Abstract
Inbred mouse strains are widely used for genetic studies because of the isogenicity within a strain or F1 hybrid and the genetic heterogeneity between inbred strains. In the process of modifying Disc1 in the mouse genome, a 25-bp deletion was discovered in exon 6 of the gene in the 129S6/SvEvTac inbred strain, and subsequently in 16 other inbred strains in the category known as ‘Castle’s mice’. The deletion (Disc1del ) induces a frame shift that introduces a premature termination codon, which has been shown to confer an impairment in working memory. To extend knowledge of the distribution of Disc1del among the various inbred strains of laboratory mouse, we investigated whether Disc1del is present in the categories known as ‘Swiss mice’ and ‘strains derived from China and Japan’. We found that the FVB/NJ, SJL/J and SWR/J strains in the ‘Swiss mice’ category and DDY/JclSidSeyFrkJ in the ‘China and Japan’ category are homozygous for the Disc1del allele, while ICR/HaJ in the ‘Swiss mice’ category is homozygous for wild-type Disc1. Since the Disc1del -positive strains FVB and SJL are commonly used for the generation of transgenic mice, and thus contribute to the genetic background of multiple transgenic lines, our results may allow scientists to avoid the potential confounding effects of the Disc1del allele in transgenic studies of learning and memory.
DISC1 (Disrupted in schizophrenia 1) was directly implicated as a candidate gene for psychiatric illness through a balanced translocation t(1;11) that directly disrupts the DISC1 open reading frame and co-segregates hemizygously with schizophrenia and affective disorder in a large Scottish family. 1,2 Two subsequent resequencing studies found that ultra-rare heterozygous missense mutations in DISC1 are associated with an estimated attributable risk of about 2% in schizophrenia 3 and 0.5% in bipolar disorder, 4 confirming DISC1 as a promising susceptibility factor for major mental illnesses. The DISC1 orthologue in mice maps to the mouse syntenic region and shows conserved genomic structure. The mouse Disc1 and human DISC1 genes show 60% homology at the DNA level and 56% identity and 14% similarity at the protein level. 5 Although the amino acid sequence of DISC1 is diverging rapidly, several protein motifs are conserved between the orthologues, including leucine zipper, nuclear localization, and coiled-coil domains. 6
Animal models can be valuable tools for exploring the underlying pathologies of human diseases and developing better therapies. Although it is difficult to model a psychiatric disorder like schizophrenia in animals (e.g. hallucinations and delusions are likely to be human specific), anatomical, behavioural and cognitive characteristics have been described in patients, and it is feasible to study these in animal models. 7 To study the consequences of altered DISC1 function on endophenotypes involved in psychiatric illness, efforts have been made to modify Disc1 in the laboratory mouse genome. In the process of a gene targeting manipulation of Disc1 in 129S6/SvEvTac-derived embryonic stem (ES) cells, a 25-bp deletion was discovered in exon 6 of the gene. 8 The deletion (Mouse Genome Informatics nomenclature: Disc1del ) induces a frame shift that introduces a premature termination codon in exon 7. 8 Disc1del does not affect Disc1 transcript abundance in 129S6/SvEvTac (129S6) mice, 8 but it is unclear what happens at the protein level. Western blotting with an antibody raised against the N-terminal region of Disc1 (amino acids 272–537) detected an ≈100 kDa band in total brain protein extracts from 10-day-old C57BL/6 J mice but not from 10-day-old 129S6 mice or 2-month-old mice of either strain, 8 suggesting that the deletion affects an isoform of Disc1 that is not expressed in adulthood. Mouse Disc1 has multiple isoforms at both mRNA and protein levels. 6,9 The absence of an ≈100 kDa protein band in brain tissue from 3-day-old Disc1del homozygotes has been confirmed by Western blotting with a C-terminal antibody (amino acids 131–170). 10 However, a collaborative study using a variety of antibodies generated by independent groups showed almost identical immunoreactivity of Disc1 around 100 kDa (corresponding in size to full-length Disc1) in brain extracts from adult C57BL/6J and 129S6 mice of an unspecified age. 11 After backcrossing of the Disc1del allele onto the C57BL/6J inbred strain, behavioural analysis showed that heterozygotes and homozygotes display a consistent impairment in working memory in a T-maze test, 8 indicating a dominant effect of the deletion.
Inbred mouse strains have long been used for genetic and immunological studies because of the isogenicity within a strain or F1 hybrid and the genetic heterogeneity between inbred strains. The genealogical chart showing the origins and relationships of inbred strains (ftp://www.informatics.jax.org/pub/datasets/misc/genealogy/genealogy_key.html) assigns each strain to one of seven categories (A–G). 12 Initial screening of several inbred strains showed that Disc1del is absent from the AKR/J, BALB/cJ, C3H/HeJ, CBA/J and DBA/2J strains within the category known as ‘(B) Castle’s mice’, which descends from mice used in breeding experiments by William Castle, and from the C57BL/6J strain within the ‘(E) C57-related mice’ category that descends from an original pair of mice bred by Abbie Lathrop. 8,12 Subsequently, we found that Disc1del is absent from C57BL/6NTac within the ‘C57-related mice’, but is harboured by 16 other inbred strains within ‘Castle’s mice’, including the 101/RI, BTBR T + tf/J and LP/J strains 13 and 13 other 129 substrains 14 that are all derived from L C Dunn’s ‘mouse house’ at Columbia University. 15 These findings have been confirmed by recent next generation genomic sequencing of 17 inbred strains. 16 Most ES cell lines for gene targeting are derived from the 129 strain, so the potential presence of a genetic lesion known to confer a cognitive impairment in the heterozygous state 8 may have adverse effects on behavioural studies of 129 ES cell-derived gene-targeted mice. Screening of two outbred stocks has revealed that the Disc1del allele is also found within colonies of Slc:ddY and Slc:ICR mice in Japan. 10
Knowledge of the distribution of Disc1del among the genealogical chart of inbred strains widely used in biomedical research is far from complete, though, as only two of the seven categories of inbred strains have been assayed to date. The objective of this study was thus to establish whether Disc1del is present in representative strains of the categories known as ‘(A) Swiss mice’ that descend from albino mice imported to the USA from Lausanne in Switzerland 17 and ‘(C) strains derived from colonies from China and Japan’. Here we report that the DDY/JclSidSeyFrkJ, FVB/NJ, SJL/J and SWR/J inbred mouse strains harbour the Disc1del allele.
Materials and methods
DNA samples
DNA samples from individuals of the FVB/NJ (FVB; Stock # 001800), ICR/HaJ (ICR; Stock # 009122), SJL/J (SJL; Stock # 000686) and SWR/J (SWR; Stock # 000689) strains in the ‘Swiss mice’ category, the DDY/JclSidSeyFrkJ strain (DDY; Stock # 002243) in the ‘China and Japan’ category, the C57BL/6NJ strain (B6N; Stock # 005304) and the 129P3/J strain (129P3; Stock # 000690) were obtained from the Mouse DNA Resource of the Jackson Laboratory (Bar Harbor, ME, USA). An ear biopsy from an individual of the 129S9/SvEvH (129S9) inbred strain maintained by MRC Harwell (Didcot, UK) was obtained from the animal facility at St James’s University Hospital, Leeds, UK. Genomic DNA was isolated from the ear biopsies using a previously described proteinase K method. 18 Following the measurement of DNA concentration by ultraviolet absorbance using a NanoDrop 1000 Spectrophotometer (Labtech International, Lewes, UK), each sample was diluted to 20 ng/µL in ultra-pure polymerase chain reaction (PCR) grade water (Bioline Reagents, Cricklewood, UK).
PCR-based genotyping assays
To genotype Disc1 in the inbred mouse strains, the forward primer (5′-GCT GTG ACC TGA TGG CAC T-3′) and reverse primer (5′-GCA AAG TCA CCG CAA TAA CCA-3′) that anneal to parts of the Disc1 gene common to both the deletion and wild-type alleles, flanking exon 6 (the location of the 25-bp deletion), were used in PCRs with template DNA from the 129P3, DDY, FVB, ICR, SJL, SWR, 129S9 and B6N strains, using a previously described method. 14 The B6N and 129P3 strains were used as Disc1 wild-type and Disc1del controls, respectively. Following thermal cycling and agarose gel electrophoresis, the deletion allele was identified by an amplicon of 171 bp and the wild-type allele by an amplicon of 196 bp.
To verify the presence of the deletion or wild-type allele of Disc1 in each mouse strain, we used two further PCR-based assays. The deletion specific assay consisted of forward primer (5′-CTG ACC TCT GCC AAC CTG AG-3′) and reverse primer (5′-AGA GGA CAC TGC AGA CAG TTA GG-3′) that were designed to amplify a fragment of 248 bp from the deletion allele of Disc1 (129P3 strain) but with no amplification from the wild-type allele (B6N strain). Conversely, the wild-type specific assay consisted of forward primer (5′-AGA GCA GGG TTA ATG GCT GA-3′) and reverse primer (5′-GTG GCT CCA CCT GGA AGG-3′) that were designed to amplify a fragment of 266 bp from the wild-type allele of Disc1 (B6N strain) but with no amplification from the deletion allele (129P3 strain). Reactions having a total volume of 20 µL were carried out inside 0.2 mL eight-well tube strips (Starlab, Newport Pagnell, UK), with each PCR consisting of 7.75 µL ultra-pure PCR grade water (Bioline), 10 µL HotShot Mastermix (containing optimized mixture of Taq polymerase, anti-Taq polymerase monoclonal antibodies in 2× reaction buffer [6 mmol/L MgCl2] with 400 µmol/L dNTPs and stabilizer) (Cadama Medical, Stourbridge, UK), 0.5 µmol/L each primer (synthesized by Integrated DNA Technologies, Louvain, Belgium), and 20 ng genomic DNA. Reactions were incubated at 95℃ for 5 min, followed by 30 cycles of 94℃ for 30 s, 62℃ for 45 s (deletion assay) or 63℃ for 30 s (wild-type assay), and 72℃ for 40 s, followed by 72℃ for 10 min in a G-Storm GS482 thermal cycler (Labtech International). Following thermal cycling, each PCR product was mixed with 4 µL blue/orange 6× loading dye (Promega, Southampton, UK) before being loaded next to 0.5 µg 100-bp DNA ladder (Invitrogen, Paisley, UK) onto a 1.5% (w/v) molecular grade agarose gel (Bioline) containing 1× SYBR Safe DNA gel stain (Invitrogen), and electrophoresed in 1× TAE buffer (40 mmol/L Tris-acetate, 2 mmol/L EDTA) at 5 V/cm for 40 min.
DNA sequencing
To corroborate the results of the Disc1 PCR-based genotyping assays, amplicons encompassing the location of the 25-bp deletion (Disc1 exon 6) were excised from the gel with a scalpel and purified using a QIAquick gel extraction kit (Qiagen, Crawley, UK). Purified DNA bands at 2 ng/µL from the DDY, FVB, ICR, SJL and SWR strains were submitted to a commercial company (Eurofins MWG Operon, Acton, UK) for Sanger sequencing. Alignment and comparisons of the resulting DNA sequences with the published Disc1 sequences of the C57BL/6J (wild-type) and 129S6 (deletion) strains 8,14 were performed using Sequencher (Gene Codes, Ann Arbor, MI, USA) and BioEdit Sequence Alignment Editor 19 software.
Results
The PCR assay using primers that anneal to parts of the Disc1 gene common to both the deletion and wild-type alleles, flanking the location of the 25-bp deletion,
14
amplified fragments of 171 bp from the 129P3 (deletion control), DDY, FVB, SJL, SWR and 129S9 strains, and 196 bp from the B6N (wild-type control) and ICR strains (Figure 1a). Application of the Disc1 deletion specific PCR primers to the same inbred strains amplified DNA fragments of 248 bp from template DNA from the 129P3 (deletion control), 129S9, DDY, FVB, SJL and SWR strains, but showed no amplification from the B6N (wild-type control) and ICR strains (Figure 1b). In parallel, the Disc1 wild-type specific PCR primers amplified DNA fragments of 266 bp from template DNA from the B6N (wild-type control) and ICR strains, but showed no amplification from the 129P3 (deletion control), 129S9, DDY, FVB, SJL and SWR strains (Figure 1b). Sequencing of amplicons encompassing Disc1 exon 6, followed by alignment with the published Disc1 sequences of the deletion and wild-type alleles,
8,14,16
confirmed that FVB, SJL and SWR in the ‘Swiss mice’ category and DDY in the ‘China and Japan’ category are homozygous for the Disc1del
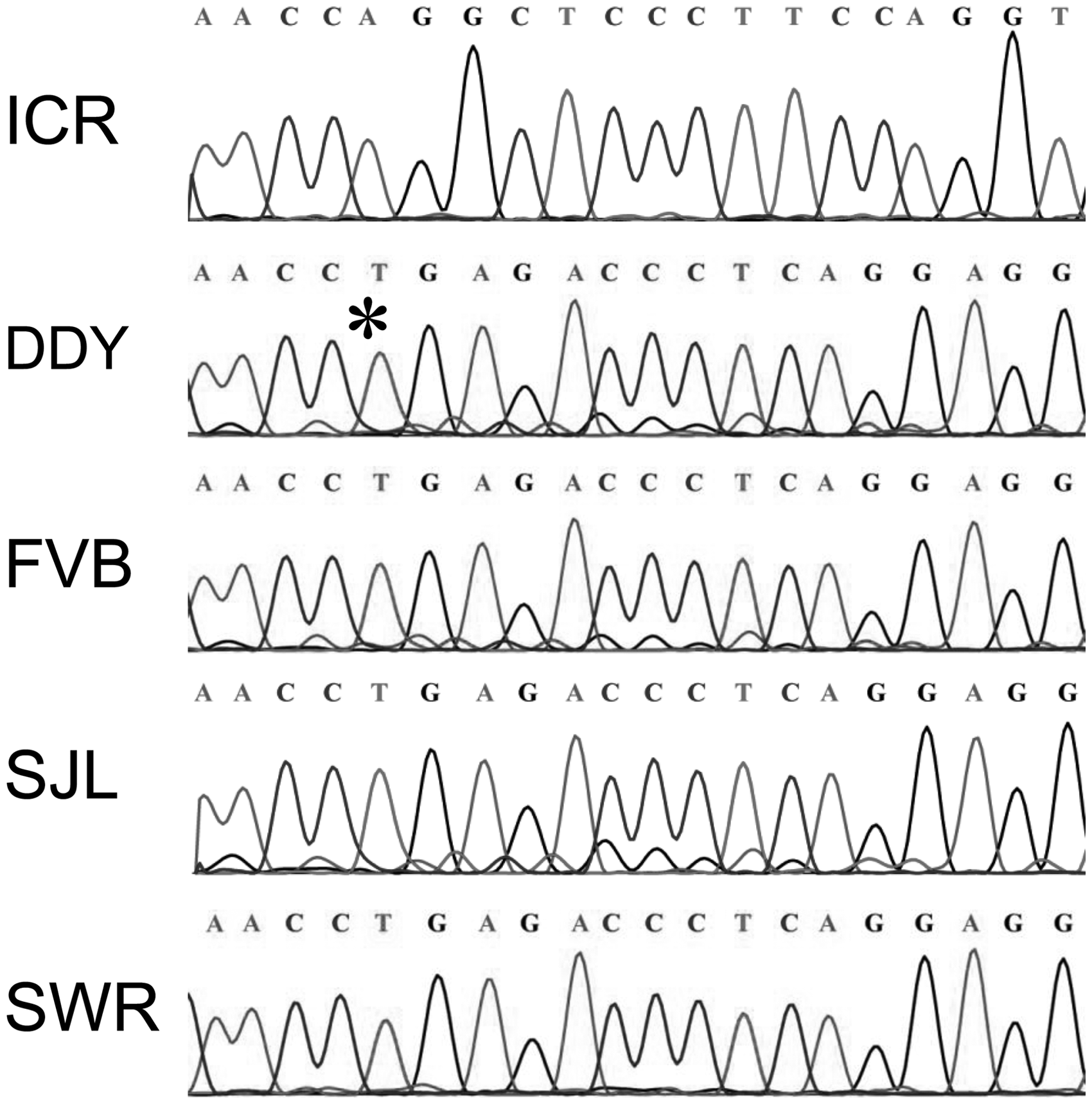
allele, while ICR in the ‘Swiss mice’ category is homozygous for wild-type Disc1 (Figure 2).
Disc1 deletion genotyping of inbred mouse strains. (a) Polymerase chain reaction (PCR) assay using primers common to both the deletion and wild-type alleles of Disc1 amplifies fragments of 171 bp from the deletion (129P3) and 196 bp from the wild-type (B6N) allele. The sizes of the DNA ladder fragments and PCR products are indicated on the sides of the figure. Ladder, 0.5 µg 100-bp DNA ladder (100 bp, 48 ng; 200 bp, 25 ng; 300 bp, 29 ng); 129P3, 129P3/J (deletion control); DDY, DDY/JclSidSeyFrkJ; FVB, FVB/NJ; ICR, ICR/HaJ; SJL, SJL/J; SWR, SWR/J; 129S9, 129S9/SvEvH; B6N, C57BL/6NJ (wild-type control). (b) PCR assays specific for the deletion or wild-type allele of Disc1. The B6N strain has a wild-type Disc1 allele, while the 129P3 strain has a 25-bp deletion (dots) in Disc1. The wild-type specific primers (F: 5′-AGA GCA GGG TTA ATG GCT GA-3′; R: 5′-GTG GCT CCA CCT GGA AGG-3′) amplify a fragment of 266 bp from the wild-type allele of Disc1 (B6N) but not from the deletion allele (129P3). The deletion specific primers (F: 5′-CTG ACC TCT GCC AAC CTG AG-3′; R: 5′-AGA GGA CAC TGC AGA CAG TTA GG-3′) amplify a fragment of 248 bp from the deletion allele of Disc1 (129P3) but not from the wild-type allele (B6N). Sequence confirmation of Disc1 deletion in inbred strains. The sequence chromatograms show the deletion in DDY, FVB, SJL and SWR, and the wild-type allele of Disc1 in ICR. An asterisk indicates the location of the deletion in the DDY sequence.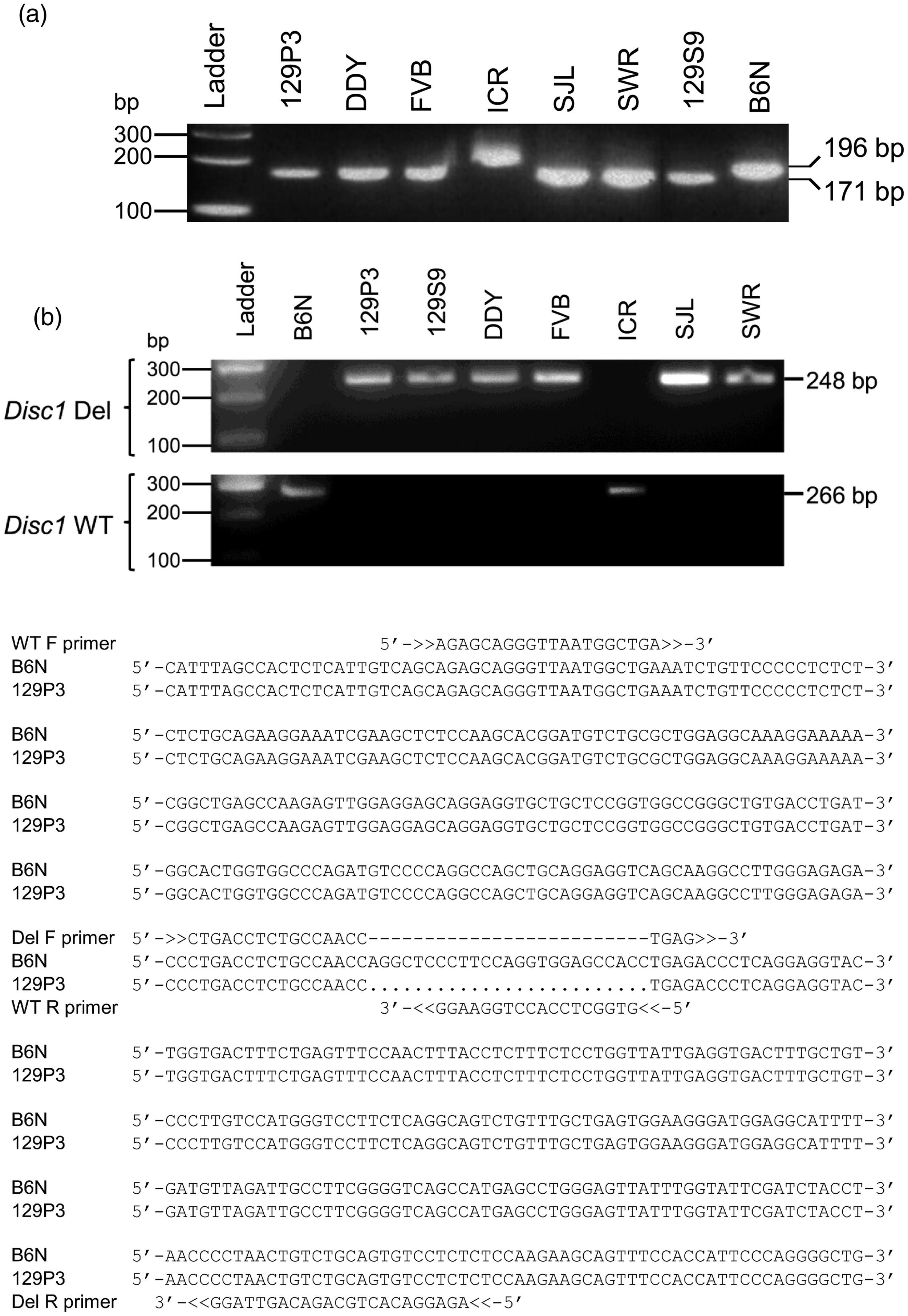
Discussion
The objective of this study was to establish whether the Disc1del allele is present in representative strains of the categories known as ‘(A) Swiss mice’ and ‘(C) strains derived from colonies from China and Japan’. Using three independent PCR-based genotyping assays, we found that FVB, SJL and SWR in the ‘Swiss mice’ category and DDY in the ‘China and Japan’ category are homozygous for the Disc1del allele, while ICR in the ‘Swiss mice’ category is homozygous for wild-type Disc1. These findings were confirmed by sequencing of PCR amplicons that encompass the location of the 25-bp deletion (exon 6) in the Disc1 gene. Since inbred strains are homozygous at every locus, and the alleles at each locus are identical by descent, we conclude that Disc1del is fixed in the FVB, SJL and SWR inbred strains, while wild-type Disc1 is fixed in the ICR inbred strain. We also found that the Disc1del allele is present in the 129S9 inbred strain, confirming fixation of Disc1del allele throughout the 129 family.
Our finding that the Disc1del allele is present in the categories of inbred strains known as ‘(A) Swiss mice’ and ‘(C) strains derived from colonies from China and Japan’, as well as in the category known as ‘(B) Castle’s mice’, 8,14,16 indicates that the Disc1del allele is more widespread than was previously thought. Although the genealogical chart of inbred strains does not indicate any links between the DDY strain, the inbred Swiss strains and Castle’s mice, 12 it is reasonable to postulate that the 25-bp deletion in the Disc1 gene arose in progenitor stock that contributed to the genetic make-up of these inbred strains as well as two commercially available outbred stocks in Japan (Slc:ddY and Slc:ICR). 10 The DDY inbred strain and the Slc:ddY outbred stock both descend from the same non-inbred ‘dd’ stock at the University of Tokyo. 12,20 This shared ancestry is consistent with the Disc1del allele being present in each of them. The FVB, SJL and SWR inbred strains and the Slc:ICR outbred stock all descend from two male and seven female albino mice derived from a non-inbred stock in the laboratory of Dr A de Coulon at the Centre Anticancéreux Romand, in Lausanne, Switzerland. These animals were imported into the USA by Dr Clara Lynch of the Rockefeller Institute in 1926. 12,21,22
The finding that the ICR inbred strain is Disc1 wild-type was unexpected in light of the other inbred Swiss strains tested (FVB, SJL and SWR) being Disc1del homozygous. The inbred ICR strain and the Slc:ICR outbred stock both descend from the same Ha/ICR progenitor stock established at the Institute of Cancer Research in 1948. 12,21,22 Since 22.5% of Slc:ICR mice tested were found to harbour the wild-type allele of Disc1 10 , it is reasonable to postulate that this allele was present in the common progenitor stock and became fixed in the inbred ICR strain during its development. Of the Disc1del -positive inbred Swiss strains, FVB and SJL are of particular importance as they are commonly used for the generation of transgenic mice, and thus contribute to the genetic background of multiple transgenic lines, 23 such as a mouse model of Alzheimer’s disease with an F1 hybrid C57BL/6J × SJL/J genetic background 24 and a model of Huntington’s disease with an FVB genetic background. 25 To avoid potential confounds resulting from the Disc1del allele, scientists using the FVB or SJL strains to generate transgenic mice for studies of brain function would be advised to genotype their transgenic lines for the Disc1 deletion.
Footnotes
Acknowledgements
This work was supported by a Medical Research Council grant (G0900625) and a Young Investigator Award from the National Alliance for Research on Schizophrenia and Depression to SJC.
