Abstract
National Health and Nutrition Examination Survey (NHANES), one of the largest curated repositories of population-level health indicators including physical examinations, blood/urine biochemistry, self-reported surveys, and dietary intake, offers rich resources for oral health research but presents challenges for machine learning analysis due to heterogeneity, missing data, and complexity. Dental caries, the most prevalent chronic disease worldwide, is a multifactorial disease and exhibits variability in clinical manifestation, calling for advanced analytical approaches for deeper understanding. Here, we develop an integrated data-cleaning and subtype discovery pipeline using unsupervised machine learning for comprehensive analysis and visualization of data patterns in the NHANES database. Our multidimensional pipeline declutters and optimizes the NHANES dataset by addressing missingness and outliers to streamline data integration and create a machine learning–ready version. Applying this pipeline reveals data patterns that led to the discovery of previously unrecognized subtypes and variables associated with the clinical heterogeneity of dental caries. We observed diverging patterns of similarity across age groups and variable subsets, identifying distinct clusters particularly in children (<5 y) and senior adults (>65 y). We also discovered unexpected associations involving lead exposure and specific laboratory markers and, importantly, identified novel dietary signatures by linking food type and co-occurring consumption patterns to caries. Altogether, we report a comprehensive data-processing and data-analysis approach that reveals significant dental caries heterogeneity in NHANES data and can support the development of more precise and robust machine learning models for dental caries and other health conditions.
Introduction
The National Health and Nutrition Examination Survey (NHANES) is one of the largest curated repositories of nationally representative population-level health-related indicators, including physical examination, blood biochemistry, self-reported surveys, and dietary intake data. Overseen by the Centers for Disease Control, NHANES data are acquired through personal interviews combined with routine clinical assessments (dental and medical examinations), dietary surveys, and laboratory tests and have been used continuously to monitor US oral and overall health since 1999. This resource provides comprehensive data for studying oral diseases such as dental caries.
Despite its breadth, NHANES presents challenges for machine learning due to data heterogeneity, varied sample sizes, missing values, and inconsistencies in data collection methods (Pfeiffer et al 2017; Dye et al 2019), all of which can affect predictive modeling. These limitations are further compounded by outliers and mixed data types, requiring dedicated pipelines for data correction (Willemink et al 2020). Moreover, complex health outcomes such as dental caries often exhibit high clinical heterogeneity, highlighting the need for more refined phenotypic subtyping as well as effective visualization and interpretation methods for high-dimensional data patterns.
In this study, we addressed these limitations by establishing a data-cleaning pipeline with a novel outlier detection algorithm and unsupervised machine learning to identify phenotype subtypes within NHANES dental caries data. The NHANES includes highly detailed oral health information, and when combined with age, environmental, laboratory, and dietary data, this dataset can be leveraged for new insights into caries determinants. A nutrition taxonomy based on the US Department of Agriculture’s dietary guidelines (US Department of Agriculture and US Department of Health and Human Services 2020) further enhances precision in identifying associations between diet and disease.
We focus on dental caries, the most prevalent oral disease affecting more than 3.5 billion people globally (Marcenes et al 2013; Richards 2013). We developed an integrated data-cleaning–subtype discovery pipeline using unsupervised machine learning with a graph-based algorithm for visualization of data-clustering patterns and outcome analysis. The pipeline declutters, optimizes, and streamlines the integration of NHANES data, producing a “machine learning–ready” dataset, while revealing previously unrecognized subtypes and variable associations as well as diverging similarity patterns underlying the clinical heterogeneity of dental caries.
Methods
Data Acquisition
We analyzed publicly available data for 8,099 samples from the 2017–2018 NHANES survey (Centers for Disease Control and Prevention 2018), which includes demographic, dietary, examination, laboratory, and questionnaire data. Full details and ethical considerations are in the Appendix.
Data Preprocessing
NHANES datasets were aggregated and processed by removing variables with >50% missing data (Appendix Fig 1), selecting domain-relevant features, and applying one-hot encoding to nonordinal categorical variables. This resulted in a final dataset of 438 variables (Appendix Table 1). The full workflow is detailed in the Appendix.
Outlier Detection and Data Imputation
We applied the Skew and Tail-heaviness Adjusted Removal of Outliers (STAR) (Verardi and Vermandele 2018; Gregg and Moore 2023) for outlier removal, followed by multivariate imputation of missing values using Scikit-learn’s iterative imputer. Detailed rationale and parameters are provided in the Appendix.
Clinical Outcome Definitions
We derived binary (any caries) and categorical (mild vs severe) outcomes from oral examination data, considering both active-only and combined active/past disease statuses. The threshold for severe caries (≥4 affected surfaces/teeth) was based on the established definition of severe early childhood caries (S-ECC) (Tinanoff et al 2019). Full details and rationale are provided in the Appendix.
Unsupervised Learning Analysis
We used bootstrap-resampled hierarchical clustering (pvclust R package; Suzuki and Shimodaira 2006) to identify significant variable clusters within subgroups stratified by age, children (≤5 y), youth (>5 to ≤18 y), adult (>18 to <65 y), and senior (≥65 y), and by caries status (severity, activity). Analysis was also performed on nutrition-only subsets stratified by sugar content. Clustering parameters were chosen based on a quantitative validation (Appendix Table 3), with full details in the Appendix.
Data Visualization
Key findings were visualized using cluster maps. Spearman correlation networks were generated for age- and lead-stratified subgroups and TooManyCells spectral clustering (Schwartz et al 2020) for sample relationships. The rationale for the lead categorization and other specific visualization techniques is detailed in the Appendix.
Results
NHANES Data-Cleaning Pipeline
We developed an integrated data-cleaning–subtype discovery pipeline for observational studies (Fig 1). Specifically, NHANES 2017–2018 datasets for caries, including demographic, disease severity/activity, laboratory, examination, dietary (total nutrient intake and dietary supplement), and questionnaire indicators, were assembled into a single dataset that, upon removal of variables with a missing data fraction greater than 50%, domain-relevant variable selection, redundancy correction, and nonordinal categorical variable encodings, contained 438 variables.
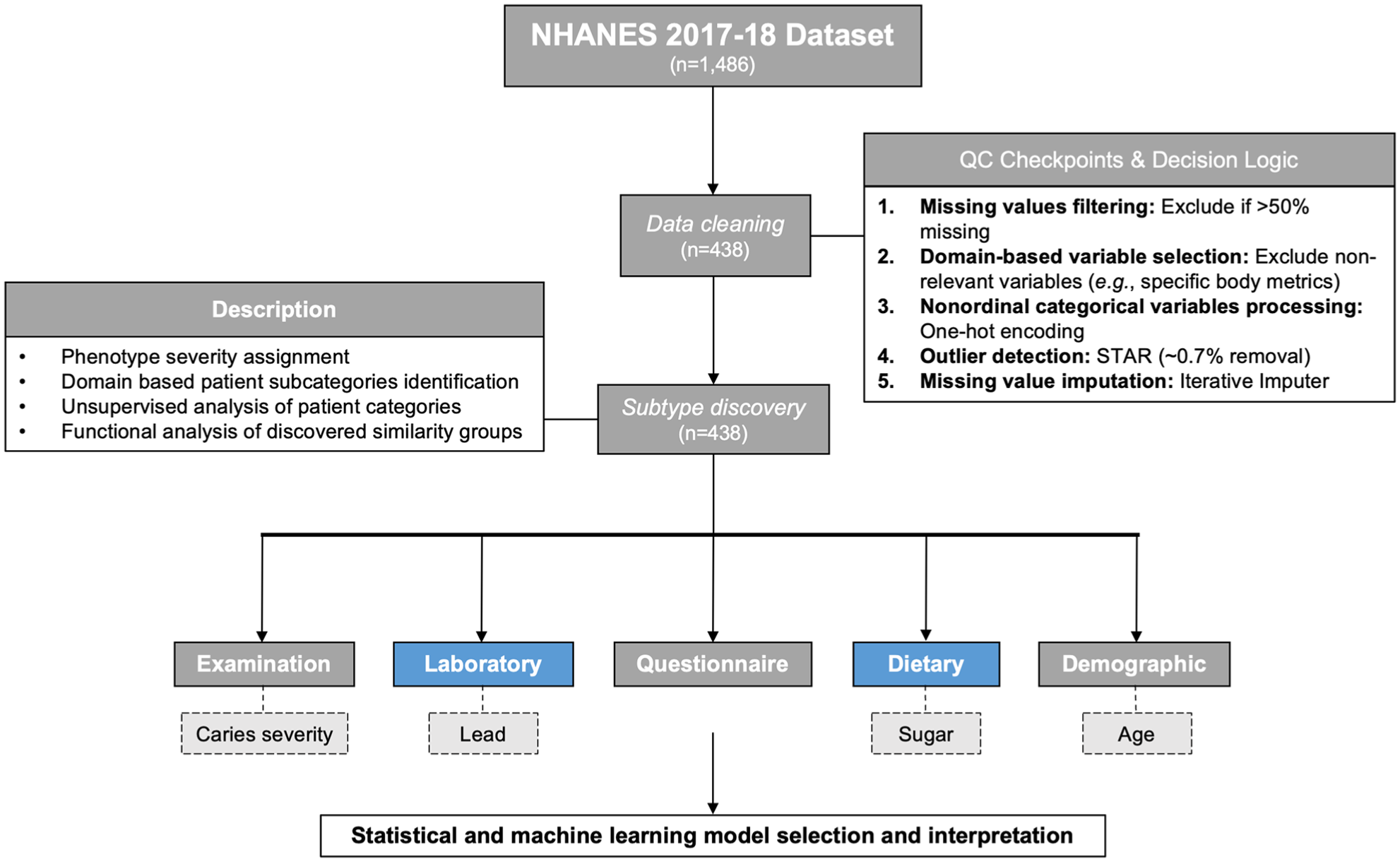
An overview of the data-cleaning and exploratory analysis pipeline components.
First, we quantified the proportion of missing values. Variables with a missingness fraction <50% were retained, which reduced the dataset size from 1,486 to below 500 variables (Appendix Fig 1). Next, oral health–relevant variable selection was performed, in which irrelevant or redundant variables were eliminated. Finally, nominal variables were processed with the OneHotEncoder operator, which transforms each variable with N categories into N binary variables. These preprocessing steps resulted in the dataset of 438 variables.
Accounting for outliers is critical for robust and generalizable analysis. This is especially important in observational studies containing self-reported evaluations (eg, nutrition diary), a common subject to measurement error. Therefore, following the multistep variable selection and transformation, the STAR outlier detection algorithm was applied, a multivariate projection-based method that does not rely on the normal distribution assumption. Appendix Figure 2 demonstrates examples of the STAR algorithm outlier detection for NHANES variables with skewed distributions and different tail heaviness. The final step of the cleaning pipeline was multivariate imputation with the iterative imputer. This method performs sequential imputation of every variable with missing values by fitting a machine learning model on all the remaining variables in the dataset and improves the missing value estimate by repeating this process multiple times.
Caries Subtype Discovery
Dental caries is a complex trait, and analyzing the entire disease population with a single model can oversimplify the true underlying mechanisms. Therefore, once the dataset was finalized, we investigated a series of clinical indicators for their role as drivers of caries subpopulation heterogeneity. We first assumed age, disease severity, and disease activity status to be the key drivers of clinical divergence and analyzed these subpopulations with unsupervised learning methods. The pvclust hierarchical clustering method assigns bootstrap resampling–derived P values to cluster dendrograms and outlines clusters supported by the data. The presence of large clusters of variables is of particular interest in the disease subtype heterogeneity analysis. Functional analysis of such patterns can provide an explanation of the driving processes within the subtype while serving as a variable selection foundation for datasets with high dimensionality. Diverging patterns between disease subtypes also affect the generalizability of machine learning model inference.
For caries age-stratified analysis performed with pvclust (Fig 2A–D), the caries subpopulation of children (≤5 y) showed the largest supported cluster among all age groups, consisting of 151 variables (Fig 2A, E). In addition, caries among the senior (≥65 y) subpopulation contained the second largest cluster discovered, consisting of 146 variables (Fig 2D and H). The remaining age groups, youth (>5 to ≤18 y) and adult (>18 to <65 y) caries subpopulations (Fig 2B, C), appeared to have a more homogenous distribution of supported clusters, with the largest clusters sizes less than 20 and 60 variables, respectively (Fig 2F, G). Interestingly, the children (CH) and senior (S) caries populations’ largest cluster content is predominantly nonoverlapping—only 46 common variables and 100 and 105 unique variables (Fig 3A). Functionally, the CH cluster content was 48% laboratory result variables, while the overall fraction of lab variables was only 28% (Fig 3C, D). The S cluster–affiliated variable categories distribution, however, is more similar to the overall categories distribution (Fig 3C and E). A color map of the scaled and aligned set of variables from the CH cluster demonstrates the discrepancy in variable patterns between the children and the senior age groups (Appendix Fig 3). This is also confirmed by the correlation network analysis in which strong linear correlation associations (Spearman’s |r| > 0.6) for children- and senior-age caries subpopulations vary dramatically (Appendix Fig 4 and Appendix Table 5).
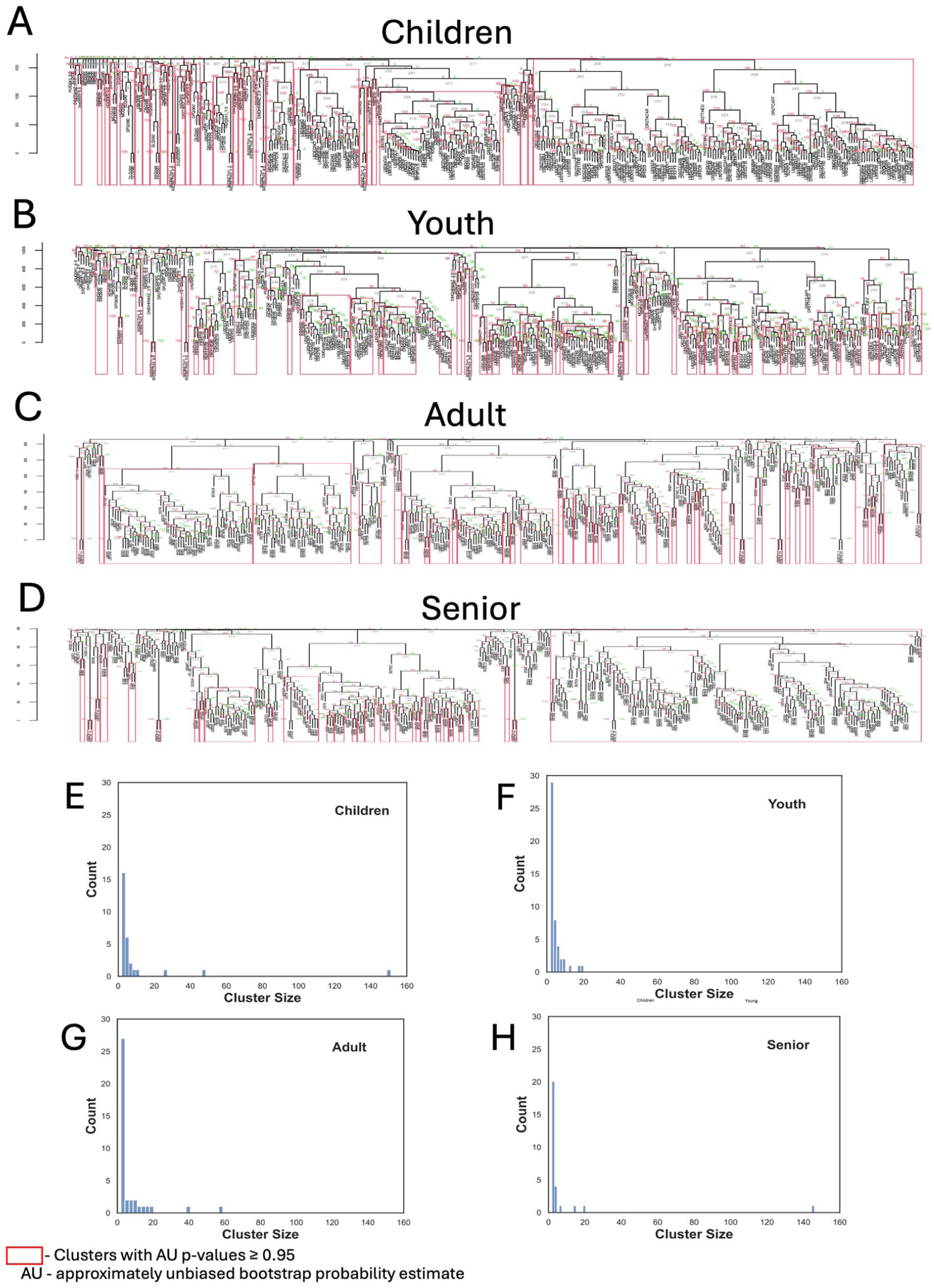
Hierarchical relationships between National Health and Nutrition Examination Survey (NHANES) variables for age-stratified subpopulations with caries identified by the pvclust clustering method. Clusters supported by the significant approximately unbiased (AU) bootstrap probability estimate are shown for (
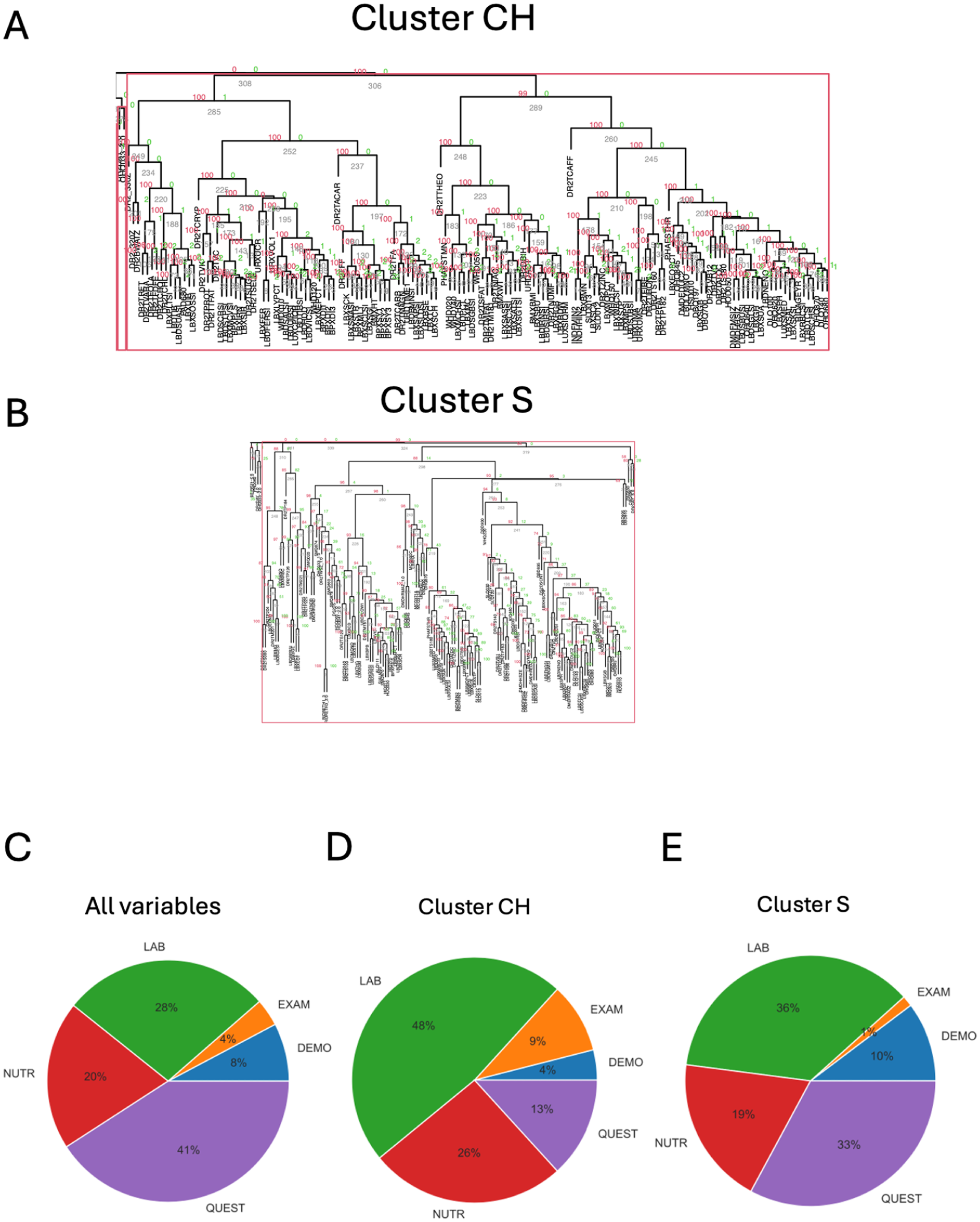
Functional analysis of the largest (>100 variables) approximately unbiased (AU) significant clusters in the pvclust analysis. Overlapping and unique features for the largest clusters for (
We investigated subpopulations based on disease severity for active and historical caries. Among these, only active caries analysis revealed the most diverging cluster patterns between severity subpopulations. Specifically, a single large supported cluster (79 variables, referred to as S1) was associated with the severe caries phenotype that represents patients with more than 4 teeth affected (Appendix Fig 5). Its functional content is almost equally divided between laboratory and nutrition variable categories (Appendix Fig 6). In addition, 4 more clusters with a size greater than 20 variables were associated with an active severe caries subpopulation. Conversely, mild and no caries subpopulations appeared to have dense distributions of small supported clusters, most with fewer than 5 variables. In summary, bootstrap resampling–powered hierarchical clustering analysis identified 2 major clusters in age-stratified caries data (children and senior) with largely unique variables that may indicate distinctive phenotypes and require separate disease models, whereas a single major cluster found in the severe active caries subpopulation may reveal novel variables associated with the disease that will not be visible in the historical caries population or in the general active caries disease population.
Spectral Hierarchical Clustering Analysis
We then performed spectral hierarchical clustering analysis with TooManyCells, a method used to visualize single-cell clades relationships, to discover if any of the disease subtype annotations we had investigated could confirm the heterogeneity of the caries population. TooManyCells analysis, performed on the entire variable set, resulted into a dendrogram with 69 terminal clusters (Fig 4A). Considering the prior discovery of patterns associated with lab and nutrition variables categories, we suggested that using a complete variable set could be dimensionally challenging to infer smaller subtype patterns, and therefore, we analyzed laboratory-only and nutrition-only variable subsets, which resulted in spectral dendrograms with 110 and 74 terminal clusters correspondingly (Fig 4B and Appendix Fig 7). We then estimated the heterogeneity of the terminal clusters for age groups by mapping them on the dendrograms and analyzing the cluster diversity index.
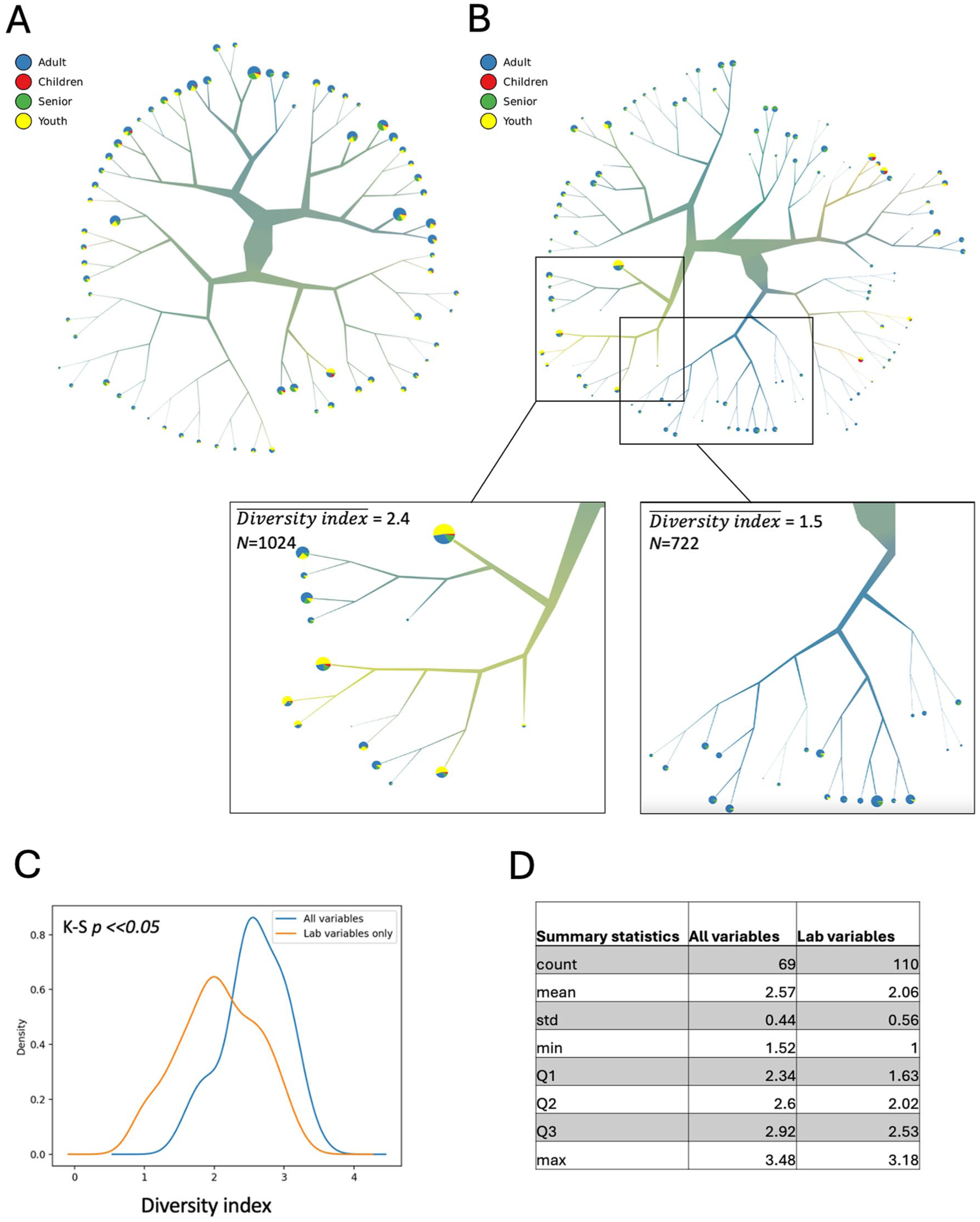
Spectral hierarchical clustering analysis of the caries subpopulation with age group annotation. (
The diversity measure was adapted from an ecological study (Heck et al 1975), where it was used to evaluate the effective number of species within a population with the minimum diversity 1 in case of a single dominant species and the maximum corresponding to a total number of evenly abundant species. Age group label diversity indexes for the entire set and nutrition-only dendrograms were substantially higher than for the laboratory-only dendrogram, which was confirmed by distribution statistics (Fig 4C, D and Appendix Fig 7). As observed in the density plot, a large portion of terminal clusters in the lab-only analysis had a diversity index less than 2 (47.3%), while this percentage was 14.5% for the entire set and 10.8% for the nutrition-only dendograms. Furthermore, in the laboratory-only dendrogram, we discovered large branches (1,024 and 722 samples) composed predominately of youth and adult caries populations, respectively, with low to medium median diversity indexes (Fig 4B). The data suggest that laboratory variables may have a discriminative effect in separating youth and adult caries subpopulations and that age stratification of the caries population has potential to improve caries risk prediction models. Age-based stratification was further confirmed by t-distributed stochastic neighbor embedding and uniform manifold approximation and projection dimensionality reduction methods conducted on the caries population with complete variable set (Appendix Fig 8).
Caries Risk Factors Targeted Analysis
We next assessed the known nutrition and laboratory variables associated with caries. Sugar is a known risk factor for dental caries (Pitts et al 2017; Lagerweij and van Loveren 2020). To evaluate its contribution to disease heterogeneity, we integrated food taxonomy data with the examined caries disease phenotypes and further stratified food variables according to their sugar content. The pvclust analysis of high-sugar foods demonstrated a more stable cluster structure associated with the caries subpopulation (Fig 5). Hierarchical clustering analysis revealed 19 bootstrap resampling supported clusters of meta-nutritional variables with high-sugar content specific to the caries group (Fig 5A). Further functional analysis revealed clusters with socially recognizable patterns such as a cluster with tea, pastries, cookies, candies, and sugar-additive variables (Fig 5B). Within the context of high-sugar-content foods, the caries population was uniquely associated with a specific set of beverages, dairy products, fruits, condiments, and desserts (Fig 5). Apple juice, energy drinks, and protein and nutritional powders are some examples of high-sugar caries–associated beverages. Dairy products such as flavored milk, milkshakes, yogurt, and ice cream also fell into this category. Protein, vegetable, and grain food groups were almost exclusively low in sugar content and not associated with caries in this analysis.
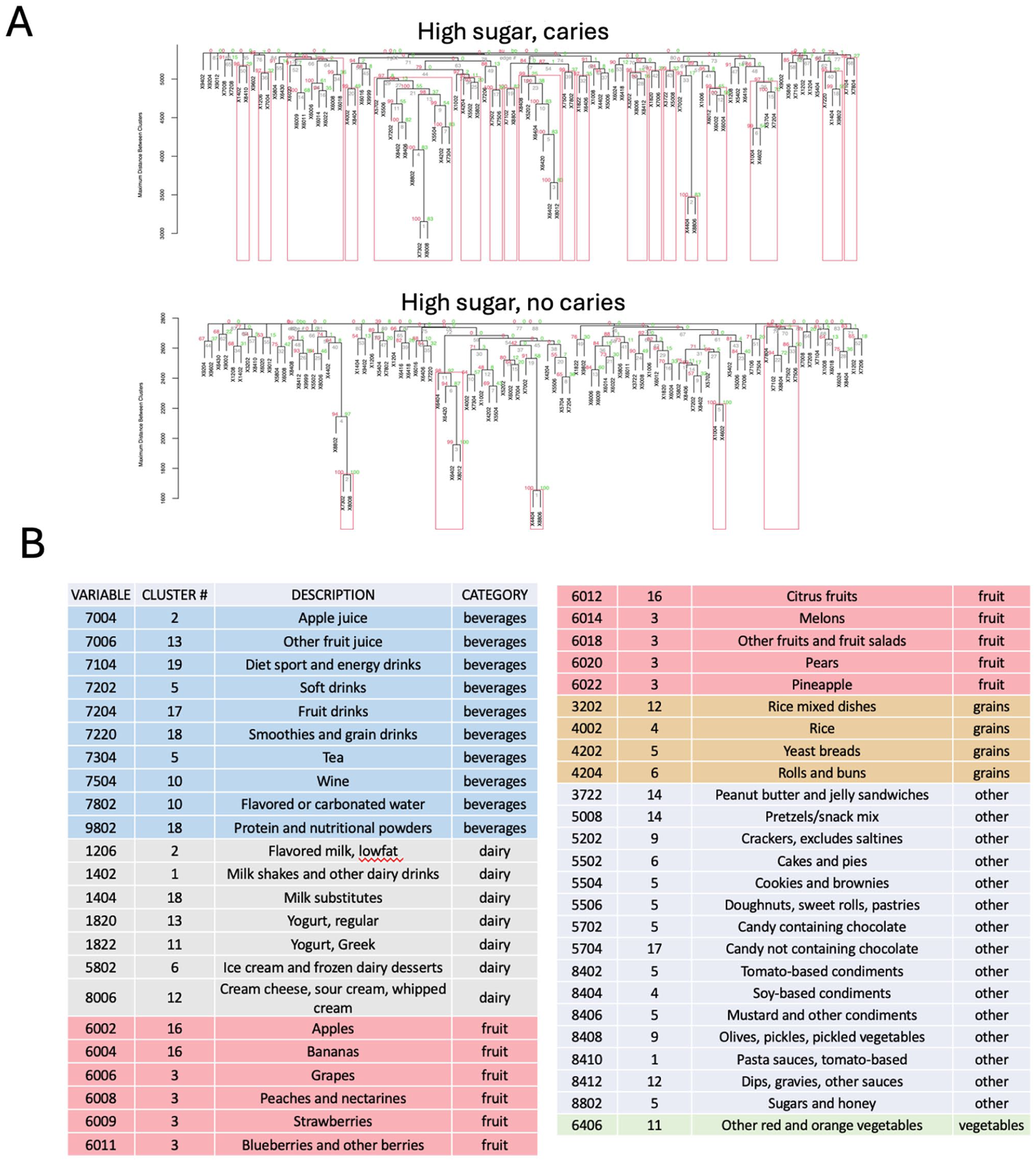
Consumption patterns of high-sugar foods in caries-stratified populations. (
Lead exposure is another known risk factor that appeared to be associated with caries occurrence (Billings et al 2004; Pradeep and Hegde 2013). For example, in pediatric patients, increased salivary and enamel lead levels have demonstrated a positive correlation with an increase in dental caries severity (Pradeep and Hegde 2013). Individuals with caries were found to have significantly higher average blood lead levels (µg/dL) than those without caries (Appendix Fig 9). In this analysis, we compared correlation structures by stratifying the population into “low-lead” versus “medium-/high-lead” groups (see Appendix Methods) (Appendix Fig 10 and Appendix Table 2). Across low-lead/high-lead groups and no-caries/high-caries groups, correlation network analysis revealed distinctive trends in continuous laboratory blood values, dietary factors, and body weight metrics for each group. Of most interest are the medium/highly correlated (Spearman’s |r| > 0.4) variables exclusive to the medium-/high-lead caries subpopulation (in relation to the low-lead no caries control). We find a positive correlation between cadmium, cotinine, and hematocrit, iron, and bilirubin (Appendix Table 2). Among both high- and low-lead caries Spearman’s rank correlation networks, a strong positive correlation exists in relation to dietary components such as fat and cholesterol in both caries and no-caries subgroups. In sum, the data show expected associations of sugars and lead with dental caries but also new possibilities and high granularity by pinpointing specific food types and laboratory variables that may lead to more precise predictive modeling.
Discussion
Large population database sources, such as NHANES, offer invaluable resources for dental epidemiology but simultaneously pose challenges for machine learning approaches due to data heterogeneity, varied sample sizes, missing data, and variations in data collection. These factors necessitate thorough preprocessing before robust machine learning frameworks can be applied. Moreover, interpreting complex patterns within high-dimensional data requires effective data visualization and pattern identification. The bioinformatics pipeline developed here provides a structured approach to address these challenges, offering utility for database users and curators. By systematically handling missingness, outliers, and variable selection, the pipeline facilitates the creation of analysis-ready datasets and, crucially, aids the discovery of clinically relevant heterogeneity including nutrition and laboratory variables that can inform more precise statistical and machine-learning models. Using the caries dataset as a case study, we find that systematic preprocessing, including addressing potential redundant variable types (eg, closely related blood biochemistry markers) can potentially streamline subsequent analyses.
The application of the pipeline combined with network-based and clustering visualization approaches provided several key insights into dental caries heterogeneity within the NHANES population. We find intriguing diverging patterns of variable similarity between different age groups, particularly comparing children (<5 y) with senior (≥65 y) populations, for the complete dataset and for nutrition- and laboratory-only subsets. Interestingly, the disease has a bimodal onset, with the children and senior groups exhibiting the largest and most distinct patterns of variable similarity, suggesting different etiological factors or disease manifestations at these life stages.
The pronounced clustering of laboratory variables within the early-childhood subgroup suggests a strong link between S-ECC and systemic health. Young children are particularly vulnerable to nutritional and metabolic deficiencies that can manifest orally. Anemia has been associated with increased caries risk in children (Schroth et al 2013; Delimont et al 2021), and our findings show that indicators for nutritional status such as iron and vitamin D are prominent in this cluster. This suggests that in young children, dental caries may not be merely a localized disease but can serve as a sentinel marker of underlying systemic health issues.
The significant large cluster primarily composed of laboratory and nutrition variables identified through pvclust was associated with the senior (≥65 y) and children (<5 y) age groups in age-stratification cluster analysis. This pronounced age-driven heterogeneity strongly suggests that age stratification is critical for developing accurate predictive models for dental caries. Furthermore, our analysis of the nutrition-only subset, stratifying foods by sugar content and examining associations within the caries population, revealed recognizable dietary patterns. The examination of consumed food taxonomy and its sugar content revealed food patterns and types associated with caries, such as the clustering of habitual consumption of pastries and specific sugary foodstuffs/beverages, expanding known dietary risk factors at a granular level.
Analysis stratified by lead exposure showed diverging correlation network patterns in the caries and no-caries subgroups. Recent systematic reviews confirm a positive association between lead exposure and caries, particularly in primary dentition (Lee et al 2024), and the NHANES database shows higher average blood lead levels in individuals with caries. However, this relationship can be confounded by socioeconomic factors. Our analytical approach did not find a direct correlation between blood lead levels alone and caries. Instead, our network-based analysis revealed a strong correlation between cadmium and cotinine specifically in the high-lead population, extending the previously established association between smoking by-products (cotinine), heavy metals (cadmium), and caries (Akinkugbe et al 2019) to a high-lead context. This tight cluster of exposures suggests common environmental sources (eg, tobacco smoke, industrial pollutants) that are known to be prevalent in lower socioeconomic settings (Hajat et al 2015). Therefore, our data indicate that lead may be a powerful indicator of high-risk environments. While plausible biological mechanisms for lead’s cariogenicity have been proposed, including disruption of enamel mineralization, impaired immune function (Billings et al 2004; Lee et al 2024), and impaired salivary function in animal models (Watson et al 1997), our findings suggest the higher caries risk is more likely explained by these co-occurring exposures rather than an independent effect of lead alone.
Beyond the insights into lead exposure, the network analysis uncovered other potentially important correlations. For instance, we identified moderate-to-high correlations between serum iron, hemoglobin, and bilirubin in the high-lead caries population. These hematologic links warrant further investigation regarding the potential interactions between lead toxicity, iron metabolism, systemic inflammation, and caries risk. Furthermore, the unexpected association between reported hours slept and age within the low-lead no-caries group opens new avenues to explore potential interactions between sleep patterns, environmental exposures, and caries susceptibility through behavioral or systemic inflammatory pathways.
Altogether, our work demonstrates a robust data-cleaning–subtype discovery pipeline that could be applied to investigate other health conditions using NHANES and similar databases for machine learning predictive modeling. This integrative approach systematically addresses data quality issues and facilitates exploratory analysis to reveal data patterns associated with subtypes and variables associated with the clinical heterogeneity of caries. The discovery of significant age-driven heterogeneity and novel factor associations highlights the value of such pipelines for generating hypotheses and guiding the development of more precise and robust machine learning predictive models in dental research. However, our analysis was limited to NHANES 2017–2018 data (the latest available release at the study conception), and future work using multiyear designs will be needed to establish longitudinal trends.
In conclusion, applying a comprehensive bioinformatics pipeline to NHANES data successfully identified substantial age-driven heterogeneity in dental caries, suggesting stratification is crucial for future predictive modeling. The approach also uncovered novel associations between caries status, lead/pollutant exposure, specific laboratory markers and food types, as well as sleep patterns, reflecting additional disease markers in susceptible populations. This work demonstrates the value of integrating data science techniques with large-scale observational data to gain deeper insights into complex, multifactorial diseases.
Author Contributions
A. Orlenko, contributed to conception and design, data acquisition, analysis, and interpretation, drafted and critically revised manuscript; J.D. Mure, J.I. Gluch, contributed to conception, data analysis and interpretation, drafted and critically revised the manuscript; J. Gregg, contributed to data analysis, drafted and critically revised the manuscript; C.W. Compher, contributed to data acquisition, drafted and critically revised the manuscript; Z. Ren, contributed to data interpretation, drafted and critically revised the manuscript; H. Koo, J.H. Moore, contributed to data conception and design, drafted and critically revised the manuscript. All authors provided final approval and agreed to be accountable for all aspects of the work.
Supplemental Material
sj-pdf-1-jdr-10.1177_00220345251398027 – Supplemental material for Uncovering Dental Caries Heterogeneity in NHANES Using Machine Learning
Supplemental material, sj-pdf-1-jdr-10.1177_00220345251398027 for Uncovering Dental Caries Heterogeneity in NHANES Using Machine Learning by A. Orlenko, J.D. Mure, J.I. Gluch, J. Gregg, C.W. Compher, Z. Ren, H. Koo and J.H. Moore in Journal of Dental Research
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by National Library of Medicine, National Institutes of Health grant LM010098. Z. Ren was supported by the National Institute of Dental and Craniofacial Research Postdoctoral Training Program under award R90DE031532.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
