Abstract
Background:
Fatty liver disease (FLD) is a serious public health problem that is rapidly increasing. Evidence indicates that the transcription factor EB (TFEB) gene may be involved in the pathophysiology of FLD; however, whether TEFB polymorphism has an association with FLD remains unclear.
Objectives:
To explore the association among TFEB polymorphism, gene–environment interaction, and FLD and provide epidemiological evidence for clarifying the genetic factors of FLD.
Methods:
This study is a case–control study. Sequenom MassARRAY was applied in genotyping. Logical regression was used to analyze the association between TFEB polymorphism and FLD, and the gene–environment interaction in FLD was evaluated by multiplication and additive interaction models.
Results:
(1) The alleles and genotypes of each single nucleotide polymorphism and haplotypes of TFEB in the case and control groups were evenly distributed; no statistically substantial difference was observed. (2) Logistic regression analysis indicated that TFEB polymorphism is not remarkably associated with FLD. (3) In the multiplicative interaction model, rs1015149, rs1062966, rs11754668 and rs2273068 had remarkable interaction with the amount of cigarette smoking. Rs1062966 and rs11754668 also had a considerable interaction body mass index and alcohol intake, respectively. However, no remarkable additive interaction was observed.
Conclusion:
TFEB polymorphism is not directly associated with FLD susceptibility, but the risk can be changed through gene–environment interaction.
Introduction
Health problem has gradually aroused people's concern with the advancement of economy and society and the improvement of living standard. Fatty liver disease (FLD) is a chronic disease and a serious public health problem that is rapidly increasing.1,2 FLD is the pathological process of excess adipose accumulation in liver cells, caused by many factors, such as disease and drugs. Simple hepatic steatosis may transform into steatohepatitis or cirrhosis as FLD progresses. According to etiology, FLD is classified as nonalcoholic fatty liver disease (NAFLD) and alcoholic liver disease (ALD). Studies have indicated that the pathogenesis of FLD is affected by abnormal fat metabolism, immune response, environmental, genetic, and other factors. 3 Currently, specific medicine for FLD is deficient; the effective prevention and control measures for FLD are early detection and intervention, including diet control and exercise; and alcohol abstinence is the chief measure for patients with ALD. 3
Similar to most diseases, FLD is likely influenced by environmental and genetic factors. Recently, transcription factor EB (TFEB) has attracted extensive attention in the study of autophagy mechanisms. TFEB is the main gene involved in lysosome biosynthesis and encodes TFEB, which is an important regulatory factor for autophagy and lysosomal biosynthesis. TFEB is considered the main activator for autophagy–lysosomal gene transcription and refers to inflammation, 4 cell autophagy, 5 lipid metabolism, 6 and other biological processes. Previous researches indicated that TFEB can regulate the expression of many genes related to lipid degradation, such as cluster of differentiation 36 (CD36), fatty acid-binding proteins (FABPs), and carnitine acetyltransferase (Crat),7,8 and TFEB can also regulate lipid degradation factors: peroxisome proliferator-activated receptor alpha (PPARα) and proliferator-activated receptor gamma coactivator 1 alpha (PGC-1α).9,10 Furthermore, pieces of evidence directly manifest that TFEB can protect mice liver from alcohol-induced damage, 11 and promoting TFEB-mediated lysosomal biogenesis using formononetin can ameliorate the fatty disease process in mice liver. 12
Single nucleotide polymorphisms (SNPs) are the most common form of mutations in the human genome. Studies have found that SNPs are associated with FLD (e.g. GCKR rs780094, pNPLA3 rs738409, and TM6SF2 rs58542926).13–15 Nevertheless, few studies have focused on the relationship between TFEB and FLD. Therefore, a case–control approach was adopted in this study to explore the association between TFEB polymorphism and FLD, and the gene–environment interaction was evaluated to provide epidemiological evidence of the genetic factors of FLD.
Materials and methods
Study design and population
This case–control study included 228 patients with FLD diagnosed by the presence of at least two of three abnormal findings on abdominal ultrasonography: liver anterior echogenicity (“bright liver”), far-field echo attenuation, and unclear intrahepatic duct structure. 16 Individuals with liver diseases, tumors, and autoimmune diseases caused by drugs and viruses were excluded. A total of 342 healthy individuals who were matched by sex and age (with a variation of ±3 years) in a proportion of 1:1.5 were selected as the control group. All the participants were permanent residents in Gongcheng County, Guilin City, Guangxi Zhuang Autonomous Region, People's Republic of China and signed the informed consent voluntarily after fully understanding the research content and importance of this project. Our research protocol was approved by the Ethics Committee of Guilin Medical University.
Data collection
All the participants were required to answer a questionnaire from a trained researcher to collect information on demography, behavior, exercise, disease history, nutritional diet, and other data. Behavioral factors include smoking and alcohol consumption. The amount of smoking is expressed in pack year, that is, the number of packs (20 cigarettes per bag) per day multiplied by the number of years of smoking. In addition to daily alcohol intake, we also assessed the daily intake of 12 types of food, including cereals and their products, potatoes, vegetables, and fruits, via a dietary survey. The participants were also examined by professional physicians to collect anthropometric indicators, such as height, weight, waist circumference, and blood pressure. Venous blood was collected for subsequent tests and experiments.
Biochemical testing
Two venous blood samples were collected from each participant. One set of samples was tested for biochemical indicators, including triglyceride (TG), total cholesterol (TC), high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein cholesterol (LDL-C), alanine aminotransferase (ALT), and uric acid (UA), by using an automatic biochemical analyzer (Hitachi 7600-020, Kyoto, Japan) at a local hospital. The other set of samples was used in the SNP typing experiment.
Selection and genotyping of SNPs
Functional and validated SNP screening strategies were utilized to screen the target SNPs. This strategy focused on important functional loci and was supplemented by susceptible loci. Finally, the rs1015149, rs1062966, rs11754668, rs14063 and rs2273068 of the TFEB gene were selected as the target SNPs for genotyping.
Genomic DNA was isolated from venous blood using a commercial DNA extraction kit (Tiangen, Beijing, China). SNP genotyping was performed using the Sequenom MassARRAY matrix-assisted laser desorption ionization time-of-flight mass spectrometry platform (Sequenom, Inc., San Diego, CA, USA). The primers were designed and synthesized by Bio Miao Biological Technology Co., Ltd (Table S1).
Statistical analysis
Descriptive statistics for continuous and categorical variables were conducted using mean ± standard deviation (SD) and frequency (proportion), respectively. Student's t-test and chi-square test were applied to compare the differences among two groups and genotype subgroups. Pearson's chi-square test was utilized to evaluate the Hardy–Weinberg equilibrium (HWE) before analyzing SNPs data. The samples were considered representative when p > 0.05. Haploview was used for haplotype analysis on the possible haplotype and mapping according to the method described by Barrett et al. 17 We performed logistic regression to estimate the effects of genotypes and gene–environment multiplicative interactions on FLD, and odds ratio (OR) and 95% confidence interval (95% CI) were calculated. The test level α = 0.05. However, logistic regression was limited to estimate additive interactions; hence, relative excess risk due to interaction (RERI), the attributable proportion of interaction (AP), synergy index (SI), and their 95% CIs were calculated. Additive interactions were considered statistically significant when the 95% CI of RERI and AP did not include 0, and the 95% CI of SI did not contain 1. 18 SPSS 25.0 (IBM, Chicago, IL, USA) and PLINK 1.90 software were used to implement general statistical analysis and gene polymorphism analysis. In addition, R software 4.0.2 and "epiR" package were utilized to complete the calculation of RERI, AP, and SI.
Results
Characteristics of the participants
The demographic and behavioral characteristics of the participants are listed in Table 1. The subjects have a total number of 570 and a roughly equal gender proportion. The age range is 30–83 years with an average of 58.15 years. The majority of the subjects (78.42%) belong to the Yao population. No remarkable differences in gender, age, ethnicity, marital status, hypertension, smoking, drinking, and other factors were observed between the control and case groups at baseline (p > 0.05). However, the proportion of subjects with a history of hypertension and the average daily sitting time were significantly higher in the case group than in the control group (p < 0.05).
Demographic and behavioral characteristics of the study population.

SD: standard deviation.
Mean ± SD; *p < 0.05 was considered statistically significant.
The dietary situation is shown in Table 2. Vegetables and cereals and their products were the main daily dietary intake of the participants. No significant difference was observed in the daily food intake of the two groups (p > 0.05).
Daily food intake of the participants.

Note: Data are expressed as mean ± SD. The unit for each food category is gram. SD: standard deviation.
The clinical indicators are exhibited in Table 3. The two groups showed no statistically significant difference in AST (t = − 1.415, p = 0.158). However, HDL-C was significantly lower in the case group than in the control group, and the other indicators were significantly higher in the case group compared with the control group (p < 0.05).
Clinical indicators of the study population.

Note: Data are expressed as mean ± SD; *p < 0.05 and **p < 0.001 were considered statistically significant. WC: waist circumference (cm); BMI: body mass index; SBP/DBP: systolic/diastolic blood pressure (mmHg); HbA1C: glycosylated hemoglobin (%); LDL-C: low-density lipoprotein cholesterol (mmol/L); HDL-C: high-density lipoprotein cholesterol (mmol/L); TG: triglyceride (mmol/L; TC: total cholesterol (mmol/L); GLU: fasting plasma glucose (mmol/L); ALB: albumin (g/L); ALT: alanine aminotransferase (U/L); AST: aspartate transaminase (U/L); UA: uric acid (μmol/L); SD: standard deviation.
Results indicate that the demographic, behavioral, and dietary variables in the two groups were matched preferably.
Basic information of SNPs
Rs1015149, rs1062966, rs11754668, rs14063, and rs2273068 of TFEB are located in chromosome 6 and at the crucial regions that may impact genetic transcription and regulation, such as exon synonymous, promoter, and 3′ untranslated region (3′ UTR). The minimum allele frequency (MAF) of each locus was larger than 0.05, and all the loci of SNPs were consistent with HWE (pHWE > 0.05). Therefore, the study subjects are representative (Table 4).
Basic information of SNPs.
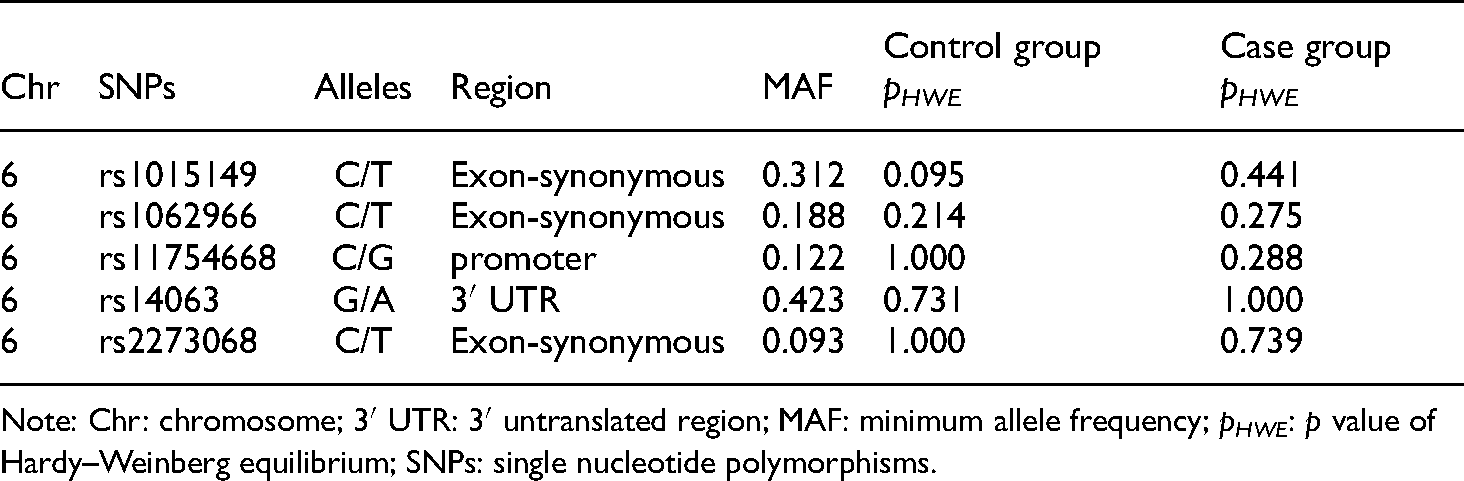
Note: Chr: chromosome; 3′ UTR: 3′ untranslated region; MAF: minimum allele frequency; pHWE: p value of Hardy–Weinberg equilibrium; SNPs: single nucleotide polymorphisms.
Genotypic frequency and the associations between genotypes and FLD
The alleles and genotypes of each SNP of TFEB in the two groups were evenly distributed. No statistically significant difference was observed (p > 0.05, Table 5).
Descriptive statistics of TEFB genotypes.

SNPs: single nucleotide polymorphisms.
FLD was regarded as the dependent variable. Co-dominant, dominant, and recessive models were used for logistic regression analysis using gender and age as adjustment factors. The result indicated that no significant correlations exist between genotypes and FLD in the co-dominant, dominant, and recessive models (p > 0.05, Table 6).
Logistic regression analysis between TEFB polymorphism and FLD.

FLD: fatty liver disease; 95% CI: 95% confidence interval; OR: odds ratio.
Haplotype analysis
Referencing the criteria of D' ≥ 0.8 and R2 ≥ 0.2, strong linkage disequilibrium (LD) was observed among the four SNPs: rs14063, rs1062966, rs2273068, and rs1015149 in one haploblock (Fig. S1). Therefore, five haplotypes (G-C-C-C, A-C-C-T, G-T-C-C, A-C-T-T, and G-C-C-T) were constructed in accordance with the LD analysis result. The result in Table 7 shows that all haplotype frequencies did not show statistically significant differences between the control and case groups (p > 0.05). Similarly, the regression analysis indicated that five haplotypes were also insignificantly associated with FLD (p > 0.05, Table 7).
Haplotype frequencies and logistic regression analysis between haplotypes and FLD.

Note: Haplotypes consisted of these four SNPs (rs14063, rs1062966, rs2273068, and rs1015149) of TFE; adjusting by gender and age; Freq.: frequencies (%); pa: p value of chi-square test; pb: p value of logistic analysis; FLD: fatty liver disease; 95% CI: 95% confidence interval; OR: odds ratio.
Interactions between environmental factors and SNP in FLD
Multiplicative and additive models were used to evaluate the interactions of each SNP locus with environmental factors, including diabetes, cigarette smoking, alcohol intake, daily sitting time, waist circumference, and body mass index (BMI). Compared with single genes, some gene–environment interactions were remarkably associated with FLD susceptibility. In the multiplicative interaction model, rs1015149, rs1062966, rs11754668 and rs2273068 had substantial interaction with the smoking amount. Among them, rs1062966 and rs11754668 also had remarkable interaction with BMI and alcohol intake, respectively.
Notably, the 95% CIs of RERI and AP contained 0, and the 95% CI of SI contained 1; thus, all gene–environment additive interactions with FLD were not statistically significant (Table 8, Table S3).
Results of gene–environment multiplication and additive interactions (only the parts with statistical significance are exhibited).
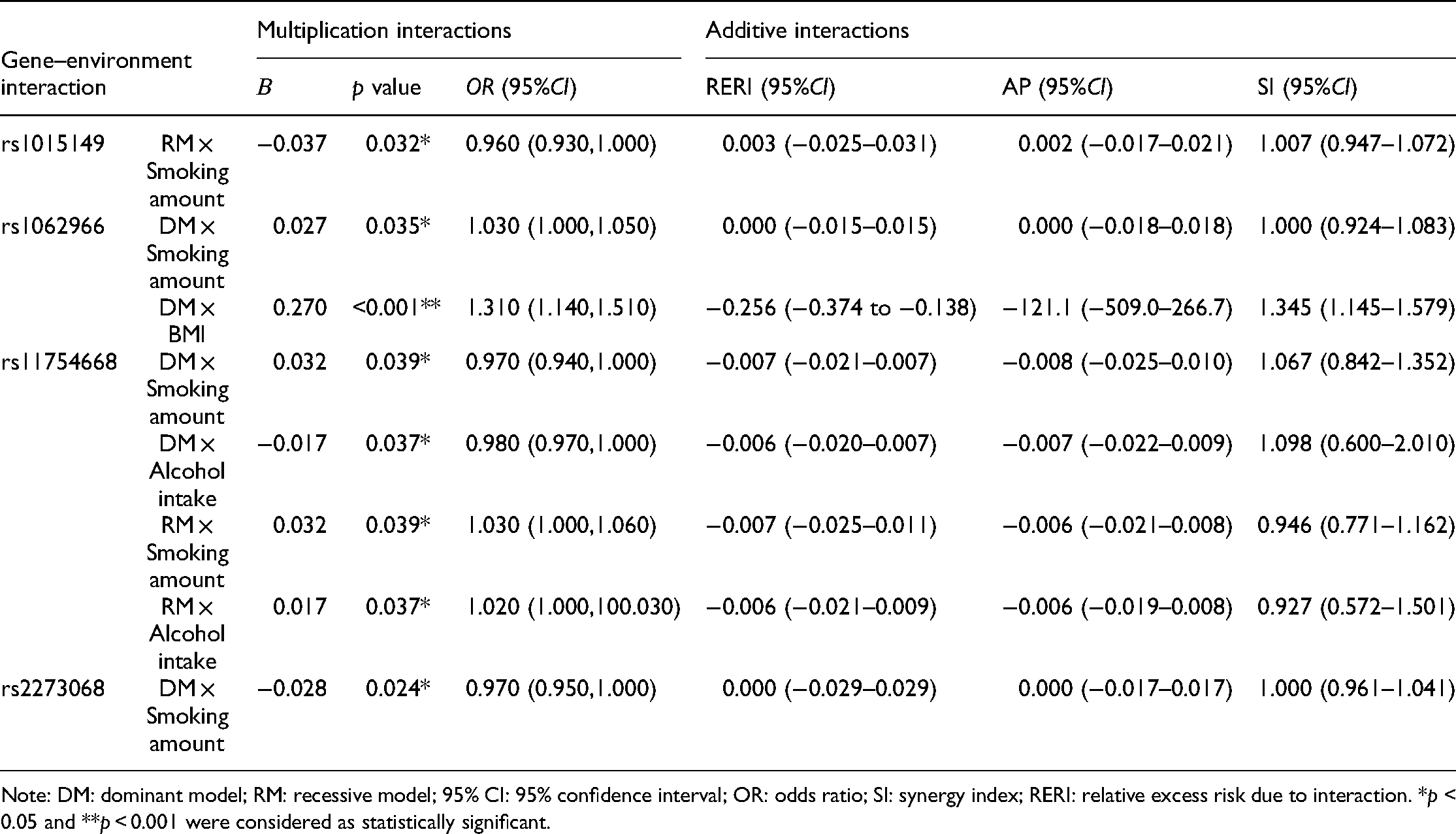
Note: DM: dominant model; RM: recessive model; 95% CI: 95% confidence interval; OR: odds ratio; SI: synergy index; RERI: relative excess risk due to interaction. *p < 0.05 and **p < 0.001 were considered as statistically significant.
Discussion
In this case–control study, we analyzed the association of TFEB polymorphisms and FLD and assessed gene–environment interactions to provide epidemiological evidence of the genetic factors related to the occurrence and development of FLD. Results showed that the alleles and genotypes of each SNP and haplotypes of TFEB in the case and control groups were evenly distributed; no statistically significant difference was observed. Logistic regression analysis indicated that TFEB polymorphism is not substantially associated with FLD. Previous studies have suggested that autophagy plays an important role in maintaining liver homeostasis. 19 TFEB knockout in mice may result in the hepatic accumulation of fatty acid-β and impaired oxidation in hepatocytes, which lead to elevated fatty acid and glycerol levels and lipid metabolism disorders in hepatocytes. 7 According to the results, TFEB may be involved in the pathophysiological basis of FLD. However, the relationship between TFEB polymorphism and FLD was not observed in this study; thus, these SNPs and haplotypes may not affect the normal expression of TFEB.
Gene–environment interaction plays an important role in the occurrence and development of complex diseases, such as FLD. Zhu et al. demonstrated that the gene–gene interaction between AGTR1 and PPARγ is associated with the occurrence of NAFLD in the Chinese population. 20 Zhang et al. found that people with 11391G/A(AA) and EC-SOD (CG + GG) genotypes suffer a higher risk of NAFLD, and these genotypes have an interaction with Helicobacter pylori infection. 21 Therefore, the analysis of gene–environment interaction might be conducive to understand etiological factors and guide the prevention and treatment of FLD. The result exhibited that some SNP loci, such as rs1062966, and rs11754668, had positive interactions with smoking, which is a risk factor for FLD susceptibility. 22 This finding is consistent with the results of Zhang et al. on the interactions between GPX-1 polymorphism and smoking in NAFLD and also agrees with the results of Oniki et al. 23 Interestingly, rs1015149 and rs2273068 had a negative interaction with smoking in FLD; thus, they are considered “protective factors” (OR = 0.96 and 0.97, respectively). Smoking is a recognized risk factor that is remarkably associated with the occurrence of many diseases. However, the relationship between smoking and FLD is not clear yet.24–27 In this study, we found that smoking might reduce the risk of FLD of individuals who carry the CT + CC genotype of rs1015149 or the TT + CT genotype of rs2273068. However, the results do not "advocate" smoking in these populations for FLD prevention. A more rational explanation for the reduced FLD risk is that compared with individuals who carry other genotypes, people with the CT + CC genotype of rs1015149 or the TT + CT genotype of rs2273068 may be more able to offset the risk of FLD caused by smoking. The same explanation can also be utilized to explain the interaction between rs11754668 and alcohol intake in this study.
No significant additive interaction was observed in this study. This result is in agreement with the result of Zhao et al. 28 In other words, additive interaction may not be remarkable even if the factors studied have substantial multiplicative interaction. In fact, the interaction between multiple factors is based on multiplication and synergism, whereas additive interaction is relatively rare. In the field of medicine, the analysis models for gene–environment, gene–gene, and gene–environment–gene interactions, such as cross-generation analysis, 29 multifactor dimensionality reduction (MDR), 30 and generalized MDR, 31 are based on multiplication. Although substantial multiplicative interaction results were not observed in the present study, this study still provided a meaningful attempt to explore the gene–environment interaction in FLD, which might be ignored.
Dietary factors are important influencing factors of FLD. Numerous works of literature have reported the association between different dietary patterns or food intake and the incidence of FLD. For example, the high intake of meat, high-fat dairy products, and refined grains may increase the risk of FLD, whereas a diet based on fruits, vegetables, whole grains, fish, and olive oil can reduce FLD risk.32,33 A cross-sectional study based on Chinese adolescents illustrated that adolescents who have a traditional Chinese diet have lower risks of FLD compared with those with the Western diet. 34 Therefore, the potential impact of diet on the result needs to be fully considered and controlled to reduce analysis error. In addition, in this study, we ensured data quality and improved the reliability of the results. The advantages are as follows: (1) This study is the first epidemiological study to uncover the associations of TFEB polymorphism and gene–environment interaction with FLD. (2) All participants were from the same area with relatively similar genetic backgrounds, living environments, and habits; these similarities were helpful to control potential confounding factors. (3) In terms of grouping, gender and age (±3 years) were adopted in matching to reduce the influence of gender and age on the results to a certain extent.
However, this study also has many deficiencies that need to be further improved. First, the sample size is relatively small, and sampling error is difficult to decrease. Second, the conclusions are based on the population from Gongcheng County. Therefore, the applicability of the conclusions to other populations is limited, and extrapolation is deficient. Third, few studies focused on the loci of SNPs selected in this study. Hence, references on the target SNP selection were insufficient. This insufficiency increases the uncertainty of the research conclusion and enhances the difficulty of interpretation simultaneously. Therefore, an optimized SNP screening strategy is required. Fourth, the degree of FLD was not classified. Thus, the effect of research factors on the process of FLD might have been ignored. Finally, the causal demonstration power is not strong because of the case–control design.
Conclusion
The polymorphisms of the rs1015149, rs1062966, rs11754668, rs14063, and rs2273068 of the TFEB gene are not directly associated with FLD susceptibility, but the risk can be changed through the gene–environment interaction.
Supplemental Material
sj-docx-1-sci-10.1177_00368504211043766 - Supplemental material for Association between TFEB gene polymorphism, gene–environment interaction, and fatty liver disease: A case–control study in China
Supplemental material, sj-docx-1-sci-10.1177_00368504211043766 for Association between TFEB gene polymorphism, gene–environment interaction, and fatty liver disease: A case–control study in China by Chunbao Mo, Tingyu Mai, Jiansheng Cai, Haoyu He, Huaxiang Lu, Xu Tang, Quanhui Chen, Xia Xu, Chuntao Nong, Shuzhen Liu, Dechan Tan, Qiumei Liu, Min Xu, You Li, Zhiyong Zhang and Jian Qin in Science Progress
Footnotes
Acknowledgments
Not applicable.
Authors’ contributions
CM, TM, and JC contributed to conception and design: HH, HL, XT, QC, XX, CN, SL, TD, QL, and MX contributed to acquisition of data; CM, TM, JC, and HH contributed to analysis and interpretation of data; CM, TM, JC, LY, and JQ contributed to writing, review, and/or revision of the manuscript; YL, ZZ, and JQ contributed to administrative, technical, or material support; ZZ and JQ contributed to study supervision. All authors approved the final manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The study was supported by the National Natural Science Foundation of China (Grant No. 81760577,81960583,and 81560523), the Guangxi Science and Technology Development Project (Grant No. AD17129003 and AD18050005), the Guangxi Natural Science Found for Innovation Research Team (2019GXNSFGA245002), the Guangxi Graduate Education Innovation Project (Grant No. YCSW2020230), and the Guangxi Scholarship Fund of Guangxi Education Department of China.
Availability of data and materials
Please contact the authors for reasonable requests.
Ethics approval and consent to participate
Our research protocol was approved by the Ethics Committee of Guilin Medical University. Informed consent was signed by all study participants.
Supplemental material
Supplemental material for this article is available online.
Abbreviations
Author biographies
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
