Abstract
Objectives:
tert-Butylhydroquinone (TBHQ) is an antioxidant and preservative used in unsaturated vegetable oils and processed foods. However, when consumed in higher doses daily, it may pose a threat to public health by potentially increasing the risk of cancer, as it has an affinity with both the aryl hydrocarbon receptor (AhR) and the estrogen receptor alpha (ERα).
Methods:
This study aimed to examine the impact of substituting the 1,4-diol of TBHQ with 1,4-dithiol, referred to as TBDT, on the carcinogenic and antioxidant systems using computational methods. The binding affinity of TBHQ and TBDT to the two carcinogenic receptors, AhR and ERα, as well as to the antioxidant receptor Keap1 alone and in connection with Nrf2 (Nrf2-Keap1) was investigated through docking analysis.
Results:
The results indicated a decrease in TBDT's binding strength to ERα and AhR when assessed using Molegro Virtual Docker (P-value: 0.0001 and 0.00001, respectively), AutoDock Vina (P-value: 0.0001 and 0.0001), and the online server Fast DRH (P-value: 0.0001 and 0.0001). However, TBDT's binding affinity to Keap1 was predicted to be significantly stronger than TBHQ's by both MVD and AutoDock Vina (P-value: 0.0001 and 0.04), while its binding to Nrf2-Keap1 assessed to be stronger only by MVD (P-value: 0.0001).
Conclusion:
These findings suggest that TBDT not only exhibits higher antioxidant activity as a better ligand for the antioxidant system but also shows lower affinity with the AhR and ERα receptors. Therefore, TBDT can be considered a safer compound than TBHQ.
Introduction
The tert-butylhydroquinone (TBHQ) is a synthetic aromatic organic compound that is considered a type of phenol. 1 It contributes to food preservation by extending the shelf life of food products and reducing food waste caused by spoilage. 2 Because of its OH content, it also has high antioxidant properties.3,4
TBHQ frequently can be found in foods like edible oils, crackers, fats, fish, meats, cereals, dairy products such as milk and cheese, mayonnaise, other snacks, pre-made frozen foods, packaged foods, and fast foods, 5 as well as cosmetic products at concentrations less than 0.1%,6,7 and pharmaceutical products.8,9 Based on US Food and Drug Administration (FDA) criteria, the maximum safe daily intake of TBHQ is 0.02% and 0–0.7 mg/kg. 10
According to previous reports, long-term consumption and high concentrations of TBHQ can cause many risks such as the development of various cancers, including gastrointestinal, 11 kidney, 12 bladder, 12 prostate, 13 and breast cancer. 14 Through the activity of glutathione transferases, the glutathione (GSH) conjugates to TBHQ and creates the 3-tert-butyl-4-hydroxyanisole as a mediated product which is a toxic substance, and its accumulation in the kidney can result in tumor development. 12 The main receptors and mediators of TBHQ in the carcinogenic pathway are the aryl hydrocarbon receptor (AhR) and estrogen receptor alpha (ERα) in various cells and tissues.14,15
The AhR is the direct receptor of TBHQ but its binding affinity is weaker than its classical ligands. 15 The ligand-binding bioactivation of AhR leads to the induction of several phase II enzymes that are involved in the metabolism of xenobiotics and endogenous compounds. These metabolites are involved in the overexpression of proteins in the cytochromes P450 superfamily including CYP1a1 and CYP1a2, CYP1b1, and CYP2S1. 16 Among the AhR-regulated genes modulated by polar-aromatic ligands, CYP1a1 is the most activated pathway related to carcinogenesis. 17 Also, ERα is another proven receptor that is influenced by TBHQ. It is unclear whether TBHQ directly impacts ERα or if its effect is just mediated by changes in the expression of ERα. ERα is a biomarker that plays a pivotal role in the classification of breast cancer subtypes since its overexpression is related to an increase in proliferation and metastasis in breast cancer.14,18,19
The substrate binding domains of AhR and ERα are PAS-B [PAS (PER: period circadian protein; ARNT: AhR nuclear translocator; SIM: single-minded protein domain] and LBD (ligand binding domain) respectively. 20 Previous research revealed the importance of residues 307–329 within the PAS -B domain, which was shown to be especially flexible, acting as an access gate to the ligand-binding pocket. 21
TBHQ acts as an inducer of the Nrf2-Keap1 antioxidant system, which is enabled by increasing the stability of Nrf2 (Figure 1). 22 Nrf2 and Keap1 detect cellular oxidative stress, and in response, modulate the expression of many phase II detoxification enzymes. 23 TBHQ and TBDT state both bind to the Cys151 in the BTB position of the Cul binding domain within Keap1. 24 Keap1 acts as a crucial Nrf2 repressor and plays a pivotal role in regulating the Nrf2 signaling pathway. 25 According to recent studies, in the presence of TBHQ, Nrf2 dissociates from Keap1 and is translocated into the nucleus, where it undergoes dimerization with small Maf (sMAF) proteins. Then this complex binds to the antioxidant response elements in the enhancer region of the gene and subsequently triggers the transcription of antioxidant genes. 15
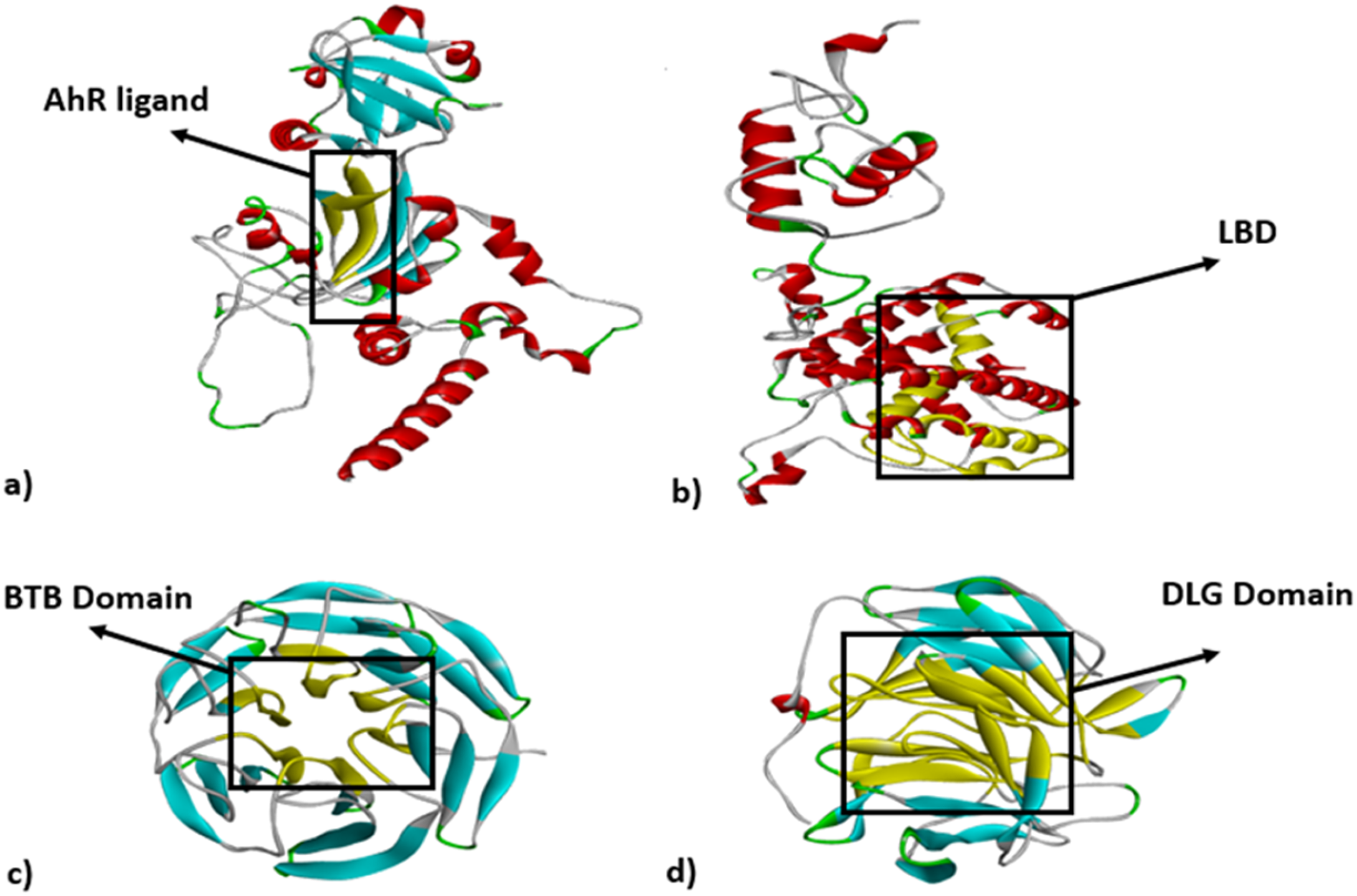
Cartoon representation of AhR, ERα, Keap1 and Nrf2-Keap1. (a) AhR complex: the PAS-B which is the ligand binding domain (yellow ribbons); (b) ERα complex: ligand binding domain (yellow ribbons); (c) Keap1: BTB domain which is the ligand binding domain; (d) Nrf2-Keap1: DLG is Keap1 binding domain. Crystal structure retrieved from (https://swissmodel.expasy.org/) at 11 November 2023 and ligand binding domains visualized by Discovery Studio 3.0.
One of the advantages of hydroxyl substitution by thiols is their advantage in antioxidation features which protects cells from oxidative damage.26,27 They possess a wide range of highly specialized and unique characteristics that enable them to serve various functions within the cell, 28 for example, these play important roles in enzymatic reactions, apoptosis, detoxification, and antioxidant protection in the body. 29 Many studies have shown that changes in the homeostasis of thiol status in various diseases such as the digestive system, respiratory system, reproductive system, urinary system, metabolic diseases, and cancers, are evident. 30 This also shows that the thiol state is important in the pathogenesis of oxidative stress-mediated diseases. 30 Some studies indicate that thiophenol should be an even more effective radical scavenger better than phenol. 31 Furthermore, their thiophenolic activity is mainly due to their electron-donating capacity, this feature increases scavenging properties compared to OH. 32 Another advantage of the SH group compared to OH is that low-molecular-weight (LMV) thiols serve as cofactors for thiol-dependent enzymes and reduce existing oxidative. 33 So, the substitution of hydroxyls in TBHQ by the –SH group can change its roles in cell signaling, antioxidant activity, and carcinogenesis (Figure 2(A)).

Structural comparison between TBHQ and TBDT and their binding affinities to AhR, ERα, Keap1, and Nrf2-Keap1. (a) Chemical structure of TBHQ and TBDT visualized by Chem Draw Pro 12.0. (b) 3D structure of AhR, ERα, Keap1 and Nrf2-Keap1 proteins and the coloring shows the QMEAN accuracy of structure (from purple color with the highest accuracy to red with the lowest accuracy) retrieved from (https://swissmodel.expasy.org/) at 13 November 2023. (c) Attachment of TBHQ and TBDT to ligand binding domain of AhR, ERα, Keap1, and Nrf2-Keap1 retrieved from Graph Pad Prism 8.0.
Since Ahr, ERα are two main TBHQ receptors active in the carcinogenic signaling pathway and Keap1, Nrf2-Keap1 are two key TBHQ systems active in the antioxidant signaling pathway, we focused on the effect of substitution of –OH in TBHQ by -SH on binding power of this ligand to AhR, ERα, Keap1, and Nrf2-Keap1. For this purpose, the docking method was employed to examine the binding affinity and electrostatic and H bonds of TBHQ and thiol-replaced TBHQ (tert-butylbenzene-1,4-dithiol: TBDT) to the mentioned proteins.
Materials and methods
Preparation of structures
The structure of TBHQ was obtained from PubChem (CID: 16043) (http://pubchem.ncbi.nlm.nih.gov) in a Structure Data File (SDF) format, which is suitable for docking programs. The 3D structure of the TBDT, with hydroxyl groups substituted by thiol groups, was drawn by Chemdraw3D and saved in SDF format. For a proper docking procedure, it is important to avoid unpredicted interventions. Therefore, the obtained 3D structures of proteins should not contain other ligands, activators, or inhibitors. The 3D structures of the selected proteins were predicted using the Swiss-MODEL web tool (http://www.Swissmodel.com). The accuracy of the models was evaluated using the QMEAN Z-score and GMQE score.
Docking analysis
To compare the binding affinity of TBHQ and TBDT with selected proteins, Molegro virtual docker (MVD) was utilized. MVD is based on guided differential evolution algorithms (http://www.molegro.com). The ligand binding sequences in proteins were assessed (Figure 1), and the chosen cavities for the docking process were aligned with these sequences.
The dockings simulations were conducted 10 times for both TBHQ and TBDT, with 50 runs for AhR, and 35 runs for ERα, Keap1, and Nrf2-Keap1. The number of runs was determined according to the number of amino acids in the selected proteins. The structures were prepared in docking mode, and the positions of cavities were calculated using expanded Van der Waals or molecular surface methods. For ERα, the ligand binding site (LBS) was aligned with one of the cavities based on expanded Van der Waals calculations. At the same time, for AhR, the PAS-B matched cavity was based on molecular surface calculations. In Keap1 and Nrf2-Keap1, the DLG matched the polar hole of the central cavity based on expanded Van der Waals, as there was only one cavity.
Upon completion of the docking process, poses with minimum energy scores were selected and their RMSD and hydrogen bonds (H-bond) were evaluated in the final comparison. The binding affinity of TBHQ and TBDT in four protein states was compared using AutoDock Vina software. Receptors and ligands were in PDBQT format and prepared using AutoDockTools. The fastDRH online software was utilized to insert the receptors, ligands, and specifications of the receptor's pocket. This was followed by calling out the receptor and ligand via AutoDock Tools to compute fit grid maps for each protein–ligand complex. Grid size, grid spacing, and grid center (Supplement S1) were determined by AutoDock Tools based on the location of the LBD with the receptor. The grid box properties were saved in Txt file format. Subsequently, docking was performed using AutoDock Vina by inserting command lines in the Command Prompt application to generate output scores. The best suitable model was then selected from the nine various conformations. Amino acid residues interacting in the binding site were visualized using LigPlot+ software. The interacting amino acid residues displaying hydrogen bonding and hydrophobic interactions with receptors are summarized in Table 1. Additionally, the fastDRH was used to determine free binding energies via molecular mechanics Poisson–Boltzmann surface area (MM/PBSA) and molecular mechanics Generalized Born surface area (MM/GBSA) approaches. The combined rescoring technique including AutoDock Vina, AutoDock-GPU docking engines, and the structure-protein MM/PB(GB)SA free energy design module featured a success rate of over 80%, surpassing AutoDock Vina's 70% success rate. The GAFF2 force field was employed for ligands, while the ff14SB force field was used for macromolecules in this study. Binding scores of TBHQ and TBDT for receptors were computed three times using AutoDock Vina and fastDRH, and the resulting P-values were recorded. This computational study was conducted from 01 November 2023 to 30 June 2024 at Mazandaran University by a personal computer.
Binding score and bond length of hydrogen bond and interaction amino acid residue (electrostatic bond) via Auto Dock Vina.
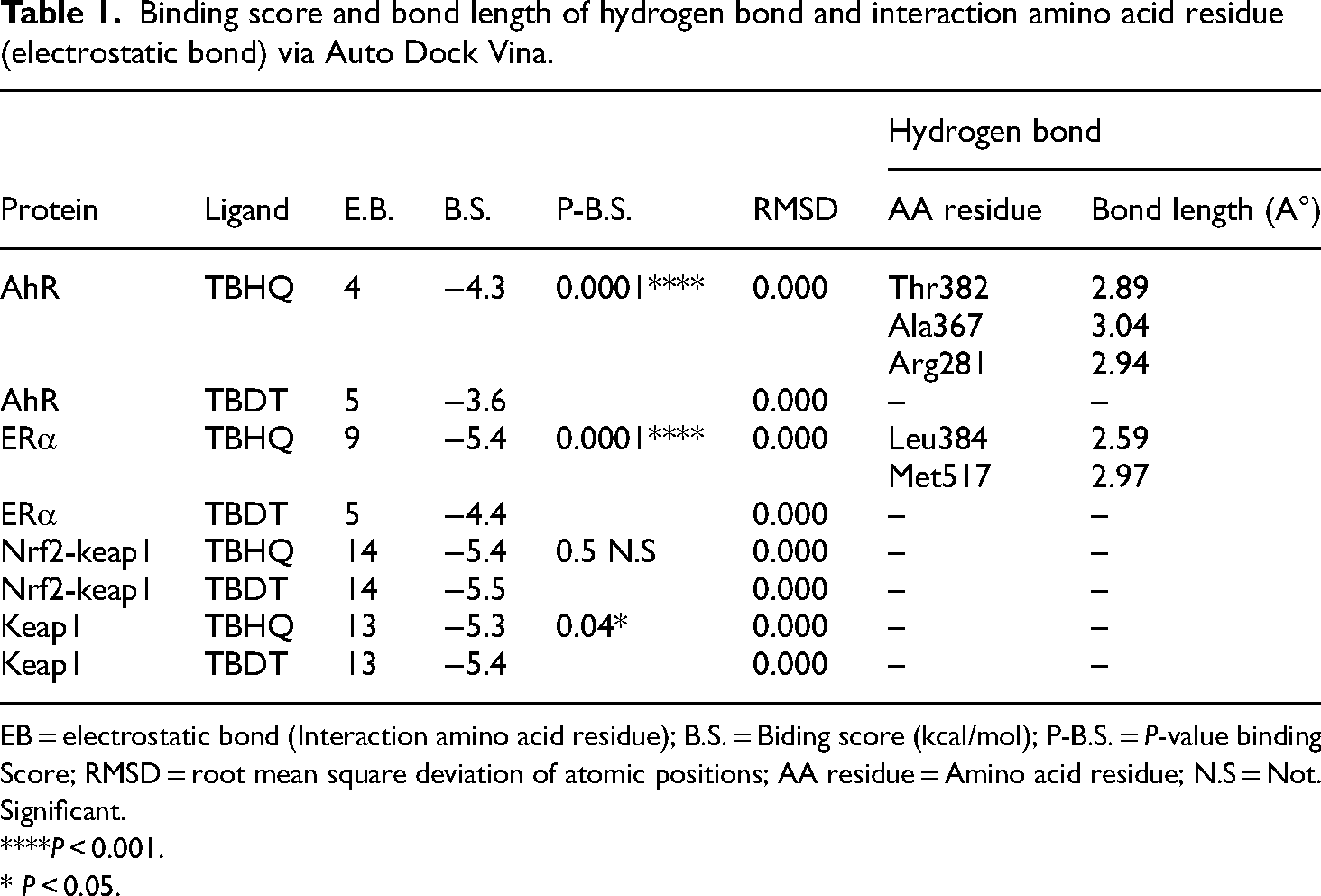
EB = electrostatic bond (Interaction amino acid residue); B.S. = Biding score (kcal/mol); P-B.S. = P-value binding Score; RMSD = root mean square deviation of atomic positions; AA residue = Amino acid residue; N.S = Not. Significant.
****P < 0.001.
* P < 0.05.
Statistical analysis
For the analysis of differences in binding energy between TBHQ and TBDT for each protein, an independent t-test was performed using SPSS 16.0, with statistical significance set at a P-value of less than 0.05. The graphical presentation of the results was generated using GraphPad Prism 8.0.2.
Results
The proteins’ 3D structures were predicted using the SWISS Model server, and the corresponding schematics are displayed in Figure 2(B). The GMQE scores for AhR and ERα were calculated as 0.26 and 0.39, respectively, and the QMEAN Z-scores of −1.94 and −3.58. For Keap1, the GMQE and QMEAN Z-scores were 0.94 and 0.33 while they were 0.94 and 0.29 for Nrf2-Keap1.
The results of the independent t-test indicate that the binding energies of TBDT to both AhR and ERα are significantly lower than TBHQ (AhR: P < 0.0001, ERα: P < 0.001), demonstrating the reduction of binding affinity of TBDT compared to TBHQ (Table 2). Conversely, the binding energies of TBDT to both Keap1 and Nrf2-Keap1 are significantly higher than TBHQ (Keap1: P < 0.001; Nrf2-Keap1: P < 0.001), indicating the improvement of TBDT binding affinity compared to TBHQ (Table 2). Figure 2(C) depicts the eight binding energies of the proteins to TBHQ and TBDT.
Comparison of TBHQ and TBDT in the binding domain of proteins and their binding energy.

CI = Confidence interval; df = Degrees of freedom; Ti = T-test independent; SD = Standard deviation; *Significant differences between TBHQ and TBDT based on independent t-test; ***P < 0.001; ****P < 0.0001.
Given the structural differences between the output of ChemDraw and the crystalized compound, it is anticipated that the root means square deviation of atomic positions (RMSD) in our docking format will range between 30 and 60 Å. Conversely, in the co-crystallized conformation of ligands, the RMSD is expected to be under 4 Å. This divergence in RMSD values is attributed to the unbound atomic base of the ligands with proteins.34,35
The mean RMSD values of the ten best poses of TBHQ and TBDT for ERα were 55 and 50.58, for Keap1 were 2.72 and 6.52, and for Nrf2-Keap1 were 32.19 and 32.31, respectively. RMSD of all types of ligands of AhR was higher. The average RMSD of TBHQ and TBDT poses for AhR were 103.89 and 104.22 Å, respectively. Because of the high RMSD of AhR bindings, the tetrachlorodibenzo-p-dioxin (TCDD) and formylindolo[3,2-b] carbazole-6 (FICZ) bindings to AhR were also evaluated to validate the findings for TBHQ and TBDT. The RMSD of these two ligands were estimated to be 102.87 and 103.06 Å, respectively. Apart from binding energy, the H-bonds have a considerable effect on the proper poses and binding strength of ligands to proteins. 36 So here the H-bonds energy and involved amino acids in H-bonds, and the changes in amino acids that have electrostatic interaction with ligands were presented in Table 3. Also, ester bond energy, energy intra, van der Walse H-bond, covalent radius H-bond, H-bond and electrostatic interactions length, H-bond and electrostatic interactions energy were presented in Supplement S2.
Comparison of TBHQ and TBDT, in the binding domain of proteins and their binding energy based on H-bond, and amino acids involved in these interactions.

H-b-E = Hydrogen bond (H-bond) energy (kCal/mol); Ei = Electrostatic interaction; H-b-aa = H-bond amino acid; L-H-B = Length hydrogen bond (A°).
The substitution position of TBHQ and TBDT in the LBD of ERα and PAS-B domain of AhR along with the binding, hydrogen bond, and surrounding amino acids obtained by MVD software, which illustrated in Figure 3.
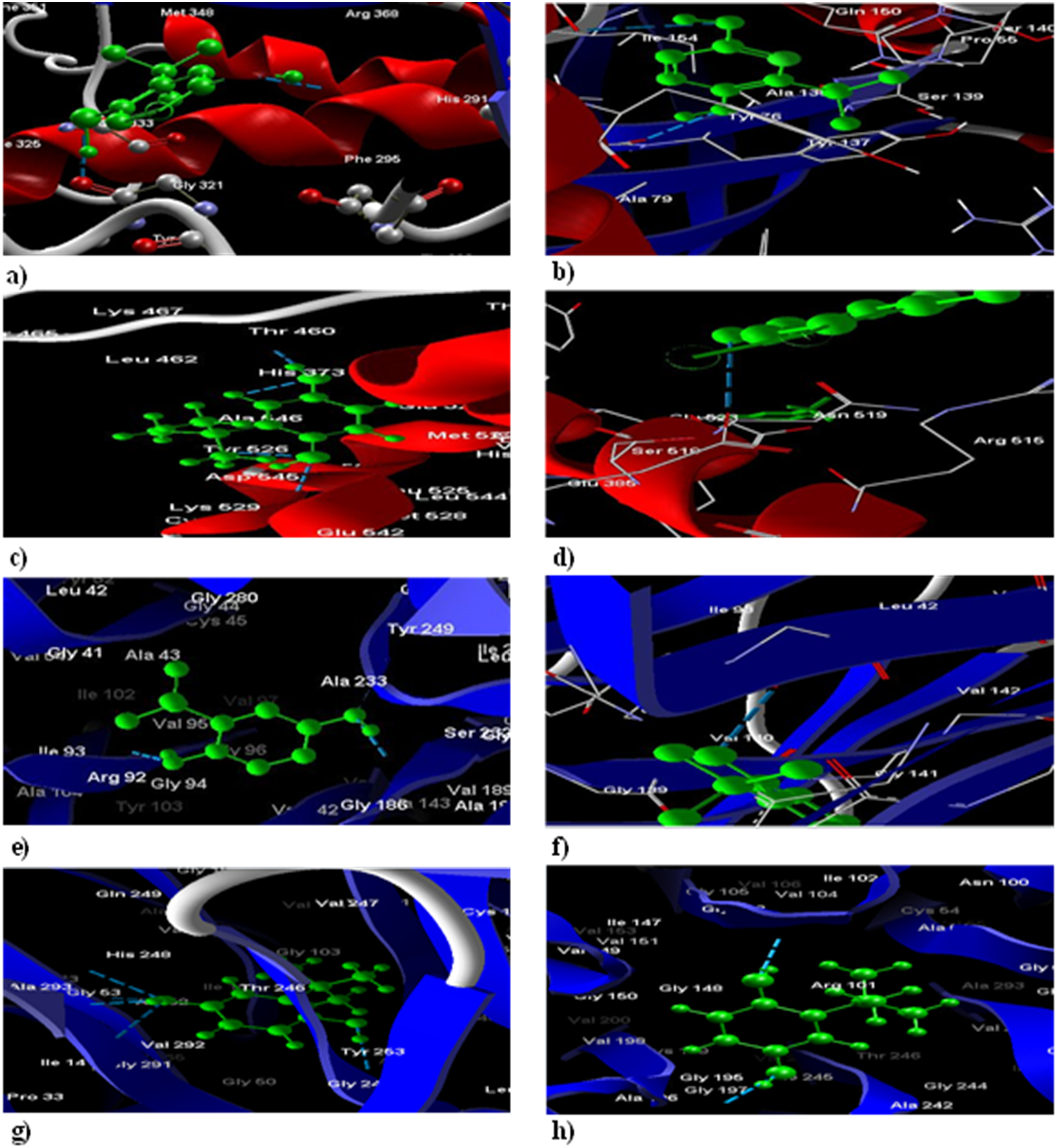
The poses of ligands and their interactions with amino acids. (a) The binding of TBHQ (green) within LBD of AhR is represented; H-bonds (blue dash-lines) between them and the closest amino acids are shown in this part. (b) The TBDT (green) within AhR's LBD is different from TBHQ and its H-bonds (blue dash-lines) are with different amino acids and this ligand is closer to α-helix (red-ribbon) than β-sheets (blue-ribbons). (c) TBHQ (green) docking with ERα has one H-bond and there are only five closes by amino acids. d) TBDT (green) shows different positions in ERα and there are more H-bonds (blue dashes) with TBDT and ERα than TBHQ. (e) TBHQ (green) docking with Nrf2-Keap1 has one H-bond and there are only five closes by amino acids. (f) TBDT (green) shows different positions in Nrf2-Keap1 and there are more H-bonds (blue dashes) with TBDT and Nrf2-Keap1 than TBHQ. (g) TBHQ (green) docking with Keap1 has one H-bond and there are only five closes by amino acids. (h) TBDT (green) shows different positions in Keap1 and there are more H-bonds (blue dashes) with TBDT and Keap1 than TBHQ retrieved from Molegro Virtual Docker 6.0.
The binding affinities of TBDT to both AhR and ERα, as determined via AutoDock Vina, were found to be lower than those of TBHQ, indicating the lower binding power of TBDT compared to TBHQ (AhR: P < 0.0001, ERα: P < 0.001) (Table 1). Conversely, the binding affinity of TBDT to Keap1 (P < 0.04) was stronger than that of TBHQ, suggesting the enhanced binding affinity of TBDT compared to TBHQ. Supplement S3 depicts AutoDock Vina binding energies of the proteins to TBHQ and TBDT by Prism.
In the secondary structures generated through LigPlot+ (Figure 4), the analysis reveals hydrophobic interactions involving a moderately diverse number of interacting amino acid residues in the altered ligand-receptor complex. Upon AhR interaction with TBHQ, three hydrogen bonds are formed, whereas with TBDT, the strength of the hydrogen bond is absent. Similarly, ERα interaction with TBHQ results in two hydrogen bonds being established, while with TBDT, the strength of the hydrogen bond is absent.

2D representation results for LigPlot+ analysis of the ligands interacting with receptors: (a) TBHQ-AhR, (b) TBDT-AhR, (c) TBHQ-ERα, (d) TBDT- ERα, (e) TBHQ, Nrf2-keap1, (f) TBDT-Nrf2-keap1, (g) TBHQ-Keap1 and (h) TBDT-Keap1. Hydrophobic interaction are represented by red spiky arcs, while hydrogen bond are indicated by green dashed lines along with calculated bond lengths retrieved from LigPlot 2.2.
In the case of Nrf2-Keap1 and Keap1, no hydrogen bond is formed with TBHQ and TBDT, and not significant difference in electrostatic bonding was observed. A complete summary of the results regarding hydrophobic interaction and hydrogen bond analysis using the LigPlot+ application is provided in Figure 4. The binding score of TBDT, as evaluated using the online software fastDRH, indicates lower affinity to both AhR and ERα compared to TBHQ (AhR: P < 0.0001, ERα: P < 0.0001) (Table 4). Additionally, the binding score of Nrf2-Keap1 and Keap1 demonstrates not significant. Furthermore, there was not significant difference observed in the MM/PBSA free energy calculation for any of the ligand/receptor interactions. However, the MM/GBSA free energy calculation shows that TBDT exhibits significantly lower free energy than TBHQ in its interaction with both AhR and ERα (AhR: P < 0.04, ERα: P < 0.01), indicating a preference for this substance over TBHQ. Conversely, the MM/PBSA free energy calculation reveals not significant difference between TBHQ and TBDT in their interactions with Nrf2-Keap1 and Keap1 (Table 4). Supplement S4 per residue energy decomposition of TBHQ-receptors complex and TBDT-receptors complex and MM/PB(GB)SA computation of the key interacting residues.
Docking and MM/PB(GB)SA score (kcal/mol) for the binding pose of the ligand.

B.S. = Biding score (kcal/mol); P-B.S. = P-value Binding Score; P-P.B = P-value Poisson−Boltzmann surface area; P-G.B. = P-value of molecular mechanics Generalized Born surface area; N. S. = Not Significant.
****P value <0.0001.
*P value < 0.05
Discussion
Investigating protein–ligand interactions and the discovery and development of therapeutic drugs are crucial aspects of molecular docking. This method provides valuable insights into the binding sites of ligands to proteins, the specific roles of individual interactions, and the binding energies of the amino acids in the protein and ligand atoms, including factors such as hydrogen bond length, electrostatic interactions, and free energies. Notably, molecular docking offers important information about the active sites of the receptor and the LBS, specifically connecting to the LBS of the receptor. In this study, we focused on the docking of TBHQ and TBDT with the AhR and ERα as carcinogenic receptors, as well as the Nrf2-keap1 and keap1 as antioxidant receptors.
Though TBHQ is used as an antioxidant in foods to prevent lipid oxidation chronic exposure to this food additive can cause inflammation and cancers. 15 The reason for TBHQ's role in cancers can be because of its dual antioxidant and pro-oxidant nature within cells. 37 AhR-dependent pathway, such as expression of CYP1A1, is one of the major pathways activated in cells by direct attachment of TBHQ. 15 Ebert's study on breast cancer resistance proteins showed that TBHQ significantly increased this group of proteins in Caco-2 cells, compared to other assessed phytochemicals. This effect was mediated through AhR-dependent pathway. 38 Based on the result of our study, the change of hydroxyl groups of TBHQ to thiols significantly reduces the binding power of this ligand to AhR in all three software.
According to the results of this study, the H-bond energy and electrostatic energy between TBDT and AhR are also in MVD weaker than TBHQ and AhR and H-bond energy and electrostatic energy between TBDT and AhR are also in AutoDock Vina missing TBHQ and AhR. Previous studies have indicated that other than binding power, the H-bond energy is an important factor in the specificity of ligand binding. This energy consists of Lennard-Jones, an electrostatic, and a H-bond term. 39 The H-bonds energy strongly contributes to ligand–protein stability and its impact is not limited to the ligand orientation which brings specificity. 40 In this context, not only the connection of TBDT to AhR is weaker than TBHQ, but it also has less specificity for AhR. Consequently, the substitution of hydroxyl groups by thiols may affect the AhR-dependent carcinogenic pathway and the MM/GBSA energy between TBDT and AhR are stronger than TBHQ.
ERα plays a key role in breast and prostate cancer, with its dysregulation serving as a significant inducer of breast cancer in the majority of patients. 41 It also counted as a biomarker for breast cancer chemoresistance. 42 While the endogenous ligands of ERα are estrogens especially estradiol which has the highest affinity among other endogenous ligands, some of the phenolic compounds also can bind to ERα as either inhibitors or activators.43,44 The impact of TBHQ on ERα protein level has been shown previously, which is leading to its decrease in breast cancer cells. 14 The result of the current study showed the direct connection between TBHQ and the LBD of ERα is possible since the binding energy was negative in three software and also in the MVD there is a H-bond between one of its hydroxyl groups and Glu 385 and the Autodock vina H-bond is missing. The MM/GBSA energy between TBDT and ERα are stronger than TBHQ.
McCullough revealed some of the phenolic compounds bind to ERα by connection of their phenolic hydroxyl with Arg394/Glu353. Based on their study, this bond and the distance between two hydroxyl groups create the high specificity of ERα's ligands and only their orientation determines them as agonists or antagonists. 45 Based on our results and those of previous studies, TBHQ may bind to LBD of ERα but it can’t trigger its activation, since this process needs a very specific interaction between specific amino acids and hydroxyl groups which TBHQ fails to establish. This result is matched with the findings of the Pfiffner study, based on which alteration of ERα by TBHQ is independent of direct connection. Because, while the decrease in protein level is happening in the presence of both estrogen and anti-estrogen there were no changes in ERα localization in the presence of TBHQ and estradiol. 14 Given that several studies proved the importance of phenolic hydroxyl in ERα ligands,43,45 the substitution of –OH groups of TBHQ by –SH would eliminate the possibility of strong binding between them. According to the results of our study, not only is binding energy significantly weakened compared to TBHQ, but the H-bonds energy and electrostatic energy are also weaker. Hence, this alteration can change any possible direct effect of TBHQ on ERα. Also, there is a crosstalk pathway between AhR and ERα. However, findings of Madak-Erdogan's research showed that the AhR itself without its ligands, could suppress the estradiol-ERα activity. 46 Hence, the indirect activation ERα by substitution hydroxyl groups to thiols in TBHQ through their influence on AhR has a low possibility too.
Li's report emphasized the antioxidant effects of TBHQ by activating the Nrf2-Keap1 pathway. 47 Under physiological conditions, Keap1 represses Nrf2 by binding it in the cytoplasm, thereby promoting its degradation by the ubiquitin–proteasome pathway. 48 However, upon expose to oxidative stress, specific reactive cysteine residues of Keap1 are modified, leading to the disruption of the Nrf2-Keap1 interaction. This leads to the stabilization of Nrf2, which then translocates to the nucleus, forming a heterodimer with small MAF (sMAF) proteins. 49
Recent reports suggest a direct relationship between AhR and Nrf2, with some reports indicating indirect interactions. Nrf2 can self-induce independently of CYC enzyme activity, while AhR ligands can directly or indirectly affect Nrf2-Keap1. 48 Almazari's research demonstrated that under normal conditions MCF7 typically has low levels of Nrf2 protein. However, upon treatment with TBHQ, the levels of Nrf2 protein increase, which results in the upregulation of the cytoprotective antioxidant enzyme gamma-glutamylcysteine synthetase (GCS). 50 The study also suggests that converting TBHQ's hydroxyl groups to thiols can significantly increase its binding power to Nrf2-Keap1 and Keap1. Additionally, in its TBDT state, the strength of the H-bond becomes stronger, leading to the loss of its connection to two amino acids, Val292 and Gly53. As a consequence, the substitution of hydroxyl groups with thiols can impact the Nrf2-Keap1-dependent antioxidant pathway.
Our findings showed that TBDT can be a safer candidate compared to TBHQ regarding its influence on carcinogenic and antioxidant pathways. But all other pathways are also influential in soft tissue cancer. Thus, further studies on the impact of TBDT compared to TBHQ on these pathways are still needed in cancer. For further studies, numerous criteria should be considered when selecting the proper preservative compound TBDT in food packaging (such as thermal and pH stability, safety, performance characteristics, etc.); toxicity studies of TBDT and its preservative compound on cells to be used as a suitable substitute in food.
Conclusion
The permissible threshold of TBHQ in foods has been carefully determined due to its strong antioxidant activity. However, long-term consumption of foods containing TBHQ can potentially lead to carcinogenicity, particularly through the AhR pathway. Our study results suggest that replacing hydroxyl groups with thiols not only enhances antioxidant activity by improving scavenging in the Nrf2-Keap1 and Keap1 pathways but also affects the binding of TBHQ to AhR. Since AhR is the main receptor for TBHQ in carcinogenic pathways, a weaker and less specific binding of TBHQ to AhR could impact this crucial pathway. This change may also eliminate the possibility of TBHQ's direct effect on the ERα pathway.
Supplemental Material
sj-docx-1-sci-10.1177_00368504241280869 - Supplemental material for 1,4-Diol Hq (TBHQ) vs 1,4-dithiol (TBDT); simulation of safe antioxidant with a lower carcinogenic activity
Supplemental material, sj-docx-1-sci-10.1177_00368504241280869 for 1,4-Diol Hq (TBHQ) vs 1,4-dithiol (TBDT); simulation of safe antioxidant with a lower carcinogenic activity by Seyed Zahra Mosavi, Abasalt Hosseinzadeh Colagar, Tahereh Zahedi and Bagher Seyedalipour in Science Progress
Footnotes
Acknowledgements
The authors are grateful to our colleagues for their critical reviews of the manuscript.
Authors contribution
All authors contributed to the writing of this article: Colagar A.H. was involved in the conceptualization and methodology. Mosavi S.Z. contributed to data curation, analysis, and preparation of the original draft. Zahedi T. contributed to the docking method, the data analysis and the editing of the manuscript. Seyedalipour B. was involved in the design of the docking plan related to the new compound of TBDT, and some data analysis.
Availability of data and materials
The datasets generated and analyzed during the current study are not available.
Consent to participate
As the corresponding author, I confirm that the manuscript has been read and approved for submission by all the named authors.
Consent to publish
All authors would like to submit this manuscript, to be considered for publication as a Research Article, in the science progress. We declare that it is original, has not been published before and is not currently being considered for publication elsewhere.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This study did not include animal or human experiments.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by a grant from the University of Mazandaran, dedicated to the PhD thesis of Seyedeh Zahra Mosavi.
Supplemental material
Supplemental material for this article is available.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
