Abstract
The coordination of dental cells is essential for tooth development. Various dental epithelial cell types are involved in tooth development, and each cell type plays a distinct role; however, the functional role of the outer enamel epithelium (OEE) remains unclear. We performed single-cell RNA-sequence (scRNA-seq) analysis of postnatal day (P) 7 incisors and embryonic day (E) 14 P1 molars of the mouse tooth germ to reveal the gene expression profile and role of OEE during tooth development. We identified keratin 15 (Krt15) as a specific marker gene of OEE in the dental epithelium. Characterization of dental epithelial clusters using scRNA-seq suggested that Krt15-negative cycling inner enamel epithelial (IEE) cells give rise to Krt15-positive OEE cells, whereas the proliferative activity of dental epithelial cells decreases toward the development of OEE cells. We performed ex vivo organ cultures of the tooth germ to examine the effects of Krt15 knockdown on tooth development. Depletion of Krt15 in the tooth germ resulted in ectopic expression of Ki67 in OEE cells, leading to the development of an abnormal dental epithelial structure. We used the dental epithelial cell line CLDE to assess the molecular mechanisms regulated by Krt15. Krt15-depleted CLDE cells showed abnormal cellular morphology and dysregulated gene expression of cytokeratin family members. Furthermore, Krt15 knockdown in CLDE cells upregulated the expression of cell proliferation marker genes, such as Mki67. Furthermore, Krt15-depleted CLDE cells exhibited activation of the p38 MAP kinase (MAPK) pathway and high proliferative activity. This suggested that Krt15 may control tooth germ size, inhibits p38 activation, and may act as a suppressor of dental epithelial cell proliferation. These findings provide new insights into the role of OEE in tooth development and contribute to a better understanding of the mechanisms underlying tooth morphogenesis.
Keywords
Introduction
The enamel organ is a typical example of a highly multifaceted epithelial organ consisting of various cell types, each of which has a specialized function that greatly influences tooth development processes. The dental stem cells (DESCs) differentiate into various types of enamel epithelial cells: ameloblast progenitor inner enamel epithelium (IEE) cells, stratum intermedium (SI) cells, stellate reticulum (SR) cells, and outer enamel epithelium (OEE) cells. Although various marker genes of ameloblasts have been identified (Yoshizaki et al. 2017; Chiba, Yoshizaki, et al. 2020; Saito et al. 2020; Chiba et al. 2023), the gene expression profile and distinct roles of OEE have not been well-characterized.
OEE cells are cuboidal epithelial cells that connect with IEE cells in the cervical loop and exhibit cytoplasmic features resembling those of IEE cells (Inage et al. 1987). Accumulating evidence suggests that the OEE layer plays an important role in tooth morphogenesis with high regenerative potential as it gives rise to successional dental lamina. The transcription factor Iroquois homeobox 1(Irx1) is specifically expressed in OEE during tooth development, and Irx1 ablation in mice results in abnormal tooth germ morphology (Yu et al. 2017). In human tooth development processes, the proliferative activity of OEE is reduced after the bell stage, while the OEE layer continuously expresses the epithelial stem cell marker Transformation-related protein 63 (Trp63) (Laurikkala et al. 2006). In Msh homeobox 2 (Msx2) null mice, the OEE became highly proliferative and transformed into a keratinized stratified squamous epithelium, resulting in impaired enamel formation (Nakatomi et al. 2018).
Recent advances in single-cell RNA-sequence (scRNA-seq) analyses have provided a deeper understanding of organ development. Notably, scRNA-seq analyses of the tooth germs have enabled us to analyze the gene expression in each dental cell type, and several genes have been identified as markers of dental cell types (Sharir et al. 2019; Chiba, Saito, et al. 2020; Krivanek et al. 2020; Chiba et al. 2022). However, the functional roles of these marker genes in tooth development are not yet well established. Here, we used scRNA-seq analysis of the tooth germ to clarify the gene expression profile and role of OEE cells in tooth development.
Materials and Methods
Detailed descriptions of the methodology are provided in the Appendix.
Results
Keratin 15 Was Highly Expressed in OEE
We first performed scRNA-seq on postnatal day (P) 7 KRT14-RFP mouse incisors to identify specific marker genes for OEE. The clustering results identified 11 clusters consisting of dental epithelial clusters, dental mesenchymal clusters, and blood-derived cell clusters according to marker gene expression (Fig. 1A, B; Appendix Fig. 1A, B and Table 1). The dental epithelium–derived clusters were classified into 4 groups: IEE, preameloblast (Pre-Am), ameloblast (Am), and OEE. Differential expression analysis revealed the DEGs in the OEE cluster; among them, previously reported OEE marker genes such as coiled-coil-helix-coiled-coil-helix domain containing 10 (Chchd10), insulin like growth factor binding protein 2 (Igfbp2), keratin 15 (Krt15), and SPARC like 1 (Sparcl1) were highly detected in the OEE cluster (Sharir et al. 2019; Hermans et al. 2022) (Fig. 1B, C). Feature plot analysis was performed to examine the distribution of these OEE marker genes. The canonical OEE markers Irx1 and Notch2 were specifically expressed in the OEE cluster, and Krt15 was highly expressed in the OEE cluster (Fig. 1D). The cytokeratin family showed a specific distribution depending on the type of epithelial cell, and several members of this family play a critical role in tooth development (Ravindranath et al. 2001; Ravindranath et al. 2003; Inada et al. 2024). Therefore, we paid particular attention to the expression of Krt15 during tooth development.
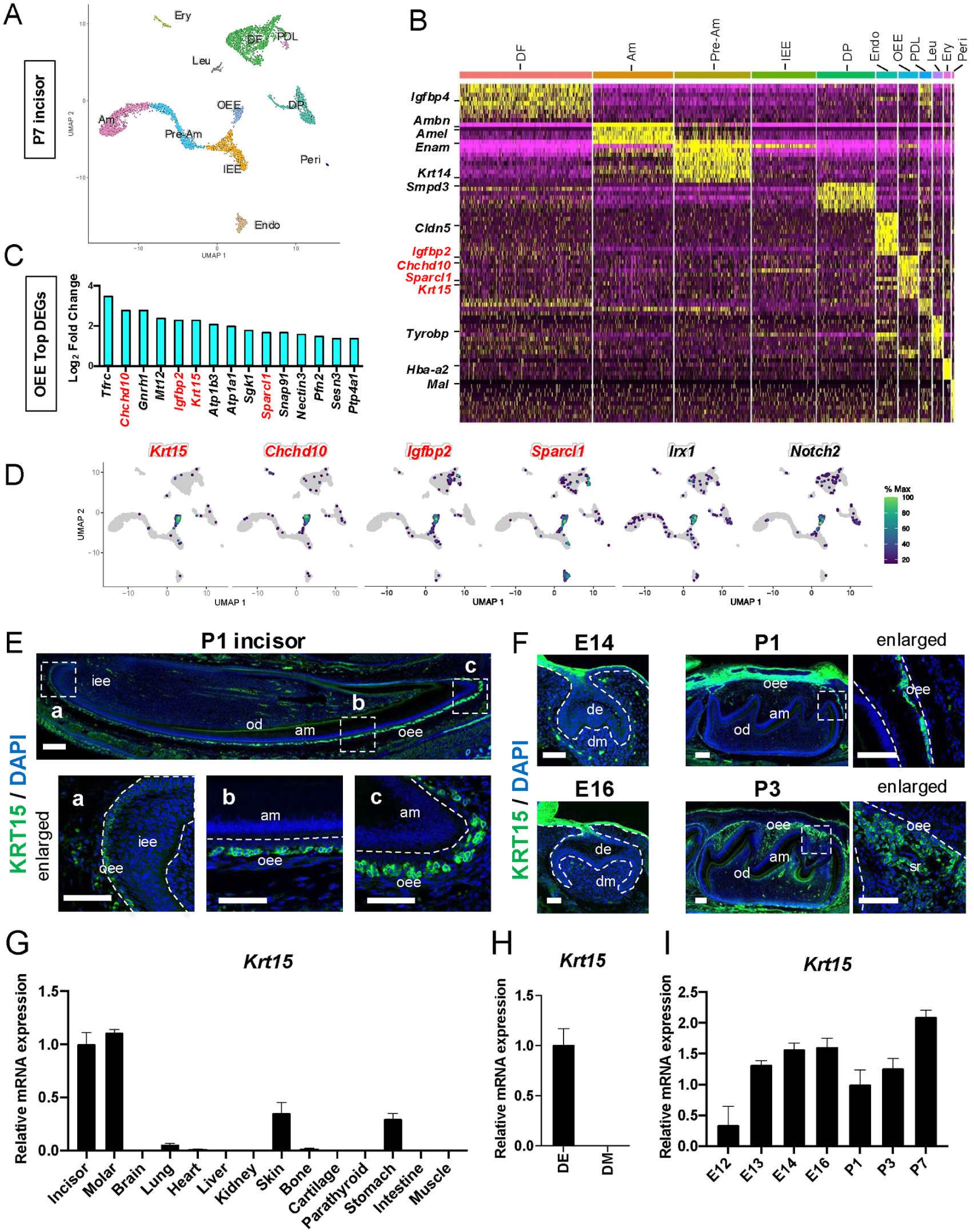
Gene expression profile of Krt15 during tooth development. (
Next, we performed immunofluorescence staining to examine the localization of KRT15 in the tooth germ sections (Fig. 1E, F and Appendix Fig. 2). KRT15 expression was restricted to OEE cells in the P1 incisors (Fig. 1E). Initially, weak fluorescence was detected in OEE cells in the cervical loop region. However, KRT15 was expressed in OEE cells in a continuous and gradually increasing manner following ameloblast differentiation (Fig. 1E). KRT15 was highly expressed in the oral epithelium and gradually extended to the dental lamina in embryonic day (E) 14 and E16 molars (Fig. 1F). Among the dental epithelial cells of P1 molars, only OEE cells expressed KRT15. Continuing to the P3 and P7 molars, it could be broadly detected in both OEE and SR cells (Fig. 1F and Appendix Fig. 2). The expression of Krt15 was high in the tooth germ among various tissue specimens and restricted in dental epithelium (Fig. 1G and H). We further examined the expression of Krt15 in the developing tooth germ; initiation stage (E12), bud stage (E13), cap stage (E14), bell stage (E16), presecretory stage (P1), secretory stage (P3), and late-secretory stage (P7) (Fig. 1I). Krt15 expression was gradually increased during the embryonic stage, and at P1, the expression was decreased. After that, the expression was increased until P7 in the dental epithelium (Fig. 1I).
Krt15-Positive OEE Cells Showed Low Expression of Cell Proliferation Marker Genes
We performed scRNA-seq using E14 and P1 molars of KRT14-RFP mice to describe the transition of the dental epithelial cell population (Fig. 2A–D). At the E14 cap stage, dental epithelial cells formed 1 cluster (dental epithelium), whereas differentiated dental epithelial cells in the P1 molar were classified into 4 clusters: IEE, OEE, Pre-Am, and Am (Fig. 2A, C, and Appendix Fig. 3A–D). We analyzed the expression of the OEE marker genes Krt15 and Trp63 (Laurikkala et al. 2006); ameloblast marker genes Amelogenin (Amel), Enamelin (Enam), and Ameloblastin (Ambn); and cycling cell marker gene Marker of proliferation Ki-67 (Mki67) in the dental epithelium subset of the scRNA-seq datasets (Fig. 2B, D). Amel, Enam, and Ambn were not detected in the E14 molar dataset, whereas Krt15, Trp63, and Mki67 were expressed in only 1 cluster. These marker genes were expressed separately in the P1 molar dataset, suggesting that ameloblasts, IEE, and OEE could be clearly distinguished during differentiation. We further analyzed these genes in the P7 incisor dataset, where we observed the continuous development of the dental cells (Fig. 2E, F). Mki67 was highly expressed in the IEE cluster, and this cluster showed a branch of development into Amel-, Enam-, and Ambn-positive ameloblasts cluster and Krt15 positive OEE cluster (Fig. 2F and Appendix Fig. 4A). Other cycling cell marker genes such as Baculoviral IAP repeat containing 5 (Birc5) and Cyclin B1 (Ccnb1) were also highly expressed in the IEE cluster (Appendix Fig. 4B). Next, we performed a trajectory analysis of the dental epithelium subset in the P7 incisor to examine the transition in gene expression through the developmental processes (Fig. 2G). Trajectory analysis indicated that the IEE progenitors of the dental epithelium and the IEE cluster give rise to 2 different branches of cell fate: ameloblasts and OEE. We extracted the IEE and OEE clusters as inputs for the pseudo-time analysis and examined the expression of the OEE marker genes Krt15 and Trp63 (Fig. 2H). Both Krt15 and Trp63 showed increased expression during IEE differentiation into OEE. Notably, the cell proliferation marker genes Mki67 and Ccnb1 showed opposite expression patterns to Krt15 and Trp63, which were highly expressed in IEE and were gradually downregulated during OEE differentiation. We examined the localization of Krt15 and cell proliferation marker genes in the P7 incisor scRNA-seq dataset (Appendix, Fig. 4C). Krt15 and Trp63 were co-localized in the OEE cluster; in contrast, Krt15 and cell proliferation markers (Mki67 and Ccnb1) did not colocalize.
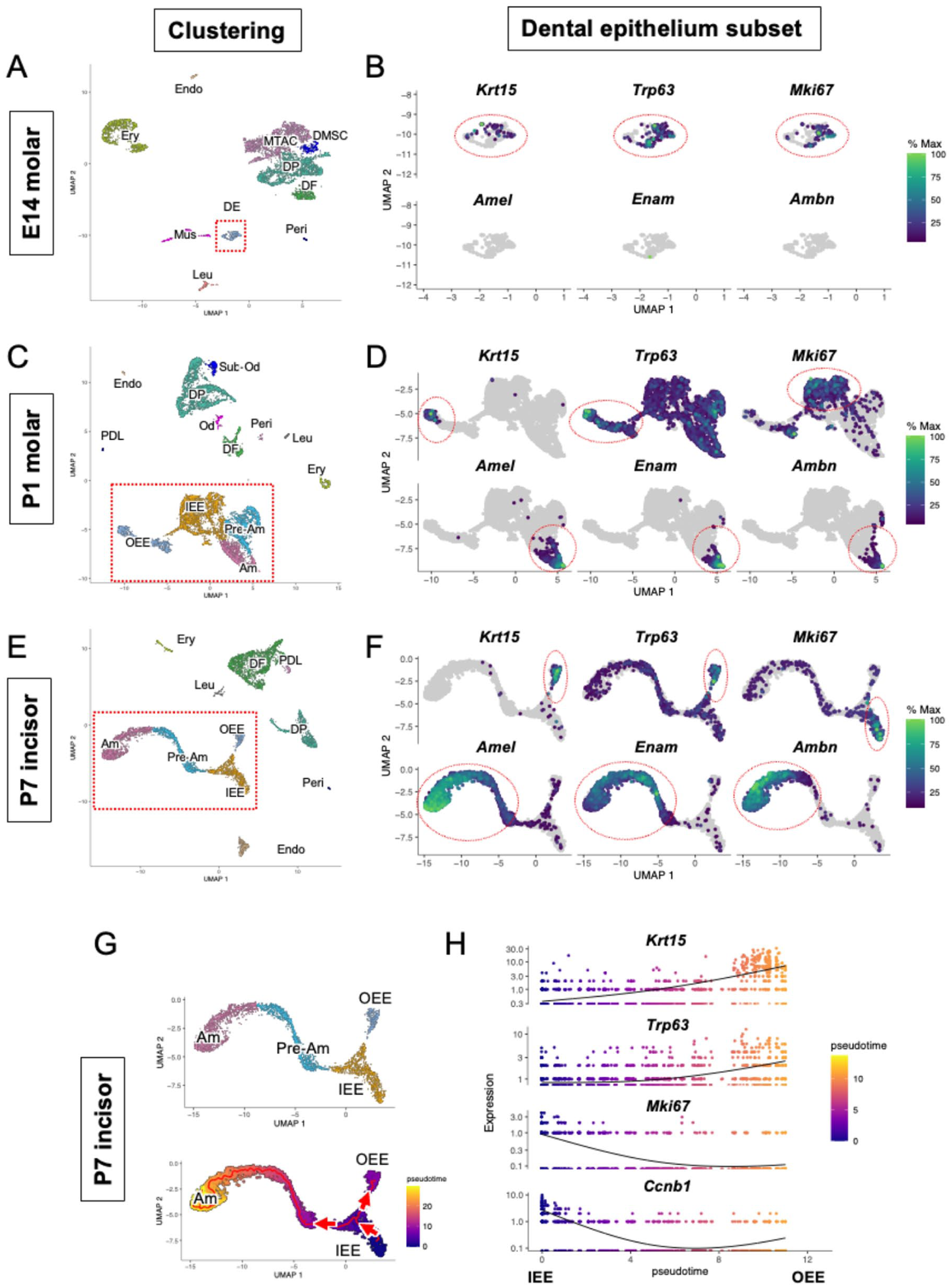
Transition of the Krt15-positive outer enamel epithelial cell population in scRNA-seq datasets. (
Krt15 Regulates Cell Proliferation of OEE during Tooth Morphogenesis
To explore the function of Krt15 in dental epithelial cells, we conducted knockdown experiments of Krt15 in the developing tooth germ using the ex vivo tooth germ organ culture method (Han et al. 2020). The mandibular first molar was dissected from E14 mice and cultured in a cell culture insert with control siRNA (siNeg) or siRNA against Krt15 (siKrt15) for 5 d (Appendix Fig. 5). We confirmed that the siKrt15 group substantially suppressed the expression of Krt15 compared with the siNeg group in the organ culture system (Fig. 3A). Five days after transfection, the dental epithelium of the siKrt15-transfected tooth germ showed elongation compared with that of the control tooth germ (Fig. 3B and Appendix Fig. 6A). We examined the shape of the cultured tooth germs using microscopy. The distance between the cervical loops and the width of the tooth germ was significantly decreased in the siKrt15-transfected group (Fig. 3C and Appendix Fig. 6B). We further performed a histological analysis of the cultured tooth germ using frozen sections of tooth germ. We used heparan sulfate proteoglycan 2 (HSPG2) as a basement membrane marker, to distinguish the border of the epithelium and mesenchyme (Fig. 3D). Notably, a greater number of Ki67-positive cells were detected in the OEE and dental mesenchyme of the siKrt15-transfected tooth germ cells (Fig. 3D). Interestingly, ectopic proliferation was observed not only in OEE but also the root epithelium. Furthermore, the ratio of Ki67-positive cells was significantly increased in both dental epithelium and mesenchyme (Fig. 3E). These results suggested that KRT15 suppresses cell proliferation in the OEE and regulates tooth morphogenesis.
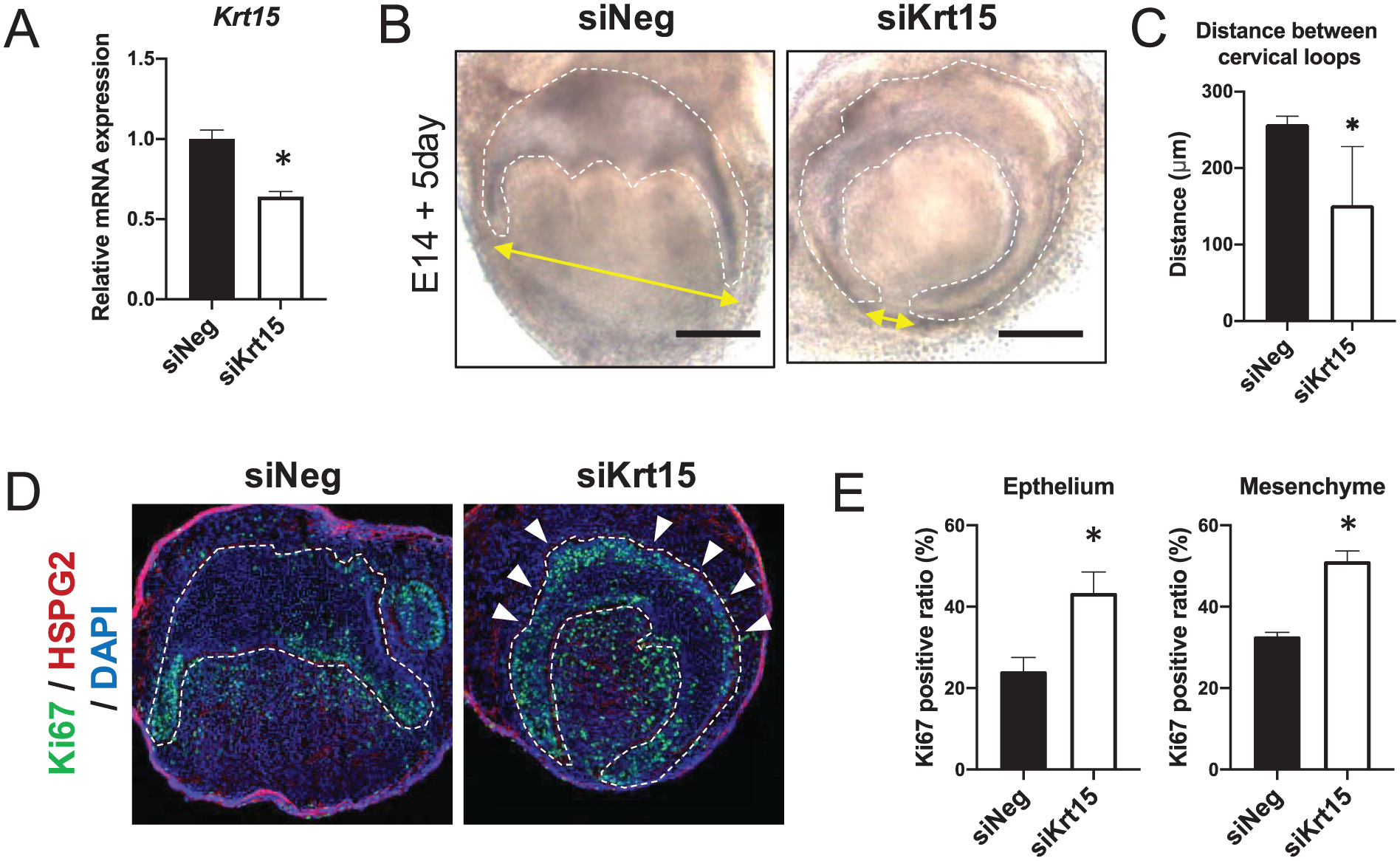
Effect of Krt15 knockdown on tooth morphogenesis. (
Depletion of Krt15 Promoted the Proliferation Activity of Dental Epithelial Cells
We further explored the molecular mechanisms regulated by Krt15 in the dental epithelium using CLDE cells. KRT15 knockdown at the protein level in CLDE cells was validated using Western blotting and immunocytochemistry (Appendix Fig. 7A, B). The results showed the suppression of KRT15 protein. Interestingly, depletion of Krt15 affected the cellular morphology of CLDE cells, suggesting that cytoskeleton formation was altered (Fig. 4A). In the siNeg-group, the phalloidin intensity peak was observed in the cell–cell contacts between dental epithelial cells, while in the siKrt15 group, the phalloidin intensity was significantly altered, suggesting that F-actin fiber is radially localized in the cytoplasm (Fig. 4B). Next, we analyzed the DEGs in siNeg- or siKrt15-transfected CLDE cells using a microarray (Appendix Fig. 8A, B). Heatmap and gene ontology analyses showed that the loss of Krt15 may induce a core set of cell proliferation genes. Furthermore, the downregulated genes were associated with the cytoskeleton. We paid particular attention to the changes in the expression of cytokeratin family members (Appendix Fig. 8C, D) and found that the expression of Krt5, Krt6a, Krt10, and Krt14 was not altered. However, the expression of Krt8, Krt17, and Krt18 was significantly altered. Reverse transcription quantitative polymerase chain reaction (RT-qPCR) results confirmed that siKrt15 depleted the expression of Krt15 at the mRNA level and that the mRNA expression levels of proliferation marker genes, namely, Birc5, Ccnb1, and Mki67, were markedly upregulated in Krt15-knockdown CLDE cells (Fig. 4C). However, the expression levels of the ameloblast (Ambn and AmeloD) and SI (Notch1 and Notch2) marker genes remained unaltered (Appendix Fig. 8E, F). These results suggest that Krt15 may suppress the expression of cell proliferation marker genes. Next, we examined the effect of Krt15 depletion on cell proliferation. The depletion of Krt15 resulted in the upregulation of Ki67 expression at the protein level (Fig. 4D). Furthermore, the ratio of EdU-positive cells increased significantly after Krt15 knockdown (Fig. 4E). We assessed the proliferative activity of CLDE cells using the CCK-8 assay and found that Krt15-depleted cells showed significantly higher proliferative activity than control cells did (Fig. 4F). In addition, phosphorylation of retinoblastoma (Rb) protein was induced by knockdown of Krt15 in CLDE cells (Appendix Fig. 9), suggesting that Krt15 may act as a suppressor of dental epithelial cell proliferation.
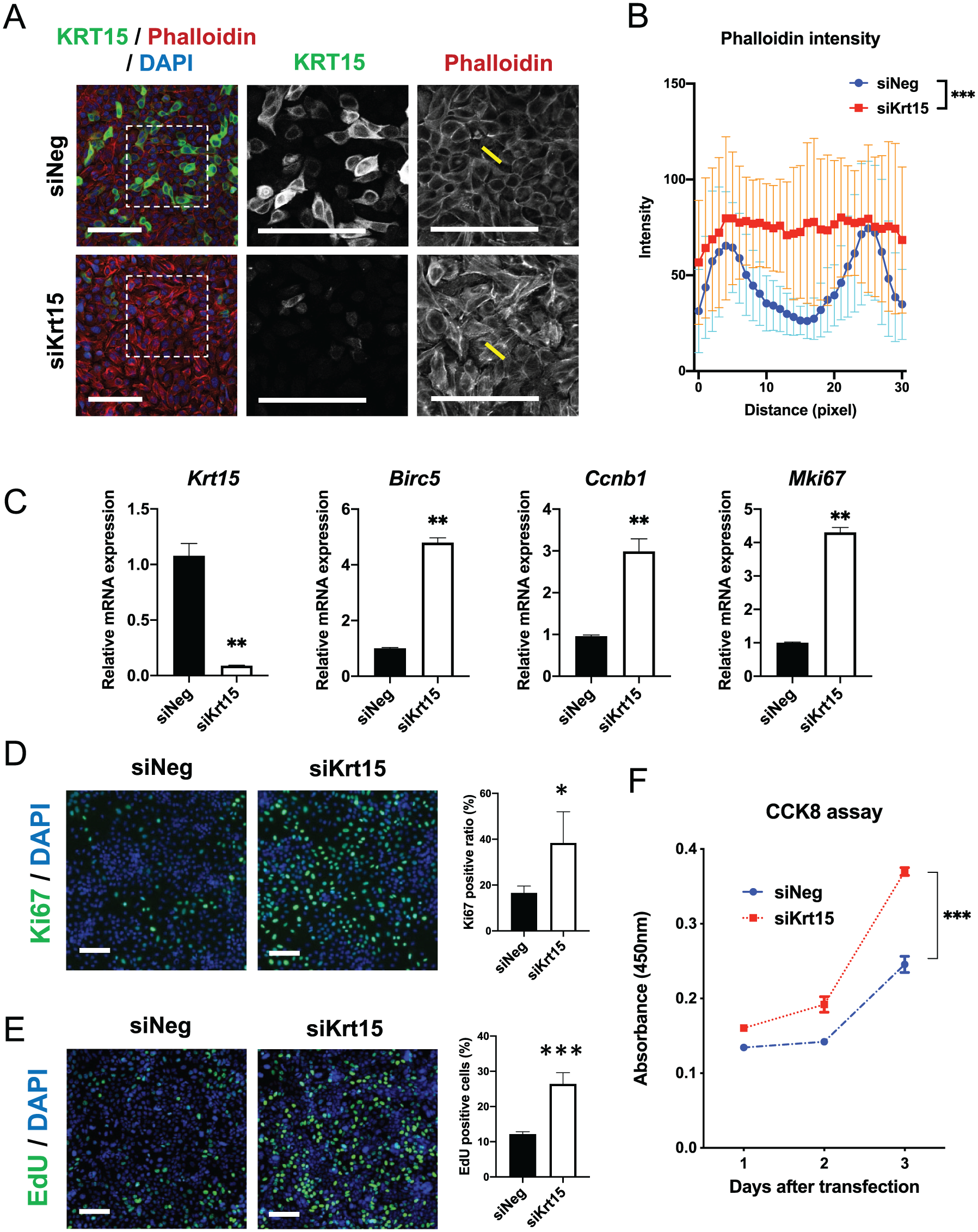
Depletion of Krt15 promotes the proliferation of dental epithelial cells. (
Depletion of Krt15 Induces Phosphorylation of p38 MAP Kinase and Upregulates Cell Proliferation
We further examined the downstream signaling pathways of KRT15 using Western blotting (Fig. 5A). CLDE cells transfected with siNeg- or siKrt15-transfected CLDE cells were stimulated with serum after 1 h of serum starvation. The phosphorylation activities of Akt, ERK1/2, and SAPK/JNK were not significantly altered by the depletion of Krt15, whereas it significantly upregulated the phosphorylation of p38 MAP kinase (MAPK) (Fig. 5A, B). Notably, the induction of phospho-p38 was observed at 48 h of siKrt15 transfection (Appendix Fig. 10). These results suggested that KRT15 may inhibit p38 MAPK activation. We further examined the effects of the p38 MAPK inhibitor, SB203580, on cytoskeleton formation, gene expression, and cell proliferation (Fig. 5C, D). We confirmed that 10 µM of SB203580 sufficiently inhibited the phosphorylation of p38 MAPK (Appendix Fig. 11). As a result, treatment of SB203580 upregulated the expression of Krt15 and inversely suppressed Mki67 (Fig. 5B). Importantly, the alteration of cytoskeleton formation observed by depletion of Krt15 was partially rescued by treatment of SB203580 in siKrt15-transfected CLDE cells (Appendix Fig. 12). Furthermore, p38 inhibition suppressed cell proliferation activity in CLDE cells (Fig. 5C). These results indicated that KRT15 inhibited the p38 MAPK pathway, which is important for the proliferation of dental epithelial cells (Fig. 5D).
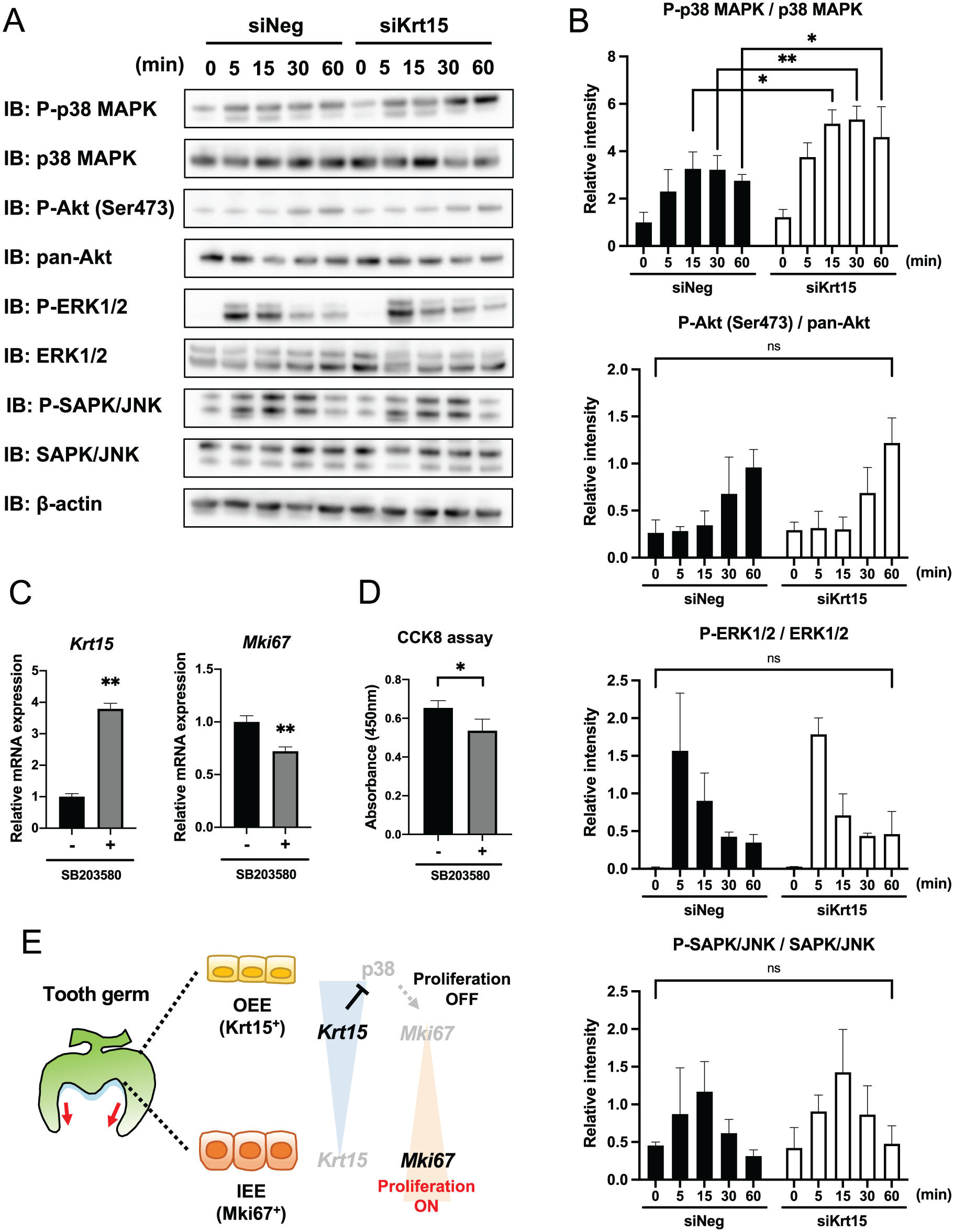
Depletion of Krt15 induces phosphorylation of the p38 MAPK pathway. (
Discussion
The cytokeratin family consists of more than 50 genes in humans and shows specific distributions depending on the type of epithelial cell. In dental epithelial cells, keratin 5 (Krt5) and keratin 14 (Krt14) are known as markers of the dental epithelium, especially highly expressed in ameloblasts, and play a chaperone role for nascent amel polypeptides during secretion of amel (Ravindranath et al. 2001; Ravindranath et al. 2003). We recently performed a single-cell RNA-sequence analysis of developing mouse molars to identify novel markers of SI cells and found that keratin 17 (Krt17) is specifically expressed in SI and SR cells and regulates SI and SR cell differentiation (Inada et al. 2024). Krt15 has recently been reported as a specific marker gene for OEE, as identified by scRNA-seq (Sharir et al. 2019; Chiba, Saito, et al. 2020; Krivanek et al. 2020; Chiba et al. 2022) and we revealed the importance of Krt15 on tooth morphogenesis. Importantly, KRT15 expression expanded to SR cells later than the P3 molar (Fig. 1F). This may reflect the expression changes of Krt15 in RT-qPCR of the tooth germ; Krt15 was increased from P1 to P7 molars (Fig. 1I). The change in Krt15 may indicate that OEE and SR cells have similar gene expression profiles after the secretory stage of the tooth germ. However, knockdown of Krt15 in CLDE cells did not affect ameloblast differentiation markers nor SI and SR marker genes (Appendix Fig. 8F), which was different from Krt17, crucial for Notch1 and Notch2 expression (Inada et al. 2024). Also, knockdown of Krt17 in the dental epithelial cell line does not affect cell proliferation activity (Inada et al. 2024), suggesting that the phenotypes observed in Krt15-depleted CLDE cells may be distinct from the role of Krt17. Thus, cytokeratin family members mark each type of dental epithelial cell and play crucial roles in tooth development.
Several cytokeratin family members play important roles in cell proliferation of epidermis. Importantly, KRT10, KRT16, and KRT19 have been reported to regulate the cell cycle through p-Rb dependence (Paramio et al. 1999; Sharma et al. 2019). Among them, KRT10 inhibits cell-cycle progression through inactivation of Akt and PKCζ (Santos et al. 2002). In this study, we found that knockdown of Krt15 in CLDE cells increased phosphorylation of Rb protein (Appendix Fig. 9). This result suggests that the modulation of cell proliferation by Krt15 is p-Rb dependent, and depletion of Krt15 promoted the cell cycle from the G1 phase to the S phase, which may initiate the transcription of cell proliferation–related genes to activate the cell cycle. During organogenesis, the mitogen-activated protein kinase pathway regulates epithelial cell proliferation and differentiation. Previous reports have suggested that p38 is essential for tooth morphogenesis and odontoblast and epidermal cell proliferation (Sharma et al. 2003; Greenblatt et al. 2015; Chen et al. 2018). In this study, we found that the depletion of Krt15 enhanced the phosphorylation of p38. In contrast, the inhibition of p38 phosphorylation by SB203580 upregulated the transcriptional activity of Krt15 in CLDE cells (Fig. 5). This indicates that p38 MAPK exerts negative feedback on Krt15 to regulate cell proliferation. The involvement of p38 in keratin filament reorganization has been demonstrated in other studies. Various types of stress and cellular stimuli induce the phosphorylation of p38 and cause recruitment of p-p38 to keratin filaments. Activated p38 regulates keratin reorganization during mitosis (Wöll et al. 2007). These findings suggest that Krt15 and p38 negatively regulate each other to modulate the proliferation of dental epithelial cells.
During tooth development, OEE and IEE play important roles in morphogenesis by regulating the proliferation and migration of the dental epithelium. With regard to the Hippo signaling pathway in tooth development, the nuclear localization of the active yes-associated protein (YAP) in IEE cells promotes proliferation and inhibits precocious differentiation and apoptosis (Hu et al. 2017). In contrast to IEE cells, OEE cells exhibited inactivated YAP localization in the cytoplasm and were resistant to YAP-driven proliferation. Thus, OEE and IEE may have different regulatory mechanisms of cellular proliferation. Interestingly, p38 MAPK regulates the TEAD transcription factor family, which mediates transcriptional activity of the Hippo pathway (Lin et al. 2017). Although OEE proliferation seems to be dependent on the nuclear localization of YAP, the relationship between p38 MAPK and the Hippo signaling pathway should be investigated. Furthermore, the depletion of Krt15 caused abnormal cellular morphology in CLDE cells, which may have resulted from the altered expression of cytokeratin members (Fig. 4A and Appendix Fig. 8C, D). Although the abnormal localization of the cytoskeleton caused by knockdown of Krt15 was partially recovered by the addition of SB203580 (Appendix Fig. 12), the disruption of cytoskeleton-associated signaling may affect cell proliferation activity as the secondary effect of depletion of Krt15. The signaling pathway and the molecular mechanism regulated by Krt15 is still unclear compared with other cytokeratin families (Pan et al. 2013). Further analyses are required on the detailed mechanism of Krt15 on cell proliferation and rational pathways of Krt15-p38 MAPK signaling.
With regard to the unique characteristic of OEE, OEE cells are more densely clustered than IEE cells are and express cell adhesion molecules such as E-cadherin, which are absent in IEE cells (Li et al. 2012; He et al. 2019). This indicates the importance of OEE in tooth morphogenesis. In addition, the proliferative activity of OEE and IEE showed drastic changes depending on the developmental stage. At the cap stage, all dental epithelial cells showed high proliferative activity, whereas once they entered the bell stage, the expression of proliferation markers was restricted in the IEE. After crown morphogenesis, OEE and IEE form HERS and change their proliferation activities (Yokohama-Tamaki et al. 2006). The modulation of cell proliferation by OEE and IEE may play an essential role in the development of proper tooth morphology. Interestingly, Krt15 depletion caused ectopic proliferation of dental mesenchymal cells in organ culture (Fig. 3D). Since Krt15 is expressed only in the dental epithelium (Fig. 1), this could be the result of epithelial–mesenchymal interactions. Thus, OEE may interact with IEE and the mesenchyme, which may balance proliferation, which is essential for proper tooth germ morphogenesis.
In the present study, we used an ex vivo organ culture system of a molar with transient siRNA transfection. This system enabled the simple evaluation of gene function and morphological changes in the tooth germ. However, there are limitations to the transfection efficiency of siRNA into the deep portions of the organ, and this method cannot be used to evaluate the long-term effects of gene functions on organ development. In fact, our results showed partial suppression of KRT15 at the protein level (Appendix Fig. 7A, B). Therefore, the conclusive effects of Krt15 on tooth morphogenesis, such as root and incisor formation and the effect of complete deletion of Krt15, should be examined in in vitro and in vivo experiments using the Krt15-KO cell line or Krt15-KO mice model in the future.
Our findings suggest a novel function of OEE in tooth morphogenesis and shed light on a new mechanism underlying tooth development. In the field of tooth regeneration research, artificial regeneration of the tooth germ from Induced pluripotent stem (iPS)–derived ameloblasts and odontoblasts has been successful. However, it is necessary to develop a method to regulate the morphology of regenerated tooth germs. Our current findings may shed light on the potential use of Krt15 as a marker gene of OEE cells to evaluate the differentiation from iPS cells and may contribute to developing a regulatory system for tooth morphogenesis using Krt15 expression.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345251368346 – Supplemental material for Keratin 15 Regulates Cell Proliferation in Outer Enamel Epithelium
Supplemental material, sj-docx-1-jdr-10.1177_00220345251368346 for Keratin 15 Regulates Cell Proliferation in Outer Enamel Epithelium by Y. Chiba, T. Tian, K. Yoshizaki, X. Wang, A. Yamada and S. Fukumoto in Journal of Dental Research
Footnotes
Acknowledgements
Author Contributions
Y. Chiba, contributed to conception, design, data acquisition, analysis, or interpretation, drafted the manuscript; T. Tian, X. Wang, contributed to data acquisition, analysis, or interpretation, critically revised the manuscript; K. Yoshizaki, contributed to conception, design, data analysis, critically revised the manuscript; A. Yamada, contributed to data analysis, critically revised the manuscript; S. Fukumoto, contributed to conception and design, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by a Grant-in-Aid from the Japan Society for the Promotion of Science (JSPS) KAKENHI (JP22H00488 to S.F., JP20K18747 to Y.C., and 23K21498 to K.Y.).
Data Availability
The datasets analyzed in this study can be found in NCBI GEO: GSE146855 and GSE207156. All other data are included in this manuscript.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
