined. Different from higher expression levels of EGS in young leaves and inflorescences of O. basilicum, Ocimum kilimandscharicum, Ocimum gratissimum, and Ocimum tenuiflorum, 39 the high relative expression levels of IvAIS1 were found in the roots and low in stems, which showed an opposite trend to their respective concentrations of t-anol and t-anethole. In addition, the relative expression level of IvAIS1 in the fruits did not reflect the levels of t-anol and t-anethole. This may be due to differences in reaction efficiencies of downstream methylations in various tissues. Another possibility is that IvAIS1 is primarily involved in the t-anethole biosynthetic pathway in the roots of I. verum, but other isozyme forms may exist in the stems, leaves, and fruits. In other studies, plants have been demonstrated to possess several PIP reductases for the catalysis of monolignol coniferyl and coumaryl alcolols.15,18 Although we found several species of mRNAs for PIP reductase in the full-length transcriptomic data, our investigations, have as yet, only confirmed the function of IvAIS1. There are currently no other reports on the PIP reductase in I. verum. In addition, the upstream and downstream catalytic enzymes of the PIP reductase in the biosynthetic pathway of t-anethole have not been published for I. verum. This limited information restricts further speculation on the mechanism of action of IvAIS1. Further studies on the potential activity of PIP reductases isolated from I. verum will be carried out.
Conclusions
We cloned a t-anol/isoeugenol synthase gene from I. verum. It is a member of the PIP family and has the typical protein sequence characteristics observed in this family. When expressed in a prokaryote, IvAIS1, it could convert coumaryl acetate and coniferyl acetate to t-anol and isoeugenol, respectively. The Km, Kcat, and Vmax values of the enzyme expressed in IvAIS1 suggest it is more active in the catalysis of coniferyl acetate. In addition, its optimal reaction temperature suggests it is sensitive to high temperatures.
Supplemental Material
sj-docx-1-sci-10.1177_00368504251325630 - Supplemental material for A t-anol/isoeugenol synthase responsible for converting coumaryl acetate to t-anol in Illicium verum
Supplemental material, sj-docx-1-sci-10.1177_00368504251325630 for A t-anol/isoeugenol synthase responsible for converting coumaryl acetate to t-anol in Illicium verum by Suqin Guo, Fa Zhang, Feng Peng, Haixia Yu, Kun Zhang and Guiyu Tan in Science Progress
Footnotes
Acknowledgments
The authors thank Dr Dev Sooranna of Imperial College in London for editing the manuscript.
Authors’ contribution
Suqin Guo: data acquisition and original draft preparation; Fa Zhang: data acquisition and investigation; Feng Peng: writing—review and editing; Haixia Yu: data acquisition and investigation; Kun Zhang: methodology; Guiyu Tan: strategic design, original draft preparation, final editing, and fund acquisition.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
There were no human or animal subjects and tissues used in this study.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The National Natural Science Foundation of China (grant no. 82304676), the Natural Science Foundation of Guangxi Province (grant no. 2021GXNSFBA196048), the Innovative Team for Traditional Chinese Medicinal Materials Quality of Guangxi (grant no. GZKJ2305), the Scientific Research Funding Project of Guangxi Botanical Garden of Medicinal Plants (grant no. GYJ202004), and the Self-Funded Scientific Research Project of Guangxi Administration of Traditional Chinese Medicine (grant nos. GZZC2020004 and GXZYA20230013) supported this work.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
