Abstract
Stigmoid bodies (SBs) are structures in the cytoplasm of neurons. SBs are mostly found in the hypothalamic region of the rat and contain a protein called huntingtin-associated protein 1 (HAP1). In a recent publication, large cytoplasmic structures were shown to be immunoreactive for a type I receptor called SorLA/LR11. By light microscopic analysis, these structures appeared similar to SBs in size and in brain regional and subcellular localization. To determine whether these large puncta correspond to HAP1-containing SBs, we used antibodies specific to various domains of the apolipoprotein receptor LR11 to perform immunocytochemistry in rat and mouse brain tissue. Transfection studies using HeLa cells were conducted to demonstrate the specificity of the antibodies. We found that, in both species, antibodies to the domain II (or VSP10 for vacuolar sorting protein 10 domain) of LR11 immunoreact with large cytoplasmic structures. Co-localization immunolabeling experiments in rat brain tissue sections and in neuron cultures showed that these LR11-immunoreactive structures correspond to HAP1-positive SBs. Electron microscopy was performed in rat hypothalamus and further demonstrated the presence of LR11 in SBs and its co-localization with HAP1. LR11-containing SBs were most abundant in the hypothalamus but were also found in many brainstem nuclei, thalamus, and hippocampus. Our data also show that sortilin, another transmembrane protein containing a VPS10 domain, localizes to large cytoplasmic puncta and is found in LR11-positive and Hap1-positive SBs in hypothalamic neuron cultures.
Keywords
S
In 1993, Shinoda and colleagues found that an antibody raised against a placental protein complex with aromatase activity (X-P2) detected SBs in the rat brain. X-P2-containing SBs were numerous in hypothalamic regions. However, the specific identity of the antigen present in the X-P2 complex that was detected in SBs was not determined. More recently, we have shown that multiple antibodies spanning the entire sequence of the huntingtin-associated protein 1 (HAP-1) detect SBs in rat brain (Gutekunst et al. 1998). HAP1 was first identified by yeast two-hybrid screening for its association with huntingtin, the protein mutated in Huntington's disease (Li et al. 1995). Using HAP1 antibodies as markers, studies have addressed the brain regional distribution of SBs in rat and mouse brains with light microscopic immunocytochemistry (Gutekunst et al. 1998; Martin et al. 1999). Similar to what had been described with X-P2, the greatest concentration of HAP1-containing SBs was found in the preoptic region, the paraventricular nucleus, the bed nucleus of the stria terminalis, the medial and central amygdala, the arcuate nucleus, and the subfornical organ (Gutekunst et al. 1998). HAP1 has been shown to have a role in microtubule transport, based on its association with motor proteins (Li et al. 1998b), and in neurite outgrowth (Li et al. 2000). More recently, HAP1 was shown to interact with hepatocyte growth factor-regulated tyrosine kinase substrate (hrs), suggesting a role in endosomal trafficking (Li et al. 2002). HAP1 proteins have been shown to bind to themselves, and transfection of HAP1A in HEK 293 cells resulted in the formation of large cytoplasmic inclusions (Li et al. 1998a). By electron microscopic analysis, these inclusions contain filamentous material but their ultrastructure does not resemble that of brain SBs (Li et al. 1998a).
A recent study determined that the apolipoprotein E receptor LR11 (lipoprotein receptor containing 11 LDL binding domains), also called SorLA (sorting protein-related receptor containing LDLR class A repeats), is localized to a large structure found in the perikarya of some neurons in the brains of rat and human (Motoi et al. 1999). Given the similarity in size and subcellular localization of these puncta, we sought to determine whether they corresponded to HAP1-containing SBs. In the present study we have developed and characterized polyclonal and monoclonal antibodies to HAP1 and have used these specific antibodies in conjunction with well-characterized monoclonal antibodies to the VSP10 domain of human LR11 in co-localization studies in brain tissue sections, transfected HeLa cells, and primary neuron cultures. We show by co-localization and electron microscopy studies that LR11 co-localizes with HAP1 in SBs and that LR11-containing SBs are distributed throughout the brain in a similar fashion as HAP1-containing SBs. Furthermore, we show that another VPS10 domain-containing protein, sortilin, is also present in large cytoplasmic puncta in rat brain tissue sections and co-localizes with LR11 and Hap1 in SBs in primary neuron cultures.
Materials and Methods
cDNA Construct and Transfection of Cells
PCR fragments of the rat HAP1A cDNA provided by Dr. Christopher Ross (nucleotides 5–1758 and 1421–1863 of GenBank U38373) were cloned into PEGFP-C1 (ClonTech; Palo Alto, CA) by XhoI and KpnI. Full-length LR11 cDNA cloned into PCDNA3 was obtained from Dr. Chica Schaller. HeLa cells were used for transfection with LIPOFECTAMINE PLUS Reagent (Life Technologies, Lexington, KY). After 24–48 hr, the cells were used for either immunocytochemistry (ICC) or Western blotting analysis.
Antibodies
A polyclonal antiserum specific for HAP1B (HAP1B-C, also called CAG1-3) was raised in rabbits against a synthetic peptide, EQQPIVPTQDSQRLE, corresponding to residues 595–609 deduced from rat HAP1B cDNA sequence. Briefly, the peptide was coupled to keyhole limpet hemocyanin (Pierce; Rockford, IL) according to the manufacturer's instructions. Conjugated peptides were sent out to Pel Freeze (Redding, CA) where they were used as antigen to immunize rabbits and generate antisera (Pel Freeze). Antibodies were affinity-purified and characterized by ICC and Western blotting experiments (Chan et al. 2002). A monoclonal antibody (MAb) detecting both HAP1A and B was also developed as follows. The recombinant glutathione-S-transferase (GST) fusion protein expression plasmid, PGEX/HapmAb, was constructed by cloning a PCR-amplified fragment corresponding to residues 290–426 of rat HAP1 cDNA into the expression vector PGEX-2T cleaved by BamHI and EcoRI. The recombination vector was transformed into E. coli DH5α. The expressed fusion protein was purified and used for immunization of several female BALB/c mice. B-cells from draining lymph nodes were fused with PUA1 (ATCC; Rockville, MD) fusion partner and anti-fusion protein MAb-secreting cells were cloned by a dilution method. One of the clones, 1B6, showing specific immunoreactivity for both HAP1 isoforms by Western blots (Chan et al. 2002) and specifically detecting SBs by ICC, was used for this study. Mouse MAbs to domain II of human LR11 (mLR11 or 5-4-30-19-2) were provided by Dr Hikeida Bujo. This well-characterized antibody was made against a GST fusion protein containing amino acids 287–679 of the human LR11 cDNA and has been used to detect LR11 in a variety of tissue and cell lines (Taira et al. 2001). Rabbit polyclonal antibodies to the fibronectin domain and to a C-terminal segment of human LR11 were provided by Dr. I. Björn Riedel and Dr. Chica Schaller (Hampe et al. 2000). Rabbit serum against sortilin (SortK) was provided by Dr. Kostya Kandror (Lin et al. 1997; Kandror and Pilch 1998).
Western Blotting
Total homogenates from cultured cells and mouse brains were separated by SDS-PAGE, transferred to nitrocellulose membrane, and probed with the following antibodies: HAP1BC (1:1000), 1B6 (hybridoma medium), and mLR11 antibody (1:50). The proteins were detected by chemiluminescence (ECL system; Pierce). To determine that the lanes were loaded with equal amounts of proteins, blots were stained with Ponceau S before immunodetection.
Perfusion
Adult Sprague–Dawley rats (150–200 g) and C57Bl6J mice (40–50 g) were obtained from Jackson Laboratories (West Chester, PA) and maintained in a 12/12 light/dark cycle with ad libitum access to food and water. Rats (n=4) and mice (n=6) were deeply anesthetized with 4% choral hydrate solution (400 mg/kg). Perfusion was performed through the left cardiac ventricle with 3–4% paraformaldehyde (Sigma–Aldrich; St Louis, MO) in 0.1 M PB. For the electron microscopic analysis, rats (n=2) were perfused with 3% paraformaldehyde and 0.1% glutaraldehyde (Electron Microscopy Science; Fort Washington, PA). Brains were removed and postfixed overnight in 4% paraformaldehyde. Brains were cut in 10 series of 50-μm coronal sections using a vibratome. Sections were rinsed in 0.05 M PBS, pH 7.2, and processed for ICC as described below. All protocols involving animals were approved by the Emory University Animal and Care Committee and conform to NIH guidelines.
Neurospheres
Neurospheres were prepared by culturing cells from the hypothalamic region and colliculi and brainstem regions of Wistar rat fetuses. Fetuses of 14 days’ gestation were obtained from pregnant rats under deep anesthesia with isoflurane. Tissue from the hypothalamus and brainstem were dissected under a dissecting microscope and dissociated into single cells by papain treatment, followed by gentle trituration with a fire-polished Pasteur pipette. The dissociated cells were cultured in suspension in 75-ml tissue culture flasks containing MEM with 10% fetal bovine serum, penicillin (100 U/ml; GIBCO, Grand Island, NY), streptomycin (100 g/ml; GIBCO). Within 3–5 days of culture at 37C with 5% CO2/95% air, cells proliferated and formed floating neurospheres. Neurospheres were then dissociated and plated on poly-lysine-coated coverslips and incubated in neurobasal medium supplemented with B27. In the differentiation medium, neurosphere cells attached to the dish, many of which extended a variety of cell processes.
Light Microscopic Single Immunolabeling
Free-floating sections were rinsed in PBS and processed for ICC. To reduce endogenous peroxidase activity and nonspecific antibody binding, sections were incubated in 3% hydrogen peroxide, rinsed several times in PBS, and incubated in PBS containing 4% normal goat serum (NGS) and 10 μg/ ml avidin for 30 min. After PBS rinses, sections were incubated for 24 hr at 4C in 2% NGS in PBS containing 50 μg/ ml biotin and either HAP1B-C (1:1000), mLR11 (1:200), anti-LR11 fibronectin domain (1:500), anti-LR11 C-terminal domain (1:1000), or rabbit anti-sortilin serum (SortK at 1:5000). Sections were then rinsed in PBS and incubated for 2 hr at 4C in biotinylated goat anti-mouse (mLR11) or goat anti-rabbit (all other antibodies) secondary antibodies used at 1:200 (Jackson Laboratories) in PBS containing 2% NGS. After rinses in PBS, sections then incubated in avidin–biotin complex (Vector ABC Elite; Vector Labs, Burlingame, CA) for 2 hr at 4C, followed by several PBS rinses. Final development was done by incubation in 0.05% in diaminobenzidine (DAB, Sigma) and 0.01% hydrogen peroxide in 50 mM Tris buffer for 5–15 min. Sections were rinsed thoroughly, mounted on glass slides, dehydrated, and coverslipped for light microscopic examination using a Leica DMRE. Brain localization was determined according to the rat brain atlas (Paxinos and Watson 1986). To better visualize regional outline some sections were counterstained using thionin or Cresyl violet. Controls included omission of primary antibodies.
Double-label Immunofluorescence
Immunofluorescence was used to co-localize HAP1B and LR11 in rat brain and in primary neuron cultures and to colocalize sortilin with LR11 and HAP1 in primary cultures. Free-floating sections were rinsed in PBS, blocked in 5% NGS and 5% normal horse serum (NHS), and avidin for 60 min at room temperature (RT) and rinsed in PBS. After rinses in PBS, sections were incubated overnight at 4C in rabbit anti-HAP1B (HAP1B-C at 1:500) and mLR11 (1:50) in PBS containing 1% NHS, 1% NGS, and biotin. Sections were rinsed in PBS and incubated in biotinylated donkey anti-mouse IgG at 1:250 (Jackson Immunoresearch) in 1% NHS and 1% NGH for 1 hr at RT. Sections were rinsed in PBS and incubated in avidin–biotin complex (ABC kit; Vector Laboratories) for 60 min at RT. After rinses in PBS, sections were incubated in Cy5 tyramide (NEN; Boston, MA) according to the manufacturer's recommendation. After rinses, sections were incubated in FITC-conjugated donkey anti-rabbit (1:100) for 60 min at RT, rinsed in PBS, incubated for 30 min in 10 mM cupric sulfate in 50 mM ammonium acetate, pH 5.0, to eliminate autofluorescence, rinsed with PBS, then mounted on glass slides with Vectashield mounting medium for fluorescence. For some sections, nuclear counterstaining was obtained by a short incubation of the sections in PBS containing bis-benzimide before mounting. Staining of the primary cultures was done as follows. Coverslips were rinsed in cold PBS and fixed for 5 min in 4% paraformaldehyde. Coverslips were rinsed and blocked in 4% NGS for 30 min, rinsed, and incubated overnight at 4C in a combination of either HAP1BC(1:500) and mLR11 (1:50) antibodies, mLR11(1:50) and SortK (1:2500) antibodies, or 1B6(1:10) and SortK(1:2500). After several washes, coverslips were incubated with both rhodamine-conjugated goat anti-mouse and FITC-conjugated goat anti-rabbit secondary antibodies used at 1:100 (Jackson Immunoresearch). Nuclear counterstaining was obtained by a short incubation in PBS containing bis-benzimide before mounting. For each double-label experiment, controls included omission of one or both primary antibodies.
Electron Microscopy
For ultrastructural analysis, we used a combination of pre-embedded immunogold and DAB labeling of HAP1 and LR11, respectively. Single and double labeling were performed using HAP1BC and mLR11 antibodies. Sections were rinsed in PBS, blocked according to manufacturer's instructions, and incubated in HAP1B-C(1:1000), mLR11 (1:100), or a combination of HAP1B-C (1:1000) and mLR11 (1:100) overnight. Sections were rinsed in PBS and incubated in a combination of biotinylated donkey anti-rabbit and ultrasmall colloidal gold-conjugated secondary antibody (Aurion; Wageningen, The Netherlands) overnight. Sections were rinsed, incubated in avidin–biotin complex, and developed with DAB as described above. After a post-fixation with 2.5% glutaraldehyde, gold particles in sections were intensified using an R-gent SE-EM silver enhancement kit (Aurion). Sections were then further fixed with 0.5% osmium tetroxide in 0.1 M PB for 15 min and processed for electron microscopy as described elsewhere (Gutekunst et al. 1998). Selected sections were then placed in 0.5% osmium tetroxide in 0.1 M phosphate buffer for 30 min. Sections were then rinsed in PB, dehydrated in 25–100% EtOH followed by propylene oxide, infiltrated, and flat-embedded in Epon between sheets of Aclar and cured at 60C for 2–3 days.
Results
HAP1 and Stigmoid Bodies
Our polyclonal antibody to HAP1BC identified a 110-kD protein in rat and mouse brain supernatant (Figure 1A), while our MAb to HAP1 (1B6) detected two distinct bands with molecular weights of approximately 110 kD and 95 kD (Figure 1A) using the rainbow molecular weight markers (BioRad; Hercules, CA). These molecular weights correspond to previously reported molecular weights for HAP1A and HAP1B respectively (Martin et al. 1999). By ICC, both polyclonal and monoclonal antibodies detected small to large cytoplasmic puncta, which we have previously identified as SBs (Gutekunst et al. 1998). We have found that antibodies developed against various regions along the HAP1A and HAP1B sequences all recognize SBs, suggesting that full-length HAP1A and HAP1B are likely to be present in these structures. As we had previously shown using HAP1 antibodies, some neurons contained only one large SB (up to 5 μm in diameter), whereas others contained up to 12 small (less than 1 μm) to medium (1–4 μm) SBs (Figures 1B and 1D). Occasionally a neuron contained one large SB and a few smaller HAP1-positive cytoplasmic puncta.
LR11 and Stigmoid Bodies
Most of the studies were performed using an LR11 MAb directed against the vacuolar protein sorting/targeting protein (VPS10) domain located in the extracellular region of LR11 (mLR11). mLR11 antibodies detect a unique protein band at a molecular weight of approximately 250 kD (Figure 1A) on Western blots of mouse brain homogenates. The characterization of this antibody is described elsewhere (Taira et al. 2001). When used for ICC on rat brain sections, mLR11 antibodies detected large cytoplasmic structures (Figures 1C and 1E) within neuronal perikarya in many different regions of the brain, including hypothalamus, amygdala, colliculi, central gray, hippocampus, and cortex. At the light microscopic level, these structures appeared very similar in size, shape, and subcellular localization to HAP1-containing SBs. These results were consistent with findings previously reported using polyclonal antibodies to LR11 (Motoi et al. 1999). When mLR11 antibodies were preabsorbed over HAP1 GST fusion protein-containing amino acids (aa) 1–578 of HAP1A, aa 578–599 of HAP1A, or aa 578–630 of HAP1B, SBs were still immunoreactive, indicating that mLR11 did not crossreact with HAP1 proteins (data not shown). Two other polyclonal antibodies generated, respectively, to the fibronectin domain and the C-terminal region of LR11 were found to detect large cytoplasmic puncta only in scattered neurons of the rat cortex (Figures 1F and 1G). In addition to the large immunoreactive puncta, all three LR11 antibodies also showed diffuse cytoplasmic staining visible in the perikarya and dendrites of neurons throughout the brain.
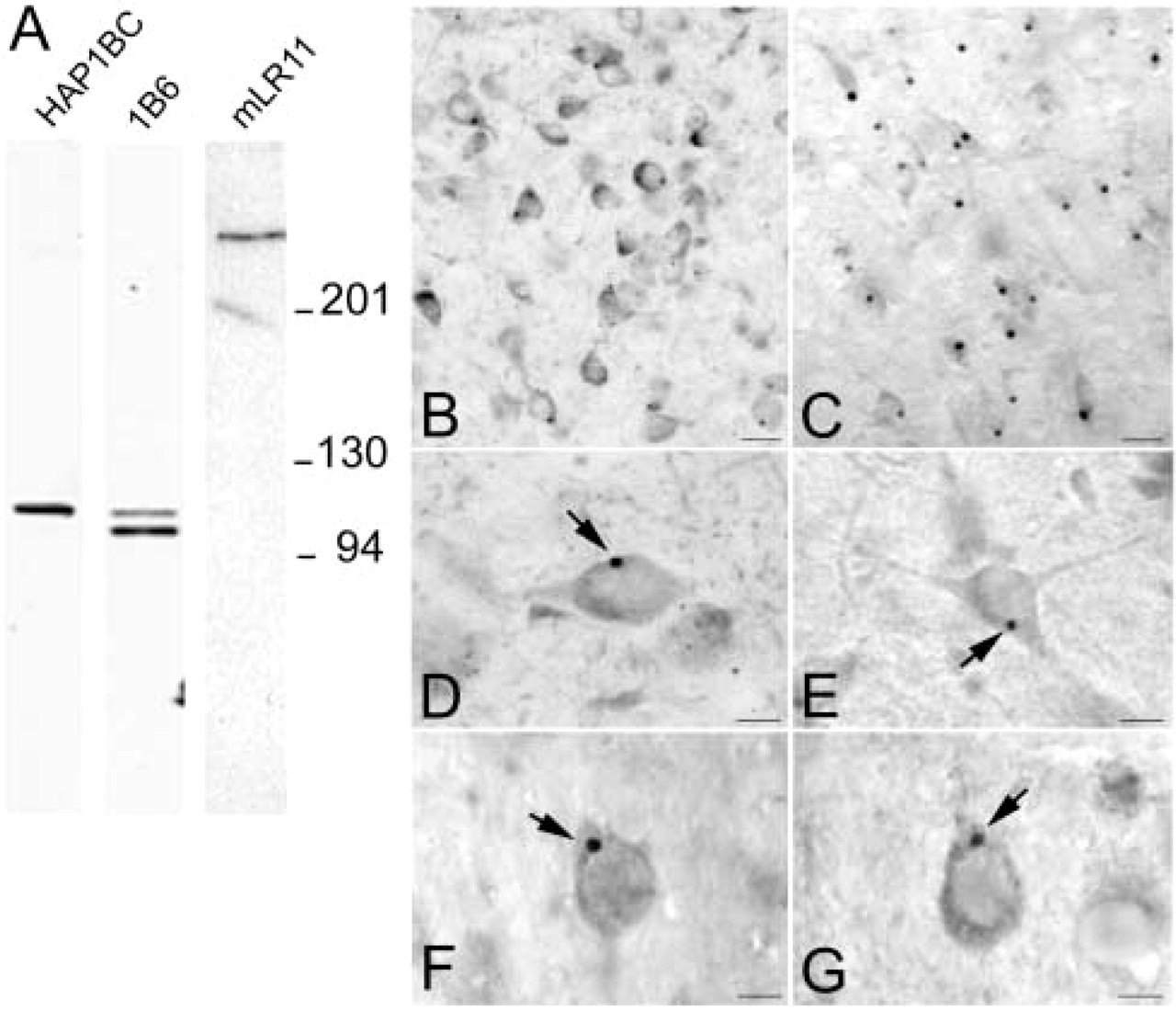
Specificity of HAP1 and LR11 antibodies in brain. (
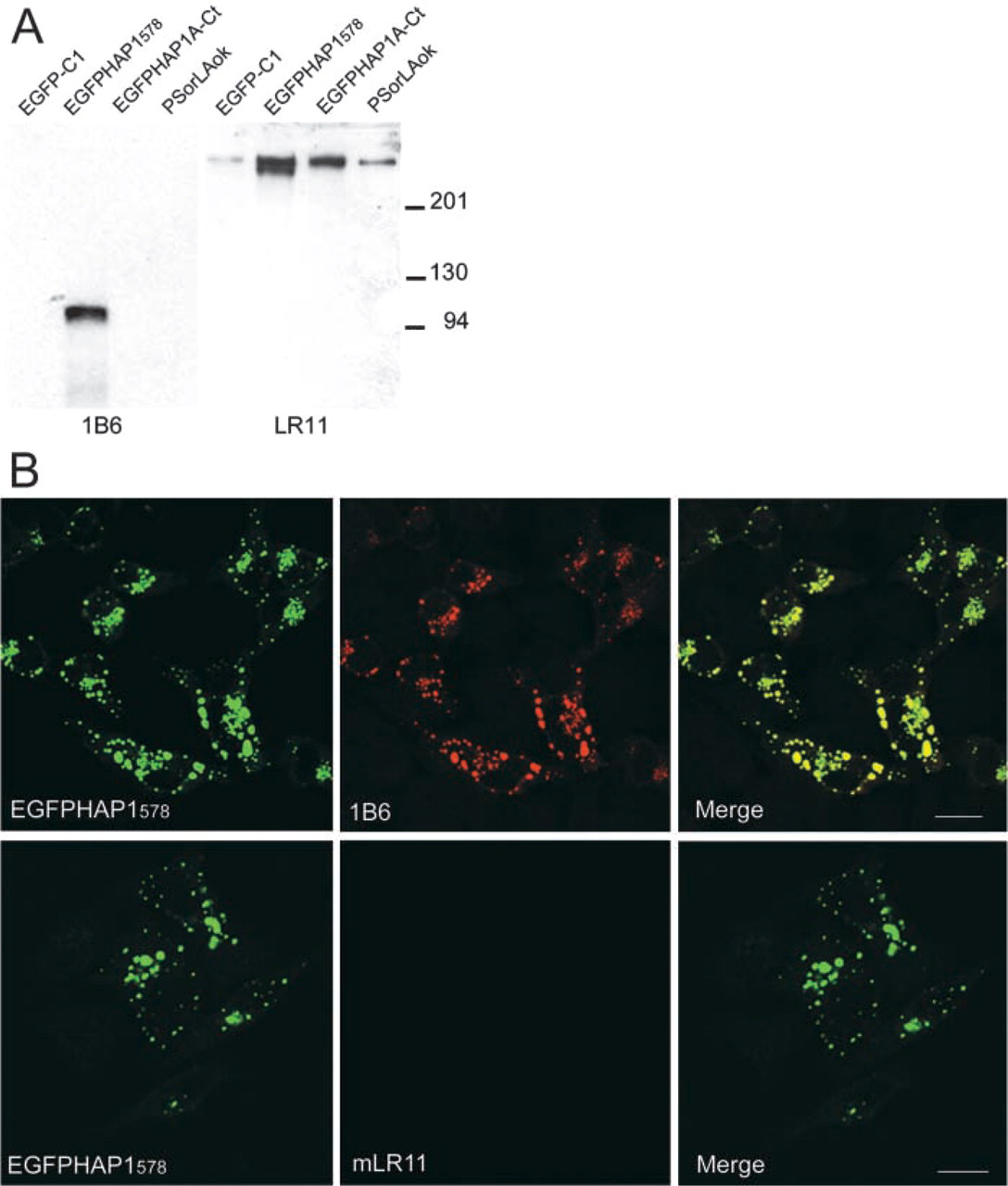
HAP1 and LR11 in transfected HeLa cells. (
To further demonstrate the specificity of mLR11 antibodies and to rule out any crossreactivity between mLR11 and HAP1, we performed a series of experiments using transiently transfected HeLa cells. PEGFP-C1 containing either the HAP1 aa 1–578 (EGFPHap1578), the HAP1A-specific C-terminal sequence (EGFPHAP1A-Ct), and PCDNA3 containing LR11 full-length cDNA (pSorLAOk) were transiently transfected into HeLa cells. After 24 hr, total homogenates from the transfected cells were blotted and immunoreacted for HAP1 or mLR11. HAP1 is mostly expressed by neurons, and HeLa cells do not show detectable levels of endogenous HAP1 expression. As expected, our 1B6 antibodies only detected EGFPHap1578 (Figure 2A) as a single band with a molecular weight of approximately 95 kD, which corresponds to the added weight of truncated HAP1A (∼60 kD) and GFP (∼30 kD). LR11 MAbs detected endogenous and overexpressed full-length LR11 (Figure 2A) and did not react with EGFP-C1, EGFPHAP1578, or EGFPHAP1A-Ct. To verify that mLR11 did not crossreact with HAP1 proteins by ICC, HeLa cells were transiently transfected with a variety of HAP1- or LR11-containing vectors, fixed, and immunostained with 1B6 or mLR11. EGFP-transfected cells showed diffuse GFP in their cytoplasm and nuclei and reacted with neither 1B6 nor mLR11. EGFPHAP1578 induced the formation of 1B6-positive cytoplasmic inclusions that were negative for mLR11 (Figure 2B). The formation of the cytoplasmic inclusions after HAP1 transfection is consistent with our previous work in HEK 293 cells (Li et al. 1998a).
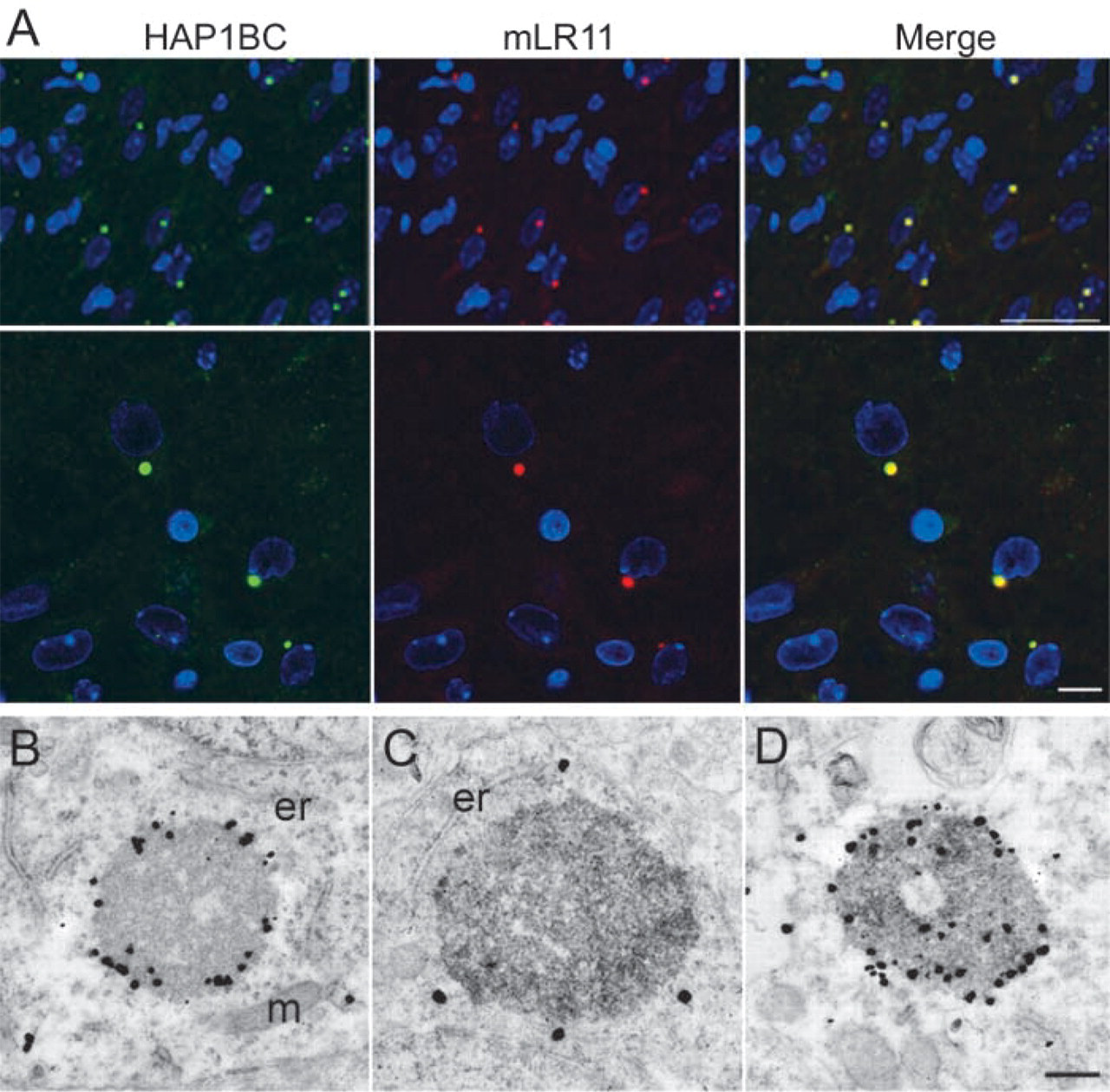
LR11 co-localizes with HAP1 within SBs. (
To determine whether the large cytoplasmic puncta immunodetected by the mLR11 antibodies did indeed correspond to the SBs detected by our various HAP1 antibodies, we performed double immunolabeling of rat brain tissue sections using HAP1BC and mLR11 antibodies. In the regions analyzed, including the lateral hypothalamus, the amygdala, the bed nucleus of the stria terminalis, and the cortical mantle, almost all of the HAP1BC-positive SBs were positive for mLR11 (Figure 3A). Double labeling was also performed at the electron microscopic level using mLR11 DAB staining and HAP1 immunogold. The ultrastructure of the mLR11 immunoreactive SBs was identical to that of HAP1-containing SBs. In double-stained sections, mLR11 DAB reaction product was present in HAP1 immunogold-positive SBs (Figure 3D).
SBs in the Rat and Mouse Brain
To further address the distribution of SBs in adult rat and mouse brain, we performed ICC using a variety of polyclonal and monoclonal HAP1 antibodies and monoclonal LR11 antibodies. Serial sections were cut in various planes (coronal, parasagittal, and horizontal) and immunoreacted. Both HAP1 and LR11 MAbs immunodetected SBs throughout the mouse and rat brain. Both markers detected very large numbers of SB-containing neurons in various nuclei of the hypothalamus, bed nucleus of the stria terminalis, amygdala, arcuate nucleus, subfornical organ, and the granule cell layer of the hippocampus. Regions in which both markers detected moderate numbers of SBs included nucleus accumbens, geniculate nucleus, septum, substantia nigra, and zona incerta. Regions with very few SBs included olfactory bulbs, striatum, globus pallidus, cortex, central gray, the habenula, and the cerebellum.
In our previous report of SBs using a fusion protein antibody against HAP1A and HAP1B, we had found little or no SBs in thalamic and hippocampal regions of the rat brain (Gutekunst et al. 1998). In the present study, however, antibodies to HAP1 isoforms and LR11 MAbs all detected many SBs in regions at which they had not been previously described, including various nuclei of the thalamus, the CA1–4 pyramidal layer of the hippocampus, and scattered neurons in layer II of cortex (Figure 4). In the hippocampus, many SBs were detected in the dendate gyrus and the CA pyramidal layer (Figure 5). In the dendate gyrus, SBs were found in neurons of the granular cell layer with only one SB visible per neuron. In the CA pyramidal layer of the hippocampus, SBs were found primarily in the CA1 region, some were seen in the CA2–CA3 region, but only rare ones were detected in the CA4 region. In the hippocampus there was only one detectable SB per neuron. Electron microscopic analysis revealed their ultrastructure to be similar to that of the small SBs found in the hypothalamus and other SB-rich regions (Figure 5). Thalamic and CA1-3 SBs had a maximum of 1 μm diameter and did not have the empty core characteristic of some of the larger SBs described in the hypothalamic and septal regions (Shinoda et al. 1993; Gutekunst et al. 1998).
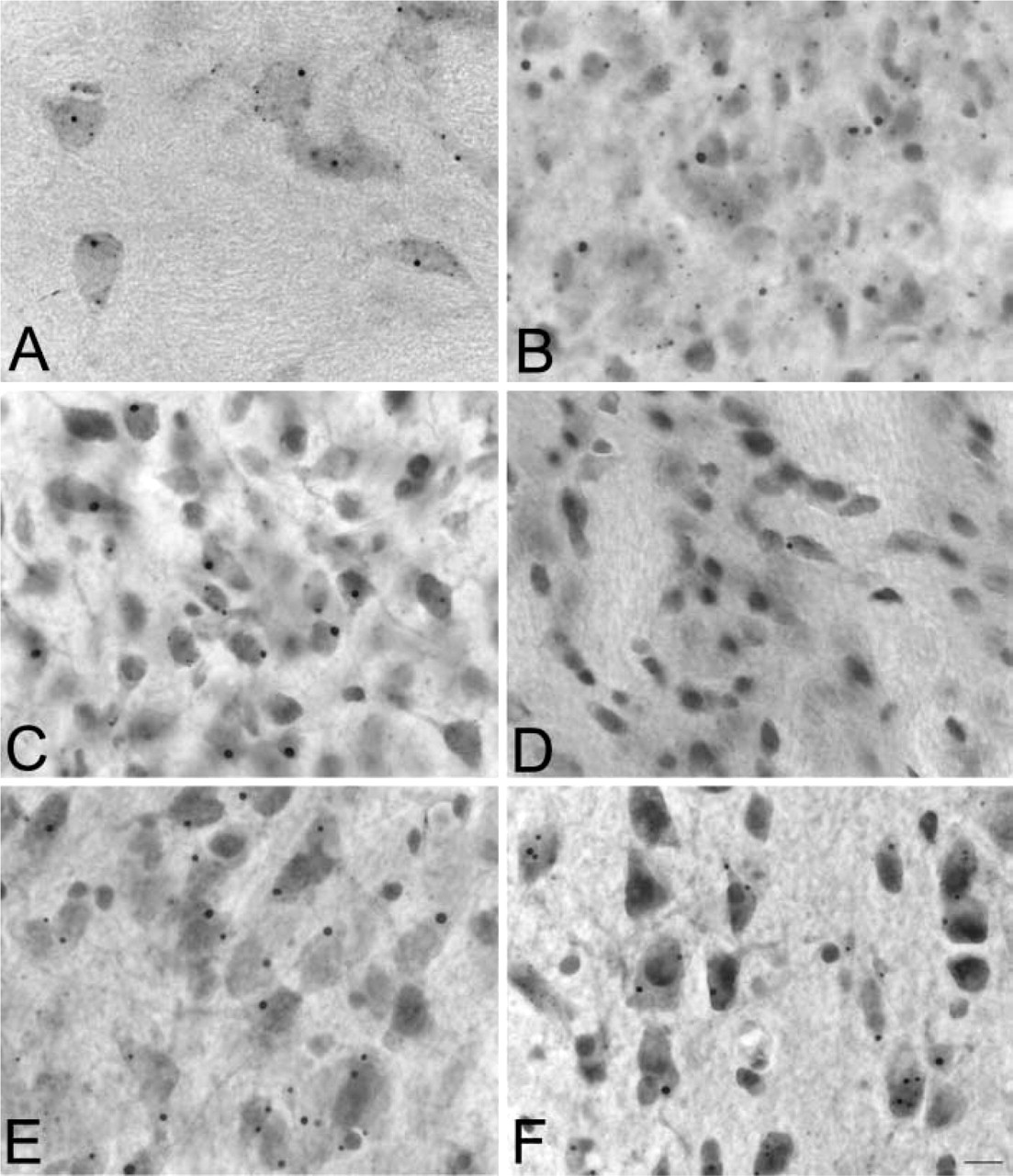
SBs throughout the rat brain. Micrographs showing SBs in various regions of the rat brain, including thalamus (
SBs in Neuron Cultures
To determine whether SBs also formed in primary neuron cultures, we performed HAP1 and LR11 ICC in primary neurosphere preparations from brainstem E14 rat embryos. Neurospheres were maintained in suspension for 5 days. They were then trypsinized and plated on poly-
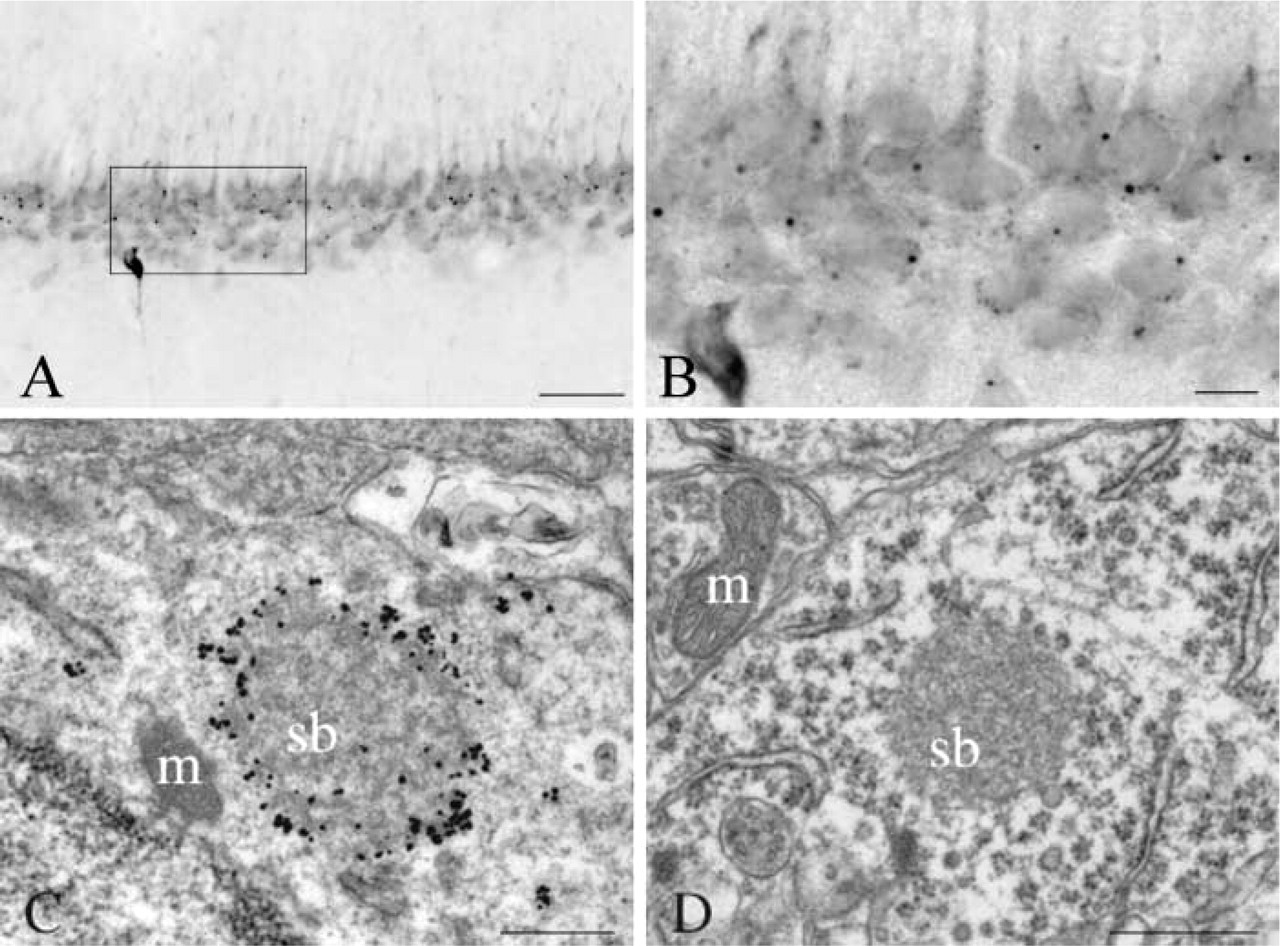
SBs in hippocampus. (
Sortilin in SBs
LR11 is a member of a recently identified type I receptor family that includes LR11, sortilin (also known as neurotensin receptor 3), SorCS1-3, and the yeast VPS10 protein (Hampe et al. 2001). All members contain a VPS10 (vacuolar protein sorting 10) domain in their luminal (extracellular) region and a C-terminal sequence with internalization signals. We sought to determine whether other members of this family were present in SBs. Antibodies to sortilin (SortK) were used to immunostain rat brain tissue sections. These well-characterized antibodies used for these studies detected only one band at around 95 kD in rat brain homogenates (Lin et al. 1997). In the hypothalamic region, SortK stained large cytoplasmic puncta in the perikarya of neurons in the arcuate nucleus and in hypothalamic neurons along the third ventricle (Figures 7A and 7B). In neuronal primary cultures from colliculi and brainstem neurospheres, sortilin was found in small puncta throughout the cytoplasm of neurons. Occasionally a neuron contained a large sortilin-immunoreactive cytoplasmic punctum. Co-localization studies revealed that these puncta also contained mLR11 (Figure 7C) and HAP1 (Figure 7D).
Discussion
In this study we set out to determine whether LR11 could be found in SBs. Here we present evidence showing that not only LR11 but also sortilin, another member of the VPS10 family of proteins, is present in SBs. Using well-characterized monoclonal and polyclonal antibodies generated against LR11 (Hampe et al. 2000; Taira et al. 2001; Zhu et al. 2002), we have found this protein in large puncta present in the cytoplasm of neurons. However, whereas the MAbs detected SBs throughout the brain, the polyclonal antibodies to LR11 detected these structures only in some cortical neurons. This is consistent with a previous report showing that polyclonal antibodies raised against a peptide from the domain II of LR11 immunoreacted with large puncta in the perikarya of rat and human cortical pyramidal neurons (Motoi et al. 1999). Both polyclonal antibodies were generated against LR11 regions different from that recognized by the MAbs. Therefore, it is possible that in extracortical SBs the specific antigenic regions recognized by the polyclonal antibodies are either masked by other proteins, absent as a consequence of protein processing, or affected by fixation. Based on their appearance, morphology and number (only one large positive punctum per neuron), the cortical puncta are likely to be SBs. This idea is further supported by the finding that similar LR11-stained structures co-localize with HAP1-positive SBs by double immunofluorescence and immunoelectron microscopy in the hypothalamus.
LR11 is a member of the LDL receptor gene family and has been shown to bind ApoE (Yamazaki et al. 1996,1997) and to mediate the uptake of ApoE-rich lipoproteins in vitro (Taira et al. 2001). LR11 is present in many organs, including the smooth muscle cells of the arterial walls, where it is markedly increased in animal models of atherosclerosis (Taira et al. 2001). LR11 overexpression in smooth muscle cells has recently been shown to enhance migration and invasion activities (Zhu et al. 2002). The ectodomain of LR11 contains binding sites for both ApoE and for the neuropeptide head activator (HA) (Hampe et al. 2000). LR11 is a member of a recently identified type I receptor family of proteins including sortilin (also known as neurotensin receptor 3 or gp95), SorCS1-3, and the yeast VPS10 protein (Hampe et al. 2001). All members contain a VPS10 domain in their luminal (extracellular) region and a C-terminal sequence with internalization signals (Nielsen et al. 2001; Dennes et al. 2002; Jacobsen et al. 2002). Because our results indicate that the VPS10 region of LR11 reaches SBs, we asked whether other VPS10 domain-containing proteins could be found in SBs. Consistent with this scheme, our results show evidence that sortilin is present in LR11- and Hap1-positive SBs. In addition, preliminary data show that a polyclonal antibody to SorCS1 (Hermey et al. 2001) also detects SBs in brain tissue and neuron cultures (manuscript in preparation).
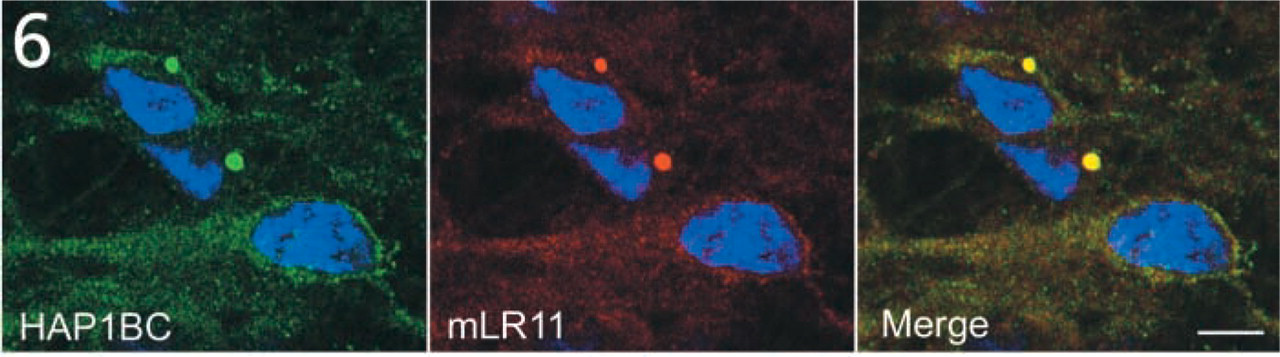
SBs in primary neurons. Micrographs showing HAP1B (green) and mLR11 (red) immunoreactivity in cultured neurospheres of colliculi and brainstem regions from E14 rats. HAP1B and LR11 immunoreactivity is visible as diffuse staining and small puncta in the neuronal cytoplasm. HAP1BC and mLR11 co-localize within two large SBs visible in the perikarya of two neurons. Nuclei are stained with bis-benzimide (blue). Bars = 20 μm.
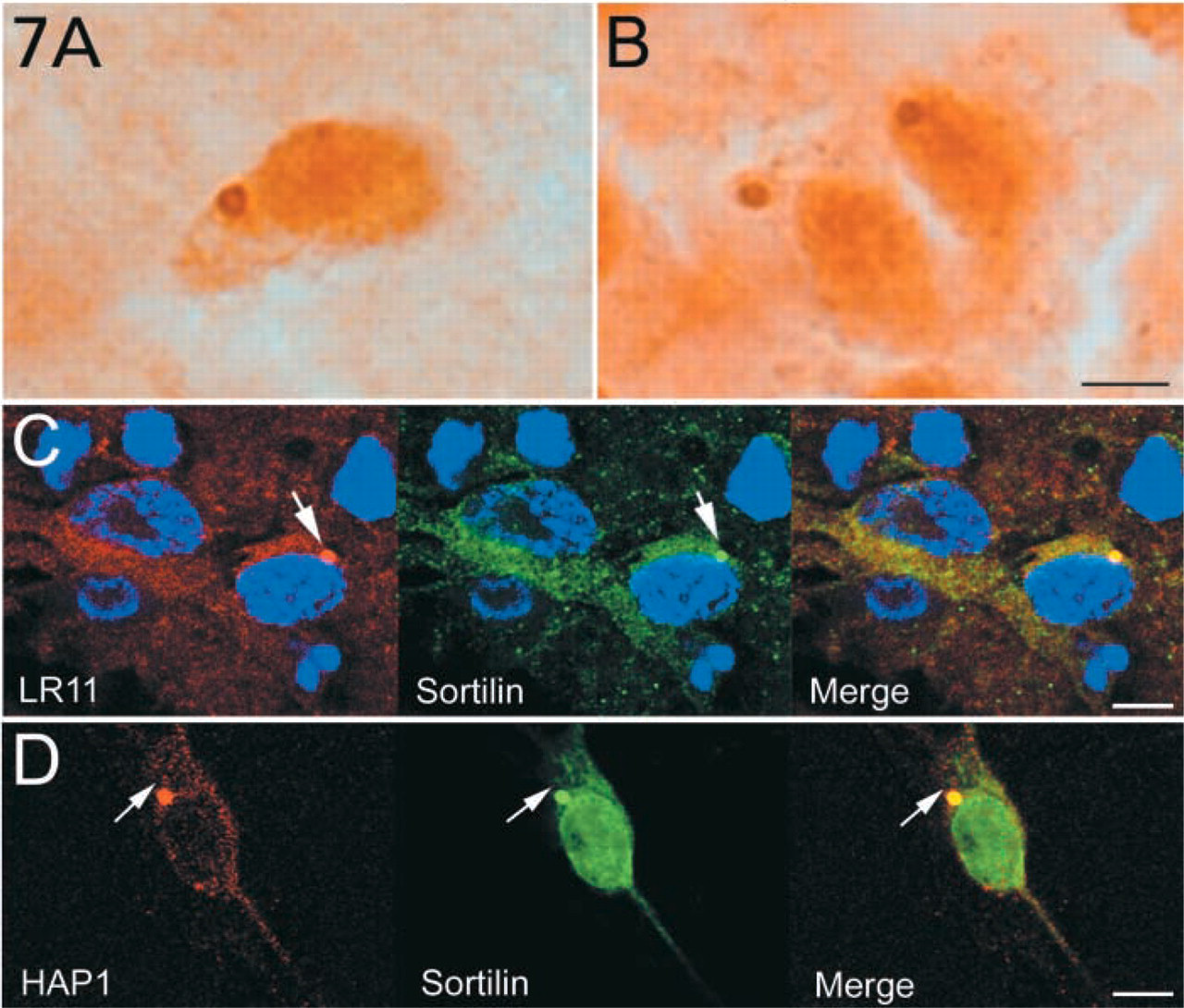
Sortilin is present in SBs. (
LR11, sortilin, and SorCS1 are type I membrane receptors with a small transmembrane spanning region mostly localized in intracellular vesicles and plasma membrane (Chabry et al. 1993; Petersen et al. 1997; Mazella et al. 1998; Hermey et al. 2001). Furthermore, it is noteworthy that a recent manuscript has described another potential membrane protein included in SB-like structures. The ionotropic glutamate receptor subunit 1 (iGluR1) has recently been localized to a large cytoplasmic punctum in neurons of postnatal day 3–10 rat spinal cord (Serrando et al. 2002). Although the authors did not perform any co-localization with HAP1 antibodies, the ultrastructural data presented suggest that these GluR1-positive puncta correspond to SBs. HAP1- and LR11-positive SBs are numerous in the spinal cord of postnatal day 10 rats (unpublished data) and this iGluR1 localization study provides further evidence for the presence of yet another transmembrane protein in SBs.
How can these transmembrane receptors reach SBs? What are the possible mechanisms leading to the presence of membrane proteins in SBs? Soluble forms of LR11 have been described in several studies. The ectodomain of mature LR11 can be shed from the transmembrane protein by a metalloprotease and has been detected in media from cell culture and brain slices using antibodies to the fibronectin domain but not to the intracellular domain (Hampe et al. 2000). In the same study, a fragment of LR11 lacking its C-terminus is detected in a soluble rat brain fraction (Hampe et al. 2000). The authors suggest that the detected fragment comes from extracellularly shed LR11. In another study, LR11 is found in a cytosolic fraction from LDLR-deficient CHO cells stably expressing human LR11 (Taira et al. 2001). Both studies used Western blotting techniques. Whether the LR11 they detected in their soluble fraction comes from a cytoplasmic pool or from shed LR11 from the extracellular compartment remains unclear. Most studies describe sortilin as a membrane receptor, and no soluble form has yet been described.
Alternative routes that would take LR11 and sortilin to the cytosol and therefore make them accessible to SBs can be speculated. Both proteins could be in the cytoplasm as a result of misfolding and subsequent translocation from the endoplasmic reticulum (ER) to the cytosol. In recent years, several studies have shown the existence of a highly efficient quality control system in the ER, which allows the elimination of misfolded proteins or unassembled subunits of protein complexes. ER degradation of membrane proteins is a cytosolic event mediated by the proteasome (Bonifacino 1996; Kopito 1997). For example, misfolded cystic fibrosis transmembrane conductance regulator (CFTR) molecules that fail to exit the ER are rapidly degraded by the proteasome (Ward et al. 1995). Other ER-restricted proteins that are degraded via the proteasome after their retrotranslocation to the cytoplasm include mutant human 1-anti-trypsin (Qu et al. 1996), yeast carboxypeptidase Y (Hiller et al. 1996), and MHC class I heavy chains (Wiertz et al. 1996a,b). Substrates destined for degradation by the proteasome are commonly “tagged” by the covalent attachment of multiubiquitin chains (Coux et al. 1996). Ubiquitination, however, is neither a necessary nor a sufficient signal for degradation (Bercovich et al. 1989; Rosenberg-Hasson et al. 1989; Johnston et al. 1995). Could SBs represent an intermediate compartment for this ER degradation pathway? We have found neither ubiquitin nor proteasome markers in SBs (unpublished observations) and therefore hypothesize that, if SBs are involved in protein degradation, they would represent a novel degradation pathway.
Alternatively, the luminal region of SorLA/LR11 or sortilin could be cleaved once inside the endocytic compartment and thereafter translocated to the cytosol. Proteases including metalloproteinase, which have been shown to cleave SorLA, have been localized to early endosomes and could participate in such a mechanism (Jiang et al. 2001). Another possibility is that the extracellular domains of these receptors enter the endocytic compartment after they have been shed into the extracellular milieu. As mentioned earlier, the extracellular region of LR11 is shed by a metalloproteinase (Hampe et al. 2000). The mechanism leading to the presence of LR11 in SBs after luminal escape or the traditional endocytic pathways would also have to be original.
In our previous studies we postulated that SBs were not aggregates but new subcellular organelles. HAP1A transfected into PC12 cells resulted in the formation of large inclusions (Li et al. 1998a). Although these resembled SBs at the light microscopic level, electron microscopic analysis revealed their ultrastructure to be quite different from that of SBs. Here we demonstrate that HAP-1 inclusions formed by overexpressing HAP-1 in HeLa cells are also different from SBs in that they are not LR11-positive despite endogenous expression of LR11. It is most likely that, as for other subcellular structures, SBs contain many other proteins. Both HAP1 and LR11 are enriched in SBs and could therefore be synthesized or stored there. We have previously suggested that HAP1 could participate in transporting SB components to and from SBs.
SBs have been identified in neurons in various regions of the brain and have been detected by electron microscopy in the rodent brain as early as embryonic day 8 (Pelaez and Alvarez-Uria 1987). In a recent study we have shown that homogyzous deletion of the HAP1 gene results in early postnatal death and have suggested an important role of HAP1 and perhaps SBs in feeding behavior (Chan et al. 2002). Whether SBs are present in the brain of HAP1 knockout mice is under examination and will determine whether this protein represents an essential component of SBs.
Whether the presence of HAP1, LR11, and sortilin in SBs indicates a role for these proteins in SB formation or in SB function remains unknown. We believe that identification of their molecular components will help shed light on their potential cellular function.
Footnotes
Acknowledgements
Supported by a National Science Foundation grant IBN 9983078 to CAG.
We would like to thank Carlos Saavedra and Julie Jun for their technical assistance. We also thank Dr Christopher Ross for providing the rat HAP1A cDNA, Dr Chica Schaller for providing the PsorLAok plasmid, and Dr Kostya Kandror for providing the anti-sortilin serum.
