Abstract
Oral candidiasis is primarily caused by Candida albicans; however, bacteria such as Streptococcus mutans can also coexist and cooperate within the same biofilm. This study evaluated the combined effects of Histatin 3 (His3) and Histatin 5 (His5) with antimicrobial photodynamic therapy (aPDT) using a 3-dimensional oral epithelium model infected with mixed biofilms of C. albicans (polyene-resistant strains) and S. mutans. The tissues, divided into groups (His3 + aPDT, His5+aPDT, His3, His5, or aPDT), were first treated with the salivary proteins His3 or His5 (2 h, 37 °C, 60 rpm/min) followed by infection with mixed biofilms. After 24 h of biofilm formation, aPDT was applied (PDZ: 200 mg/L; LED light: 660 nm with 50 J/cm²) using intermittent irradiation cycles (n = 9/group). Biofilm viability (CFU/mL), total proteins (intracellular and extracellular), and proteomic analysis were performed. Biocompatibility and tissue invasion were also assessed by MTT assays and histological analysis, respectively. His3+aPDT and His5+aPDT reduced mixed biofilm viability by 67% (P = 0.0038) and 65% (P = 0.0062), respectively. His3 and His5 alone promoted a 35% to 45% (P = 0.018; P = 00.24) reduction, while aPDT alone reduced by 50% to 55% (P = 0.022; P = 0.037). Combined treatments enhanced efficacy by ~30% compared with individual treatments. Histological evaluation also confirmed reduced biofilm penetration and hyphal formation, particularly with His5+aPDT. Proteomic analysis of the proteins from C. albicans revealed the downregulation of structural and transport proteins, which can likely weaken the biofilm. The combination of His3 and His5 with aPDT enhances antimicrobial efficacy against resistant multispecies biofilms while maintaining biocompatibility. By reducing biofilm viability and compromising proteins, this approach provides a promising alternative for treating oral candidiasis associated with polyene-resistant C. albicans species.
Introduction
Oral candidiasis is the most common opportunistic fungal infection encountered in oral tissue, and the fungus Candida albicans is the main pathogen (Vila et al 2020). Although the oral mycobiome is diverse and includes fungi beyond Candida spp (Ghannoum et al 2010; Krom et al 2014), C. albicans remains the primary etiological agent of this disease, being particularly prevalent in immunocompromised patients (Patel 2022). Its ability to form hyphae, penetrate mucosa, and persist in biofilms makes it the most clinically relevant fungal species in this context (Talapko et al 2021).
The bacteria Streptococcus mutans also coexists with C. albicans in multispecies biofilms (Kim et al 2021). C. albicans can induce S. mutans to produce extracellular polysaccharides, which promote biofilm organization and stability (Lobo et al 2019). This interaction is reinforced by sucrose, which increases adhesion and biofilm formation by enhancing glucan production by S. mutans. Both microorganisms act synergistically, increasing the likelihood of individual survival in microenvironments and, consequently, promoting the progression of the infection (Kim et al 2021). Although other oral bacteria may also coexist with fungi, they do not establish the same consistent, virulence-enhancing relationship as these two species (Metwalli et al 2013).
The first choice for controlling oral candidiasis is conventional drugs derived from antifungals. However, antifungals show limited action to control C. albicans, and they are not effective in inhibiting bacterial growth (Gad and Fouda 2020). To overcome this limitation, new therapeutic approaches such as antimicrobial peptides/proteins (AMPs) and antimicrobial photodynamic therapy (aPDT) have emerged as alternatives (Hu et al 2018; Boparai and Sharma 2020). aPDT uses a photosensitizer, which, when activated by light in the presence of oxygen, generates reactive oxygen species (ROS). These ROS effectively inactivate a variety of microorganisms, including bacteria and fungi (Alfei et al 2024). AMPs such as Histatin 3 (His3) and Histatin 5 (His5), extracted from human saliva, have demonstrated outstanding antimicrobial performance (Dias et al 2025). His3 has shown strong antibacterial activity against S. mutans by disrupting cell membranes and impairing energy metabolism (Xu et al 1999), while His5 can lyse C. albicans cells and other bacteria by permeating membranes, it modulates the host immune system (Mochon and Liu 2008). Based on this, the combination of AMPs and aPDT may potentially enhanance the treatment efficacy against resistant pathogens that colonize the oral epithelium, offering an integrated solution to address oral candidiasis.
Thus, the aim of the present study was to investigate the effect of the proteins His3 and His5 combined with aPDT on a 3-dimensional (3D) oral epithelium model infected with polyene-resistant C. albicans and S. mutans mixed biofilms. This investigation assessed the biocompatibility of the combined treatment, its histological aspects, and biofilm protein quantification. Moreover, proteomic analysis was conducted.
Materials and Methods
The complete Materials and Methods are provided in the Appendix. In brief, Figure 1A summarizes the experimental workflow applied in the present study: biofilms of C. albicans (polyene-resistant strain) and S. mutans were established on a 3D oral epithelium model by incubating tissues with microorganisms for 24 h, followed by treatment with His3 or His5, aPDT, or their combination. After treatment, the following assays were performed: biofilm viability, tissue biocompatibility, histological analysis, protein quantification, proteomic profiling, and subsequent bioinformatics analyses.
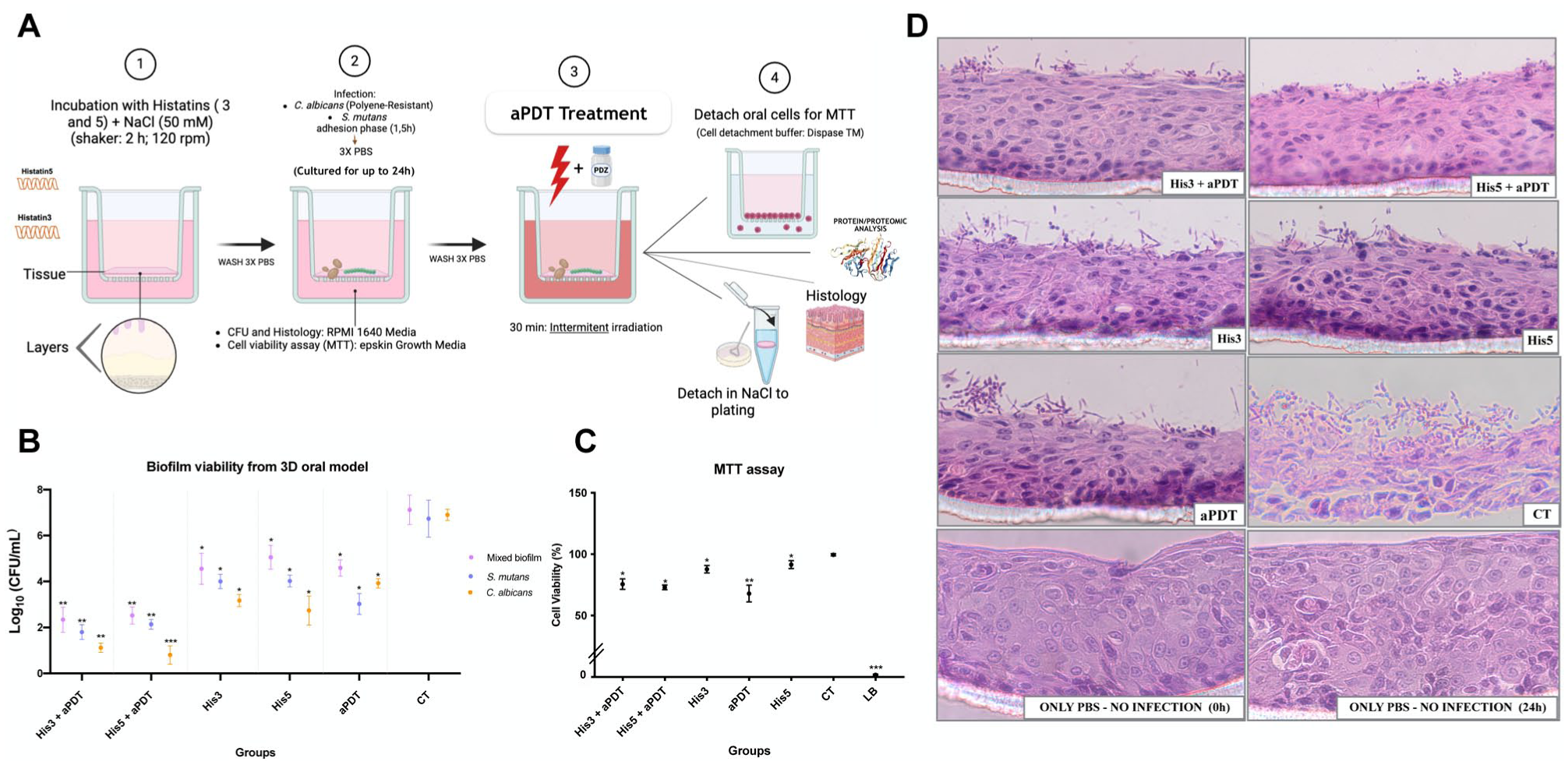
(
Results
Biofilm Cell Viability Determination (by CFU/mL)
For the mixed biofilm of C. albicans and S. mutans cell viability, His3+aPDT and His5+aPDT reduced 67% (P = 0.0038) and 65% (P = 0.0062) of CFU/mL, respectively, both significantly different from the CT (P < 0.001) (Fig 1B). These effects were ~30% enhanced over aPDT or Histatin alone (also compared with the CT). Individually, His3 and His5 reduced viability by 35% to 40% (P = 0.018) and 40% to 45% (P = 0.0024), respectively. aPDT alone achieved 50% to 55% reduction (P = 0.022; P = 0.037) over the CT.
MTT Assay
The MTT assay on the 3D model showed that His3+aPDT and His5+aPDT reduced cell viability by 24% (P = 0.034) and 25% (P = 0.022) compared with the control (CT), respectively, while still being classified as noncytotoxic by ISO guidelines (Fig 1C). His3 and His5 produced minor reductions (10% and 6%; P < 0.05) relative to the CT and were also noncytotoxic. In contrast, aPDT treatment alone had the most pronounced effect, with a reduction in cell viability of about 32% (P = 0.013). The lysis buffer, used as the death control, reduced cell viability to nearly 0%.
Histological Analysis
In the 3D tissue model, the His3+aPDT and His5+aPDT groups showed only a slight presence of mixed biofilm on the surface, with limited penetration into deeper layers (Fig 1D) compared with the other groups. His3 or His5 alone also restricted biofilm invasion compared with CT, with His5 producing fewer invasive hyphae than His3, consistent with the CFU results. aPDT alone resulted in moderate biofilm damage with fewer invasive hypha than CT. In contrast, the CT group showed extensive biofilm growth and deep tissue invasion, whereas the PBS control (0 h and 24 h) groups maintained an intact tissue structure.
Proteins Quantification
The BCA total protein analysis revealed that His3+aPDT had the lowest mean in the analysis (67%; P = 0.0021) compared with the CT (Fig 2A). The His5+aPDT group also showed a significant reduction (40%; P = 0.014) compared with the CT, while the His3 and His5 groups individually decreased proteins by 38% (P = 0.034) and 13% (P = 0.041), respectively. The aPDT group also exhibited a reduction of 28% (P = 0.033) in protein levels compared with the CT. As expected, the CT group maintained the highest protein concentration, serving as the reference baseline.
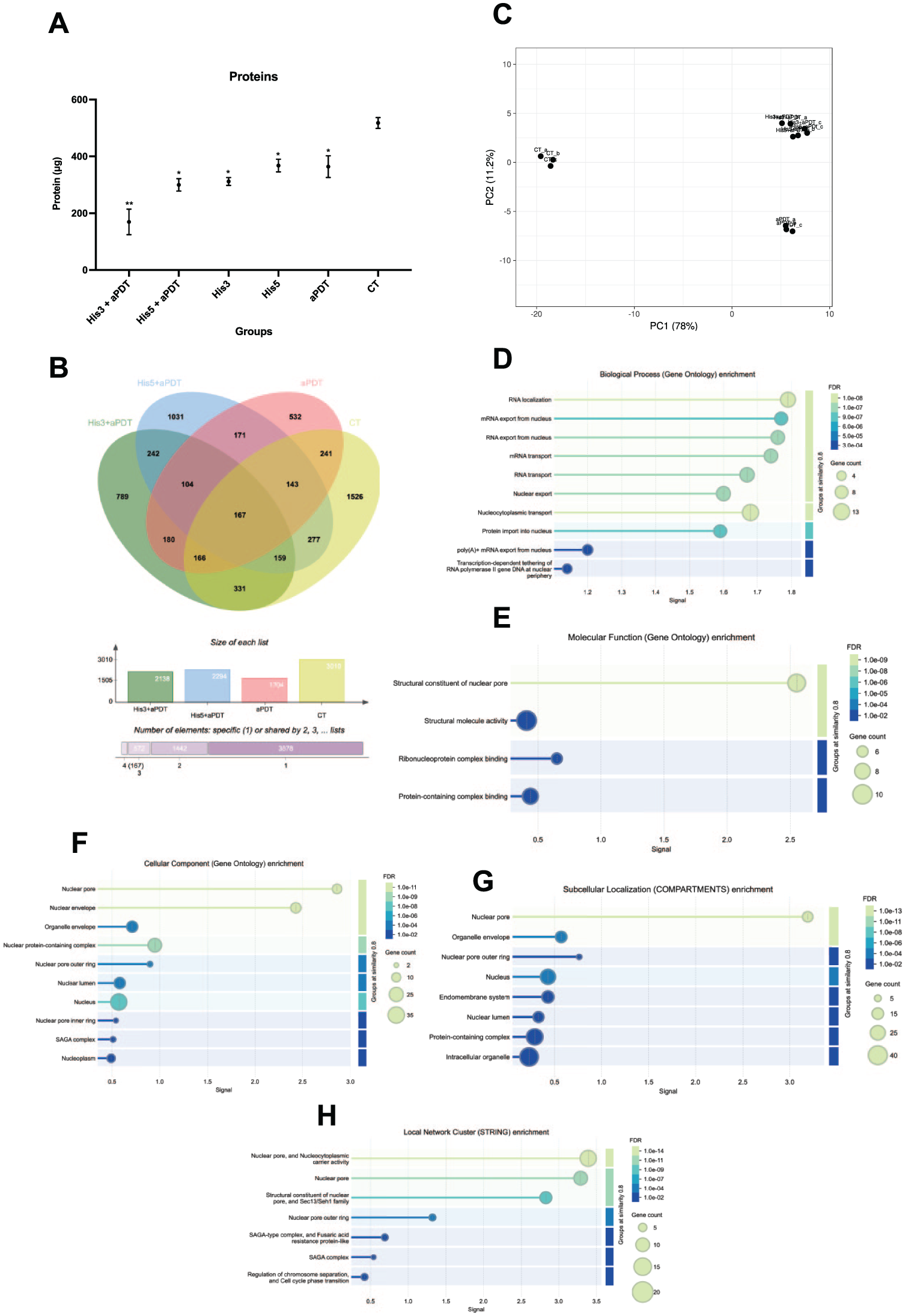
Mean and 95% confidence interval of (
Bioinformatics and Systems Biology Analyses from the Proteomic Profile
Protein overlap and shared expression pattern
To assess the proteins shared across treatments, a Venn diagram was generated (Fig 2B). It illustrates the overlap in protein identification among the 4 experimental groups, with 167 proteins commonly detected across all conditions.
Principal component analysis
Principal component analysis (PCA) was applied to assess replicate consistency and overall separation between conditions. The PCA plot (Fig 2C) demonstrates the clustering of biological replicates and the separation between treatment conditions. PC1 explained 78% of the variance, separating the CT samples into their own cluster, while all aPDT-treated groups (including His3+aPDT and His5+aPDT) grouped together into a single cluster.
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis
To categorize the proteomic profile, Gene Ontology (GO) enrichment classified the proteins into biological processes (BP), molecular functions (MF), and cellular components (CC). Additionally, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis mapped these proteins into biological pathways using STRING-db (Yang et al 2019). The protein–protein interaction (PPI) network showed 60 nodes and 414 edges, significantly exceeding the expected 220 interactions (PPI enrichment P value < 1.0e-16), indicating a strong functional connectivity.
Functional categorization and pathway mapping of the proteome
Among the three GO categories, most enriched terms were found in the BP category. These were strongly associated with RNA transport and localization, including RNA localization (GO:0006403; false discovery rate [FDR] = 8.07e-10), mRNA transport (GO:0051028; FDR = 5.55e-08), RNA transport (GO:0050658; FDR = 5.55e-08), mRNA export from nucleus (GO:0006406; FDR = 1.20e-06), and RNA export from nucleus (GO:0006405; FDR = 7.32e-07). Additional enriched BP terms included protein import (GO:0006606; FDR = 3.53e-08), intracellular transport (GO:0046907; FDR = 2.16e-06), chromosome organization (GO:0051276; FDR = 0.0100), and regulation of biosynthetic processes (GO:0031326; FDR = 0.0165) (Fig 2D). In the MF category, proteins were mainly enriched in structural roles linked to nuclear transport, including structural constituents of the nuclear pore (GO:0017056; FDR = 7.28e-12), ribonucleoprotein complex binding (GO:0043021; FDR = 0.0119), and structural molecule activity (GO:0005198; FDR = 0.0075) (Fig 2E). Finally, the CC category localized these proteins predominantly to nuclear-associated structures such as the nuclear pore (GO:0005643; FDR = 5.21e-12), nuclear envelope (GO:0005635; FDR = 6.63e-12), nuclear membrane (GO:0031965; FDR = 8.82e-06), nucleoplasm (GO:0005654; FDR = 0.0044), intracellular organelle lumen (GO:0070013; FDR = 0.00035), and chromosome organization sites (GO:0005694; FDR = 0.0364) (Fig 2F).
Subcellular localization (COMPARTMENTS) and functional network associations
The analysis of subcellular localization confirmed that the identified proteins were mainly associated with nucleus-related compartments. Enriched terms included the nuclear pore outer ring (GO:0031080, FDR = 0.0064), nuclear lumen (GO:0031981, FDR = 9.43e-06), peribosome and large subunit precursor (GO:0030687, FDR = 0.0238), which were identified and supports the involvement of both enriched terms in nuclear–cytoplasmic transport and ribosomal processing (Fig 2G). Consistently, STRING-db clustering highlighted functional subgroups related to nucleocytoplasmic transport and chromatin remodeling. The most significantly enriched cluster included the nuclear pore and nucleocytoplasmic carrier activity (CL:4692, FDR = 6.88e-17) (Fig. 2H). KEGG pathway enrichment identified RNA transport (cal03013, FDR = 1.61e-06) as the most significantly enriched pathway, reinforcing the GO results.
Heatmap-based clustering of protein expression profiles
The heatmap (Fig 3A) depicts the hierarchical clustering of the 167 most abundant proteins detected across all experimental conditions. All treatment groups showed varying degrees of protein downregulation. The group treated with aPDT alone (triplicates: aPDT_a, aPDT_b, aPDT_c) showed a moderate reduction in protein expression compared with the CT (triplicates: CT_a, CT_b, CT_c). The combination of His3+aPDT (triplicates: His3+aPDT_a, His3+aPDT_b, His3+aPDT_c) led to a stronger reduction in protein abundance than aPDT alone. However, the greatest decrease in protein levels was observed in the His5+aPDT group (triplicates: His5+aPDT_a, His5+aPDT_b, His5+aPDT_c). The 15 proteins showing the greatest downregulation between His5+aPDT and CT are highlighted in Figure 3B, with the full protein list provided in the Appendix.
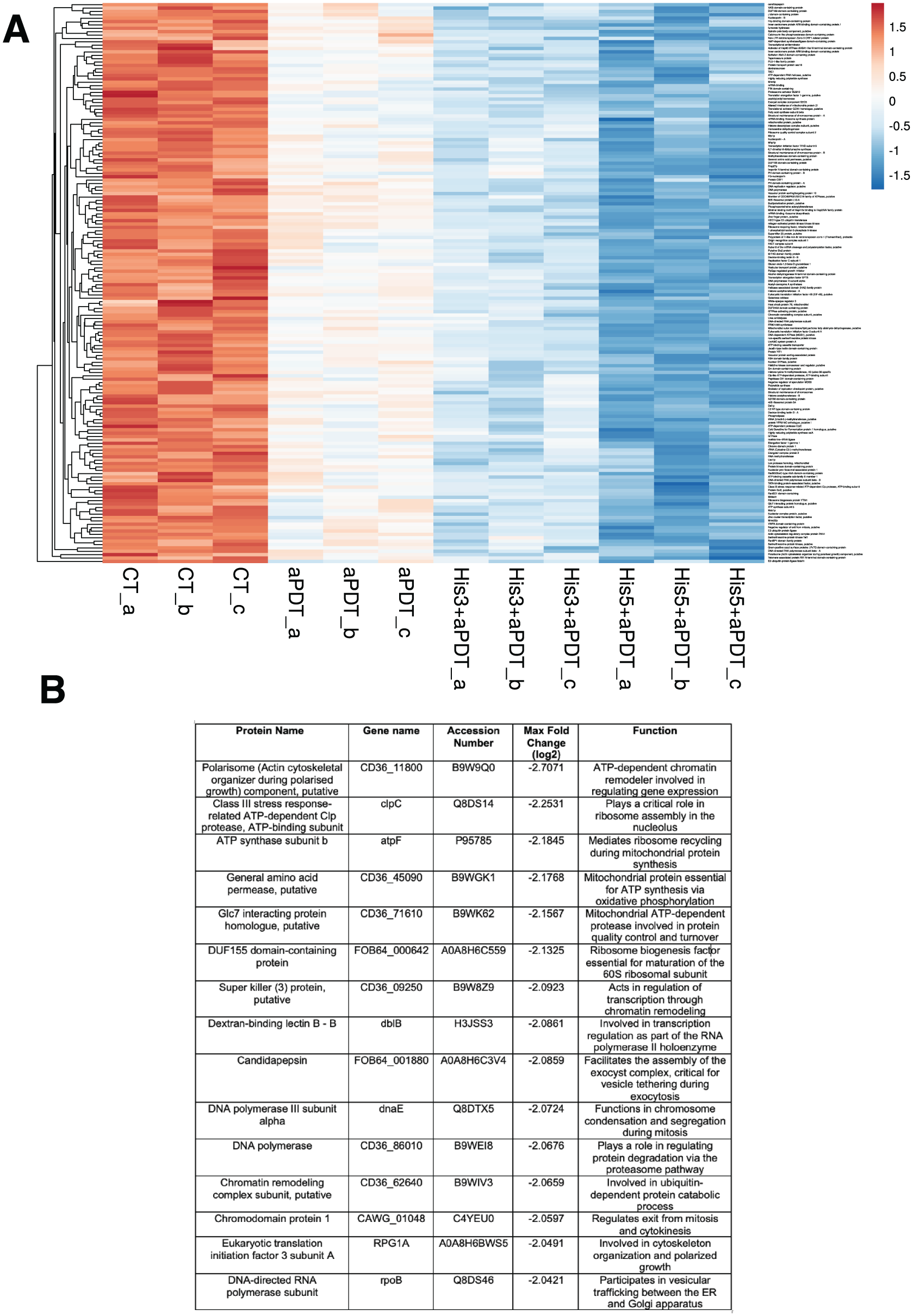
(
STRING-based interaction network analysis
The PPI network analysis identified 60 nodes and 414 edges, exceeding the expected 220 edges (PPI enrichment P value < 1.0e-16) (Fig 4A). The average node degree (13.8) and clustering coefficient (0.572) suggest a well-structured network with interconnected subnetworks (Fig 4B and C).
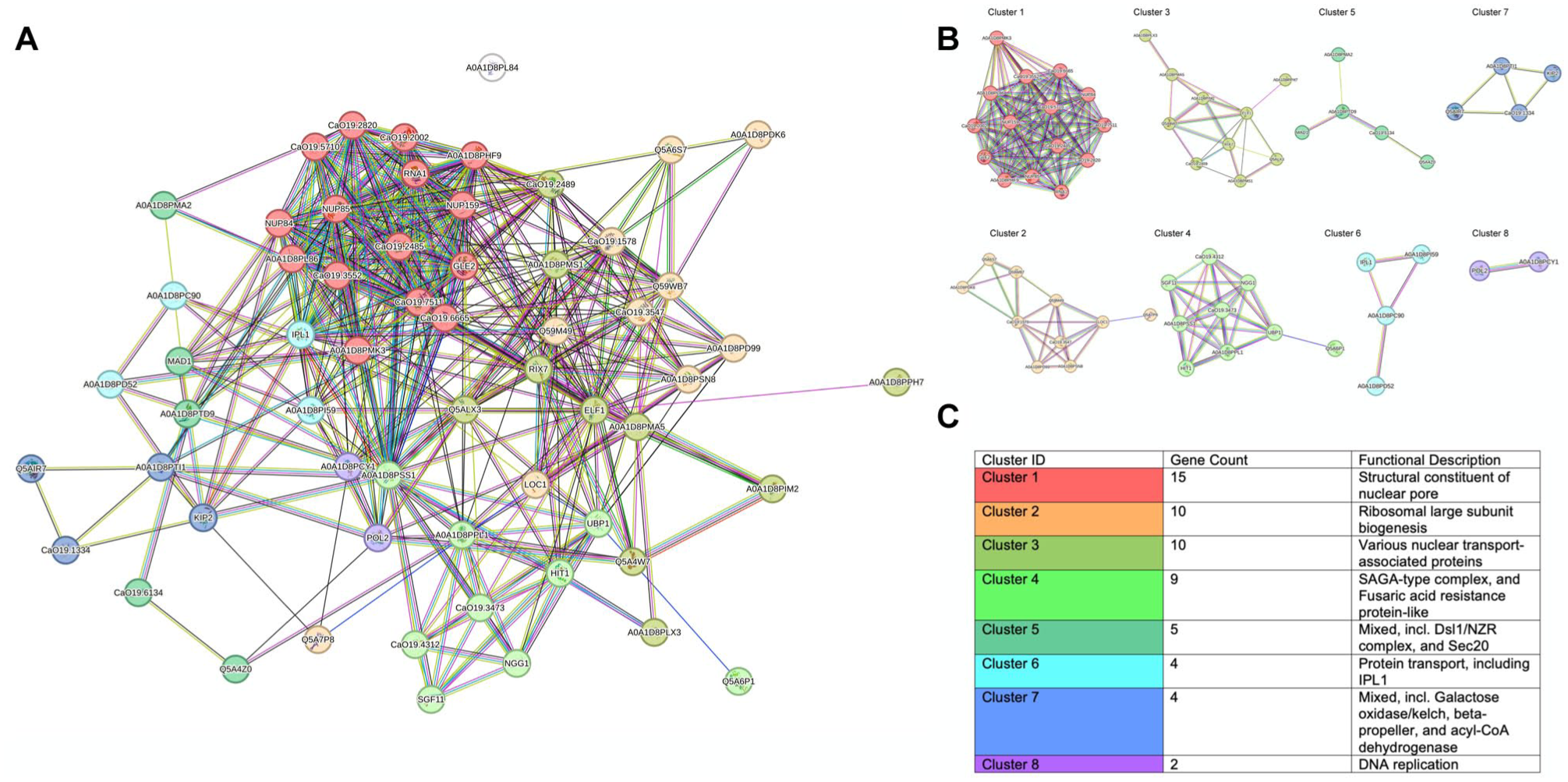
(
Network topology and hub proteins
According to network topology analysis, ELF1 emerged as the protein with the highest connectivity level with a node degree of 33, followed by A0A1D8PSS1 with 31 connections, CaO19.6665 with 28 connections, and CaO19.2489 with 26 connections (Fig 5A).
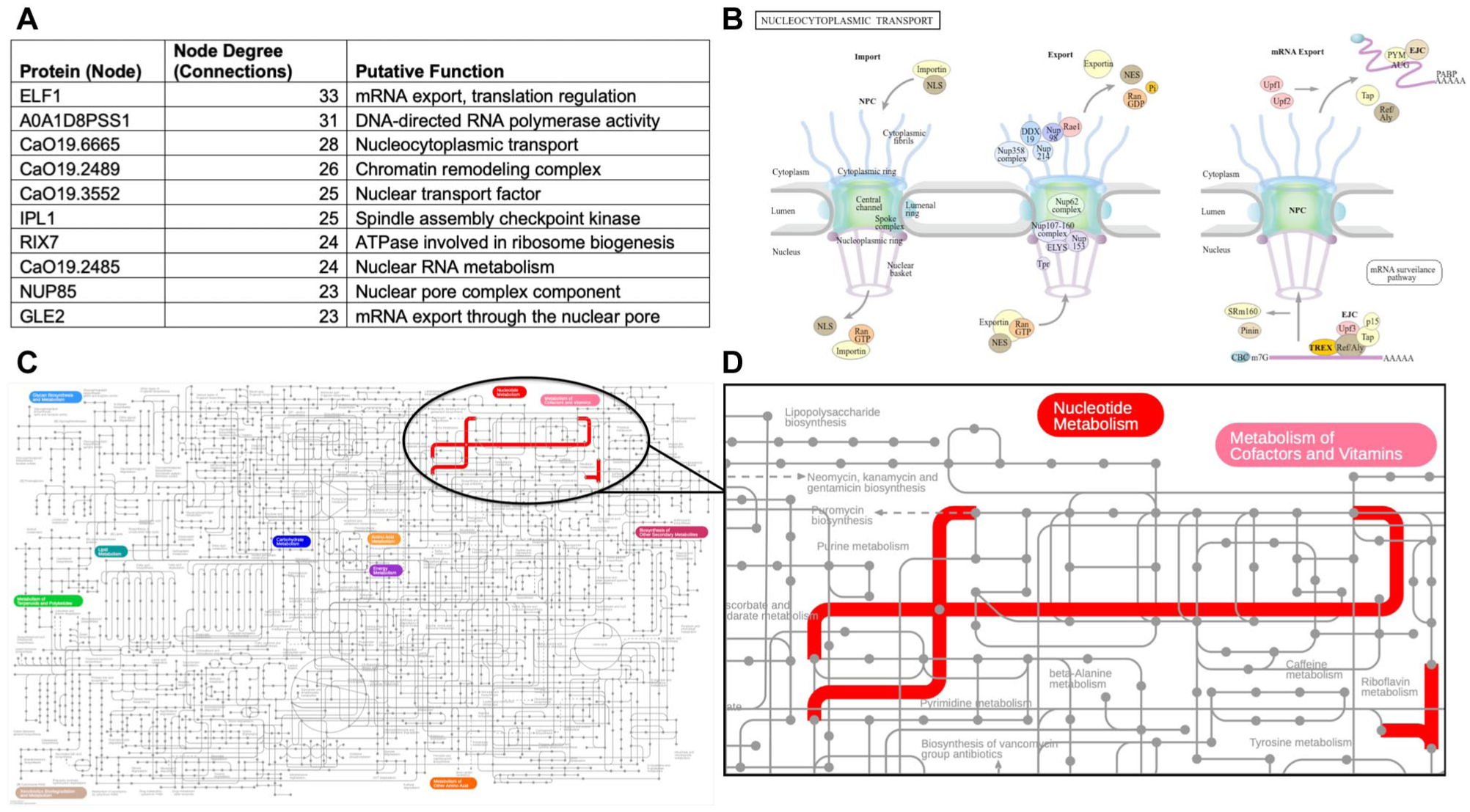
Functional and pathway analysis of identified proteins. (
Nucleocytoplasmic transport pathway analysis
Analysis of protein functions revealed the KEGG nucleocytoplasmic transport pathway as a primary functional process (Fig 5B).
Metabolic pathway analysis
Metabolic pathway mapping using iPath 3.0 revealed a predominance of nucleotide metabolism—particularly purine and pyrimidine pathways—alongside cofactor and vitamin metabolism (Fig 5C and D).
Discussion
This study demonstrates the effectiveness of Histatin with photodynamic therapy against a 3D oral epithelium model infected with mixed biofilms of C. albicans (polyene-resistant strains) and S. mutans. By focusing on a clinically relevant resistant strain, our findings highlight both the antimicrobial and host-protective effects of this therapy. The study of a 3D oral epithelium model is crucial for understanding the complexities of oral infections, particularly in the context of multispecies biofilm interactions (Last et al 2021). This model is configured to create gradients of oxygen, nutrients, and antimicrobials, which are inherent to biofilm behavior and resilience (Fusco-Almeida et al 2023). This investigation assessed for the first time the behavior of the SkinEthic™ 3D oral epithelium model infected by mixed biofilm. The primary mechanism suggested by the results is that both AMPs penetrated the outermost tissue layer, protecting it and avoiding the complete maturation of the biofilm.
In this study, the effects of two different Histatins, His3 and His5, were evaluated in the 3D tissue model, highlighting their dual functions as antimicrobial agents and epithelial protectants. When both proteins were combined with aPDT, their antimicrobial effects were potentiated, achieving greater biofilm reduction than either treatment alone. His5 specifically reduced C. albicans hyphal formation while preserving the epithelial structure. Previous studies have shown His5 to be effective in reducing the metabolic activity (>50%) of C. albicans at lower concentrations (195 µg/mL); however, these did not include mixed biofilms or 3D models (Moffa et al 2015). Here, His5+aPDT produced the greatest reduction in C. albicans viability within the mixed biofilm, with an 88% decrease, followed by the His3+aPDT group with an 84% decrease, both compared with the control group. Considering the microbial interplay, the results indicate that C. albicans was more affected by Histatin-aPDT treatments than S. mutans was. This distinct susceptibility may be due to differences in their cell walls and biofilm matrix composition. The fungal wall, composed of mannoproteins and β-glucans, allows Histatin binding and action, whereas the thick peptidoglycan layer of S. mutans may restrict protein entry, reducing its effectiveness (Xiao et al 2023). In addition, the coexistence of S. mutans with C. albicans in the same biofilm typically produces abundant glucan-based extracellular polysaccharides that shield the bacterial cells from external agents (Kim et al 2017).
aPDT combined with His3 and His5 demonstrated favorable biocompatibility in a 3D oral model. The addition of His3 and His5 decreased cell viability by 24% and 25%, respectively. Interestingly, aPDT alone resulted in the greatest reduction in cell viability and is classified as slightly cytotoxic although still within the acceptable range for in vitro assays (International Organization for Standardization 2009; Last et al 2021). In contrast, when combined with His3 or His5, the treatments modulated this effect, resulting in viability levels that were considered noncytotoxic. These findings suggest that Histatins may help protect host cells not only from microbial infection but also from cytotoxic stress. This behavior is supported by the unique action of Histatins, whose effects vary between microorganisms and host cells, exhibiting selective activity depending on the type of surface with which they are interacting (Macri et al 2025). One potential mechanism for this protective action is that Histatins act as scavengers of ROS generated when excited by light (Komatsu et al 2019).
The histological examination of the tissue confirmed that the groups treated with His3 or His5 combined with aPDT had improved protection of the epithelium from the biofilm infection, showing minimal adherence and penetration of C. albicans and S. mutans. This aligns with other studies that have indicated the antimicrobial effect of AMPs can influence the composition of biofilms when used with adjuvant therapies (Dias et al 2022). Notably, the His5 group alone exhibited fewer invasive hyphae compared with His3, showing the distinct bioactivity of this specific type of Histatin against fungal invasion as well as supporting the CFU viability results from the current study (Mochon and Liu 2008). Moreover, considering that hyphae are the invasive form of Candida ssp., their absence, or even the reduction of this structure, suggests that this fungus is less invasive in this type of mixed biofilm, with less tissue damage and reduced infection severity (Gulati and Nobile 2016).
In terms of total proteins (intracellular and extracellular), the BCA analysis showed that the His3+aPDT group had the lowest mean levels of protein levels compared with the CT. Proteins are essential components of the matrix, playing multiple roles that can enhance biofilm resilience (Flemming et al 2007). The decrease in proteins disrupts crucial cell-to-cell contacts and its adhesion to surfaces (Panariello et al 2018, 2021). Based on these findings here, the reduction in protein content weakens the biofilm architecture, making this structure more permeable and consequently facilitating the photodynamic action that promotes cell lysis. Although His3+aPDT showed the greatest reduction in total proteins based on BCA quantification, the proteomic evaluation revealed that His5+aPDT promoted a more pronounced downregulation of specific proteins in the mixed biofilm. This apparent discrepancy likely reflects two complementary mechanisms: a broader, nonselective degradation of proteins in the His3+aPDT group (bulk protein reduction) versus targeted suppression of key structural and regulatory proteins in the His5+aPDT group (Wu et al 2014).
The heatmap-based proteomic analysis revealed that Hist5+aPDT significantly disrupts protein-associated expression. Previous research has demonstrated that antimicrobial treatments are able to reduce protein biosynthesis and damage regulatory mechanisms in C. albicans biofilm (Song et al 2022). The pronounced downregulation observed in the His5+aPDT group reflects a synergistic effect involving oxidative and proteolytic stress, which can contribute to the biofilm resilience (Zolin et al 2021). In addition, the 15 most downregulated proteins in the analysis include components involved in ribosomal structure, metabolic processing, and membrane transport—key elements for biofilm maintenance and virulence. These protein-level changes are further contextualized by GO enrichment and STRING-based network analyses. Chromatin remodelers are crucial for modifying chromatin structure, allowing the transcription of genes related to stress response, adhesion, and biofilm maintenance (Hefny et al 2024), potentially compromising biofilm survival and adaptive responses in the present investigation.
In the PCA plot, combination therapies formed distinct clusters separated from the CT along PC1, which explained 78% of the variance, indicating marked differences in protein expression profiles due to treatment effects (David and Jacobs 2014). These proteomic findings complement CFU/mL and histological data, reinforcing that combining Histatins with aPDT elicits a broader biological response at both molecular and structural levels compared with individual treatments. Among the 167 shared proteins, 60 were selected for PPI and GO enrichment analysis, confirming that core cellular functions—such as RNA transport, nuclear pore activity, and chromatin regulation—remained active across all conditions, suggesting a conserved molecular framework essential for C. albicans biofilm survival. This aligns with studies identifying chromatin remodeling complexes as critical regulators of fungal morphogenesis and stress response. In addition, network topology analysis highlighted hub proteins linked to mRNA export, translation, and nucleocytoplasmic transport, which are also related to biofilm maintenance (Jeong et al 2001). For instance, ELF1, in particular, is a transcription elongation factor that binds RNA polymerase II and helps maintain chromatin structure in actively transcribed regions, contributing to gene regulation under stress conditions (Prather et al 2005).
Network clustering analysis revealed 8 subclusters, with cluster 1 containing 15 genes and the most enriched function being the structural constituent of the nuclear pore, a key element in nucleocytoplasmic transport (Petrovic et al 2022). The enrichment of this pathway may indicate disruption of regulatory mechanisms related to RNA trafficking and stress response, which are relevant for biofilm persistence under treatment conditions. Nucleocytoplasmic transport is a vital process for regulating the stress response through transcription regulators and RNA, specifically between the nucleus and the cytoplasm (Yang et al 2017). The activity of this system enables C. albicans to regulate essential genes and rapidly adapt to therapeutic stress, thus maintaining the stability of fungal cells (Enjalbert et al 2003). Supporting this, the metabolic pathway mapped by iPath 3.0 showed enrichment in purine and pyrimidine metabolism, both of which are directly linked to nucleic acid synthesis and RNA processing.
Although this 3D model effectively mimics the in vivo environment for initial evaluation, it lacks the ability to replicate long-term outcomes, such as cell turnover or tissue regeneration. Moreover, this model does not replicate key physiological factors such as salivary flow, shear forces from mastication, or the presence of immune cells (Last et al 2021). We limited the biofilm development to 24 h based on evidence indicating that longer infection durations in oral mucosa models can severely impair epithelial viability and consistency (De Ryck et al 2014). This limitation suggests that complementary in vivo studies are also important to fully understand the therapeutic potential of Histatin-mediated aPDT. Although the aPDT parameters used here ensured the delivery of the optimal therapeutic dose and have been considered acceptable in clinical contexts (Alves et al 2020), future investigations should explore strategies to shorten the irradiation period, such as optimizing light delivery systems or using higher-intensity light sources, to facilitate translation into routine clinical practice.
Antimicrobial strategies such as Histatin-aPDT remain highly relevant, as they provide immediate therapeutic activity while maintaining compatibility with ecological approaches. The combination of His3 or His5 with aPDT proved effective against resistant mixed-species biofilms, suggesting it could be translated into a promising delivery system, such as a hydrogel, a mucoadhesive paste, or an oral rinse, that contains Histatins and a photosensitizer. The dual-phase approach (Histatin application followed by light activation of the photosensitizer) could be adapted as an adjunctive therapy in clinical dentistry.
Conclusion
This study demonstrates that His3 and His5, when combined with aPDT, exhibit antibiofilm effects against mixed biofilms of polyene-resistant C. albicans and S. mutans while also protecting the 3D oral epithelium model. Proteomic analysis revealed reduced expression of proteins critical for cell stability, highlighting the treatment’s molecular impact.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345251392876 – Supplemental material for Histatin–Photodynamic Therapy for Polyene-Resistant Biofilms in a 3D Model
Supplemental material, sj-docx-1-jdr-10.1177_00220345251392876 for Histatin–Photodynamic Therapy for Polyene-Resistant Biofilms in a 3D Model by L.M. Dias, A.C. Pavarina and W.L. Siqueira in Journal of Dental Research
Authors Contributions
L.M. Dias, contributed to conception and design, data acquisition, drafted and critically revised the manuscript; A.C. Pavarina, contributed to conception, data acquisition, analysis, and interpretation, critically revised the manuscript; W.L. Siqueira, contributed to conception, data acquisition and interpretation, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Footnotes
Acknowledgements
We gratefully acknowledge the Histology Core Facility at the Health Sciences Building, University of Saskatchewan, for their valuable support with the histological procedures essential to the development of this study.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Canadian Institutes of Health Research #PJT-159760 and #MOP-106657 and the Natural Sciences and Engineering Research Council #RGPIN 2020−06119 to W.L.S.; the São Paulo Research Foundation (Fundação de Amparo a Pesquisa do Estado de São Paulo FAPESP# 2013/07276-1 [CePID CePOF]) and National Council for Scientific and Technological Development CNPq #306533/2023-5 to A.C.P.; and FAPESP# 2020/16227-8 and 2022/12059-9 and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES) – finance code 0001, under the CAPES-Pring program to L.M.D.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
