Abstract
Pasteurellosis is a well-recognized disease with similar pathology in all laboratory rodent species. Pasteurella pneumotropica is the most frequently mentioned member of the Pasteurellaceae isolated from mice and rats. Numerous other Pasteurellaceae taxa have been obtained from mice, rats, and other rodent species. Recently, rodent Pasteurellaceae have been submitted to comprehensive genetic and phenotypic (polyphasic) taxonomic studies. As a result they are now classed within six validly published new genera, namely Cricetibacter, Mesocricetibacter, Mannheimia, Muribacter, Necropsobacter, and Rodentibacter. All previously used names such as P. pneumotropica have become obsolete. The new classification forms a firm basis for the correct phenotypic identification of Pasteurellaceae from laboratory animals and for the selection of strains for pathogenicity studies. Consequences of taxonomic changes notably involve molecular methods used for the detection of Pasteurellaceae infection in laboratory animal colonies. Testing may be done using primer sets that detect all Pasteurellaceae taxa or sets developed to detect host-specific members of the family.
Detection
Pasteurellosis is a well-recognized disease with similar pathology in all laboratory rodent species. Pasteurella pneumotropica is the most frequently mentioned member of the Pasteurellaceae isolated from mice and rats. P. pneumotropica has been separated into the biotypes Jawetz and Heyl, which later have been recognized as two separate species. 1 Other Pasteurellaceae taxa isolated from healthy and diseased laboratory rodents include Actinobacillus muris, 2 Haemophilus influenzae murium, 3 SP-group Pasteurella, 4 and Bisgaard Taxa 17, 5 22, 6 23, 7 24, 25,6,8 and 41. 8
All early publications claiming the isolation of a “new” taxon as a member of the Pasteurellaceae family were invariably based on phenotypic characteristics.
From around 1980, DNA–DNA hybridization studies have delineated the Pasteurellaceae and at that time three genera were recognized (Haemophilus, Actinobacilius, and Pasteurella).1,9 After the introduction of 16S ribosomal RNA (16S rRNA) sequencing into phylogenetic studies, 10 the structure of the Pasteurellaceae has been explored in more detail. 11
Based on a variety of genotypic and phenotypic tests, 12 at present more than 25 genera have been described. 13
In the last 6 years, Pasteurellaceae taxa from rodents have also been submitted to comprehensive genetic and phenotypic (polyphasic taxonomic) studies. As a result, rodent isolates have been classed within six new genera, namely Cricetibacter, 14 Mesocricetibacter, 14 Mannheimia, 15 Muribacter, 16 Necropsobacter, 8 and Rodentibacter. 17 All these publications refer to the many reports describing the isolation of the bacteria from healthy and diseased rodents.
Rodent Pasteurellaceae classified in the genera Cricetibacter, Mannheimia, Mesocricetibacter, Muribacter, Necropsobacter, and Rodentibacter.
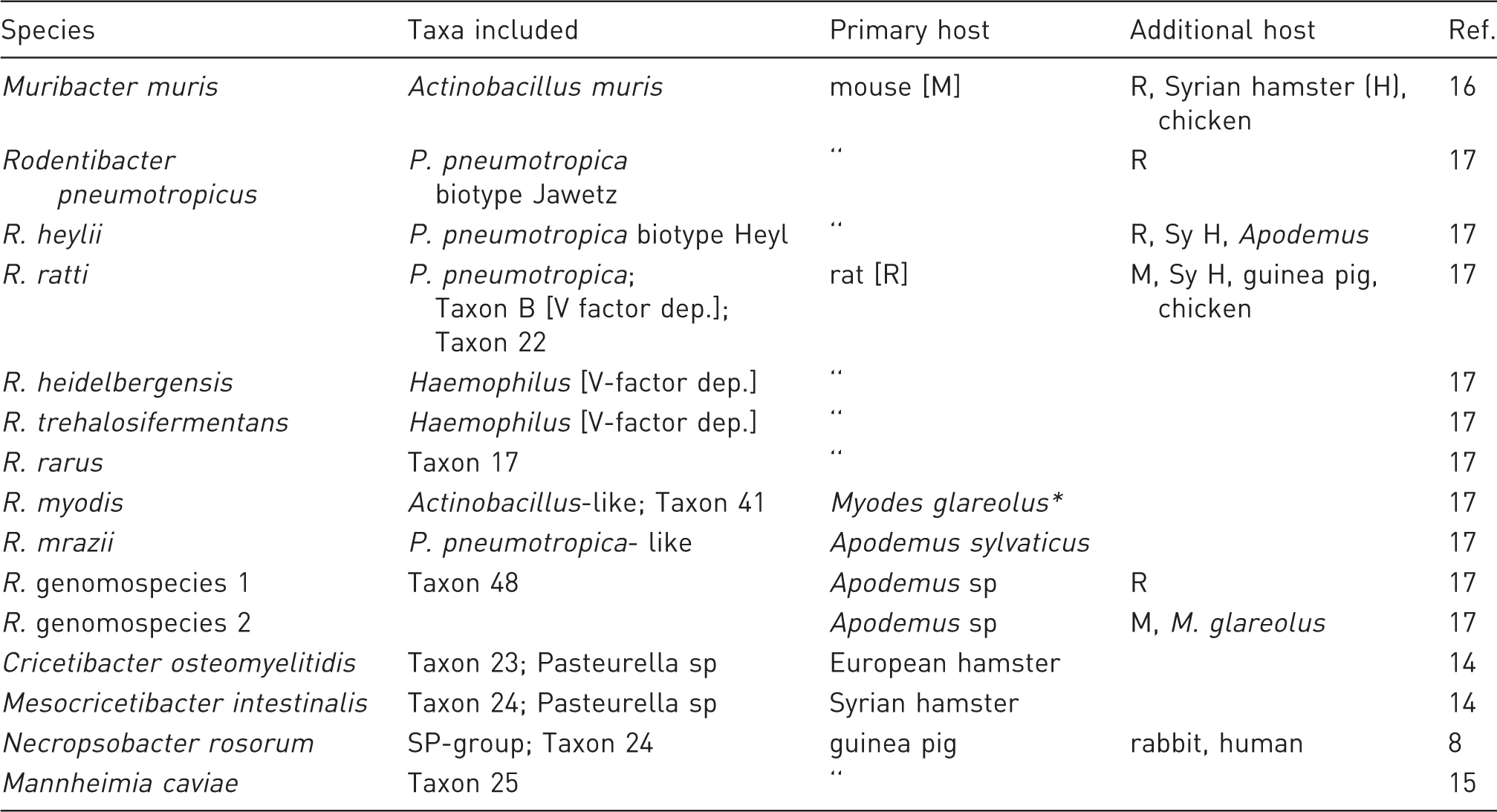
Previously Clethrionomys glareolus.
The new classification forms a firm basis for the correct phenotypic identification of Pasteurellaceae from laboratory animals and for the selection of strains for pathogenicity studies.
Consequences of changes in the taxonomy of rodent Pasteurellaceae involve methods used for the detection of Pasteurellaceae infection in laboratory animal colonies.
All “new” species can be cultured and differentiated from close relatives using classic biochemical tube testing methods. 19 Biochemical identification systems such as API, Rapid-ID, Vitek, etc. will be inferior for proper identification of rodent Pasteurellaceae as long as the newly described species are lacking in databases.
Recently, matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF) has been found a fast and cost-effective tool for the proper identification of a collection of 15 genera and 40 species of clinically relevant Pasteurellaceae. 20 Isolates representing R. pneumotropicus, R. heylii, R. myodis, R. ratti, R. rarus, R. heidelbergensis, and Rodentibacter genomospecies 2 could be identified to the species level. 17
Major consequences exist for the outcome of molecular detection, notably by polymerase chain reaction (PCR) analysis. Primer sets have been described for the detection of all Pasteurellaceae, 21 and separate species such as A. muris, now M. muris, 5 and both P. pneumotropica biotypes, now R. pneumotropicus and R. heyli, respectively.22,23
Most animal testing laboratories test samples from all rodent species for the presence of P. pneumotropica and occasionally for P. multocida, a bacterium that does not naturally infect rodents, which is fully clear from the fact that none of the around 1000 rodent Pasteurellaceae isolates studied by Adhikary et al. was classed as P. multocida. 17 Some laboratories offer testing for biotypes Jawetz and Heyl P. pneumotropica, but none offers testing for A. muris (M. muris). Until now testers do not take into account that different rodent species host different Pasteurellaceae species (Table 1). Likely laboratory rodents may be tested using Bootz’s primer set, which is suitable for the detection of all Pasteurellaceae, 24 or using primer sets developed to detect the taxa that are known to infect the different rodent species. Primer sets developed to detect both P. pneumotropica biotypes (now R. pneumotropicus and R. heyli) will not detect other taxa. Testing laboratories need to specify the PCR primer sets used as these will determine which of the newly described rodent Pasteurellaceae may be detected during health monitoring screening.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
