Abstract
Vision-language models (VLMs) have demonstrated significant potential in medical image analysis, yet their application in intraoral photography remains largely underexplored due to the lack of fine-grained, annotated datasets and comprehensive benchmarks. To address this, we present MetaDent, a comprehensive resource that includes 1) a novel and large-scale dentistry image dataset collected from clinical, public, and web sources; 2) a semistructured annotation framework designed to capture the hierarchical and clinically nuanced nature of dental photography; and 3) comprehensive benchmark suites for evaluating state-of-the-art VLMs on clinical image understanding. Our labeling approach combines a high-level image summary with point-by-point, free-text descriptions of abnormalities. This method enables rich, scalable, and task-agnostic representations. We curated 60,669 dental images from diverse sources and annotated a representative subset of 2,588 images using this meta-labeling scheme. Leveraging large language models (LLMs), we derive standardized benchmarks: approximately 15,000 visual question answering (VQA) pairs and an 18-class multilabel classification dataset, which we validated with human review and error analysis to justify that the LLM-driven transition reliably preserves fidelity and semantic accuracy. We then evaluate state-of-the-art VLMs across VQA, classification, and image captioning tasks. Quantitative results reveal that even the most advanced models struggle with a fine-grained understanding of intraoral scenes, achieving moderate accuracy (e.g., less than 70% in VQA) and producing inconsistent or incomplete descriptions in image captioning. These findings underscore the gap between general-purpose VLMs and the demands of specialized models, highlighting the need for domain-adapted training and more sophisticated evaluation protocols to assist professional dental practice and community oral health efforts. We publicly release our dataset, annotations, and tools to foster reproducible research and accelerate the development of vision-language systems for dental applications.
Keywords
Introduction
Dental photography plays a crucial role in diagnosis, treatment planning, patient education, and documentation across dental disciplines (Ding et al 2023; Caron et al 2025; Mania et al 2025). In this context, recent advancements in vision-language models (VLMs) offer a compelling opportunity for automated and scalable image interpretation. By leveraging large-scale annotated datasets, VLMs have demonstrated remarkable capabilities in understanding and reasoning about visual content in natural images (Radford et al 2021; Liu et al 2023).
In health care research, VLMs have been increasingly utilized in analyzing visual and textual data, aiding professionals in detection, diagnosis, and reporting to improve efficiency (Li et al 2023; Ghosh et al 2024; Ryu et al 2025). Despite these advances, few studies report VLM performance for dental image understanding. A concurrent study established a benchmark for panoramic radiograph analysis and found no significant difference between general-purpose and medical-specific VLMs in dentistry (Hao et al 2025). Another study collected a mixture of large-scale X-ray and intraoral images and fine-tuned a VLM using derived data mainly from categorical labeling (Meng et al 2025). While the result looks promising, the dataset and the model are not publicly available.
Based on our preliminary observations, even state-of-the-art VLMs struggle with a fine-grained understanding of intraoral images. We attribute this limitation to 2 interrelated challenges:
Dental diagnosis often requires nuanced, hierarchical interpretations that go beyond closed-set classification. Current VLMs, trained primarily on categorical labels or vague text descriptions, lack the capacity to reason over such fine-grained and clinically contextual features.
The development and evaluation of VLMs for dentistry are severely constrained by the scarcity of well-annotated, diverse, and publicly available datasets. Most existing datasets for intraoral images focus on specific categories (Dot et al 2024; Nguyen et al 2025; Wang et al 2025) and lack diversity across different data sources (Huang et al 2024; Uribe et al 2024).
To address the above limitations, we curated a dedicated dataset and benchmarked state-of-the-art VLMs, introducing innovations in 2 key aspects: annotation strategy and data sourcing. First, we proposed a novel semistructured labeling strategy that formulates the annotation as an open-set weak labeling task for abnormality detection. This approach provides a compact yet comprehensive representation of each image, which can be reliably translated into various downstream task-specific formats. Second, we constructed a large-scale image dataset with substantial diversity by filtering and curating images from web-scraped data, and then we labeled a subset with the proposed semistructured scheme to enable both precise annotation of abnormalities and holistic scene understanding.
In this work, we introduce MetaDent, a semistructured annotation framework and large-scale resource for vision-language understanding of intraoral images. MetaDent is designed to support diverse downstream tasks—including visual question answering, multilabel classification, and image captioning. The name “MetaDent” reflects its meta-annotation structure, broad data diversity, and adaptability across a wide range of dental artificial intelligence (AI) applications.
To encourage reproducibility and further research, we publicly release the dataset, annotation interface, and benchmarking tools at https://menxli.github.io/metadent/.
Materials and Methods
Ethics Statement
The study protocol was approved by the institutional ethics committee of the Hospital of Stomatology, Wuhan University (No. WDKQ2025[C02]). The study adhered to the ethical considerations outlined by the committee, including participant privacy and data protection measures.
Meta Dataset Curation
We collected images from 3 sources: in-house collected, public dataset, and web-crawled. Specifically, we collected 4,373 clinical photographs from the Department of Prosthodontics at the School of Stomatology, Wuhan University (Data Source 1, DS1). For the public component, we included 9,390 images from the Teeth or Dental image dataset (Data Source 2, DS2) (Chaudhary et al 2024), given its relatively large quantity and good quality. Lastly, we filtered the COYO-700M (Byeon et al 2022), a large-scale web-crawl image corpus, using a fine-tuned ViT-L/16 binary classifier to retain only dental images and an image hasher (Haviana and Kurniadi 2016) for duplicate removal (details in Appendix Section 1). This step resulted in 46,906 images (Data Source 3, DS3). In total, the collected dataset comprises 60,669 dental images.
From this combined image dataset, we randomly sampled a subset of 3,576 images for human review. We excluded images lacking clinical relevance (e.g., artificial images) or sufficient quality (e.g., blurry images), resulting in a final set of 2,588 images for annotation. The annotation follows a semistructured format, where each image is assigned an overall descriptive summary and a list of identified abnormalities (Fig. 1). The overall description is a concise paragraph that introduces the main content of the image as well as the shooting perspective. Abnormalities were defined as any clinically relevant deviations from normal dental anatomy or healthy tissue appearance. Annotators were instructed to list the abnormalities point-by-point using natural, unstructured language focused on visual appearance, including diagnostic interpretations where applicable. Meanwhile, the annotators also draw a rough contour for each abnormality in the image for its corresponding entry.
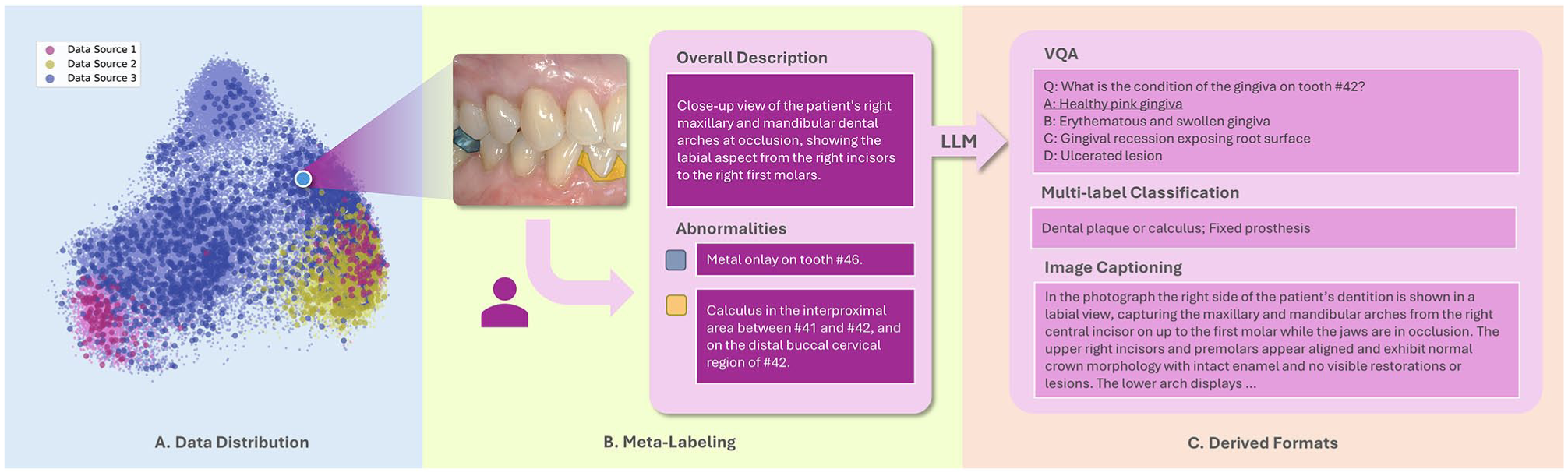
Data processing pipeline. (
All images were annotated by 2 dentists with 7 and 4 y of clinical experience, respectively. Prior to formal annotation, the annotators underwent a 1-mo training, during which they received guidance from a senior dentist with 10 y of clinical experience. Training sessions included experimental annotation and corrective feedback. During the formal annotation, any ambiguous or challenging cases were discussed among all 3 experts. When the visual evidence was inconclusive but the annotators had moderate confidence, descriptions were used with cautious wording. When confidence was very low, the entry was flagged as “uncertain.” To assess the consistency between the raters, interrater reliability was evaluated. Cohen’s κ coefficient, calculated on 100 images randomly drawn from the dataset, was 0.83 (Appendix Section 10), indicating a high level of agreement. To facilitate labeling all abnormalities exhaustively, after the initial annotation phase, the 2 annotators cross-verified each other’s labels, ensuring that every image was reviewed by at least 2 raters. The initial annotations were in Chinese and translated into English via large language models (LLMs); all LLM-assisted steps in the study were carried out using GPT-OSS-120B (OpenAI et al 2025). Of the 2,588 images processed, the labeling effort yielded 6,314 confidently annotated records and 138 entries flagged as uncertain.
Secondary Dataset Generation
Following the meta-dataset labeling, we converted the semistructured data into standardized formats with the help of an LLM. In particular, we chose 3 tasks: visual question answering (VQA), multilabel classification, and image captioning. This process effectively expands its size and applicability for diverse downstream tasks. The integration of LLM is based on 2 motivations: 1) our semistructured label is, in essence, a compact and complete representation of the intraoral image, which makes the deduction of unmentioned normal structures possible, and 2) current VLMs still face limitations in reliability, particularly in specialized domains (Jeong et al 2024; Nath et al 2025). In contrast, LLM is far more reliable in terms of reasoning and less hallucination when given an appropriate context (Li et al 2024). We harnessed linguistic reasoning to achieve more robust and scalable data processing.
For the VQA generation, we allowed the model to reason about nonmentioned common abnormalities and questioning beyond explicit labels. We generated 5 to 10 questions per image depending on the number of labeled entries. When fewer than 2 abnormalities were labeled in an image, we generated 5 questions; otherwise, 10 would be raised. The question types included judgment (true/false) and multiple-choice. To enhance the quality of the generated VQA pairs, a self-refinement step was applied (Madaan et al 2023). Entries marked as “uncertain” were handled with exclusion from downstream analyses to minimize ambiguity (Appendix Section 5).
For the classification, we established 18 classes based on visual appearance, with clinical pathology serving as a guiding reference (Fig. 4C and Appendix Table 1). For example, “chalky patches” was grouped into “tooth color abnormality,” regardless of whether they stemmed from early caries, enamel hypoplasia, fluorosis, or postorthodontic lesions. Likewise, plaque and calculus were sometimes visually indistinguishable in photographs and merged into 1 class. This image-centric approach balances practicality and medical relevance and is presumably more effective than a strict clinical diagnosis from the perspective of image analysis.
For the image captioning, the LLM was prompted to generate free-form descriptions from the meta-labels, which were used as reference captions.
To ensure data quality and to better understand the sources of errors, we analyzed errors arising during the transition, focusing on VQA and multilabel classification. During dataset generation, the LLM was prompted to provide explanations for its answers to support human review. Two annotators evaluated the entire classification dataset and randomly sampled 2 VQA pairs per image to analyze errors and make necessary revisions or deletions. We identified and defined 6 common error types; detailed descriptions of each are provided in Appendix Table 2.
The above steps resulted in 18,416 VQA pairs and 2,588 multilabel classification questions for subsequent assessment. Prompts for generation are supplied in Appendix Section 5.
Dataset Summary
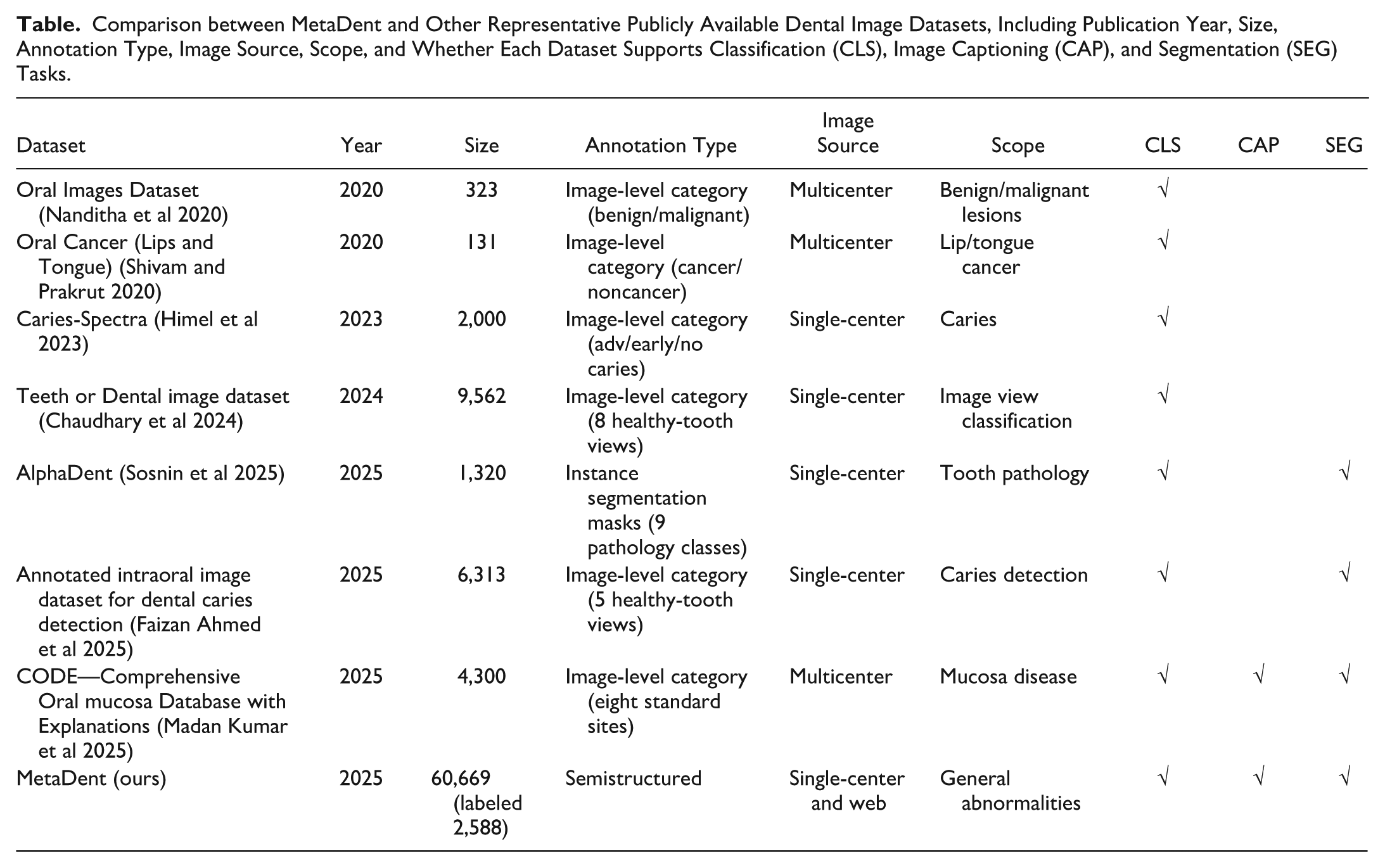
Figure 2 characterizes MetaDent’s visual properties. Most images span 105 to 106 pixels (width: ~200–1,000 px), with aspect ratios varying by source: DS3 shows the broadest spread, with DS2 the most uniform (Fig. 2A, B). Analysis by Ovis (Lu et al 2024) reveals 80% depict humans; among these, eyes and nose are detected for fine-grained categorization (Fig. 2C). Intraoral views comprise 63.5% of the dataset and facial images 5.0% (deidentified upon release). In the labeled subset, most images contain 1 to 3 annotated abnormalities (Fig. 2D). In the Table, we compare MetaDent against existing intraoral image datasets. The key gap that MetaDent uniquely fills is not merely dataset size but the combination of large-scale coverage, semistructured meta-annotation, and broad task generality within a single unified framework.
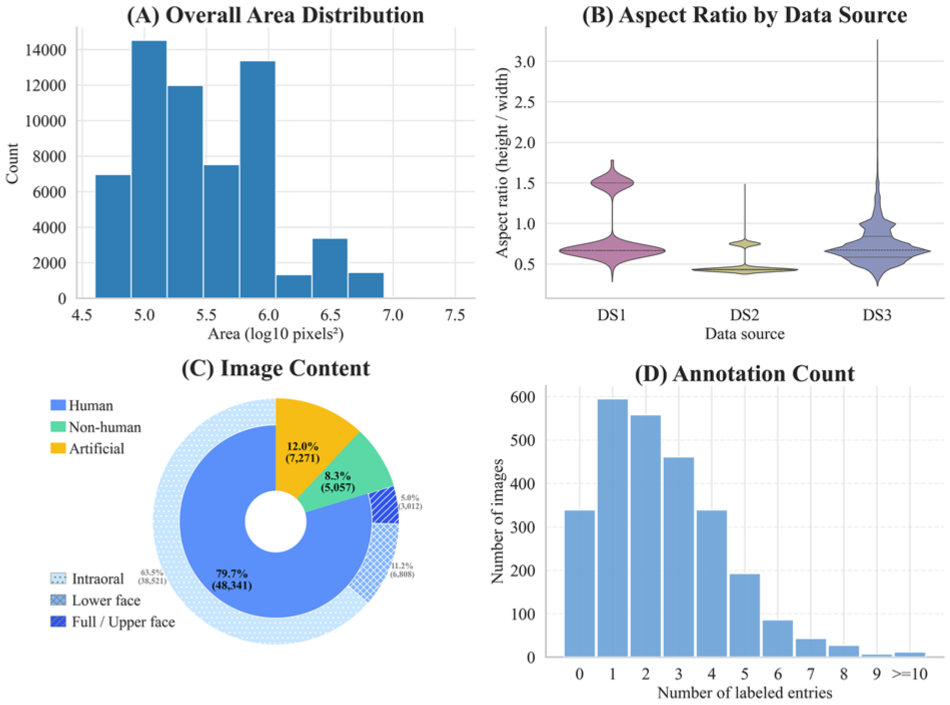
Dataset statistics. (
Comparison between MetaDent and Other Representative Publicly Available Dental Image Datasets, Including Publication Year, Size, Annotation Type, Image Source, Scope, and Whether Each Dataset Supports Classification (CLS), Image Captioning (CAP), and Segmentation (SEG) Tasks.
Evaluation of the VLMs
We evaluate VQA performance using accuracy, defined as the proportion of questions answered correctly. This evaluation is performed separately for multiple-choice and true/false question types. For multilabel classification, we report precision, recall, and F1-score, along with Exact Match—the predicted labels exactly match the ground truth. The F1-score, defined as the harmonic mean of precision and recall, reflects the balance between false positives and false negatives. In image captioning, we evaluate the generated captions using LLM-as-a-judge from semantic and diagnostic consistency perspectives. Semantically, we prompt the LLM to generate a reference caption and compare it with the outputs from the VLMs using BERTScore (Zhang et al 2020), which computes token-level contextual embeddings and measures how well the generated caption semantically aligns with the reference description. At the diagnostic consistency level, we instructed the LLM to evaluate whether the generated caption correctly identifies the key diagnostic findings and anatomical location of abnormalities, as specified in the meta-label. This essentially transforms the evaluation into an open-set multilabel classification task. Metrics were computed both per data source and over the full dataset. Detailed definitions of each metric are provided in Appendix Section 2.
The VLMs included in our study represent the current frontiers in vision-language understanding, including proprietary models: GPT-4o-2024-08-06 (OpenAI et al 2024) and Gemini-2.5-flash (Comanici et al 2025), as well as leading open-source models: Qwen3-VL-235B-A22B-Thinking (Yang et al 2025), Ovis2-34B (Lu et al 2024), and Baichuan-Omni-1.5 (Li et al 2025). Several models occasionally produced nonresponses or format-inconsistent outputs; these cases were excluded from analysis, and the exclusion rate is reported in Appendix Table 13. To ensure VLMs did not benefit systematically from sharing linguistic priors with the LLM used in benchmark construction, we repeated the evaluation using an alternative LLM—results showed no noticeable bias (Appendix Section 9).
Results
Datasets
In Figure 1A, image features were extracted using DINOv3 (Siméoni et al 2025) and subsequently dimensionally reduced using principal component analysis (PCA). In the plot, labeled data points are represented by darker, larger dots, while unlabeled raw images appear as lighter, smaller dots. The visualization reveals that the Internet-scraped images (DS3) dominate the dataset and occupy a broader region, indicating they introduce greater visual diversity and cover a wider range of scenarios. Moreover, the labeled data were sampled uniformly from the entire distribution. This justifies that the annotated subset is representative of the overall dataset without significant sampling bias.
To assess benchmark quality, we conducted a human review of the VQA subset (5,176 randomly sampled question-answer pairs from 18,416 total) and the entire classification dataset (2,588). Annotators rated 94.2% (4,875) of VQA pairs as correct; for classification, 90.5% of images (2,340) had perfectly correct labels, while 248 (9.5%) contained 1 or more errors. This evaluation confirms the high overall fidelity of the LLM-generated datasets from the meta-labels. The error analysis for VQA and classification dataset generation is illustrated in Appendix Figure 4; the incorrect responses spanned a mix of the error types.
Figure 4B shows the manual classification of the randomly sampled labeled subset, reflecting overall pathology prevalence: “malocclusion or dental malalignment” is most common, while “residual root” and “oral ulcer” are less frequent.
VQA
All evaluated VLMs achieve moderate accuracy in VQA (Fig. 3A, B). True/false questions (TFQ) consistently receive higher accuracy than multiple-choice questions (MCQ). Gemini-2.5-Flash led performance (MCQ 64.1%, TFQ 67.9%), slightly ahead of GPT-4o (60.6% MCQ, 67.4% TFQ). Open-source models lagged behind: Ovis-2 (61.7% MCQ, 67.3% TFQ), Qwen3-VL (60.4% MCQ, 63.9% TFQ), and Baichuan-Omni (57.5% MCQ, 64.5% TFQ). Performance varied by image source: all models performed slightly better on DS3. Nevertheless, results remained suboptimal, as no model surpassed 68% accuracy on any question type, highlighting the difficulty of fine-grained intraoral VQA and the modest advantage of proprietary over open-source models.
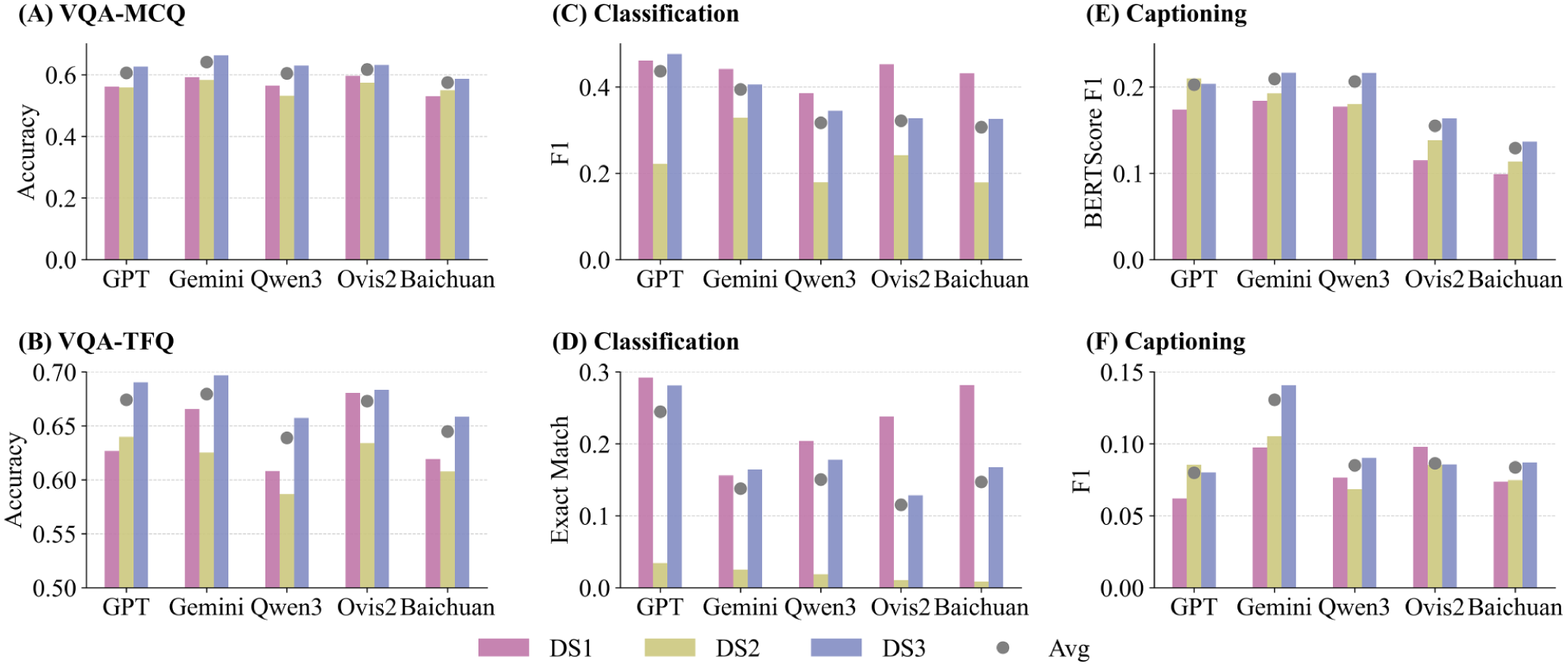
Performance of 5 vision-language models on MetaDent across 3 tasks. (
Classification
Multilabel classification was challenging for all models (Figs. 3C, D and 4). GPT-4o achieved the highest F1-score (0.437) with balanced precision and recall (0.475/0.438), outperforming Gemini-2.5 (F1 ≈ 0.394, higher recall 0.444 but lower precision 0.398). Open-source models scored lower (F1 ≈ 0.30–0.33). Exact Match accuracy was very low (GPT-4o 24.5%; others <16%), showing models rarely predicted all labels correctly. Overall, even the best model detected less than half of all findings.
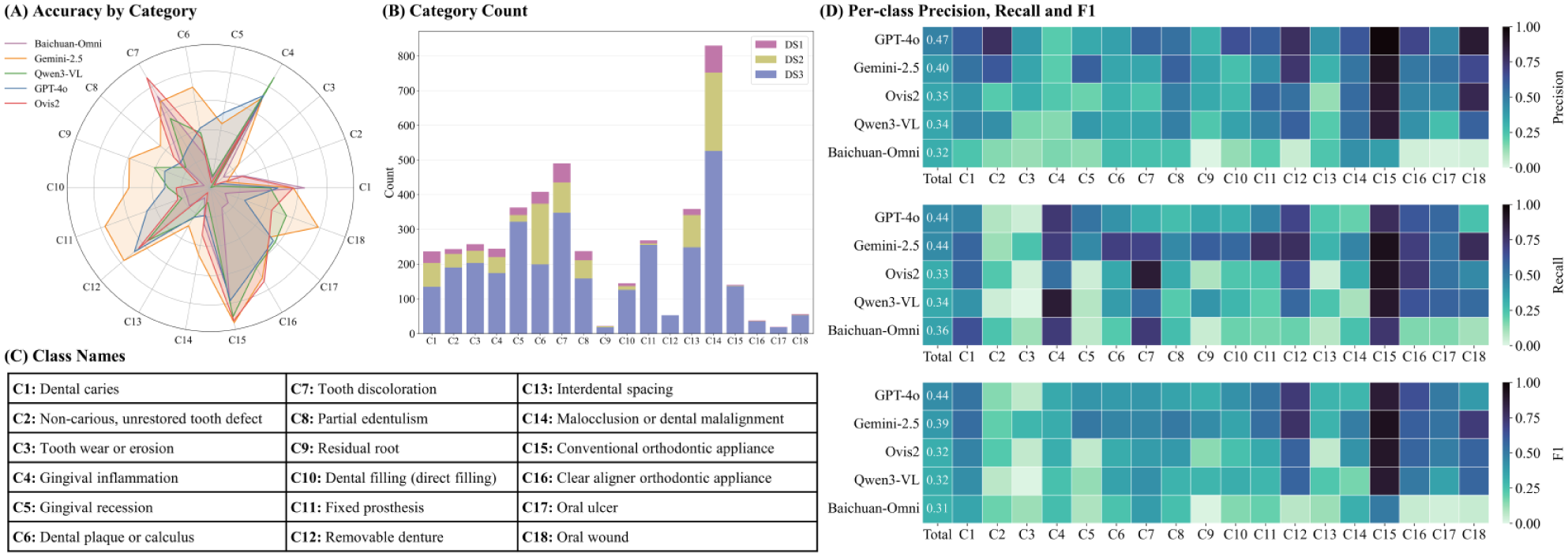
Category-level performance for the multilabel classification task. (
Image Captioning
Models struggled with accurate free-form captions (Fig. 3E, F). Gemini-2.5 achieved the highest semantic similarity to references (BERTScore-F1 ≈ 0.209), followed by Qwen3-VL (0.206), GPT-4o (0.203), Ovis-2 (0.155), and Baichuan-Omni (0.129). Appendix Section 9 reveals slightly different styles: GPT-4o (P ≈ 0.112, R ≈ 0.072) and Ovis (P ≈ 0.108, R ≈ 0.084) favored precise captions but led to less sensitivity (low recall), while the others were more balanced. Consistency remained low across all models (F1 ≈ 13% at best), indicating most key findings were missed or incorrectly interpreted. In summary, captions generated by VLMs are often unreliable for clinical interpretation.
Discussion
In this study, we focus on the limitations of VLMs for intraoral image understanding and propose a semistructured meta-labeling technique to support analysis of clinical images. Unlike conventional categorical or textual labels, our approach generates rich, hierarchical representations that capture semantic meaning at arbitrary levels of detail, enabling effective transfer to downstream tasks. To support this work, we collected a diverse dataset of dental images primarily from online sources and carefully annotated a subset to evaluate representative VLMs. Our results show that state-of-the-art models struggled across all tasks—barely reaching about 65% to 70% accuracy in VQA and around 0.4 F1 in multilabel classification—and their image captions often miss important findings. We further observe a sharp performance degradation on DS2, suggesting the presence of a domain shift. A more detailed analysis of the challenges associated with DS2 indicates that the performance drop may be attributed to shifts in image tone and demographic differences between datasets (detailed analysis in Appendix Section 3). These findings highlight that current VLMs, even cutting-edge systems such as GPT-4o, are not yet reliable for deployment in clinical settings without further refinement.
In contrast to prior studies that typically rely on photographs from a single institution or a limited number of clinical centers, we adopted a different approach by primarily utilizing web-scraped images. Specifically, we used a filtered subset of the COYO-700M dataset, which was originally sourced from Common Crawl (2008)—a large-scale web archive. This set of images contains diverse photos compared to single-center or public single-source datasets, as shown in Figure 1A. This diversity in imaging conditions, lighting, and patient demographics enhances the generalizability of the dataset, reducing bias and enabling better real-world applicability.
Another contribution of this study lies in the labeling protocol. The advantages of the proposed technique are as follows:
It effectively captures the hierarchical complexity of intraoral conditions that conventional categorical labels cannot represent. For example, a full crown may vary in material, color, and defects, which cannot be documented comprehensively with a single category and can be described simultaneously within our structure (Appendix Section 7). This enables richer, more precise supervision signals for VLMs.
Compared with free-form captions commonly used in general-domain VLMs, the proposed scheme offers a more efficient and clinically meaningful annotation process. By focusing on abnormalities rather than exhaustive descriptions, it reduces redundancy. Normal conditions can be inferred from the absence of abnormalities, minimizing annotation workload without compromising interpretability. Moreover, the point-by-point label structure also facilitates the attachment of metadata (e.g., bounding boxes, segmentations) for downstream applications.
By balancing comprehensiveness with conciseness, the proposed method provides a compact yet comprehensive annotation of the image by a brief summary of the main visual content and a structured list of free-form abnormalities. By leveraging the strong reasoning capabilities and dental knowledge of LLMs, we can easily scale the dataset to different formats for downstream tasks. These models can not only assist in diagnosis to reduce errors but also enable applications such as smart health care management, early-stage oral disease screening, and self-conducted oral health checks at home. Some of the potential use cases are listed in Appendix Section 8.
The primary limitation of this work lies in the relatively small size of the labeled dataset, which is insufficient to claim broad coverage of intraoral conditions and is hard to support fine-tuning of large, domain-specific VLMs. Additionally, most of the images are web-scraped, introducing variability in quality and uncertain provenance. The labels combine human annotations with LLM-generated outputs, which may introduce noise. That said, LLM-assisted data generation and quality control are common in related research, and we quantitatively assessed error sources to support a certain level of confidence in the dataset’s overall reliability. As large models continue to advance, these techniques are expected to yield even higher-quality data. Lastly, while our benchmark is in large quantity and reasonable quality, we acknowledge the absence of direct comparison to human expert performance.
Despite these constraints, our benchmark offers a robust and challenging evaluation platform for multimodal models in dentistry. We release the dataset, an initial set of annotations, together with the labeling tools, hoping this work will encourage community-driven expansion, supporting fine-tuning, regulatory validation, and ultimately aid the development of more robust and practically useful models in oral health.
Conclusion
In this work, we proposed a semistructured annotation framework for intraoral image analysis that enables rich, scalable, and task-agnostic representations. By curating a diverse dataset and standardized benchmarks, we evaluated state-of-the-art vision-language models and revealed their limitations in fine-grained dental understanding. The result reveals that the gap between current VLM capabilities and dental requirements remains wide. Closing this gap requires concerted efforts and cross-disciplinary collaboration between AI researchers and dental practitioners. By providing the community with a roadmap and tools, we aim to help such interdisciplinary synergy to advance multimodal AI in dentistry—ultimately moving us closer to practically useful multimodal AI in oral health care.
Author Contributions
M.-X. Li, W.-H. Deng, contributed to conception and design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; Z.-X. Wu, C.-X. Jin, contributed to data acquisition, analysis, and interpretation, critically revised the manuscript; J.-M. Wu, J. K. H. Tsoi, contributed to data conception and design, critically revised the manuscript; Y. Han, contributed to data analysis, drafted and critically revised the manuscript; G.-S. Xia, C. Huang, contributed to data conception and design, drafted and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345261424242 – Supplemental material for MetaDent: Labeling Clinical Images for Vision-Language Models in Dentistry
Supplemental material, sj-docx-1-jdr-10.1177_00220345261424242 for MetaDent: Labeling Clinical Images for Vision-Language Models in Dentistry by M.-X. Li, W.-H. Deng, Z.-X. Wu, C.-X. Jin, J.-M. Wu, Y. Han, J. K. H. Tsoi, G.-S. Xia and C. Huang in Journal of Dental Research
Footnotes
Acknowledgements
The authors gratefully acknowledge Chao Pang for helpful discussions and Zhong-Shi Zhang, Yu-Jie Wu, and Mu-Qi Jiang for their valuable contributions to data processing and their unwavering support throughout this work.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the National Natural Science Foundation of China (82401200), Natural Science Foundation of Hubei Province (2024AFB033), Key R&D Program of Hubei Provincial Department of Science and Technology (2023BAB058), and National College Students Innovation and Entrepreneurship Training Program (202510486171).
Data Availability
A supplemental appendix to this article is available online.
