Abstract
As sessile photoautotrophs, plants constantly encounter diverse environmental stresses. Recent research has focused on elucidating sugar and energy signaling mediated by hexokinase (HXK), sucrose non-fermenting 1-related protein kinase 1 (SnRK1), and the target of rapamycin (TOR) and assessing its intricate interplay with hormones and secondary metabolism. HXK serves as a pivotal regulator of glucose sensing and metabolism. It affects plant growth and development in response to nutrient availability. SnRK1 acts as a vital energy sensor that regulates metabolic adjustments during stress to bolster plant resilience. Moreover, TOR integrates nutrient signals to finely modulate growth and development, balancing cellular metabolism and resource allocation. Understanding the functions of HXK, SnRK1, and TOR can provide profound insights into plant adaptation mechanisms and open promising avenues for leveraging biotechnological strategies to enhance the stress tolerance and nutritional value of crops. This narrative review focuses on recent advancements in the molecular mechanisms of HXK, SnRK1, and TOR and explores their potential applications in agricultural biotechnology.
Keywords
Introduction
Plants encounter diverse environmental stresses, including nutrient deficiencies, extreme temperatures, drought, and high salinity, throughout their life cycles. These stresses cause constant fluctuations in the levels of endogenous sugars, which not only serve as the initial carbon backbone for biological molecules but also represent the main energy source for both autotrophic and heterotrophic tissues.1,2 Plants can detect sugar levels using specialized sensors. These sensors trigger a signaling cascade that leads to changes in gene expression and protein modification. 3 These alterations enable plants to respond to specific stresses, thereby enhancing their chances of survival. Hence, understanding the mechanisms underlying sugar and energy signaling is crucial to understand these regulatory processes in plants. Recent advancements in molecular biology, genomics, and metabolomics have provided unprecedented insights into the regulatory networks and molecular mechanisms underlying the sugar-mediated control of secondary metabolism. 4 However, despite these advances, our understanding of the intricate interplay between sugar and energy signaling and secondary metabolism remains fragmented. Key questions about the specificity, redundancy, and evolutionary conservation of these signaling pathways across different plant species remain unanswered. Furthermore, the potential application of sugar and energy signaling modulation for enhancing the resilience, nutritional value, and medicinal properties of plants remains largely unexplored.
In this narrative review, we discuss current knowledge and recent breakthroughs in the sugar and energy signaling-mediated regulation of stress response and secondary metabolism. Specifically, we focus on the role of hexokinase (HXK), sucrose non-fermenting 1-related protein kinase 1 (SnRK1), and the target of rapamycin (TOR) signaling in plant stress response and secondary metabolism (Figure 1). We also discuss the potential biotechnological applications of sugar and energy signaling manipulation for improving plant stress tolerance and secondary metabolite production.
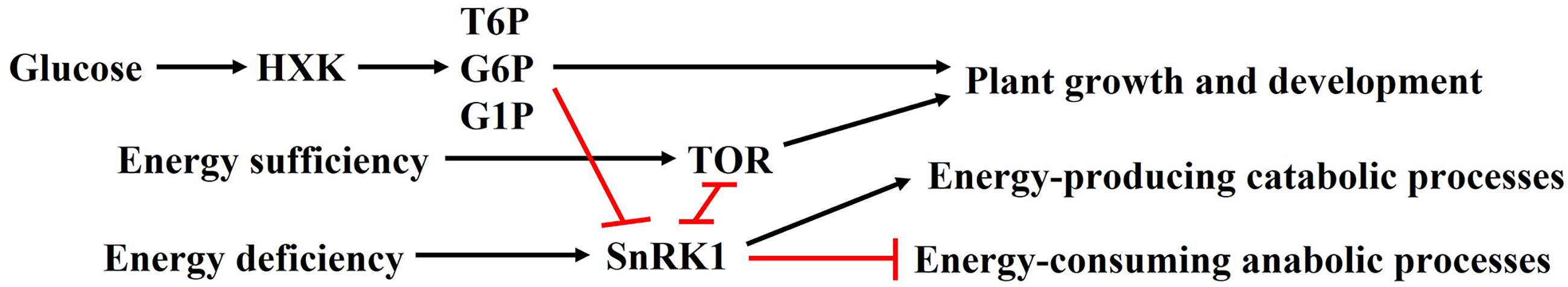
Hexokinase (HXK)- and glycolysis-dependent signaling pathways in plants. HXK and the target of rapamycin (TOR) exhibit growth-promoting effects,11,46 whereas sucrose non-fermenting 1-related protein kinase 1 (SnRK1) serves as a vital growth regulator under energy deprivation conditions. 46 T6P: trehalose-6-phosphate; G6P: glucose-6-phosphate; G1P: glucose-1-phosphate.
HXK signaling pathways in plants
Sugar perception in plants occurs through sophisticated sensor mechanisms that trigger intricate signaling cascades, resulting in profound changes in gene expression and post-translational modifications. While hexose sugars, such as glucose (Glc) and fructose (Fru), have been traditionally acknowledged as pivotal regulators of plant growth and development in response to diverse internal and external stimuli, their specific signaling pathways remain unknown. 5
The Glc and Fru signaling pathways are notably mediated by HXK, which catalyzes the formation of Glc-6-phosphate and Fru-6-phosphate, thereby providing substrates for the oxidative pentose phosphate pathway and promoting starch degradation and fat storage.6,7 In Arabidopsis, AtHXK1 can form a Glc signaling complex core with two unconventional partners: vacuolar proton pump subunit B (VHA-B) and the 19S regulatory particle of the proteasome subunit (RPT5B). This complex then binds to the target regulatory protein, which mainly consists of putative transcription factors.8,9 Although the downstream target genes are largely unknown, the aforementioned results suggest a multifaceted role of AtHXK1 in both sugar metabolism and signaling processes. The sensing function of AtHXK1, in which the catalytic activity of HXK is not essential for Glc sensing, 10 is evolutionarily conserved and is shared by HXK in other species, including potato (StHXK1 and 2), rice (OsHXK5, 6, and 7), and tobacco (NtHXK1). 11 Thus, the Glc sensing function of HXKs may be mediated by their affinity to Glc. However, how the conformational changes induced by Glc binding in HXKs transmit the signal of Glc abundance remains unclear. Understanding this mechanism is crucial not only for advancing the existing knowledge but also for agricultural applications.
HXKs (Table 1) can be classified into four distinct types, A–D, based on their structural and subcellular localization attributes.11,12 Type A HXK isoforms, such as AtHXK3 (Arabidopsis), OsHXK4 (rice), and LeHXK4 (tomato), are predominantly localized in the plastid stroma. Moreover, they contain a conserved chloroplast transit peptide consisting of approximately 30 amino acid residues. 5 In contrast, type B HXKs share a consensus hydrophobic membrane anchor domain and are predominantly associated with mitochondrial membranes. Representative members include AtHXK1 and 2 (Arabidopsis), SlHXK1, 2, and 3 (tomato), NbHXK1 (Nicotiana benthamiana), and OsHXK2, 3, 5, 6, and 9 (rice). 5 Emerging classifications have identified a type C HXK subgroup. Representative members include OsHXK1, 7, and 8 (rice), which predominantly reside within the cytoplasmic and nuclear compartments. Notably, these variants lack both plastid transit peptides and the characteristic membrane anchor domains. 13 Type D HXKs, encoded by a triad of distinct genes, also feature membrane anchor domains; however, the sequences diverge from those of their type B counterparts. 5 A prototype of this category is the Physcomitrella patens HXK PpHXK1, which is predominantly localized in the chloroplast envelope. 14 Thus, the diversity of subcellular localizations of HXKs in plants reflects their roles in various metabolic pathways.
HXK, SnRK1, and TOR family members in Arabidopsis, rice, and maize.

Rice Genome Annotation Project: http://rice.uga.edu/index.shtml.
Under stress conditions, the HXK pathways play a crucial role in inducing plant stress resistance by regulating sugar induction signals and glycolysis. Plants activate their innate immune systems through two pathways: pathogen-associated molecular pattern-triggered immunity (PTI) and effector-triggered immunity (ETI). In Arabidopsis, AtHXK1 was found to significantly boost plant defense against Pseudomonas syringae pv. tomato DC3000 infection by positively modulating PTI and partially engaging in ETI through Glc-induced effects mediated by AtHXK1-related pathways. 15 In addition, the overexpression of AtHXK2 enhanced plant resistance to P. syringae pv. tomato DC3000, correlating with increased H2O2 production and the upregulation of defense-related genes. 16 Similarly, OsHXK1-overexpressing rice exhibited enhanced reactive oxygen species (ROS)-mediated basal resistance to rice black-streaked dwarf virus infection. 17 Moreover, the overexpression of MdHXK1 improved disease resistance in apple calli, leaves, and fruits by upregulating genes associated with salicylate synthesis and signaling, in addition to augmenting ROS production. 18 These results indicate that HXKs play an important role in ROS-induced programmed cell death to enhance disease resistance. Extensive research has also been conducted to elucidate the pivotal signaling role of plant HXK genes in response to diverse environmental stresses, including alkaline, cold, drought, and salt stresses.19–23 The phosphorylation of the tonoplast-localized sodium (Na+)/proton (H+) exchanger by apple HXK1 was found to enhance salt tolerance. 24 Similarly, the overexpression of soybean HXK2 significantly improved salt stress tolerance by preserving Na+ and potassium (K+) homeostasis. 21 This indicates that the HXK Na+/H+ exchanger regulatory module maintains ion homeostasis for enhancing Glc-induced salt tolerance. Moreover, the overexpression of Prunus HXK3 in Arabidopsis disrupted carbon and energy metabolism, resulting in the accumulation of starch and organic acids and consequently enhancing tolerance to salt and drought stresses. 12 The analysis of the downstream mechanisms and physiological changes mediated by kinase-induced phosphorylation events has been facilitated by the development of small synthetic molecules that can specifically inhibit particular protein kinases. Under drought conditions, the enzymatic antioxidant mechanism, drought-induced abscisic acid (ABA) accumulation, and drought-responsive genes (RD29A, DREB2A, and PP2C genes) were affected by treatment with the HXK inhibitor N-acetylglucosamine (NAG). 25 Taken together, these findings indicate that phosphorylation events mediated by HXKs play a crucial role in the stress response machinery. However, further investigations and identifications of HXK substrates are warranted.
Effect of HXKs on secondary metabolism
Sugar serves as the main molecule in secondary metabolic pathways, such as shikimic acid, malonic acid, mevalonic acid, and methylerythritol phosphate pathways. 26 Treatment with sucrose improved the biosynthesis of polyphenols, flavonoids, γ-aminobutyric acid, phytosterols, and vitamins, thereby increasing the biological activities of mung bean sprouts. 27 Moreover, in various plants, treatment with sucrose and the accumulation of endogenous sugar were found to elevate the expression levels of both early- and late-stage anthocyanin biosynthetic genes, including those encoding chalcone synthase, dihydroflavonol reductase, and anthocyanidin synthase.28–31 However, the effect of sugars on anthocyanin accumulation varies depending on the plant species. In Arabidopsis seedlings, sucrose is significantly more efficient than Glc in inducing anthocyanin biosynthesis. 32 In contrast, in Paeonia suffruticosa, Glc treatment is more efficient than sucrose treatment in increasing anthocyanin contents. 33 This suggests that sugar regulates the expression of anthocyanin biosynthetic genes through different signaling pathways in different species. The anthocyanin biosynthetic pathway is transcriptionally controlled by the MYB transcription factor/basic helix–loop–helix (bHLH) domain protein/WD repeat (MYB/bHLH/WD40) protein complex, which is highly conserved among higher plant species. 34 MdHXK1 phosphorylates MdbHLH3 to modulate anthocyanin accumulation in response to Glc in apples. 35 Interestingly, under Glc deficient conditions, the overexpression of HXK fails to enhance anthocyanin accumulation. On the other hand, increased anthocyanin accumulation is induced by the activation of HXK upon exposure to a high level of Glc. 36 This indicates that the activation or function of HXK is required for Glc-induced anthocyanin accumulation (Figure 2). Similarly, PTMs, such as phosphorylation and ubiquitination, have been suggested to regulate HXK activity in human cells.37,38 However, our understanding of HXK-PTMs in plants remains limited. Based on in silico analysis, potential SUMOylation sites have been predicted in plant HXKs. 39 Therefore, it would be intriguing to ascertain whether small ubiquitin-like modifier proteins (SUMO proteins) are necessary for the activation of the HXK-dependent signaling pathway.

Glucosinolates are nitrogen- and sulfur-rich secondary metabolites that exhibit diverse biological activities, including antimicrobial, antioxidant, and anticancer activities. They also play an important role in plant defense mechanisms. 40 In the Glc-insensitive mutant gin2-1 (a null mutant of HXK1), the total aliphatic glucosinolate content was lower than that in wild-type plants after Glc treatment. 41 Moreover, genetic studies have suggested that Glc enhances the biosynthesis of aliphatic glucosinolates by inducing the transcription factors MYB28, MYB29, and ABA-insensitive protein 5 (ABI5) through HXK1-mediated signaling (Figure 2). 41 Concurrently, Glc promotes the accumulation of indolic glucosinolates by inducing MYB34, MYB51, and MYB122 expression (Figure 2). This process is regulated by HXK1 independently of ABI5. 42
Plant hormones play a crucial role in regulating the production of secondary metabolites. Interestingly, sugars can influence the synthesis, conjugation, and transport of hormones, in addition to influencing hormone response pathways. In turn, various hormones, such as ABA, gibberellins, and cytokinins, can affect sugar metabolism, transport, and response pathways.43,44 HXK1-deficient plants exhibit reduced cytokinin levels and enhanced expression levels of genes involved in strigolactone signaling. 45 This suggests that HXK is crucial for the crosstalk between sugar and hormone signaling pathways and is necessary for hormone-induced secondary metabolism. Further analysis of the function of HXK in hormone signaling can provide additional insights into the roles of HXK signaling and its importance in secondary metabolism.
Glycolysis-dependent SnRK1–TOR pathway
Sugar signals can be transmitted by protein kinases involved in energy metabolism, specifically through the glycolysis-dependent SnRK1–TOR pathway. 46 This pathway includes the protein kinases SnRK1 and TOR (Table 1), which likely sense cellular energy levels instead of directly sensing sugars. 47 TOR is activated to facilitate growth in response to favorable nutrient and energy conditions, whereas SnRK1 is triggered to suppress growth under nutrient and energy starvation conditions (Figure 1), such as prolonged darkness, hypoxia, and photosynthesis inhibition. 48 This suggests a close interaction between TOR and SnRK1, which act in coordination to regulate nutrient-driven processes.
TOR signaling in response to abiotic and biotic stresses
TOR is a conserved eukaryotic Ser/Thr protein kinase characterized by five conserved domains: the HEAT repeat domain, FAT domain, FRB domain, Ser/Thr kinase domain, and FATC domain (Figure 3). 49 Each domain has a distinct function. In particular, the HEAT domain participates in rRNA synthesis, the FRB domain is the binding site for the macrocyclic immunosuppressant drug rapamycin, and the FAT and FATC domains are involved in scaffolding and mediating protein–protein interactions, respectively. Glc activates TOR through glycolysis–mitochondria–electron transport chain relays, thereby promoting root meristem activation during the transition from heterotrophic to photoautotrophic growth in mitotically quiescent seedlings. 49 On the other hand, the depletion of Glc abolishes TOR activity and increases the expression of genes associated with autophagy and protein degradation. Transcriptome analysis has revealed that the Glc–TOR signaling pathway governs the transcriptional reprogramming of diverse gene sets associated with central and secondary metabolism, cell cycle regulation, transcriptional regulation, signaling cascades, cellular transport, and protein folding. 50 This finding underscores the role of TOR as a master regulator that integrates nutrient and energy signals to facilitate cell proliferation and growth. TOR also plays multifaceted roles in response to various environmental stresses (Figure 3). During the response to cold stress, plants often accumulate soluble sugars, which can profoundly affect their sensitivity to low temperatures, 51 indicating the potential involvement of TOR signaling in cold acclimation in plants. In fact, the TOR RNAi line (two-week-old soil growth plants) exhibits a cold-sensitive phenotype, providing direct evidence that an active TOR is essential for inducing cold tolerance in plants. 52 However, the analysis of estradiol-induced tor mutant seedlings (four-day-old seedlings) suggested that the inactivation of TOR is necessary for the inhibition of protein translation, a crucial step for inducing cold tolerance. 53 These conflicting results may be attributed to differences in silencing efficiencies between the estradiol-inducible promoter and the ethanol-inducible promoter. Another possibility is that the varying ages of the plants used in the experiments contributed to these discrepancies, as the cold response and cold acclimation in Arabidopsis are known to be age-dependent. 54 Although the transient inhibition of TOR activity by cold stress suggests that TOR inhibition plays a crucial role in enabling plants to resist cold stress, 53 future experiments should aim to clarify these conflicting findings by standardizing plant age across studies and directly comparing the effects of different inducible promoters on TOR activity under cold stress conditions. Additionally, examining the role of TOR in cold tolerance across various developmental stages could provide deeper insights into its function and regulation. TOR inhibitors and activators may serve as useful tools for assessing biological processes and understanding the physiological roles of TOR signaling in the response of plants to cold stress. In addition to cold stress tolerance, the overexpression of TOR enhances thermotolerance. However, TOR RNAi plants exhibit a reduced thermotolerance response. 55 The transcription factor E2Fa is a phosphorylation substrate of TOR. Under heat stress conditions, heat stress genes, including HSFA2, HSFB2B, HSFB1, HSP70, and HSP20-1, were significantly downregulated in the e2fa-1 mutant, indicating that the TOR–E2Fa pathway plays a crucial role in thermotolerance. 55 Similarly, TOR can enhance plant tolerance to drought and osmotic stresses. In Arabidopsis, TOR overexpression lines exhibited longer primary roots than control lines when exposed to a high concentration of potassium chloride. 56 Moreover, the ectopic expression of the Arabidopsis TOR gene in rice enhanced water use efficiency, growth, and yield under drought conditions. 57 Under non-stress conditions, TOR phosphorylates PYR/PYL/RCAR receptors, which function as ABA sensors, thereby negatively regulating ABA signaling. In contrast, under stress conditions, ABA inhibits TOR activity through the SnRK2-mediated phosphorylation of regulatory protein of TOR (RAPTOR B). 58 This indicates a crosstalk between TOR signaling and ABA signaling in response to stresses.

Domain structure of plant target of rapamycin (TOR) and its regulation during abiotic and biotic stresses. The HEAT domain is crucial for rRNA synthesis, whereas the FRB domain functions as the binding site for rapamycin. 49 The FAT and FATC domains play essential roles in scaffolding and mediating protein–protein interactions. 49 The C-terminal regions contain a Ser/Thr kinase domain that exhibits sequence similarity to the catalytic domain of phosphoinositide-3-kinase. 49 P denotes phosphorylation.
Emerging evidence has highlighted the role of TOR in plant immunity. In rice, TOR was found to function as a negative regulator of immunity by negatively interacting with salicylic acid (SA) and jasmonic acid signaling pathways. 59 Similarly, TOR acts as a negative regulator of JA biosynthesis and the JA-dependent response in both Arabidopsis and cotton. 60 Moreover, TOR expression was downregulated following NB-LRR protein activation. TOR suppression correlated with enhanced disease resistance, while TOR overexpression reduced resistance. 61 In addition, estradiol-induced tor mutant exhibited enhanced resistance to Fusarium graminearum, 62 suggesting a crucial role of TOR in modulating plant defense mechanisms. The identification of aliphatic 3-hydroxypropylglucosinolate as a natural endogenous TOR complex inhibitor suggests a mechanism through which plants regulate both defense and growth. 63 By inhibiting TOR, plants can enhance their defense responses while still managing growth, highlighting the key role of TOR in balancing these essential processes. Furthermore, the inhibition or silencing of TOR in tomato and N. benthamiana enhanced resistance to Xanthomonas euvesicatoria, tobacco mosaic virus, Alternaria alternata, and Botrytis cinerea by activating defense responses in a SA-dependent manner. 64 Recent studies have identified TOR as a pivotal regulator, integrating local auxin-dependent pathways with the translational control of auxin-responsive genes. 65 Moreover, plant hormones such as auxin and ABA have been implicated in modulating plant–pathogen interactions, particularly through their influence on the SA/JA signaling network. 66 These findings suggest that TOR is a central hub in the crosstalk between hormone signaling pathways, playing a crucial role in coordinating hormonal responses under diverse stress conditions. Future research should focus on unraveling how TOR integrates signals from biotic and abiotic stressors to regulate these networks.
Effect of TOR on secondary metabolism
While the physiological role of TOR signaling in secondary metabolism remains largely unexplored, evidence suggests that Glc–TOR signaling orchestrates the transcriptional reprogramming of several genes involved in secondary metabolic processes, including the biosynthesis of glucosinolates, lignins, and flavonoids. 50 Inducible artificial microRNA lines targeting TOR exhibited the upregulation of transcripts related to flavonoid synthesis. 67 In Salvia miltiorrhiza, the inhibition of TOR signaling through the TOR inhibitor AZD8055 increased the expression of genes encoding key enzymes involved in the production of the terpene skeleton, which is crucial for the biosynthesis of terpenoid and isoprene units. 65 Furthermore, the transcription factors SmWRKY2, SmMYB98, and SmMYB36, which positively regulate the synthesis of tanshinone and salvianolic acid, were significantly upregulated following TOR inhibition. 68 These findings indicate that TOR may act as a negative regulator of secondary metabolism. Thus, TOR may play a central role in Glc and energy signaling by reprogramming the transcriptome, which is essential for remobilizing carbon and nitrogen via secondary metabolic pathways.
Role of SnRK1 signaling pathway in plant response to various stresses
In plants, SnRK1 acts as a crucial metabolic sensor that is activated under low energy conditions and inhibited by various sugars, such as trehalose-6-phosphate (T6P), Glc-6-phosphate, and Glc-1-phosphate (Figure 1). 69 In Arabidopsis, SnRK1 functions as a heterotrimeric complex consisting of a catalytic α subunit and regulatory β and γ subunits, which are encoded by distinct genes (SnRK1α1/α2, SnRK1β1/β2/β3, and SnRK1βγ genes). 70 The α subunit contains a highly conserved N-terminal Ser/Thr kinase domain and a C-terminal β-subunit interaction domain. The N-terminal domain contains a conserved phosphorylation site within a T-loop structure, which is crucial for kinase activity across various species. 70 The catalytic α subunit is essential for the kinase activity of the SnRK1 complex, while the regulatory β and γ subunits are crucial for modulating the stability, substrate specificity, subcellular localization, and overall kinase activity of the SnRK1 complex.71,72 Unlike TOR, SnRK1 triggers catabolic activation and suppresses energy-consuming anabolic processes in plants to promote energy conservation and nutrient remobilization under energy deprivation conditions. 46 In the presence of sufficient energy, both type 2C protein phosphatase (PP2CA) and ABA-insensitive protein 1 (ABI1) can interact with the catalytic subunit of SnRK1, resulting in the dephosphorylation and subsequent inactivation of SnRK1.1 (SnRK1α1) and the eventual inhibition of the SnRK1 signaling pathway. However, under conditions of energy deprivation, ABA binds to pyrabactin resistance 1-like (PYL) receptors, inhibiting PP2C protein expression and activating SnRK1 signal transduction. 73 Furthermore, sugar phosphates, key intermediates in sugar metabolism, can directly inactivate plant SnRK1. 74 Thus, SnRK1 functions as a crucial energy sensor through crosstalk with ABA signaling in plants. SnRK1 can be activated by various environmental stresses, such as extended darkness, hypoxia, drought, or cold conditions, which directly or indirectly result in energy deficits and impact critical processes, including photosynthesis, respiration, and carbon allocation. 75
The availability of sugars to sustain glycolysis is essential for plant survival during submergence. 76 However, the induction of sugar-utilizing enzymes under hypoxic conditions is an energy-demanding process that must be precisely regulated to avoid disrupting the balance between energy consumption and production. Under hypoxic conditions, the energy-sensing mechanism involving SnRK1 is activated through the turnover of metabolites indicative of low energy status, such as the reduction of T6P. 77 The phosphorylation of the translation initiation factor eIFiso4G by SnRK1 is crucial for the enhanced translation of genes involved in fermentation, sucrose degradation, and fatty acid biosynthesis, which are vital for energy production under hypoxic conditions. 78 Moreover, the overexpression of KIN10 (Arabidopsis SnRK1α1) activates the formation of autophagosomes through the phosphorylation of autophagy-related gene 1 (ATG1), resulting in increased tolerance to drought and submergence. 79 Under drought conditions, SnRK1 is released from sequestration by the SnRK1–PP2C–SnRK2 complex. It then inhibits the TOR complex to trigger stress responses and delay growth. 80 Furthermore, SnRK1 inhibits TOR activity by phosphorylating RAPTOR, leading to the activation of autophagy. 81 These findings imply that SnRK1 plays a role upstream of TOR in regulating autophagy under energy and nutrient deprivation conditions. 82 Thus, SnRK1 may exhibit diverse regulatory effects on energy metabolism to ensure plant survival under stress-induced energy deficiency (Figure 4). Interestingly, while TOR and SnRK1 typically function antagonistically in regulating nutrient-driven processes, TOR also plays a crucial role in enhancing drought tolerance. In rice overexpressing AtTOR, drought-responsive transcription factors such as OsNAC1, OsNAC2, OsWRKY72, OsDREB2B, and OsbZIP23 were significantly upregulated, 57 contrasting with SnRK1-mediated drought tolerance, which is primarily linked to the activation of autophagy. 79 These findings suggest that TOR and SnRK1 modulate plant drought responses through distinct mechanisms. Further investigation into the temporal activation patterns of these kinases and the changes in cellular sugar levels following drought stress could provide valuable insights into the coordination between TOR and SnRK1 signaling networks. The salt-induced activation of SnRK1 phosphorylates SRRM1L, a splicing factor that regulates plant resistance to salt stress mainly by fine-tuning the ratio of functional intron retention nuclear factor Y subunit A 10.1 (NFYA10.1) and non-functional NFYA10.3 variants. 83 In Arabidopsis, the activation of SnRK1 and treatment with ABA trigger largely overlapping transcriptional responses, suggesting that these crucial stress pathways share common target genes. 84 SnRK1 interacts with the ABA pathway at various levels, including metabolism, perception, and response, potentially regulating ABA-mediated stress responses. 85

Simplified summary of effects of sucrose non-fermenting 1-related protein kinase 1 (SnRK1) on plant stress response through the direct phosphorylation of target proteins. SnRK1 could directly phosphorylate autophagy-related gene 1 (ATG1), 82 the regulatory protein of TOR (RAPTOR), 81 and geminiviral proteins.86–89 P denotes phosphorylation.
At the core of plant immune responses lies the ability of plants to integrate diverse signaling pathways, resulting in extensive transcriptional reprogramming. Recent evidence increasingly highlights the significant role of SnRK1 in plant immune responses. In N. benthamiana, the expression of the KIN11 (Arabidopsis SnRK1α2) antisense increases susceptibility to geminivirus infection, while its overexpression enhances resistance to geminivirus infection. 86 Moreover, SnRK1 can directly phosphorylate geminiviral proteins to mitigate viral infections (Figure 4). It can target the βC1 protein from tomato yellow leaf curl China virus β-satellite and the AL2/C2 protein from cabbage leaf curl virus and tomato mottle virus, thereby delaying plant infection.87–89 Interestingly, viral proteins, such as AL2 from tomato golden mosaic virus and L2 from beet curly top virus, can inactivate SnRK1 to increase viral pathogenesis, 90 suggesting that SnRK1 interactions vary across plant viruses and play a crucial role in the plant antiviral defense response.
The hypersensitive response (HR) in plants is characterized by rapid localized cell death at the infection site of the pathogen, contributing to disease resistance. 91 In pepper, SnRK1 can induce a Xanthomonas avirulence protein (AvrBs1)-specific HR. 92 In rice, the SnRK1A-mediated phosphorylation of cytosolic ATPases positively regulates innate immunity against Ustilaginoidea virens infection. 93 Moreover, in wheat, TaSnRK1 interacts with the Triticum aestivum Fusarium resistance orphan gene (TaFROG) to positively regulate resistance to Fusarium head blight. 94 SnRK1 also directly confers immunity against Blumeria graminis f. sp. hordei by phosphorylating and destabilizing WRKY3 in barley. 95 Intriguingly, bacteria, fungi, and oomycetes produce significant amounts of trehalose during their symbiotic and pathogenic interactions with plants. 96 Trehalose and its intermediate, T6P, are identified as SnRK1 inhibitors,69,97 suggesting that microbial trehalose enhances pathogenicity by inhibiting SnRK1 activity. Taken together, these findings indicate the importance of SnRK1 in plant responses to biotic stress. Further characterization of microbial trehalose will be essential to understand how SnRK1 is involved in plant immunity.
Role of plant SnRK1 in modulating secondary metabolic pathways
Unlike HXK signaling, SnRK1 plays a crucial role in downregulating specific secondary metabolic pathways to potentially conserve carbon and energy or redirect carbon fluxes toward other essential compounds. 96 For instance, SnRK1 phosphorylates WRINKLED1, a key activator of fatty acid synthesis, leading to its rapid degradation and the consequent downregulation of fatty acid biosynthesis under low cellular sugar conditions. 98 In response to carbon and energy deprivation, Arabidopsis SnRK1 inhibits phenylpropanoid biosynthesis via Kelch domain-containing F-box proteins that mediate the ubiquitination and degradation of phenylalanine ammonia lyase. 99 Similarly, SnRK1 inhibits sucrose-induced anthocyanin biosynthesis by suppressing the activity of the MYB–bHLH–WD repeat (MBW) complex.100,101 Moreover, research on mutants with enhanced or suppressed SnRK1 activity has indicated that the inactivation of SnRK1 by sugar is necessary for the activation of anthocyanin biosynthesis under high light conditions. 102 In contrast, MdSnRK1.1 phosphorylates MdJAZ18, promoting its degradation via the 26S proteasome and consequently releasing MdbHLH3 to induce the expression of regulatory and structural genes in sucrose-treated apple calli. This ultimately enhances anthocyanin biosynthesis. 103 These variations are likely attributable to differences in plant species and developmental stages. In both Arabidopsis and apple calli, there is a threshold sucrose concentration that optimizes anthocyanin accumulation. While 1% sucrose enhances anthocyanin biosynthesis in MdSnRK1.1 transgenic Arabidopsis and apple calli, higher concentrations, such as 9% in Arabidopsis and 12% in apple calli, inhibit this process. 103 A more detailed assessment of these discrepancies will be instrumental in elucidating the relationship between SnRK1 and sucrose-induced anthocyanin biosynthesis. As another example, 3-hydroxy-3-methylglutaryl-CoA reductase (HMGR), a key enzyme in the biosynthesis of isoprenoids and sterols, is phosphorylated and inhibited by SnRK1 in Arabidopsis. 104 Therefore, the SnRK1-mediated inactivation of HMGR under stress conditions may affect the production of several secondary metabolites and plant hormones. Although extensive experimental evidence indicates that SnRK1 plays a role in reconfiguring metabolism, several fundamental questions remain unanswered. In particular, the mechanisms by which SnRK1 senses and adjusts the balance between primary and secondary metabolism remain unclear. However, evidence from maize, where reduced T6P levels in reproductive tissue downregulate primary metabolism and upregulate secondary metabolism, suggests that the T6P/SnRK1 pathway plays a central role in this regulation. 105 Understanding these mechanisms could offer valuable insights into SnRK1-mediated secondary metabolism and pave the way for enhancing the production of bioactive compounds with significant pharmaceutical and nutritional value.
Future perspectives
Sugar and energy signaling, mediated by the HXK, SnRK1, and TOR signaling pathways, is a crucial regulatory mechanism underlying plant stress responses and secondary metabolism. The HXK signaling pathways highlight the pivotal role of Glc sensors in mediating complex gene expression and metabolic reprogramming. Furthermore, the interaction between SnRK1 and TOR provides a nuanced understanding of how plants orchestrate their metabolic and stress response networks. Although substantial progress has been made, significant gaps remain in our understanding of the precise molecular mechanisms and regulatory networks. Future research should assess the specific downstream targets of these kinases and PTMs that modulate their activity. Moreover, it is essential to investigate the impact of sugar signaling on plant-microbe interactions, as understanding how sugar influences relationships with both beneficial and pathogenic microbes could uncover new strategies for enhancing plant health and resilience. This exploration could lead to significant advancements in agricultural practices and crop management. Additionally, assessing the crosstalk between HXK/SnRK1/TOR signaling and other hormonal pathways will be crucial for understanding the broader implications of sugar-mediated stress responses. The potential biotechnological applications of modulating sugar signaling to enhance plant resilience and productivity are promising yet underexplored. By addressing these gaps, we can better harness the power of HXK/SnRK1/TOR signaling to improve agricultural sustainability and crop performance in the face of environmental challenges. Tailoring these pathways could facilitate the development of crops with improved stress tolerance, higher nutritional value, and enhanced medicinal properties. The use of CRISPR/Cas9 and other gene editing technologies could enable precise modifications in key components of HXK/SnRK1/TOR signaling pathways, driving innovations in sustainable agriculture and food security.
Conclusion
The interactions between HXK, SnRK1, and TOR signaling pathways play a crucial role in how plants respond to environmental stresses. HXK is essential for Glc sensing and metabolism, contributing to stress resilience. SnRK1 serves as an energy sensor that adjusts metabolism during challenging conditions, while TOR integrates nutrient signals to influence growth and development. Understanding these pathways reveals the complex regulatory networks that enable plant adaptation to changing environments. Furthermore, this knowledge has promising implications for biotechnology, offering avenues to enhance crop resilience and nutritional value. Future research should focus on elucidating the roles of these pathways across different plant species and their interactions with other regulatory mechanisms, ultimately leading to the development of crops better suited to withstand climatic challenges and support global food security.
Footnotes
Author contributions
All authors contributed to the study's conception and design. SHE and TKH conceptualized and supervised the work. SHE and EK prepared the outline. All authors contributed to the drafting of the manuscript and were involved in the writing, reviewing, and editing of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the research grant from the National Research Foundation of Korea (grant number RS-2024-00333380).
