Abstract
Compartmentalization of the interphase nucleus is an important element in the regulation of gene expression. Here we investigated the functional organization of the interphase nucleus of HeLa cells and primary human fibroblasts. The spatial distribution of proteins involved in transcription (TFIIH and RNA polymerase II) and RNA processing and packaging (hnRNP-U) were analyzed in relation to chromosome territories and large-scale chromatin organization. We present evidence that these proteins are present predominantly in the interchromatin space, inside and between chromosome territories, and are largely excluded by domains of condensed chromatin. We show that they are present throughout the active and inactive X-chromosome territories in primary female fibroblasts, indicating that these proteins can freely diffuse throughout the interchromatin compartment in the interphase nucleus. Furthermore, we established that the in vivo spatial distribution of condensed chromatin in the interphase nucleus does not depend on ongoing transcription. Our data support a conceptually simple model for the functional organization of interphase nuclei.
T
Second, a direct relationship exists between large-scale chromatin organization and transcriptional activity. In chromosome territories, chromatin is folded in such a way that transcriptionally active sites are located near the surface of condensed chromatin domains (Fakan 1994; Cmarko et al. 1999; Verschure et al. 1999). Most of this condensed chromatin is different from classical heterochromatin because it is closely associated with transcriptionally active loci at its surface. Heterochromatin is a special type of compact chromatin that is able to silence genomic loci when they become associated with it (Henikoff and Dreesen 1989; Brown et al. 1997; Francastel et al. 1999). When large clusters of genes are activated, condensed chromatin domains have been shown to decompact (Tumbar et al. 1999; Volpi et al. 2000; Ye et al. 2001). This is compatible with the idea that only loci in direct contact with the interchromatin compartment are transcriptionally active. The interchromatin compartment [also called interchromatin domain (ICD)] (Zink and Cremer 1998) is largely devoid of DNA and consists of a continuous set of highly convoluted channels inside and between chromosome territories. Because transcription takes place at the interface between condensed chromatin and the interchromatin compartment, i.e., in the perichromatin compartment (Fakan 1994), it is likely that newly synthesized RNA is deposited directly into the interchromatin compartment.
Finally, there is a growing list of subnuclear domains (nuclear bodies) that contain little or no DNA. These domains are enriched in specific sets of nuclear factors involved in RNA synthesis and/or RNA processing. Examples are nucleoli (Pedersen 1998), clusters of interchromatin granules (also called speckles) (Mintz et al. 1999; Mintz and Spector 2000), Cajal (coiled) bodies (Gall et al. 1999), promyelocytic leukemia (PML) bodies (Zhong et al. 2000), cleavage bodies (De Jong et al. 1996), and nuclear domains that are enriched in histone deacetylase activity (Downes et al. 2000). There is growing evidence that several of these domains are somehow involved in the expression of specific classes of genes. Together, these three major aspects of nuclear organization, i.e. (a) chromosome territories, (b) large-scale chromatin organization in relation to gene activity, and (c) subnuclear domains that are involved in gene expression, impose considerable constraints on the spatial organization of the genome in the interphase nucleus. The underlying molecular principles of nuclear compartmentalization are still largely unknown.
Here we investigate the functional organization of the interphase nucleus of HeLa cells and primary human female fibroblasts. The spatial distribution in the nucleus of proteins involved in transcription and RNA processing and packaging is analyzed. We present evidence that these proteins are largely excluded from condensed chromatin and occur predominantly in the interchromatin compartment. Strikingly, we find that proteins involved in transcription and RNA processing are present in the interchromatin space of both the active and the inactive X-chromosome territories in female fibroblasts. This is in line with the notion that most nuclear proteins freely diffuse through the nucleus (Misteli 2001). Furthermore, we establish that the in vivo spatial distribution of condensed chromatin is not dependent on ongoing transcription. Our data support the idea that transcriptionally active loci are in close contact with the interchromatin compartment, which provides genes with the necessary transcriptional machinery and components required for RNA processing and packaging. The interchromatin compartment allows channeling of ribonucleoprotein particles (RNPs) to the nuclear pore complexes or to other sites inside the nucleus.
Materials and Methods
Cell Culture
Human primary fibroblasts (kindly provided by Dr. L.H.F. Mullenders; University of Leiden, The Netherlands) with a normal female karyotype (46, XX) were grown at 37C in a 2.5% CO2 atmosphere in Ham's F-10 (Gibco; Breda, The Netherlands), supplemented with 15% (w/v) heat-inactivated fetal calf serum (FCS; Boehringer, Mannheim, Germany), 2 mM
A HeLa cell line (cell line 2–12-HeLa) that stably expresses H2B-GFP was used (Kanda et al. 1998). Cells were grown at 37C in a 10% CO2 atmosphere in DMEM supplemented with 10% FCS, and were kept continuously under 0.35 mg/ml G418 (Sigma; St Louis MO) drug selection.
To inhibit RNA synthesis, cells were cultured either with 50 μM 5,6-dichloro-α-
Immunofluorescent Labeling
Immunofluorescent labeling was performed as described previously (Verschure et al. 1999). Cells were fixed for 10 min at 4C in 2% (w/v) formaldehyde in PBS. After fixation, cells were permeabilized with 0.5% (w/v) Triton-X 100 in PBS for 5 min and incubated in PBS containing 100 mM glycine for 10 min. Then the cells were incubated for 1 hr at 37C with primary antibodies diluted in PBG, PBS containing 0.5% (w/v) bovine serum albumin (BSA) and 0.1% (w/v) gelatin (Sigma). The following primary antibodies were used: mouse monoclonal antibody 2HP-1H5 recognizing HP-1 α (Taddei et al. 1999, 2001), a mouse monoclonal antibody recognizing the p62 subunit of general transcription factor TFIIH (Schaeffer et al. 1994), mouse monoclonal antibody 8WG16 (Thompson and Burgess 1989) or H14 (Bregman et al., 1995) against RNA polymerase II, rabbit antibodies R232 and R252 against histone H4 that is acetylated at lysine 8 and at lysine 16, respectively (Turner and Fellows 1989; Turner et al. 1989), and a rabbit antibody against hnRNP-U (Fackelmayer and Richter 1994). After several washes with PBS, cells were incubated with the appropriate secondary antibodies, using Cy3- or FITC-conjugated donkey anti-rabbit antibody, goat anti-human antibody, or donkey anti-mouse antibody (Jackson; West Grove, PA). Secondary antibodies were diluted in PBG. Incubations were performed for 1 hr at room temperature (RT). Cells were then rinsed with PBS at RT and DNA staining was performed with 0.4 μg/ml Hoechst 33258 (Sigma) in PBS. Slides were mounted in Vectashield (Vector Labs; Burlingame, CA). Slides were stored at 4C and analyzed within 24 hr. As a control, the primary antibody was omitted. For double-labeling experiments, slides were subsequently washed in PBS and used for in situ hybridization (ISH).
Chromosome-specific Probes
Labeling of the human X chromosome was achieved using chromosome-specific DNA library probes directly conjugated to either Cy3 or FITC (Cambio; Cambridge, UK). Probes were prepared as described previously (Verschure et al. 1999).
ISH Procedure
The fluorescence ISH procedure has been described previously (Verschure et al. 1999). Briefly, cells were rinsed with PBS and fixed for 10 min at 4C in 4% (w/v) formaldehyde diluted in PBS. To facilitate probe penetration, cells were treated with 0.1 M HCl for 10 min and subsequently with a mixture of 0.5% Triton and 0.5% (w/v) saponin in PBS for 10 min. After washing in PBS, genomic DNA was denatured by incubating cells in 2 X SSC containing 70% formamide at 37C for 4 min, followed by incubating cells in 2 X SSC containing 50% formamide and subsequent washing with 2 X SSC performed at 42C.
Cells were rinsed with PBS at RT and DNA staining was performed with 0.4 μg/ml Hoechst 33258 (Sigma) in PBS. Slides were mounted in Vectashield. Slides were kept at 4C until evaluation and were analyzed within 24 hr.
Confocal Laser Scanning Microscopy
All experiments were performed at least three times in duplicate. For each experiment, 10 nuclei were visualized. Images were recorded with a Zeiss LSM 510 (Zeiss; Jena, Germany) confocal laser scanning microscope equipped with a X100/1.23 NA oil-immersion objective. We used an argon laser at 488 nm in combination with a helium neon laser at 543 nm to excite green and red fluorochromes simultaneously. Emitted fluorescence was detected with a 505–530-nm bandpass filter for the green signal and a 560-nm longpass filter for the red signal. Pairs of images were collected simultaneously in the green and red channels. 3D images were scanned as 512 × 512 × 32 voxel images (sampling distance 49 nm lateral and 208 nm axial).
Image Processing
Images were corrected for optical crosstalk (Manders et al. 1992). Image analysis was performed using SCIL-IMAGE software (Ten Kate et al. 1990; Van Balen et al. 1994). Images were subjected to 3D image restoration to correct for diffraction-induced distortion using the Huygens System 2 software package (Scientific Volume Imaging; Hilversum, the Netherlands) (Van der Voort and Strasters 1995; Verschure et al. 1999). The 3D image restoration procedure uses a point spread function that was obtained under exactly the same conditions as used for image acquisition. To this end, 200-nm-diameter fluorescent beads (FluoSpheres; Molecular Probes, Eugene, OR) were imaged. 3D image restoration procedures significantly improved the quality of the 3D images by removing Poisson noise and deblurring the images. For a semi-quantitative analysis of the spatial relationship between the relative spatial distributions of components in dual-labeled cells, we made line scans. The signal intensities of the two labels are plotted along a line through a x-y section through the nucleus. This method provides a rapid and straightforward way to determine the extent to which two components co-localize.
Analysis of the Images
All experiments were repeated at least two times. For all experiments we analyzed at least 10 cells. To compare the localization of the double-labeled images, we made line scans measuring the local intensity distribution of the two labels. Of all the analyzed cells we evaluated at least 20 line scans. In the figures we show only one of these evaluated line scans.
Results
Condensed Chromatin Domains Are Accessible for Antibodies
Histone H2B-GFP expressing HeLa cells (Kanda et al. 1998) display a typical reticular chromatin distribution. Intensely labeled condensed chromatin domains and the interchromatin compartment, which contains little or no GFP label, can be clearly distinguished, as was shown previously by Verschure et al. (1999). To decide whether a nuclear antigen is located in the interchromatin compartment or inside condensed chromatin domains, or in both, one must make sure that components inside condensed chromatin are accessible for antibodies. To test this, we used an antibody that recognizes heterochromatin protein 1-alpha (HP-1α). HP-1α is confined to highly compacted pericentric heterochromatin (Taddei et al. 1999, 2001). The spatial distributions of GFP-labeled chromatin and of HP-1α-immunolabeled domains in HeLa cells were compared after dual-color 3D imaging. Figure 1A shows that there is a close match between the intensely GFP-labeled compact heterochromatin domains and the HP-1α signal, showing that the anti-HP-1α antibody had access to condensed chromatin domains. We conclude that under our labeling conditions antibodies are able to recognize antigens in condensed chromatin domains.
The General Transcription Factor TFIIH, RNA Polymerase II and the hnRNP-U Protein Are Predominantly Localized in the Interchromatin Compartment
We have analyzed the spatial distribution of the general transcription factor TFIIH (using an antibody recognizing the p62-subunit) and that of RNA polymerase II (using antibodies H14 and 8 WG16, which recognize all phosphorylation states of RNA polymerase II (Thompson and Burgess 1989; Bregman et al. 1995; Verschure et al. 1999). We compared the spatial distribution of these antigens to the distribution of GFP-labeled chromatin in HeLa cells and to FISH-labeled × chromosome territories in primary human female fibroblasts. Results are shown in Figure 1 and 2. TFIIH labeling shows a typical punctated nuclear distribution (Figures 1B and 1C) that has been found to significantly, but far from completely, co-localize with sites of transcription in the interphase nucleus (Grande et al. 1997). The strongly TFIIH-labeled domain in Figure 1C (arrow) most likely is a Cajal (coiled) body, which is known to be enriched in TFIIH (Matera 1999). Figure 1D shows the distribution of RNA polymerase II-rich domains scattered throughout the nucleoplasm. RNA polymerase II has previously been shown to partially co-localize with transcription sites (Grande et al. 1997). The line scans in Figures 1B-IV, C-IV, and D-IV show that TFIIH and RNA polymerase II predominantly localize in the interchromatin compartment. At some positions there is partial overlap with condensed chromatin in these HeLa cells. Because the limited resolution of light microscopy causes a systematic overestimation of the overlap between any two spatial distributions, we conclude that most TFIIH and RNA polymerase II is excluded from condensed chromatin domains. This conclusion is in agreement with observations at the ultrastructural level obtained by Cmarko et al. (1999).
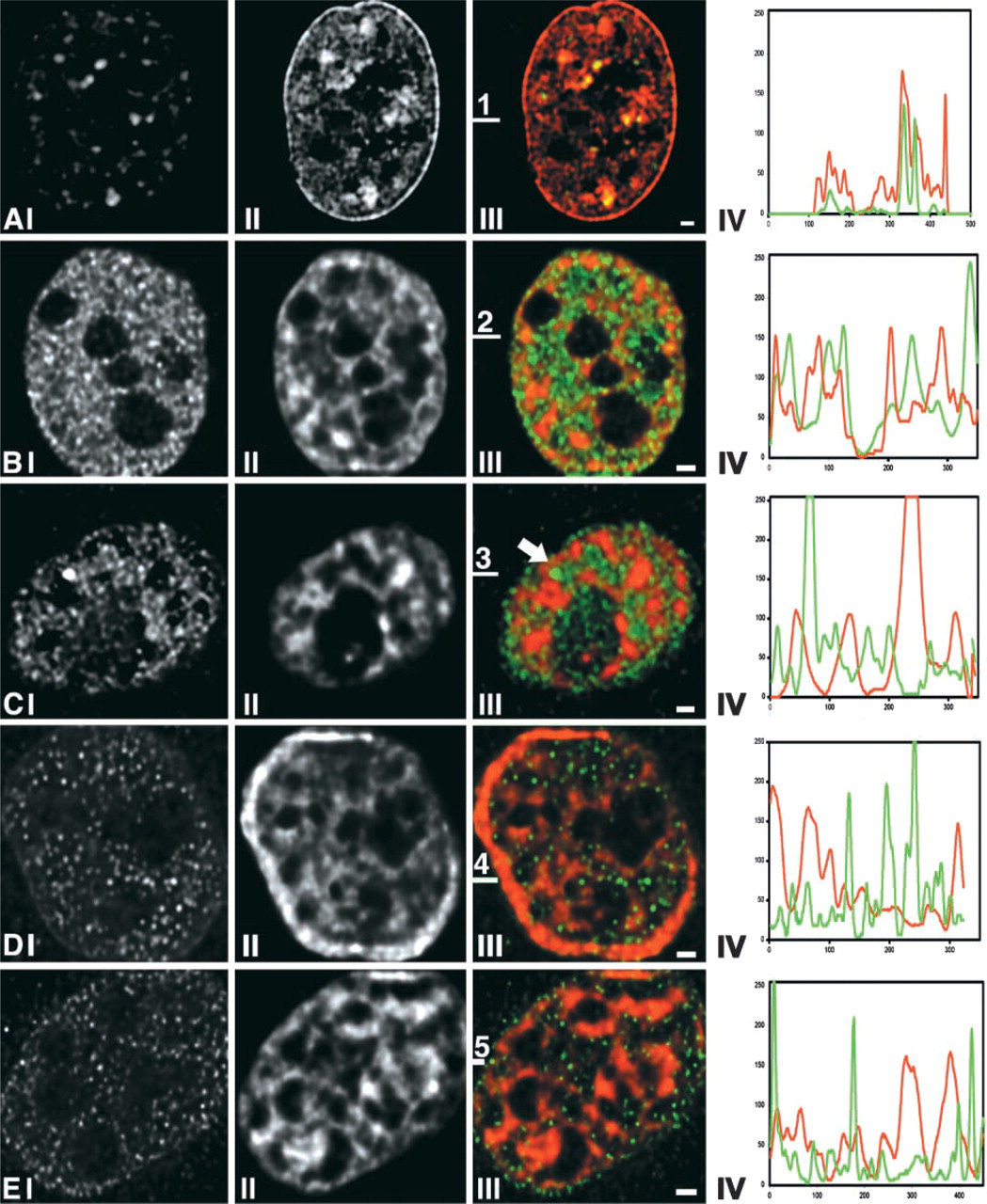
Relationship between large-scale chromatin organization and the spatial distribution of the general transcription factor TFIIH, RNA polymerase II, and hnRNP-U. HeLa cells expressing GFP-tagged histone H2B were immunofluorescently labeled with antibodies against nuclear components involved in gene expression: TFIIH (
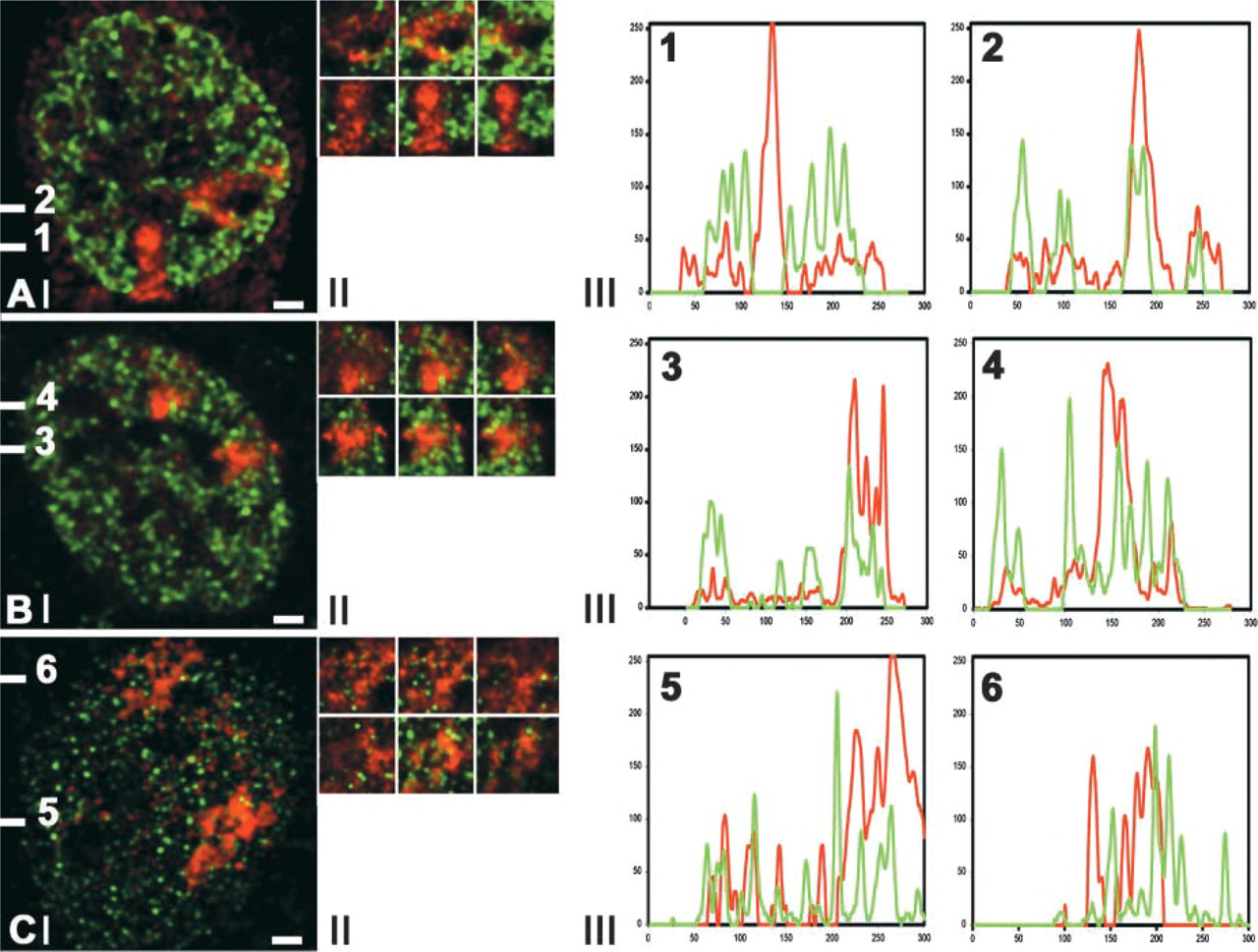
Spatial relationship between X-chromosome territories in primary female fibroblasts and RNA polymerase II, histone H4 acetylated at lysine 8, and hnRNP-U. In primary human female fibroblasts the distributions of histone H4 acetylated at lysine 8 (
In primary human female fibroblasts we analyzed the distribution of RNA polymerase II and of acetylated histone H4 with respect to both × chromosomes, of which one is transcriptionally active and the other is inactive (Figure 2A). For the labeling of acetylated histone H4 we used antibodies that recognize histone H4, which is acetylated at lysine 8 (antibody R232) or at lysine 16 (antibody R252). These histone H4 isoforms are associated with transcriptionally competent chromatin (Jeppesen 1997; Utley et al. 1998). Figure 2A shows that histone H4 acetylated at lysine 8 occurs scattered throughout the chromosome territory of only one of the two × chromosomes. The same result was obtained for histone H4 acetylated at lysine 16. This is in agreement with results of others demonstrating a lack of histone H4 acetylation in the inactive × chromosome (Jeppesen and Turner 1993; Sadoni et al. 1999).
In contrast to acetylated histone H4, we find that RNA polymerase II is present throughout both × chromosome territories (Figure 2B), indicating that this protein has access to the interchromatin space in the inactive X-chromosome territory equally well as to that of its transcriptionally active counterpart. A similar result was obtained for hnRNP-U, which is a member of the large group of hnRNP proteins that are involved in RNA packaging, processing, and transport (PinolRoma 1997). Double-labeling experiments of hnRNP-U and × chromosomes showed the presence of hnRNP-U throughout both X-chromosome territories (Figure 2C). Moreover, we observed that the distribution of both RNA polymerase II and hnRNP-U is largely excluded from the distribution of the condensed chromatin regions of the active and inactive × chromosomes (Figures 2B and 2C). In Figure 1E we compare the distribution of the hnRNP-U protein with that of GFP-tagged chromatin in HeLa cells, showing that in these cells also hnRNP-U is largely excluded from condensed chromatin. Evidently, hnRNP-U-containing complexes are unable to access most of the condensed chromatin domains.
The Chromatin Distribution in the Interphase Nucleus Does Not Require Ongoing Transcription
Results thus far are consistent with the idea that the interchromatin space is a depot for factors required for transcription, RNA processing and transport. Does this mean that RNA synthesis and processing are essential for maintaining the spatial distribution of chromatin in transcriptionally active nuclei? To address this question, we compared the distribution of chromatin in HeLa cells that express GFP-tagged histone H2B before and after inhibition of transcription with 100 μg/ml α-amanitin, which blocks RNA polymerases I and II but does not affect RNA polymerase III. Alternatively, we inhibited transcription by exposing cells to 50 μM 5,6-dichloro-α-
In addition, we analyzed whether inhibition of RNA polymerases changes the amount and the distribution of acetylated histone H4 (Jeppesen 1997; Utley et al. 1998). Labeling was performed with an antibody recognizing histone H4 acetylated at lysine 8, a marker for transcriptionally competent chromatin. The distribution of acetylated histone H4 also showed no major changes (Figure 3C). Moreover, the amount of acetylated H4 in the nucleus did not change detectably after inhibition of transcription. We conclude that the overall nuclear distribution of chromatin does not depend on ongoing transcription by RNA polymerases I and II. In addition, it appears that the distribution of silent chromatin (hypo-acetylated histone H4) and of transcriptionally competent chromatin (hyper-acetylated histone H4) does not change. Line scans of these findings are shown in Figure 3.
Discussion
Two major compartments can be distinguished in mammalian interphase nuclei. One is composed of apparently interconnected condensed chromatin domains and the other is the interchromatin compartment, i.e., the space between the compact chromatin domains, which is largely devoid of chromatin (Zink and Cremer 1998). The interchromatin compartment consists of a highly convoluted network of channels between and inside chromosome territories. This notion is mainly based on light and electron microscopic observations (Fakan 1994; Zink et al. 1998; Cmarko et al. 1999; Verschure et al. 1999). Importantly, chromatin is folded so that transcriptionally active loci are located exclusively at the interface between condensed chromatin domains and the interchromatin compartment, i.e., in direct contact with the interchromatin space (Fakan 1994; Cmarko et al. 1999; Verschure et al. 1999).
In the present study we focused on the spatial distribution of nuclear components involved in RNA synthesis and RNA processing in relation to the large-scale organization of chromatin in the nucleus. We systematically analyzed the spatial distribution of the following nuclear components involved in transcription and RNA processing and transport in relation to condensed chromatin domains and the interchromatin compartment: (a) the general transcription factor TFIIH, (b) RNA polymerase II, and (c) hnRNP-U, which is involved in RNA packaging and processing. Chromatin was visualized in HeLa cells that express GFP-tagged histone H2B (Kanda et al. 1998; Verschure et al. 1999). Results show that these proteins are present in the interchromatin compartment and are largely excluded from condensed chromatin domains (Figures 1B–1E). We demonstrate that this is not due to the inaccessibility of antibodies for antigens inside compact chromatin domains. Using primary human female fibroblasts, we show that these proteins have access to the interchromatin space in both the transcriptionally active and the transcriptionally inactive × chromosome. The two × chromosomes could be discriminated on the basis of the difference in their histone H4 acetylation level (Figure 2) (Jeppesen and Turner 1993; Sadoni et al. 1999). This observation is consistent with the notion that most nuclear proteins can diffuse freely throughout the nucleus (Phair and Misteli 2000), most likely via the interchromatin compartment. Evidently the interchromatin space is continuous and accessible in nuclear areas that are transcriptionally active, as well as in areas that are transcriptionally silent. Moreover, in these primary cells RNA polymerase II and hnRNPU proteins are excluded from the condensed chromatin regions of both the active and inactive × chromomosomes, as was observed in HeLa cells expressing histone H2B-tagged GFP. This suggests that the exclusion of nuclear proteins from condensed chromatin regions is a common organization principle.
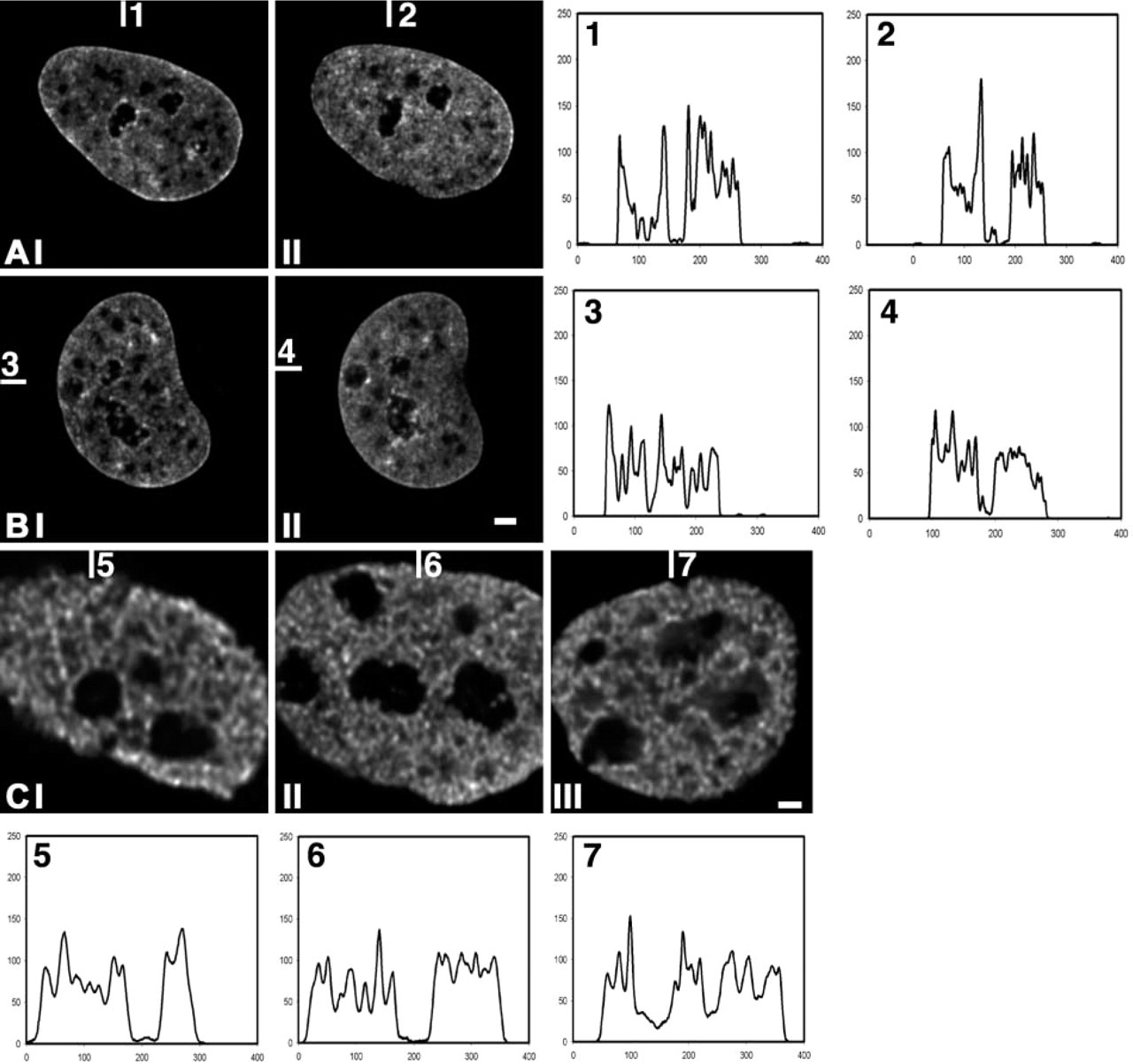
The spatial distribution of chromatin after inhibition of RNA polymerase II. (
Our results are in line with recent results from Cmarko et al. (1999), who showed by electron microscopy that RNA polymerase II and TFIIH are associated with perichromatin fibrils, the ultrastructural form of nascent transcripts, and that TFIIH is not present in condensed chromatin domains. Similarly, the TAF250 protein, which is part of the transcription factor TFIID, localizes to regions distinct from both the highly acetylated chromatin and the condensed heterochromatin (Hendzel et al. 1998). Like transcription and RNA processing components, poly-A RNA has also been found to be confined to the interchromatin space (Politz et al. 1999). Our data support the idea that the interchromatin compartment gives the transcriptional and RNA processing machinery access to the sites of RNA synthesis that are localized at the surface of condensed chromatin domains (Fakan 1994; Cmarko et al. 1999; Verschure et al. 1999).
Several groups have presented evidence that large-scale chromatin organization in the nucleus depends on transcriptional activity. Haaf and Ward (1996) showed that inhibition of RNA polymerase II results in chromatin decondensation in human fibroblasts and in lymphocytes, whereas others (Croft et al. 1999) reported just the opposite, i.e., that inhibition of transcription results in chromatin compaction. Moreover, priming chromatin for transcription by binding of an activating transcription factor, or by hormonal stimulation, results in a dramatic decondensation of chromatin (Tumbar et al. 1999; Volpi et al. 2000; Ye et al. 2001). Here we show that inhibition of transcription by α-amanitin or DRB has no detectable effect on large-scale chromatin organization in HeLa cell nuclei. Comparison of the chromatin distribution before and after inhibition of transcription in one and the same living cell gave no evidence for a change in large-scale chromatin structure. In addition, we found that in HeLa cell nuclei the amount and the distribution pattern of acetylated histone H4 did not change after inhibition of transcription. We suspect that the chromatin expansion and condensation after inhibition of RNA synthesis that was observed by others (Croft et al. 1999; Haaf and Ward 1996) was due to the fixation and/or labeling procedures that were used. Our results show that, within the limits of resolution of light microscopy, large-scale chromatin structure does not depend on ongoing transcription.
What is the reason that factors involved in transcription and in RNA processing do not enter condensed chromatin domains? One obvious answer is that they are excluded by steric hindrance due to the tight packing of nucleosomes. This argument is often taken as the explanation for the inactivity of genes that are present in a heterochromatin environment. However, heterochromatin-specific genes have been described that are actively transcribed inside heterochromatin domains (Hilliker et al. 1980; Lu et al. 2000), indicating that transcription factors must be able to enter condensed heterochromatin. Our observations showing that factors such as RNA polymerase II and TFIIH are predominantly present in the interchromatin space are consistent with the results of Politz et al. (1999), showing that poly-A RNA is found exclusively in the interchromatin compartment. Our results appear to contradict the fact that at least some active genes are found inside heterochromatin. However, it should be noted that the number of such genes is relatively small. Therefore, they are probably difficult to visualize.
Summarizing, our results support a simple model for the functional organization of the interphase nucleus. Transcriptionally active genes are localized at the surface of relatively condensed chromosomal domains in the perichromatin compartment in contact with the interchromatin compartment. In this way, components of the transcriptional machinery that are present in the interchromatin space have unhindered access to regulatory sequences of chromatin. Moreover, newly synthesized RNA can be deposited directly into the interchromatin space, where factors that are required for processing, packaging, and transport through the interchromatin channels are readily available. Thus, large-scale folding of chromatin and compartmentalization of nuclear components create a functional nuclear organization that supports efficient gene expression.
Footnotes
Acknowledgments
Supported by the EU Biomed II program (project number BMH4-CT95-1139) and by a grant to PJV from the Biological branch (ALW) of the Dutch Research Council (NWO) (project number 805–48011).
We thank Dr B.M. Turner (University of Birmingham, UK) for kindly providing us with antibodies recognizing acetylated histone H4 and Prof Dr P. Chambon (Université Louis Pasteur, Strassbourg, France) for the generous gift of the anti-HP1 antibodies. We are grateful to Dr G. Wahl (Salk Institute for Biological Studies, La Jolla, CA) for kindly providing us with HeLa cells transfected with histone H2B-GFP fusion protein. We thank Prof Dr C.J.F. van Noorden (University of Amsterdam, The Netherlands) for critical reading of the manuscript.
