Abstract
Staphylococcus aureus nasal carriage is considered a risk factor for infections, and the development of nasal decolonization strategies is highly relevant. Despite they are not naturally colonized by Staphylococcus, mice are a good model for S. aureus nasal colonization. Murine models are easy to manipulate, and inter-laboratory reproducibility makes them suitable for nasal colonization studies. Strategies using bioluminescent bacteria allow for the monitoring of infection over time without the need to sacrifice animals for bacterial quantification. In this study, we evaluated S. aureus nasal colonization in three mouse strains (BALB/c, C57BL/6, and Swiss Webster) using a bioluminescent strain (SAP231). In vitro, a visible Bioluminescent Signal Emission (BLSE) was observed until 106 bacteria and detected by IVIS® imaging system up to 104 cells. Animals were inoculated with one or two doses of approximately 109 colony-forming units (CFU) of SAP231. Swiss Webster mice showed the longest colonization time, with some animals presenting BLSE for up to 140 h. In addition, BLSE was higher in this strain. BALB/c and C57BL/6 strains showed consistent BLSE results for 48 h. BLSE intensity was higher in Swiss Webster inoculated with both doses. Three different positions for image capture were evaluated, with better results for the lateral and ventrodorsal positions. After the loss of BLSE, bacterial quantification was performed, and Swiss Webster mice presented more bacteria in the nasal cavity (approximately 105 CFU) than the other strains. Our results demonstrate that bioluminescent S. aureus allow monitoring of nasal colonization and estimation of the bacterial burden present in live animals until 48 h.
Introduction
Staphylococcus aureus is ubiquitous as a natural colonizer of human skin and mucosal surfaces, but it can also be found in a variety of domestic, livestock and wild animals. It is recognized as one of the most dangerous pathogens, responsible for life-threatening infections due to its high virulence and ability to develop antibiotic resistance, explaining the morbidity and mortality frequency worldwide. 1 Staphylococci nasal carriage is considered a risk factor for Staphylococci infections. 2 One in three people (up to ∼33%) are natural and asymptomatic intranasal carriers of S. aureus, providing an important form of dissemination, especially in hospitalized patients with low immunity which may worsen its condition. 3 Methicillin-resistant S. aureus (MRSA) is a multidrug-resistant (MDR) bacterium implicated in community and nosocomial infections, due to its low affinity to beta-lactams and resistance to the other classes of antibiotics. 4 MRSA poses a formidable clinical threat, with persistently high morbidity and mortality. 5 MRSA colonization increases the risk of infection, and infecting strains match colonizing strains in as many as 50–80% of cases.6,7 Due to the growing interest in this matter, extensive efforts have been directed to the research of new approaches to MRSA decolonization, therefore justifying the importance of developing an easily reproducible model of nasal colonization.
Although there have been several studies on nasal colonization in humans,2,8 animal models are indispensable for vaccine development and other nasal decolonization strategies. The mouse is the most common animal model used for preclinical testing of any drug or vaccine. Although mice are not natural colonizers of MRSA, the easier acquisition and interlaboratory reproducibility made it a suitable model for nasal colonization optimization. 9
Bioluminescent imaging (BLI) has allowed the detection of small numbers of cells in rodents and facilitates the assessment of disease evolution without euthanasia, resulting in a reduction in the number of animals required in experimental studies. 10 Studies of infections using animal models are facilitated by BLI use, which allows tracking the evolution of infection in individual animals over time. BLI also allows the detection of signals in several animals simultaneously, decreasing costs and the time required for data acquisition when compared with computed tomography or magnetic resonance imaging assessment. 11 Furthermore, in BLI approaches, mice do not need to be sacrificed to estimate bacterial load in infected tissues.
Gonzalez-Zorn et al., using a bioluminescent MRSA strain (Xen31) and BLI, were able to demonstrate nasal colonization in a murine model. However, luminescence was consistently seen until 12 h after inoculation. 9
In this study, using a bioluminescent MRSA strain, three different murine strains (C57BL/6, BALB/c, and Swiss Webster) were evaluated to compare the time to nasal colonization by BLI. Therefore, the appropriate strain of mice chosen for better nasopharyngeal transport can be used as a model in studies of intranasal decolonization strategies.
Animals, materials and methods
Animals
Three different murine strains (BALB/c, C57BL/6, and Swiss Webster) aged 8–10 weeks old were used in this study. Only females were used in this study, due to supplier availability, which corroborated the study since females have less weight variability. All the animal procedures were approved by the Ethics Committee on the Use of Animal (CEUA) and Fiocruz (license number LW-06/21). Animals were provided by ICTB/Fiocruz and maintained in closed cages (dimensions: 37 cm in length, 24.2 cm in width and 24 cm in height) in a microisolator ventilated rack system (ALESCO1) with autoclaved pine flakes as a substrate. The cage changes were performed weekly. Mice were allocated to groups with a maximum of four animals per cage and maintained under a 12 h/12 h (day/night) light cycle with unrestricted access to food and filtered water. The experimentation laboratory has an Environmental Enrichment Program in which a polysulfone igloo is used as shelter and cotton, shredded paper, and cardboard rolls are interspersed monthly as a form of environmental enrichment. Using a clinical evaluation based on the ARRIVE guidelines, the humane endpoints were performed based on the severity of clinical signs such as severe hunching in the back, severe respiratory disorder change, animal isolated from cage mates, minimally active; and does not readily move when the cage is disturbed. When nudged, reluctantly moves. Immobile, not moving when nudged, and consequently reduction or cessation of access to water and food will be euthanized by anesthesia at doses of 200 mg/kg ketamine hydrochloride and 20 mg/kg xylazine hydrochloride administered intramuscularly. After a few minutes of anesthetic administration in the absence of reflexes, sodium thiopental was administered intraperitoneally at a dose of 150 mg/kg. Mice of varying weights were assigned to each cage so that the samples were heterogeneous within each group. The animal health was assessed by a veterinarian. Sanitary monitoring of the colony was performed quarterly based on a list of pathogens as recommended by the Federation for Laboratory Animal Science Associations. All the animals included in this study are considered specific pathogen free. The Mice Health Monitoring Report can be seen in Supplementary material Figures S1, S2 and S3 online.
Bioluminescent bacterial strain
Bioluminescent methicillin-resistant S. aureus USA300 strain NRS384-lux (SAP231) was obtained from the FDA – Center for Biologics Evaluation and Research and donated by Dr Roger Plaut. SAP231 consists of an epidemic strain of MRSA (USA300) cloned with the lux operon, stably expressing luminescent signal, being indicated for the monitoring of infections in murine model using this bacterium. 10
Bacterial cultivation
S. aureus (MRSA) SAP 231 was cultivated using the streaking method in Luria Bertani agar medium plus 10 µg/ml of chloramphenicol overnight at 37°C. Isolated colonies were selected and added to Luria Bertani broth with 10 µl/ml of chloramphenicol at 200 rec/min at 37°C until turbidity was reached. Afterward, 200 µl of the pre-inoculum (1:100) was transferred to a new medium under the same conditions as previously described, until reaching D.O.600 0.8–1.0. The inoculum was centrifuged (5 min at 10,000 rev/min), the supernatant was discarded, the pellets were washed in phosphate-saline buffer (PBS) and the process was repeated. After the final step, the pellet was resuspended in PBS (1:1), corresponding to approximately 1011 bacteria/ml.
Bacterial quantitation
At the end of the experiment, the nose cavities of all mice were surgically removed for bacterial quantification. Each organ was macerated and a serial dilution was performed, followed by the seed of all dilutions by the streak method and the plates were incubated in plates with Luria Bertani agar medium plus 10 µg/ml of chloramphenicol. The plates were incubated at 37°C overnight and the colonies were quantified after 24 h.
Nasopharyngeal murine colonization protocol
Freshly prepared inoculum was administered intranasally (5 µl per nostril) through droplet aspiration, totaling 10 µl per dose (corresponding to approximately 109 colony-forming units (CFU)). One group per model received one dose of the inoculum, while the other group received two doses within 1 h apart, with a maximum of 20 µl administered to the animal. Animals were anesthetized via the inhalatory route with 2% isoflurane before inoculum administration and acquisition of each bioluminescent image. An intramuscular mix of ketamine 10% (200 mg/kg) and xylazine 2% (20 mg/kg) was administered before euthanasia and organ collection at the end of the experiment. To follow the 3Rs principle, the initial tests were carried out with three animals per group and subsequently submitted for confirmatory tests. The group with the best results was subjected to a trial with 10 animals for statistical validation of the results.
Bioluminescent in vitro imaging
To estimate the number of bioluminescent signal-emitting bacteria detected using the IVIS® Lumina XRMS Series III (Perkin Elmer®) equipment, in vitro serial dilutions were performed using SAP 231. The bacteria were grown as previously described. Serial dilutions were performed and 100 µl of each dilution (102–108 CFU) was added in duplicate in a 96-well black dish. Images were captured using IVIS equipment. The images were analyzed using IVIS® in vivo Imaging Software and Aura Software (Spectral Instruments Imaging©), and the regions of interest (ROIs) acquired from the bioluminescent areas were plotted in graphs using Excel (Microsoft). A graph relating the number of bacteria to signal strength (ROI) was constructed.
Bioluminescent in vivo imaging
The mice were imaged after intranasal inoculation with S. aureus (MRSA) SAP231. The images were obtained using the IVIS LuminaXRMS Series III (PerkinElmer®) in vivo imaging system, as previously established (1, 26, 48, 50, 68, 120, 140 h after the last inoculum). The images were analyzed by IVIS® Living Image 4.7.3 (64 bits) and Aura Software (Spectral Instruments Imaging©) the ROIs acquired from bioluminescent areas were plotted in graphs using Excel (Microsoft). Images were acquired in dorsoventral, laterolateral and ventrodorsal positions.
Statistical analysis
Statistical analyses were performed using GraphPad Prism software, version 5. The non-parametric Mann–Whitney U test was used for bacterial burden analysis. Statistical significance was set at p < 0.05 value.
Results
Bioluminescent signal emission (BLSE) in vitro
To establish the minimal bacterial number capable of emitting a bioluminescent signal detectable in vitro by the IVIS system, serial dilutions of S. aureus (MRSA) SAP231 were performed, and luminescence (ROI) was plotted against the bacterial count. A graph shows the correlation between the bacterial population and signal emission (BLSE) captured by the IVIS system (ROI) (Figure 1). A visible bioluminescent signal was observed up to approximately 106 bacteria; however, the IVIS equipment was able to detect ROIs up to approximately 104 bacteria (Figure 2). Wells G11 and H11, containing PBS, were used as negative controls.
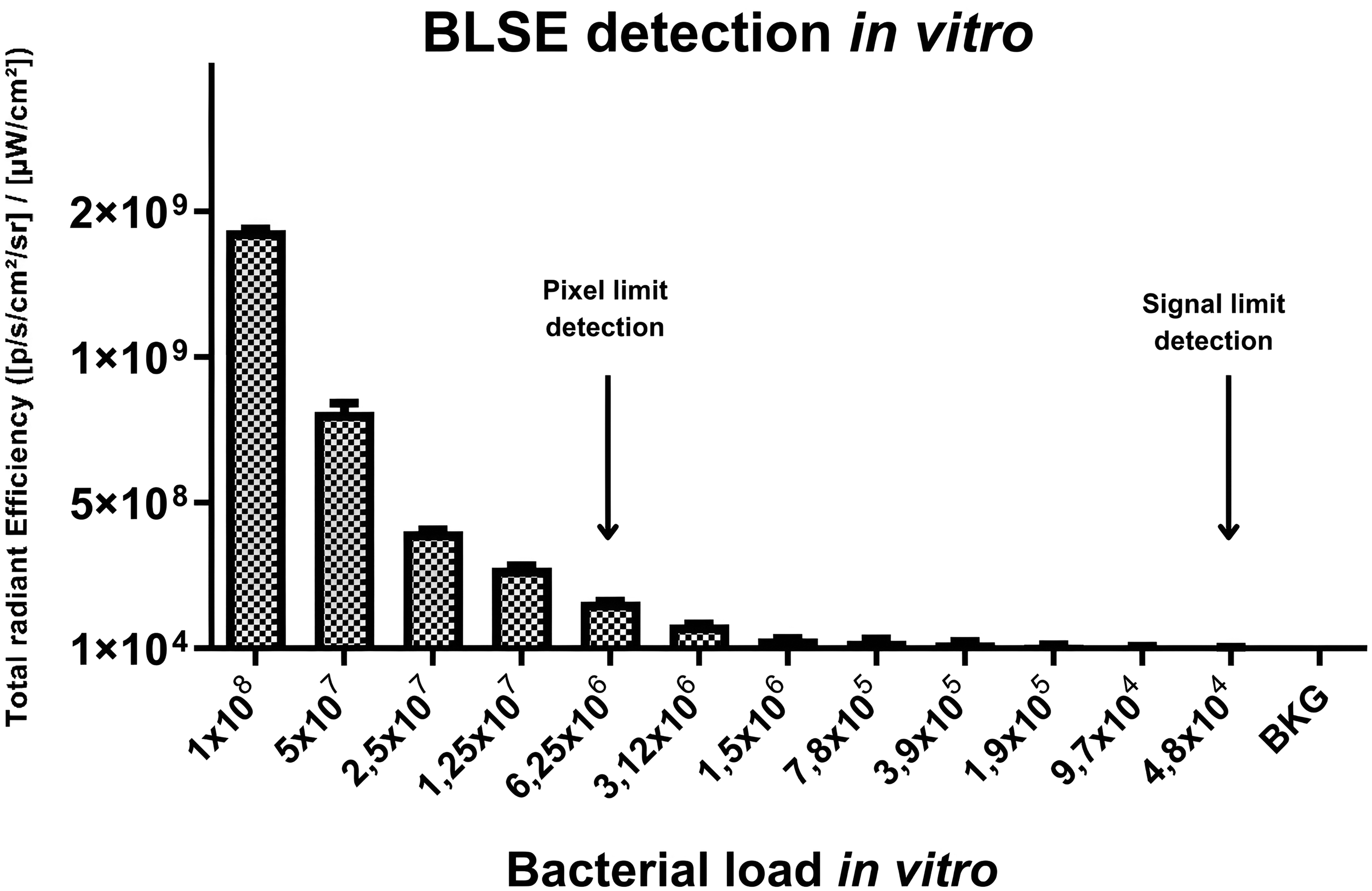
In vitro assessment of the bioluminescence signals emitted by bacteria. Relationship between bioluminescence emitted by bacteria grown in vitro. Pixel (visible) and signal detection are indicated by rows. BLSE: bioluminescent signal emission; BKG: background (phosphate-saline buffer sample).
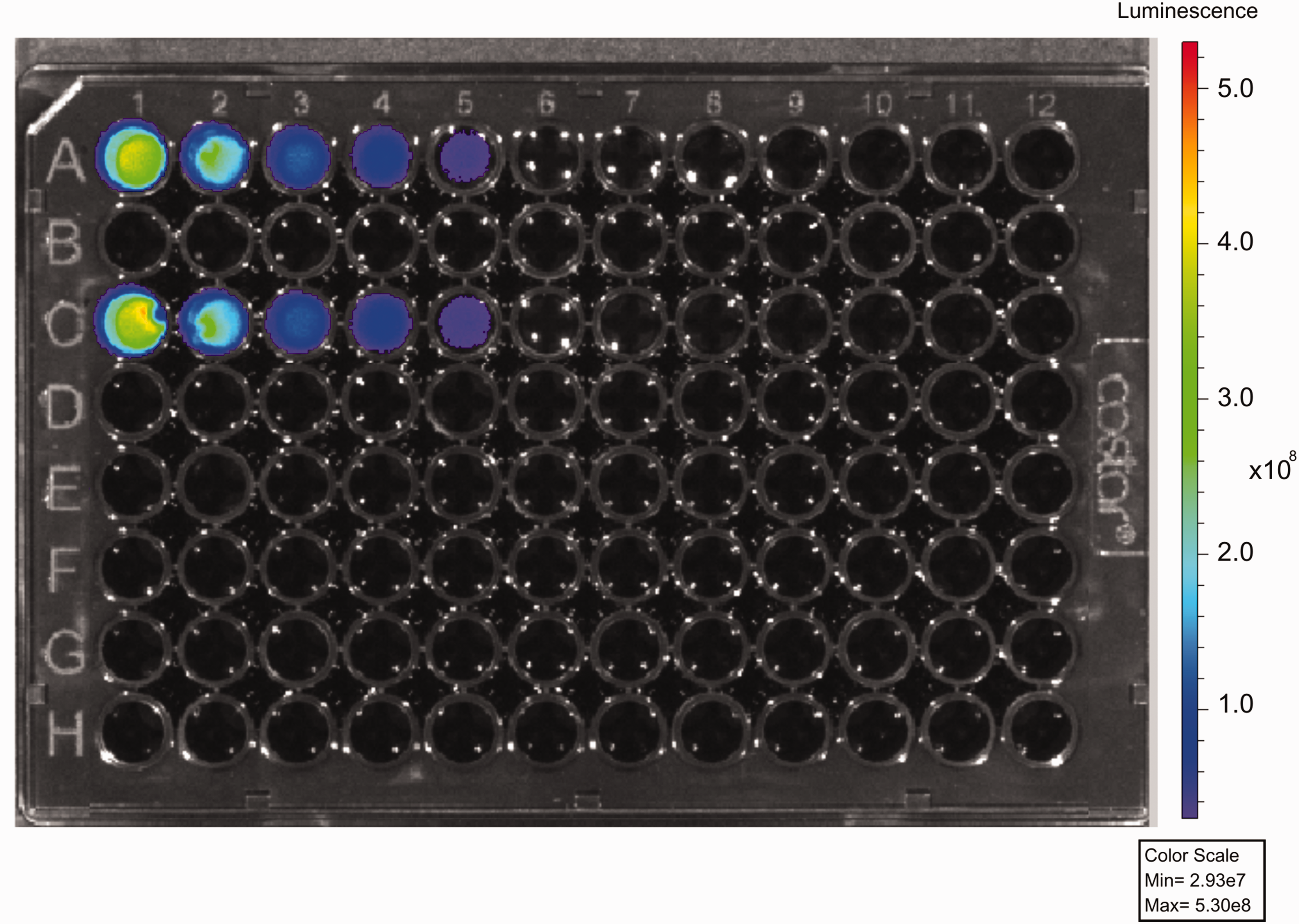

Image of a 96-well plate demonstrating the bioluminescent signal at different bacterial loads in duplicate (lines A and C). A1/C1: 1 × 108 bacteria; A2/C2: 5 × 107 bacteria; A3/C3: 1.2 × 107 bacteria; A4/C4: 6.2 × 106 bacteria; A5/C5: 3.1 × 106 bacteria; A6/C6: 1.6 × 106. H11/G11: negative control. Min: minimum; Max: maximum.
BLSE in vivo
BLSE by the murine strain was investigated after nasal colonization with SAP231 by image capture at different times until signal extinction. Swiss Webster animals showed the longest colonization, with some animals showing BLSE up to 140 h with two doses, and 120 h with one dose. However, we did not observe a homogeneous colonization pattern; some animals of this strain had decolonized before this period. In the first assay (n = 3), 100% of the animals exhibited BLSE after 140 h (Figure 3). In addition, BLSE was higher in this strain, with one or two doses. BALB/c and C57BL/6 strains did not show a long period of nasal colonization, presenting BLSE consistent until 48 h, independent of the number of doses administered. BLSE was consistently observed for approximately 26 h for all mouse strains, independent of the number of doses. The intensity of BLSE by ROIs was higher in Swiss Webster inoculated with the two doses (Figure 4). A second assay using Swiss Webster strains was performed with 10 animals to validate the long colonization time and for statistical analysis. In this study, we observed statistical significance (p < 0.05) for BLSE at 48 h and non-significance until 180 h. (Supplementary Figure S1). This data provides insights into the fold changes observed between the groups with one dose (1D) and two doses (2D) across different murine lines and time points. The number of fold changes is represented in the Supplementary file (Table S1). All animals showed BLSE at 48 h, and only two animals maintained BLSE at 180 h.
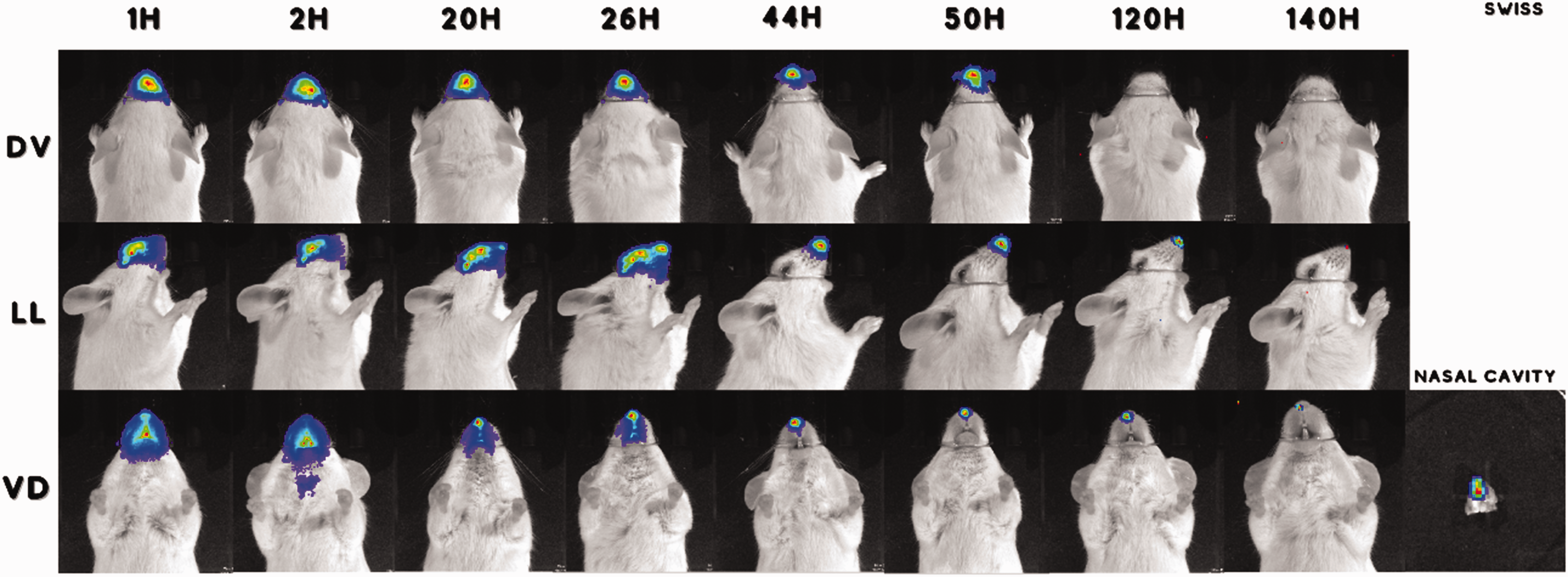
Representative IVIS images captured from Swiss Webster animals inoculated with two doses of SAP 231 at different times and positions. Nasal cavity excised after euthanasia, showing bioluminescent signal emission. DV: dorsoventral; LL: laterolateral; VD: ventrodorsal; H: hours.
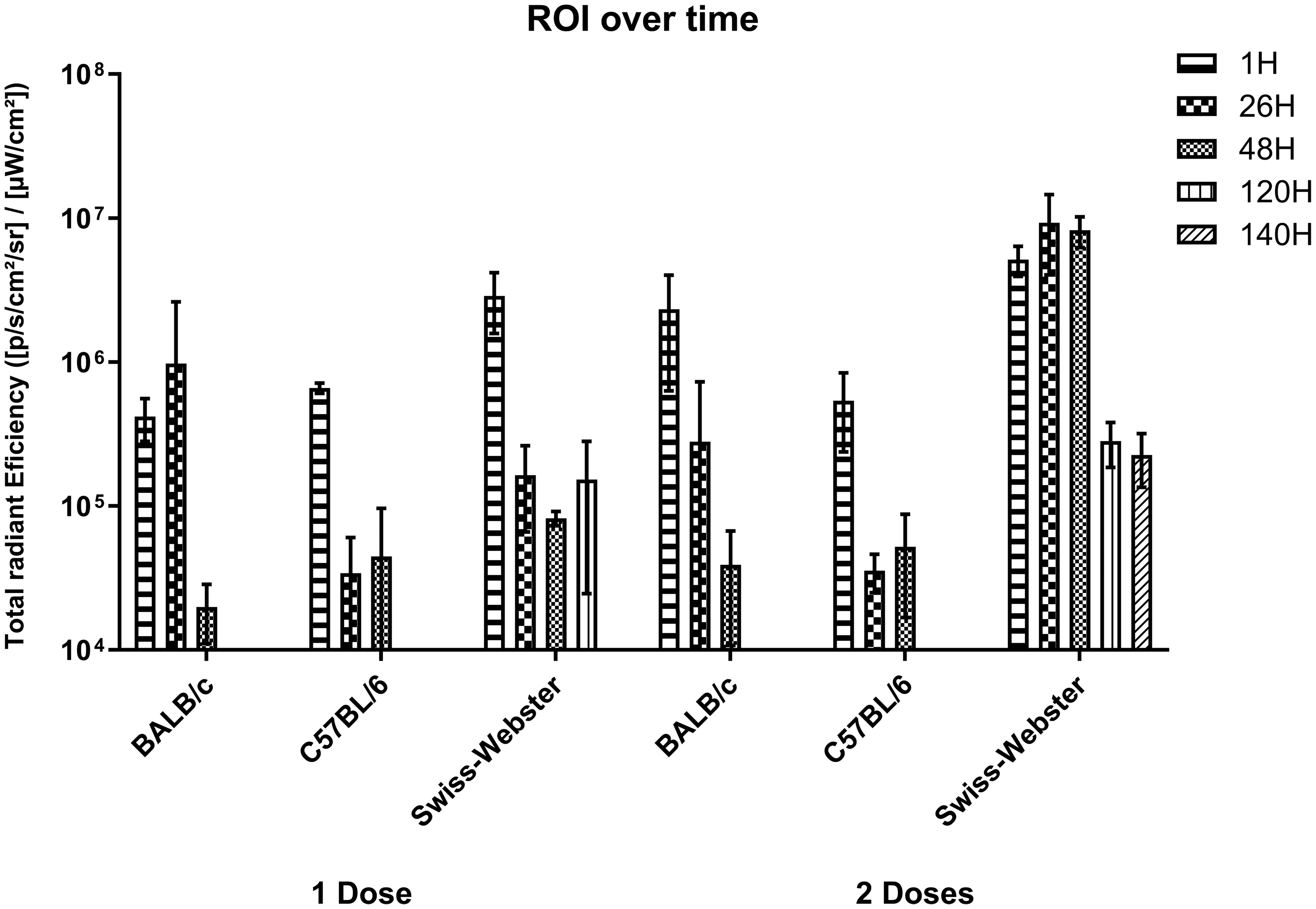
Bioluminescence by region of interest (ROI) for different murine strains (three animals per group), doses and times of nasal colonization estimated by bioluminescent signal emission; the standard deviation is represented in each bar. Swiss Webster line showed a consistent increase in values for the two-doses group across all time points. At 26h, the two-doses group showed values 56.59 times the one-dose group, peaking at 99-fold by 48h and maintaining a 22.60-fold increase at 140h.
In vivo BLSE by different animal positions
To evaluate the importance of the position in image capture and BLSE, three different positions were evaluated at different image capture times. We observed that the images taken in the dorsoventral position emitted the lowest signal and, in addition, it was the first position to completely lose the bioluminescent signal. These results were observed in all animal strains evaluated in this study. An example of BLSE captured at different positions is shown in Figure 2.
Bacterial load after in vivo BLSE
To determine the number of bacteria present in the nasal cavity after loss of BLSE, bacterial quantitation was performed after euthanasia. A mean of 104 CFU was recovered from BALB/c and C57BL/6 mice with one dose; Swiss Webster strain showed fewer bacteria (102 CFU) recovered with one dose. However, with two doses, Swiss Webster mice presented more bacteria recovered from the nasal cavity (mean of approximately 105 CFU) than C57BL/6 (mean of 104 CFU) and BALB/c (mean of 103 CFU) strains. The results are shown in Figure 5.
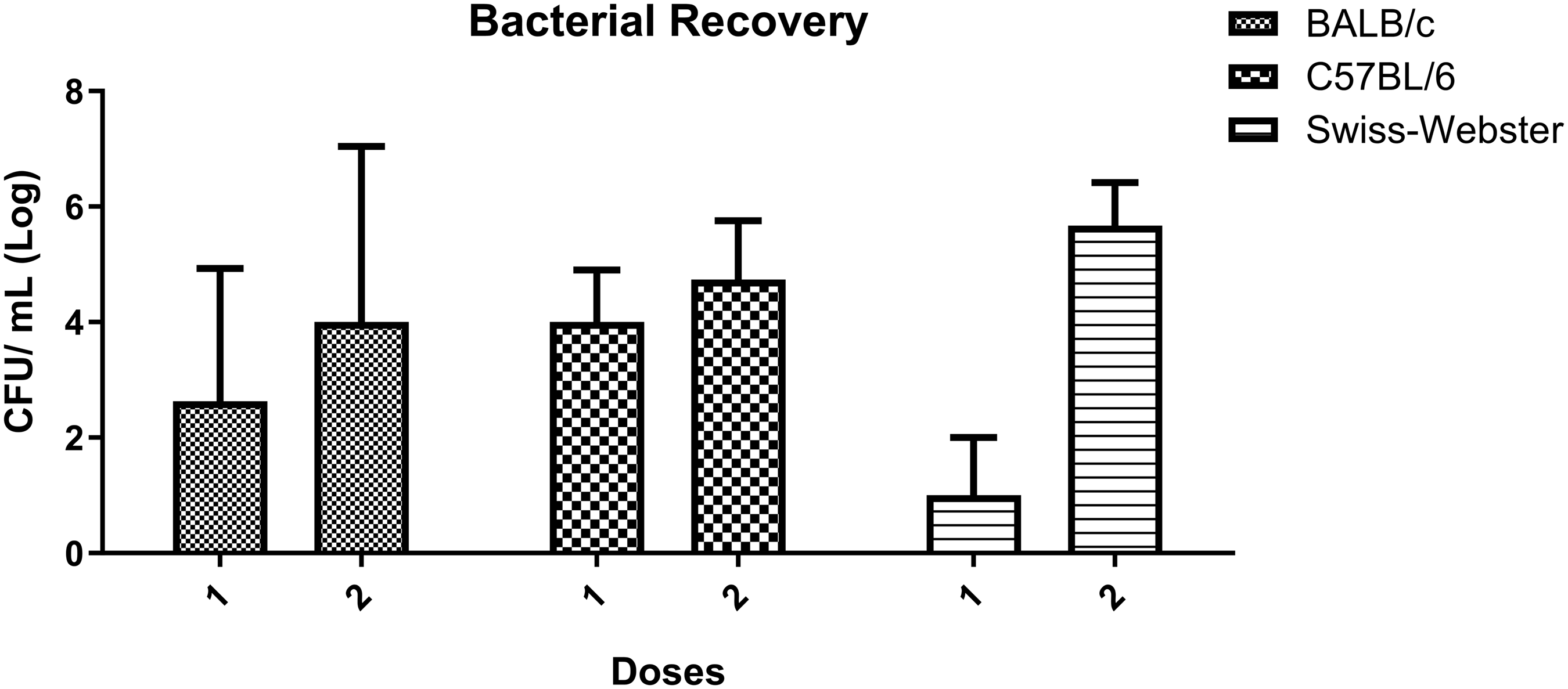
Bacterial recovery from the nasal cavity following bioluminescent signal emission loss. Each bar shows the log mean of the bacteria recovered from the three animals/groups with each standard deviation represented. BALB/c 1D (mean 4.0 one dose (1D), and 2.8 two doses (2D)), C57BL/6 (mean 4.0 1D, and 4.7 2D) and Swiss Webster (mean 2.0 1D, 5.7 2D). CFU: colony-forming units
Discussion
Mouse models serve as an essential component of preclinical research, offering economical, practical and ethical means of examining intricate biological processes and evaluating potential therapeutic strategies. Among the extensively employed mouse strains in scientific inquiry, BALB/c, C57BL/6 and Swiss Webster mice are distinguished by their meticulously established genetic profiles and distinct physiological properties, rendering them suitable for a broad spectrum of studies. These mouse strains have demonstrated efficacy in advancing our understanding of disease pathogenesis and in facilitating the development of innovative drug discoveries.
The concepts of bioethics make imperative an alternative approach to the use of animals in research to replace them when possible, reducing the number to the minimum necessary, and refining strategies to minimize animal suffering. 12 The use of in vivo bioluminescent MRSA strain (SAP 231) allowed us to visualize bacteria and estimate bacterial load in live mice and in real time. This feature provides the ability to track the progression of nasal colonization in each animal over time. Moreover, we managed to obtain a measurement of the bacterial quantity at any predetermined time, reducing the number of animals that needed to be euthanised. 10
Based on previous studies using bioluminescent imaging in mice to assess nasal colonization persistence, 9 we evaluated two isogenic (BALB/c and C57BL/6) and one non-isogenic mouse (Swiss Webster) strain and different inoculum levels.
Differences in nasal colonization patterns were observed between mouse strains and protocols using one or two doses. Since the murine nasal cavity has a volume of approximately 30 µl, 13 no more than 20 µl was used to prevent the bacterium from reaching other tissues, such as the lung. In all group protocols using two-dose mice, more bacteria were recovered and were able to increase the time and intensity of BLSE compared with the one-dose group in this study.
Swiss Webster mice presented with the longest time of nasal colonization and BLSE, but in a heterogeneous manner. Some animals remained colonized for a long time, while others lost bacteria quickly. In the second assay, two animals showed BLSE for up to 260 h. BALB/c and C57BL/6 mice presented a shorter time of nasal colonization than Swiss Webster, but in a more homogeneous pattern, staying consistently colonized for approximately 48 h.
Swiss Webster outbred mice are used widely in drug and vaccine development because they share many genetic properties with humans, which provides susceptibility to develop genetic conditions and diseases, as well as a comparative immune response. Heterogenic strains are composed of multiple genetic backgrounds and are more representative of the general population than inbred strains. In addition, its high reproductive rate, short life span and easy handling make it a suitable model for nasal decolonization strategies. 14 Inbred mice strains (BALB/c and C57BL/6) allow experimenters to vary only the parameters of interest and measure their effects (excluding genetic variation), which is important for uncovering causal factors and allowing for experimental reproducibility. 15
A previous study on nasal colonization using the BLI approach was conducted by evaluating only one image capture position. 9 In this study, we evaluated three different positions, with increased signal (BLSE) intensity and colonization time for the laterolateral and ventrodorsal positions. These results are probably because thicker bone tissue from the cranial cavity makes it difficult to capture bioluminescent signal in the dorsoventral position, as previously demonstrated by Chenchen et al., analyzing brain images after intranasal inoculation of fluorescent-labeled PEG-PLGA nanoparticles. 16
Although a similar model of nasal colonization using a bioluminescent S. aureus strain has been previously reported, 9 our study provides a more detailed protocol for evaluating mouse strains, doses and image capture positions. These results allowed us to consistently monitor nasal colonization by BLSE, estimating the bacterial burden present in live animals until 48 h in all strains evaluated. In addition, after the loss of signal emission, the bacterial load can be measured by bacterial quantification to validate the results obtained by BLSE.
Despite the long period of nasal colonization observed in some Swiss Webster strains, the heterogeneity of the nasal colonization period does not allow for the use of this strain as a model of choice. An alternative to improve Swiss Webster mice as a S. aureus nasal colonization model would be the mating of animals previously identified as colonizers, in order to select daughter lineages capable of maintaining a longer time of nasal colonization. Mouse-adapted S. aureus strains have been studied and proposed to be used as a tool for studying mucosal colonization and for screening novel vaccines and therapies directed at preventing colonization; 17 however, to our knowledge, no study aiming to select colonizer mouse strains has been described until now.
Supplemental Material
sj-pdf-1-lan-10.1177_00236772231209790 - Supplemental material for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA)
Supplemental material, sj-pdf-1-lan-10.1177_00236772231209790 for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA) by Juliana G da Silva, Juliana PC Boechat, Bruno DJ Silva, Rodrigo Müller and José PM Senna in Laboratory Animals
Supplemental Material
sj-pdf-2-lan-10.1177_00236772231209790 - Supplemental material for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA)
Supplemental material, sj-pdf-2-lan-10.1177_00236772231209790 for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA) by Juliana G da Silva, Juliana PC Boechat, Bruno DJ Silva, Rodrigo Müller and José PM Senna in Laboratory Animals
Supplemental Material
sj-pdf-3-lan-10.1177_00236772231209790 - Supplemental material for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA)
Supplemental material, sj-pdf-3-lan-10.1177_00236772231209790 for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA) by Juliana G da Silva, Juliana PC Boechat, Bruno DJ Silva, Rodrigo Müller and José PM Senna in Laboratory Animals
Supplemental Material
sj-pdf-4-lan-10.1177_00236772231209790 - Supplemental material for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA)
Supplemental material, sj-pdf-4-lan-10.1177_00236772231209790 for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA) by Juliana G da Silva, Juliana PC Boechat, Bruno DJ Silva, Rodrigo Müller and José PM Senna in Laboratory Animals
Supplemental Material
sj-pdf-5-lan-10.1177_00236772231209790 - Supplemental material for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA)
Supplemental material, sj-pdf-5-lan-10.1177_00236772231209790 for Monitoring Staphylococcus aureus nasal colonization murine model using a bioluminescent methicillin-resistant S. aureus (MRSA) by Juliana G da Silva, Juliana PC Boechat, Bruno DJ Silva, Rodrigo Müller and José PM Senna in Laboratory Animals
Footnotes
Acknowledgements
The authors would like to thank all of the technicians of the LAEAN (Animal Experimentation Laboratory) for their collaboration and the LATER (Recombinant Technology Laboratory), where bacterial samples were prepared. We are grateful to Dr. Renato Marchevsky, from Laboratório de Neurovirulência, at Instituto de Tecnologia em Imunobiológicos – Bio-Manguinhos/Fiocruz, who kindly reviewed the final revision of this manuscript. We are also grateful to Dr. Roger Plaut (FDA, USA), who kindly provided the bioluminescent bacteria (SAP231) used in this study.
Data access
We declare, to who may be interested, that access to all of the data of this manuscript can be obtained from the corresponding author, José Procópio Senna, by email
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Bio-Manguinhos/Fiocruz.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
