Abstract
Current therapeutic practices have limited effectiveness at targeting acute pain locally. This is in part due to the lack of understanding of the neuroinflammatory pathways active within pulpitis. Increased activity of inadoleamine 2,3 dioxygenase (IDO) was linked to hypersensitivity and pain in several settings. IDO is the rate-limiting enzyme of the kynurenine pathway, but changes in IDO’s activity and expression in pulpitis have not been investigated. Tooth samples with a diagnosis of symptomatic irreversible pulpitis (SIP) were collected, and levels and activity of IDO and other markers of the kynurenine (KYN) pathway were examined using high-performance liquid chromatography (HPLC), quantitative polymerase chain reaction (PCR), and immunocytochemistry (ICC). Subsequently, odontoblast-like cells (OLCs) derived from dental pulp stem cells were stimulated in vitro with bacterial lipopolysaccharide and a synthetic proxy for DNA cytosine-phosphate-guanine (CpG) oligonucleotide 2006 (ODN 2006). The effects of these compounds on KYN pathway marker levels and transcription were then analyzed using quantitative PCR and HPLC mass spectrometry. IDO activity was significantly raised in SIP compared with healthy controls. ICC fluorescent imaging showed that IDO and KYN were colocalized with the markers of macrophages and neuronal fibers. Quantitative PCR data of SIP indicated that increased IDO may direct the KYN pathway toward the generation of the neuroinflammatory catabolite quinolinic acid, a metabolite that is opposite to the production of neuroprotective kynurenic acid. Administration of OLCs with ODN 2006 in vitro induced changes similar to those developed in SIP induced by bacterial infection. To our knowledge, this article is the first to demonstrate links between IDO activity, neuroinflammation, and algogenic pathways in SIP and localize this process to neuronal fibers. These observations highlight the potential role of the kynurenine pathway in SIP, but further investigation is required to determine if these changes are mirrored in the levels of neuroexcitatory metabolites in vivo.
Introduction
Symptomatic irreversible pulpitis (SIP) and apical periodontitis (AP) are the most common causes of orofacial pain worldwide, and their severity is such that toothache was the fifth most common reason for 20- to 29-y-olds to attend an emergency department (Lewis et al. 2015) and was the most common reason for unintentional paracetamol overdose in a UK study (Siddique et al. 2015). SIP and AP are commonly treated via root canal treatment or tooth extraction (Lipton et al. 1993; Joury et al. 2018), despite neither treatment specifically addressing the inflammation and nociplastic changes present, other than by amputating the nerve. New and more precise therapeutic approaches are therefore required to manage acute pulpitic pain (Coulter and Nixdorf 2020).
Indoleamine 2,3 dioxygenase (IDO) is the initial and rate-limiting enzyme in the kynurenine (KYN) catabolism pathway of tryptophan (TRP) (Fig. 1) (Mellor et al. 2017). Increased IDO activity increased pain hypersensitivity in mice. This hypersensitivity was absent in mice lacking the IDO1 gene but could be restored following systemic administration of KYN, the catabolite of IDO (Huang et al. 2016). Inhibitors of IDO have shown promise in reducing neuroinflammation. Administration of IDO inhibitors in a murine model of Parkinson’s disease showed decreased striatum levels of proinflammatory cytokines TNF-α, IL-6, and INF-γ and increased levels of brain-derived neurotrophic factor, a protein responsible for neuronal function and survival (Sodhi et al. 2021).
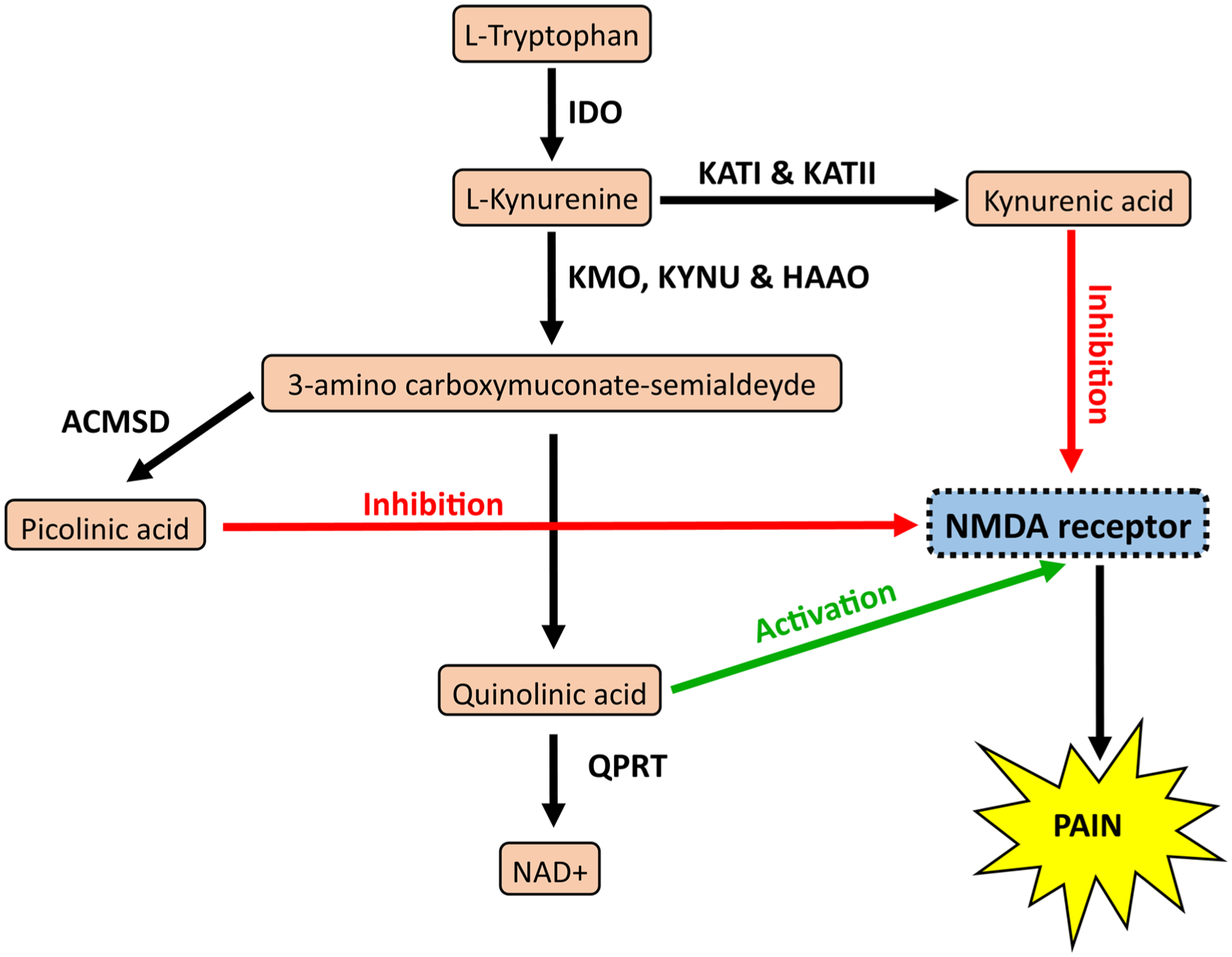
The kynurenine pathway. Flow diagram showing the catabolites produced in pink and enzymes in bold. Pathway highlights the potential for activation or inhibition of N-methyl-D-aspartate (NMDA) receptors and subsequent pain. Enzymes: kynurenine aminotransferase (KAT), kynurenine 3-monooxygenase (KMO), kynureninase (KYNU), 3-hydroxyanthranilate dioxygenase (HAAO), aminocarboxymuconate-semialdehyde-decarboxylase (ACMSD), and quinolinate phosphoribosyl transferase (QPRT).
The kynurenine pathway (Fig. 1) is associated with nociception as elevated IDO activity can ultimately lead to the production of quinolinic acid (QUIN), an agonist of N-methyl-D-aspartate (NMDA) receptors (Stone and Perkins 1981). NMDA receptors are expressed in the trigeminal ganglion (Lee and Ro 2007) and endogenously activated by the peripheral release of glutamate (Guerrero-Toro et al. 2022). Once activated, NMDA receptors can cause an influx of Ca2+ ions with consequent disbalancing of Ca2+ homeostasis, potentially leading to augmented neuronal firing and excitotoxicity, which might affect neuronal plasticity and give rise to the neurodegenerative processes of apoptosis and necrosis (Blanke and VanDongen 2009; Simões et al. 2018). The KYN pathway can also produce kynurenic acid (KYNA) and picolinic acid (PIC), which inhibit NMDA receptors (Beninger et al. 1994; Majláth et al. 2016). Inhibition of NMDA receptors would likely protect the peripheral nociceptive neurons from the glutamate released during pulpal inflammation (Yang et al. 2014) and may modulate any QUIN present. The KYN pathway has the potential to modulate neuroinflammation if directed toward QUIN or KYNA and PIC.
Despite the links between IDO and neuroinflammation, few studies have examined the KYN pathway in the context of orofacial pain. Animal models of orofacial pain induced via complete Freund’s adjuvant found elevated levels of KYN and KYNA in the trigeminal nucleus caudalis 24 h after injury (Cseh et al. 2020). A study examining the plasma concentrations of TRP and KYN in patients with myalgia temporomandibular disorder (TMD) identified that pain intensity positively correlated with circulating KYN/TRP ratios (Barjandi et al. 2020). Immunohistochemical (ICC) studies of dental pulp tissue have also shown increased expression of IDO in teeth with SIP, although IDO activity was not quantified (Takegawa et al. 2014). Western blot analysis has shown that stimulation of human dental pulp stem cells (DPSCs) cultured in vitro with lipopolysaccharide (LPS) and TNF-α increased IDO expression, and ICC demonstrated colocalization of IDO with monocyte and macrophage markers in the pulp (Lee et al. 2016), but further cell types have not been investigated.
Therefore, the aim of this study was to use an ex vivo tissue to quantify the changes in IDO activity/KYN pathway during SIP and investigate the pulpal cell types involved in the KYN pathway while an in vitro model would explore which processes may drive any changes detected during SIP.
Materials and Methods
Ex Vivo Investigations of the Kynurenine Pathway
Teeth were collected at Newcastle Dental Hospital (NHS ethics 15/WA/0129). Healthy samples were collected from those extracted for orthodontic reasons and were excluded if they had a history of pain or clinically detectable signs of caries. Painful samples were included on the basis of a diagnosis of SIP (AAE 2013). Samples from both groups were excluded if they were primary teeth or required surgical removal. Pulp tissue was obtained by propagating a crack through either side of the tooth (see Appendix for details).
High-Performance Liquid Chromatography
Following a power calculation (see Appendix for details), 27 healthy and 27 SIP samples were analyzed via high-performance liquid chromatography (HPLC). IDO enzyme activity was determined using a method adapted from Hoshi et al. (2010) (details in the Appendix).
Reverse Transcription Quantitative Polymerase Chain Reaction of Kynurenine Pathway Enzymes for Dental Pulp Samples
Reverse transcription quantitative polymerase chain reaction (RT-qPCR) data were exploratory and used 28 healthy and 24 SIP samples. Methods, primers, and thermocycling conditions can be found in the Appendix. Relative expression levels of the genes of interest were determined by normalizing with transcription levels of GAPDH and β Actin using the geNorm method (Vandesompele et al. 2002; Hellemans et al. 2007). Samples were removed from analysis if the messenger RNA (mRNA) of the gene of interest could not be detected, leading to differences in group sizes: IDO (n = 18 healthy vs. 16 inflamed), KMO (n = 25 healthy vs. 22 inflamed), alpha-aminoadipate aminotransferase (AADAT) (n = 19 healthy vs. 9 inflamed), KYNU (n = 14 healthy vs. 13 inflamed), 3-hydroxyanthranilate dioxygenase (HAAO) (n = 20 healthy vs. 12 inflamed), aminocarboxymuconate-semialdehyde-decarboxylase (ACMSD) (n = 8 healthy vs. 11 inflamed), and quinolinate phosphoribosyl transferase (QPRT) (n = 28 healthy vs. 24 inflamed).
Immunofluorescent Imaging of Dental Pulp Tissue
Teeth were obtained at UT Health at San Antonio following institutional review board approval. Teeth diagnosed with either normal pulp or SIP were included in the study. Teeth were split, and pulp tissue was dissected and fixed using 4% (v/v) paraformaldehyde. Samples were embedded in OCT freezing medium and sectioned using a MICROM MH 550 cryostat, then stained with primary antibodies consisting of rat anti-IDO 1:200 (Santa Cruz Biotechnology) with either mouse anti-KYN 1:200 (Santa Cruz Biotechnology) or mouse anti-CD68 1:200 (Serotec) and either chicken anti–neurofilament heavy (NFH) 1:500 (Abcam) or rabbit anti–von Willebrand factor (vWF) 1:200 (Dako). Secondary antibodies consisted of 1:500 donkey anti-rat 488 (Dako), 1:500 donkey anti-mouse 647 (Dako), 1:500 donkey anti-chicken 568 (Dako), and 1:500 donkey anti-rabbit 568 (Dako). Samples were imaged using a C1si laser scanning confocal microscope (Nikon) and analyzed in FIJI ImageJ2 software.
Culture of Odontoblast-Like Cells
Lonza human DPSCs differentiated into odontoblast-like cells (OLCs) using a medium comprising knockout Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 1% L-glutamine, 1% penicillin/streptomycin, 10% fetal bovine serum (FBS), 0.1 μM dexamethasone, 10 mM β-glycerophosphate, and 50 µg/mL ascorbic acid. Differentiation was confirmed using von Kossa and alizarin red staining as well as qPCR analysis of expression of dentine sialophosphoprotein and dental matrix protein 1 gene transcription (see Appendix).
Stimulation of OLCs
DPSCs were seeded at 100,000 cells per well in 24-well plates and differentiated for 4 wk. The cells were exposed to differentiation medium ± either 1 µg/mL ultrapure Escherichia coli K12 LPS (InvivoGen) or 1 µM ODN 2006 (InvivoGen) for 4 h and 24 h. Supernatant was collected for liquid chromatography–mass spectrometry (LC-MS) analysis using a method adapted from Schwieler et al. (2020) (see Appendix for details). The concentration of KYN, KYNA, QUIN, and PIC was determined as described previously for HPLC. Total RNA was also isolated and converted into complementary DNA (cDNA) for RT-qPCR analysis of KYN pathway enzyme expression, using the same primers and methods as for the pulp tissue samples. Fold expression of each gene was calculated using the 2–ΔΔCT method with GAPDH as the reference gene (Schmittgen and Livak 2008).
Statistical Tests
Student’s t-tests were performed to compare 2 groups in parametric tests, and the Mann–Whitney test was used to compare 2 groups in nonparametric tests. In parametric tests, a 1-way analysis of variance (ANOVA) was used to compare multiple groups for differences, and the Holm–Sidak method was used to determine if any groups were different from the control condition. In nonparametric tests, a Kruskal–Wallis ANOVA on ranks was used to compare multiple groups for differences, and Dunn’s post hoc test was used to compare groups for differences between themselves and the control condition. Pearson correlation coefficients were used to determine if continuous variables were associated. All tests and analyses were performed using SigmaPlot software (Grafiti) with significance set at P < 0.05.
Results
Increased Expression of IDO, KMO, and QPRT in Symptomatic Irreversible Pulpitis
Analysis with HPLC demonstrated significant increases in IDO activity within inflamed dental pulp tissue compared with healthy dental pulp (P ≤ 0.001) (Fig. 2A). Furthermore, IDO mRNA expression was significantly higher in the inflamed tissue compared with healthy controls (P < 0.01) (Fig. 2B). Expression of KMO mRNA was significantly higher in the inflamed tissue compared with healthy controls (P < 0.01) (Fig. 2C). Expression of AADAT, the gene encoding KATII, was significantly lower in the inflamed tissue compared with healthy controls (P < 0.001) (Fig. 2D). No difference in the relative expression between inflamed tissue compared with healthy controls was verified for the expression of KYNU (Fig. 2E), HAAO (Fig. 2F), and ACMSD (Fig. 2G) mRNA. Expression of QPRT mRNA was significantly higher in the inflamed tissue compared with healthy controls (P < 0.01) (Fig. 2H).
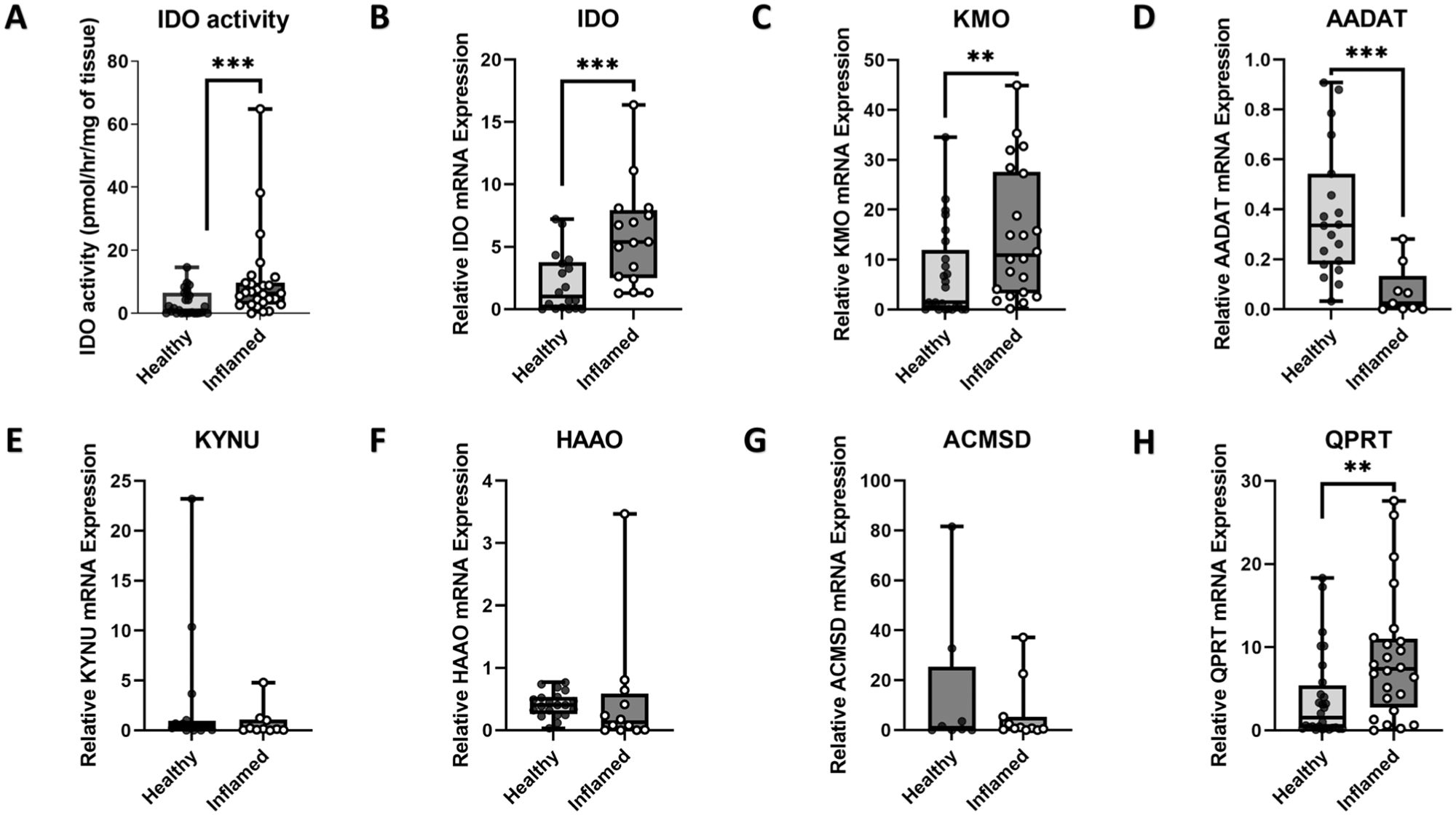
IDO activity and expression of IDO, KMO, AADAT, KYNU, HAAO, ACMSD, and QPRT in symptomatic irreversible pulpitis and healthy pulp tissue. Samples of inflamed (symptomatic irreversible pulpitis) and healthy pulp tissue showing (
IDO and KYN Expression in Neurons and Macrophages of Dental Pulp Tissue
Immunofluorescence imaging (Fig. 3) demonstrated the presence of KYN and IDO in SIP tissue. Elongated structures could be seen within the merged composite image that stained heavily for both KYN and IDO, which did not colocalize with vWF, implying the source was not vascular in origin. However, KYN and IDO showed colocalized expression with elongated structures that stained positive for NFH, consistent with neuronal staining. Colocalized staining could also be seen for both CD68 (macrophage marker) and IDO in SIP tissue, which was absent from healthy tissue.
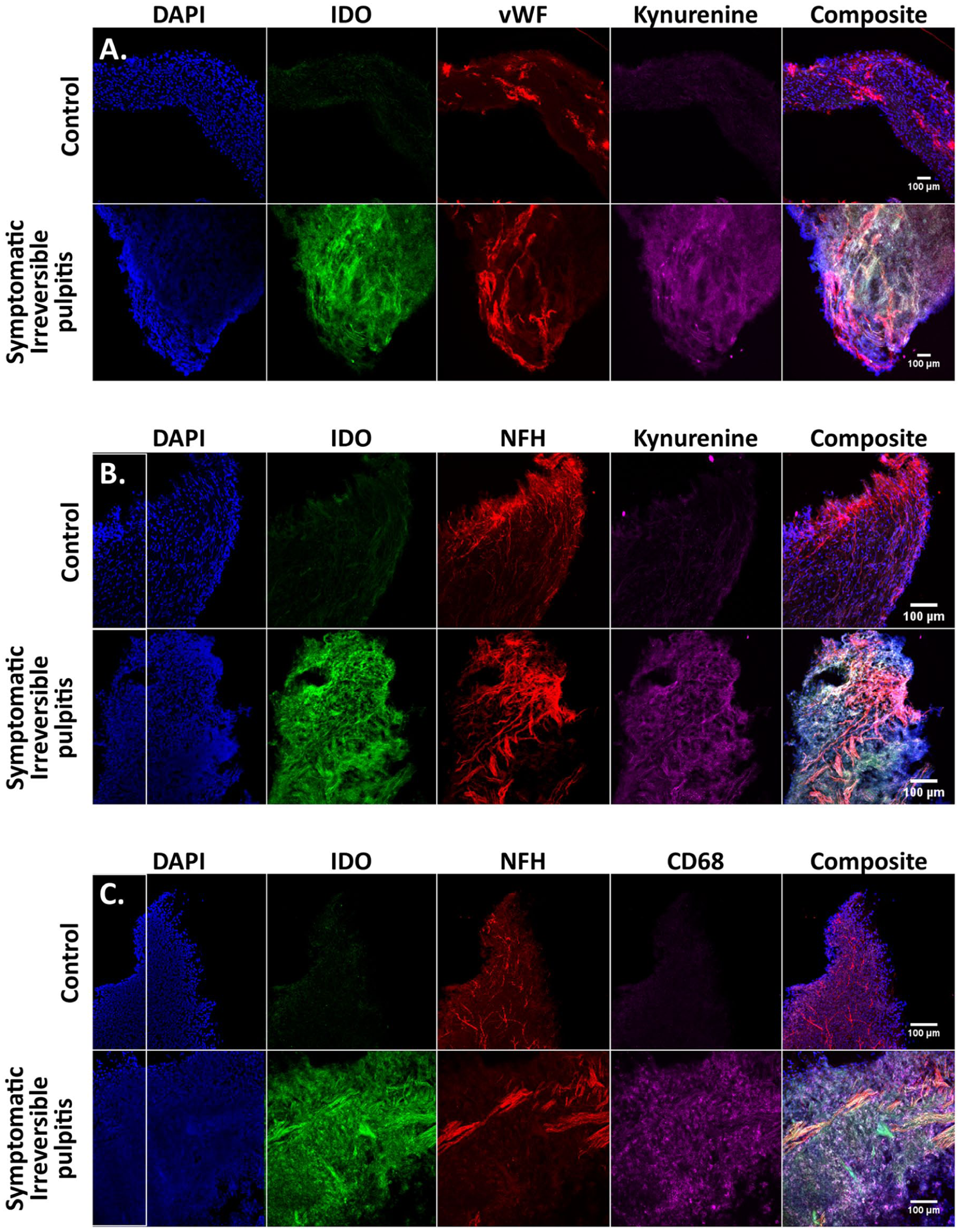
Immunostaining of IDO, KYN, vWF, NFH, and CD68 in dental pulp tissue. Representative images of a section from symptomatic irreversible pulpitis and healthy control dental pulp tissue. Coronal orientation is to the top of each slide, DAPI is in blue, and a composite panel shows a merged image of all signals. (
Activation of TLR-4 and TLR-9 Induced Increased Expression of IDO and KMO
To explore if TLR4 and TLR9 agonists, LPS, and ODN 2006 induced changes in the expression of enzymes of the KYN pathway, in vitro stimulation experiments were performed. Stimulation with LPS for 4 h induced significantly elevated expression of IDO (Fig. 4A) (P < 0.01), KMO (Fig. 4B) (P < 0.05), and KYNU (Fig. 4D) (P < 0.05), with no changes in expression of AADAT, HAAO, or QPRT (Fig. 4C, E, F). Conversely, stimulation with ODN for 4 h induced no changes in expression of any of the enzymes tested (Fig. 4A–F).
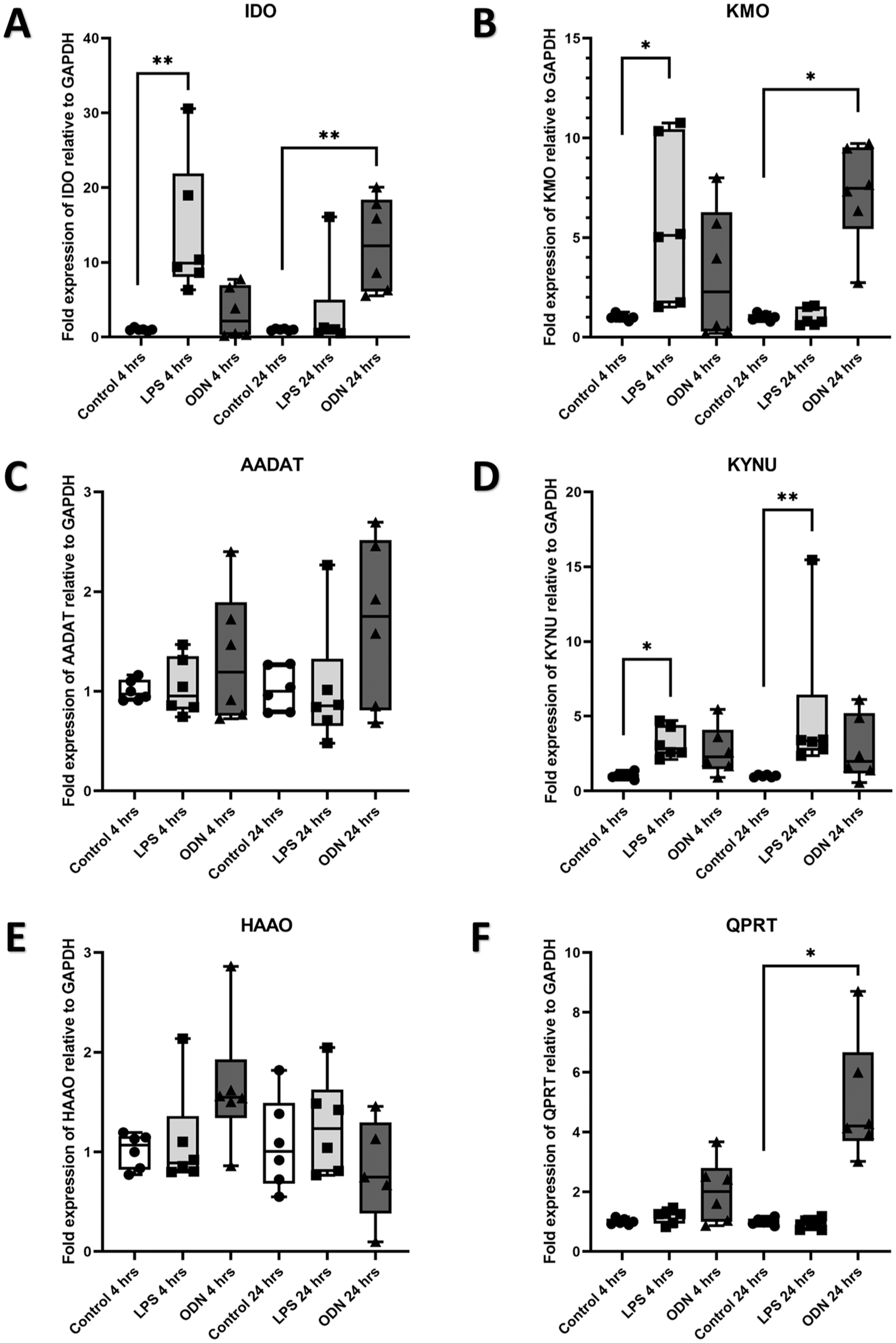
Expression of IDO, KMO, AADAT, KYNU, HAAO, and QPRT by odontoblast-like cells in response to TRL4 and TRL9 agonists. Relative expression of (
Stimulation with LPS for 1 d induced significantly elevated expression of KYNU (Fig. 4D) (P < 0.01) and no changes in expression of IDO, KMO, AADAT, HAAO, or QPRT (Fig. 4A–C, E, F). Stimulation with ODN for 1 d induced significantly elevated expression of IDO (Fig. 4A) (P < 0.01), KMO (Fig. 4B) (P < 0.05), and QPRT (Fig. 4F) (P < 0.05) and no changes in expression of AADAT, KYNU, or HAAO (Fig. 4C–E).
Activation of TLR-9 Induced Increases in KYN, KYNA, and QUIN Levels
LC-MS analyzed the supernatant from TLR-4– and TLR-9–activated OLCs. Four hours after stimulation with LPS induced no changes in levels of KYN, KYNA, PIC, or QUIN (Fig. 5A–D). Four hours after stimulation with ODN induced significantly elevated levels of KYN (Fig. 5A) (P < 0.05), KYNA (Fig. 5B) (P < 0.001), and QUIN (Fig. 5D) (P < 0.05) with no changes in levels of PIC (Fig. 5C). Twenty-four hours poststimulation with LPS induced significantly elevated levels of KYN (Fig. 5A) (P < 0.01), significantly decreased levels of PIC (Fig. 5C), and no changes in levels of KYNA or QUIN (Fig. 5B, D). Twenty-four hours poststimulation with ODN induced significantly decreased levels of KYNA (Fig. 5B) (P < 0.05), PIC (Fig. 5C) (P < 0.01), and QUIN (Fig. 5D) (P < 0.01), with no changes in levels of KYN (Fig. 5A).
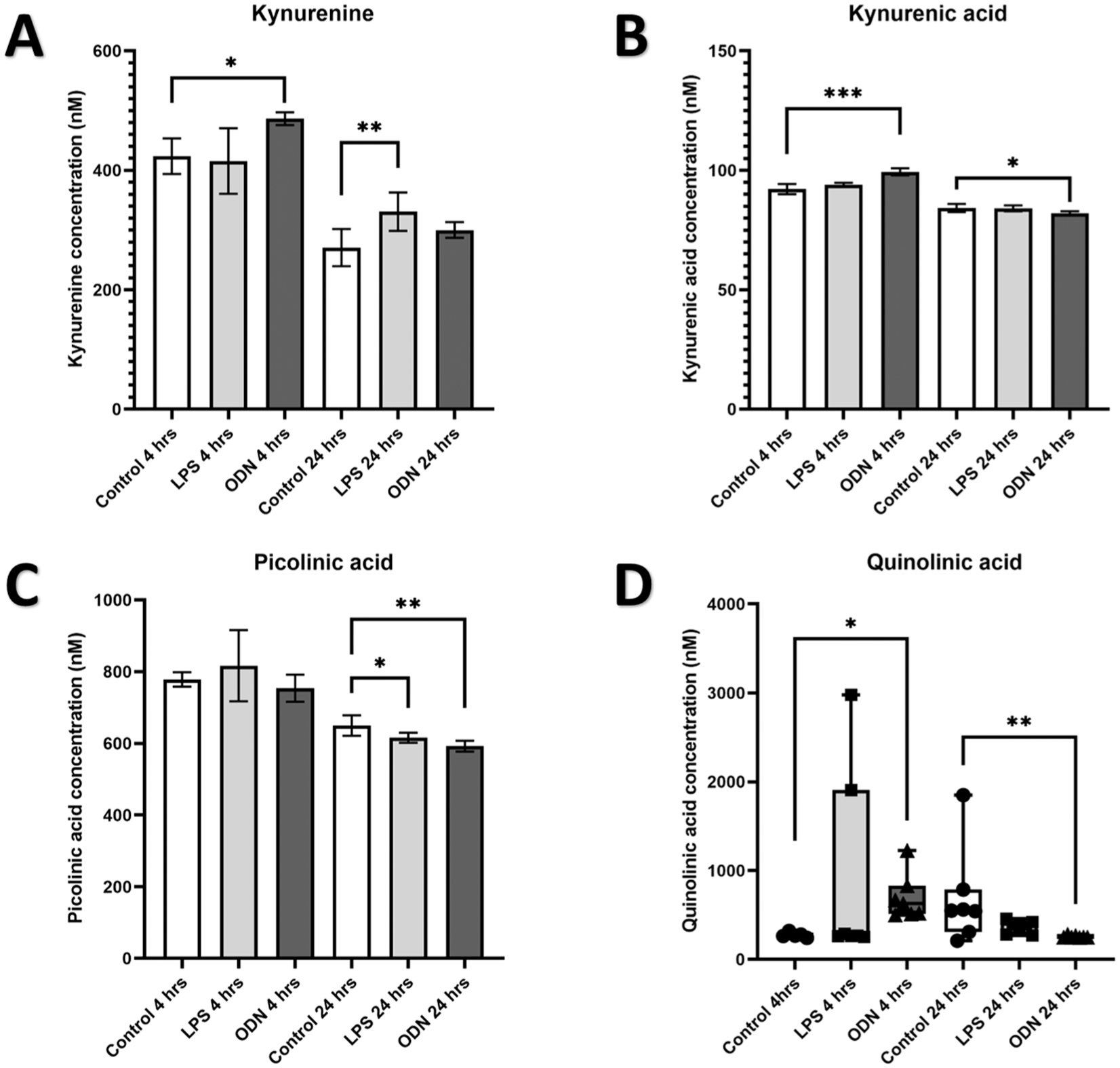
Quantification of KYN, KYNA, PIC, and QUIN in supernatant from odontoblast-like cells (OLCs) in response to TRL4 and TRL9 agonists. High-performance liquid chromatography–mass spectrometry measurement of (
Discussion
To our knowledge, this study is the first to quantify IDO activity in healthy and SIP samples. The increase in IDO activity in SIP samples may be linked to gene transcription, as RT-qPCR results showed an increase in IDO mRNA expression. Subsequently, this showed increased mRNA expression of KMO and QPRT and decreased mRNA expression of AADAT in SIP tissue compared with healthy tissue, with no changes in mRNA expression of the other enzymes tested. These results indicate that for SIP, AADAT mRNA expression is markedly reduced, potentially affecting the amount of neuroprotective KYNA generated. In health, the KYN pathway is in equilibrium with QUIN and KYNA production. Variations in the production of QUIN and KYNA could be associated with an imbalance in the activation of NMDA receptors and are linked to several chronic conditions, including depression, bipolar disorder, and Alzheimer’s disease (Guillemin et al. 2003; Myint et al. 2007; Birner et al. 2017). QUIN and KYNA equilibrium was demonstrated in a Huntington’s disease model of fruit flies, which showed that when KMO was inhibited, it led to a significant shift toward KYNA production and an associated reduction in neurodegeneration (Campesan et al. 2011). However, these results illustrate mRNA expression, and future research is required to determine if the increase in IDO activity and changes in transcription mirror changes in protein and metabolite levels.
During analysis of immunofluorescence images, fiber-like structures were highlighted in both KYN and IDO. Staining with NFH colocalized signals for both KYN and IDO to NFH, staining the sensory neuronal fibers in yellow (Fig. 3). Although NFH is usually associated with myelinated sensory neuronal fibers in other tissues, most of these sensory neuronal fibers in the dental pulp are positive for NFH and are unmyelinated (Henry et al. 2012). This indicates that NFH could be considered a generic marker for the sensory neuronal fibers in dental pulp tissue. The localization of KYN/IDO to sensory neuronal fibers has not been reported previously. The intimacy of this KYN/IDO association suggests that further neuroactive metabolites of the KYN pathway could be in proximity to neurons, but this will require further investigation to confirm. Fluorescent imaging demonstrated that IDO was associated with CD68, a marker for macrophages, as reported previously (Lee et al. 2016), but also showed an association between KYN and CD68, indicating that further parts of the KYN pathway may take place in these cells. Even with the merging of signal associated with CD68 and NFH, there was a much more widespread increase in IDO and KYN than just macrophages and neurons, indicating many cell types are involved and potentially contributing to the extracellular release of the KYN pathway metabolites.
An overview of LC-MS and qPCR data suggests that stimulation of OLCs with LPS and ODN 2006 largely mimics the transcriptional changes to the KYN pathway observed in SIP with elevated transcription of IDO and KMO, but no significant changes to HAAO transcription. ODN 2006–administered OLCs had an increase in QPRT transcription, which was also detected in SIP. These results indicate that stimulation of OLCs with LPS and, in particular, with ODN 2006 can induce changes in transcription like those observed in the KYN pathway during SIP, and stimulation of OLCs with these microbial toxins may be useful in investigating the KYN pathway in the future. However, LPS and ODN 2006 stimulation of OLCs was not an ideal model for SIP as AADAT, the gene for KATII, did not show a decrease in transcription during ODN 2006 or LPS stimulation, whereas such a reduction was observed in SIP. In addition, LPS stimulation displayed an increase in KYNU transcription at both 4 h and 24 h, which was not found in SIP. These differences may be due to the longer time course of pulpitis or the increased variety of cell types within the pulp, and factors that could influence its transcription during SIP more than can be modeled with 1 cell line.
LC-MS data support the suggestion that the altered transcription of the KYN pathway observed in SIP could stimulate the KYN pathway toward QUIN production. There were increases in KYN and a reduction in PIC levels for both ODN 2006 and LPS-administered groups (Fig. 5). In addition to this, ODN 2006–administered OLCs demonstrated an increased concentration of QUIN and KYNA after 4 h, which was subsequently decreased after 24 h compared with the healthy control. Interestingly, even though an increase in KYNA was detected for LC-MS analysis of ODN 2006 stimulation at 4 h and a decrease at 24 h (Fig. 5), the difference in KYNA concentrations identified was 7 nM at 4 h and 2 nM at 24 h, which is unlikely to be biologically relevant when the half-maximal inhibitory concentration (IC50) for KYNA on NMDA receptors is 135 µM (Weber et al. 2001). In contrast, the increase in QUIN detected in ODN 2006 at 4 h was 360 nM at the same time point. This suggests that the increase in the neuroexcitatory agent QUIN outstrips the increased neuroprotective potential of KYNA and is above the ~300-nM lower limit for saturation of QUIN’s catabolic enzyme QPRT in neurons (Rahman et al. 2009), supporting evidence toward activation of NMDA receptors within the trigeminal ganglion of sensory neurons, as hypothesized. PIC was found to have a small decrease in both LPS- and ODN-stimulated OLCs at 24 h, reducing its potential to function as a neuroprotectant and supporting the theory that increased KYN produced is moving the pathway toward QUIN production.
Limitations of the study must be acknowledged. First, ex vivo qPCR results were exploratory, with no prior power calculation and sample sizes varying between primers. In addition, as discussed previously, these results mostly represent transcriptional levels of the enzymes associated with the KYN pathway rather than the levels of the enzymes or the metabolites themselves. This was due to challenges in detection within clinical samples of small metabolites, which can vary by as little as the presence or absence of a hydroxyl group. Such measurements will be a target for future research. Limitations of the in vitro model must also be acknowledged. Dental pulp tissue is a complex mixture of odontoblasts, nerve cells, blood vessels, immune cells, and so on in a specific architecture that is not present within our model, and so there may be differences between our results and what is seen in vivo. Such differences would be challenging to investigate in ex vivo tissue due to degeneration, and a complex 3-dimensional model is likely required but is beyond what is currently available within the literature. These results therefore present the KYN pathway as an area of interest within pulp biology, which requires further research to unpick the mechanisms behind and confirm any association with pulpal pain.
Conclusion
This study demonstrates that SIP is associated with an increase in IDO activity, which appears to be driving the KYN pathway toward neuroinflammation and algesia. Imaging of pulp sections of SIP revealed that IDO and KYN were colocalized with neurons and macrophages, supporting this theory. Stimulation of OLCs with LPS and ODN 2006 produced changes that mimicked the changes observed in SIP, indicating that this process may be driven by bacterial infection (i.e., caries). This will require further investigation to verify the mechanisms and to determine if the changes in metabolite levels found in vitro are mirrored in vivo.
Author Contributions
J. Coulter, contributed to conception and design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; C.J. Nile, contributed to design, data analysis and interpretation, critically revised the manuscript; H. Lemos, contributed to conception and design, data acquisition, analysis, and interpretation, critically revised the manuscript; P. LoCoco, contributed to data acquisition, critically revised the manuscript; J. Whitworth, A. Diogenes, contributed to conception and design, data analysis and interpretation, critically revised the manuscript; R.A. Valentine, contributed to conception, data interpretation, critically revised the manuscript; C. Chang, contributed to design, data acquisition and analysis, critically revised the manuscript; A. Mellor, V. Telezhkin, J. Durham, contributed to conception and design, data analysis and interpretation, critically revised the manuscript. All authors gave their final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345261416437 – Supplemental material for The Role of the Kynurenine Pathway in Irreversible Pulpitis
Supplemental material, sj-docx-1-jdr-10.1177_00220345261416437 for The Role of the Kynurenine Pathway in Irreversible Pulpitis by J. Coulter, C.J. Nile, H. Lemos, P. LoCoco, J. Whitworth, R.A. Valentine, A. Diogenes, C. Chang, A. Mellor, V. Telezhkin and J. Durham in Journal of Dental Research
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research was funded by Newcastle Biomedical Research Centre as a Clinical PhD Fellowship and by the Royal College of Physicians and Surgeons of Glasgow via the TC White Research Grant.
A supplemental appendix to this article is available online.
