Abstract
Peri-implantitis is an increasingly prevalent chronic inflammatory disease of the peri-implant tissue. A key etiologic factor for peri-implantitis is the submucosal biofilm, which may further drive clinical severity and accelerate disease progression. The present cross-sectional study aimed to characterize the compositional (full-length 16S rRNA gene amplicon sequencing) and functional patterns (metatranscriptomics) in the microbiome as an indicator of peri-implantitis severity. For this purpose, submucosal biofilm samples were collected from 49 peri-implantitis–diagnosed implants in 34 patients. Notable microbial signatures were associated with increased probing depth (PD), a measure for disease severity. Multivariate linear regression analysis, adjusted for patient variability, showed that the genera
Keywords
Introduction
Peri-implantitis is a biofilm-associated pathologic condition affecting the tissue around dental implants. It is characterized by inflammation of the peri-implant mucosa and progressive loss of supporting bone (Berglundh et al. 2018). Depending on the extent of inflammation and tissue damage, peri-implantitis can present at varying levels of severity, which correlate with clinical indicators such as probing depth (PD), bleeding on probing, and/or suppuration (Schwarz et al. 2018).
A key etiologic factor for peri-implantitis is submucosal microbial dysbiosis (Daubert and Weinstein 2019; Belibasakis and Manoil 2021). However, until today, the specific microbial taxa and their functional profiles in dysbiosis that correlate with disease severity have not been fully elucidated. Yet, a clear ecologic and mechanistic understanding of the relationship between dysbiotic changes in the submucosal microbiome and peri-implantitis severity is crucial for the further development of microbiome-based diagnostics. Moreover, understanding this relationship may support effective disease stratification, enabling development of targeted preventive and treatment strategies, particularly for severe forms of peri-implantitis that often present with unpredictable clinical outcomes (Lagervall and Jansson 2013).
In recent years, advances in next-generation sequencing technologies have improved our understanding of the submucosal microbiome around dental implants (Kumar et al. 2012; Zheng et al. 2015; Sanz-Martin et al. 2017; Ghensi et al. 2020; Polymeri et al. 2021). Yet, most of these omics-based studies have focused on differences in the microbiome composition between health and peri-implantitis or between periodontitis and peri-implantitis, not on peri-implantitis severity. For periodontitis, subgingival microbiome profiles have been described for various disease stages (Tonetti et al. 2018; Lafaurie et al. 2022; Iniesta et al. 2023), but comparable evidence for peri-implantitis is either limited or indirectly derived (Maruyama et al. 2014; Kröger et al. 2018; Polymeri et al. 2021; Yu et al. 2024). Although studies have provided initial clues about associations between submucosal microbial dysbiosis and the severity of peri-implant disease, a comprehensive functional characterization is still lacking. However, a functional characterization of the submucosal microbiome is essential for elucidating the underlying molecular mechanisms and is therefore more informative than the composition alone, given the polymicrobial complex nature of peri-implantitis biofilms (Komatsu et al. 2020).
Hence, the aim of the present study was to characterize the peri-implantitis–associated submucosal microbiome with high-throughput full-length 16S rRNA gene amplicon sequencing (full-16S) and metatranscriptomics (RNAseq) to identify the microbial taxa and functional bacterial and enzymatic activities associated with different levels of PD, as an indicator of disease severity. Furthermore, we aimed to develop an easy-to-interpret quantitative index integrating microbial taxa and functions to stratify peri-implantitis severity.
Material and Methods
Cohort Characteristics and Clinical Examination
The present cross-sectional study is a part of the interdisciplinary consortium Safety Integrated and Infection Reactive Implants, which aims to characterize the etiopathogenesis of peri-implant diseases and develop early detection and prevention strategies. The study was conducted at the Department of Prosthetic Dentistry and Biomedical Materials Science, Hannover Medical School, in accordance with the STROBE guidelines (Strengthening the Reporting of Observational Studies in Epidemiology). The study protocol was approved by the ethics committee of Hannover Medical School, Germany (No. 9477).
The Appendix provides a detailed description of patient selection criteria (Appendix Table 1), clinical examination, biofilm sample collection, DNA-RNA co-isolation, sequencing, and bioinformatic and statistical methods. The sequencing data for this study have been deposited in the NCBI SRA as BioProject PRJNA1192962.
Results
The study population comprised 34 patients (17 male, 17 female) with a mean ± SD age of 73 ± 9 y and 49 implants diagnosed with peri-implantitis. The population- and site-specific characteristics of the study cohort are presented in Appendix Table 2. Implants were in function for 9.8 ± 6 y. The mean PD of the implants was 7.1 ± 2.3 mm (range, 5 to 11 mm). All implants showed bleeding on probing, and suppuration was observed in 41%.
Diversity and Variability of Full-16S Microbiome Profiles in Peri-implantitis Samples
Full-16S yielded 715,134 sequences (mean, 14,594 sequences per sample) after filtering and preprocessing of reads. Following removal of typical contaminants (Appendix Methods), we identified 599 species-level taxa belonging to 13 phyla, 20 classes, 33 orders, 53 families, and 93 genera. High taxonomic diversity was captured with the included number of samples (Appendix Fig. 1), with no change in alpha diversity across PD levels. Figure 1A illustrates that Bacteroidia was the most abundant class across all PD levels, followed by Bacilli, Negativicutes, and Fusobacteriia. Figure 1B displays the distribution of genera within these classes. The distinct variation in microbial composition at the species level was seen along the MDS1 axis as visualized by a non-metric multi-dimensional scaling (nMDS) ordination plot (Fig. 1C). We tested different PD thresholds and selected the cutoff based on permutational multivariate analysis of variance pseudo-
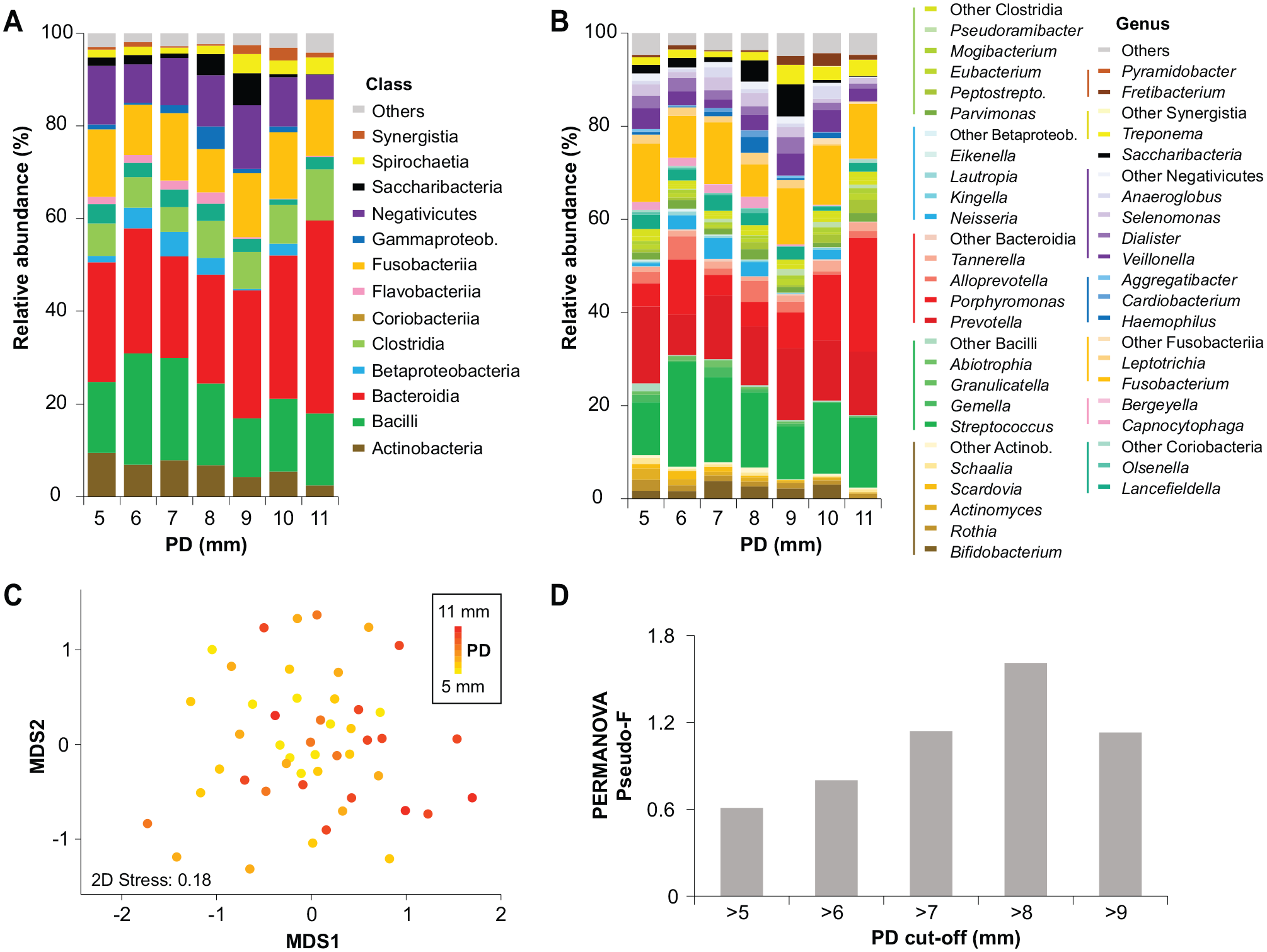
Microbial community distribution and variability across peri-implantitis samples. (
Correlation of Full-16S Microbial Taxa with Peri-implantitis Severity
Microbiome differences between groups 1 and 2 were analyzed by MaAsLin2 and DESeq2. MaAsLin2 showed that classes Gammaproteobacteria, Flavobacteriia, Mollicutes, Actinobacteria, and SR1 were significantly associated with group 1, while Deltaproteobacteria and Synergistia were significantly associated with group 2 (Fig. 2A). At the genus level, several genera were significantly associated with group 1 (Fig. 2B). Most significant of those were
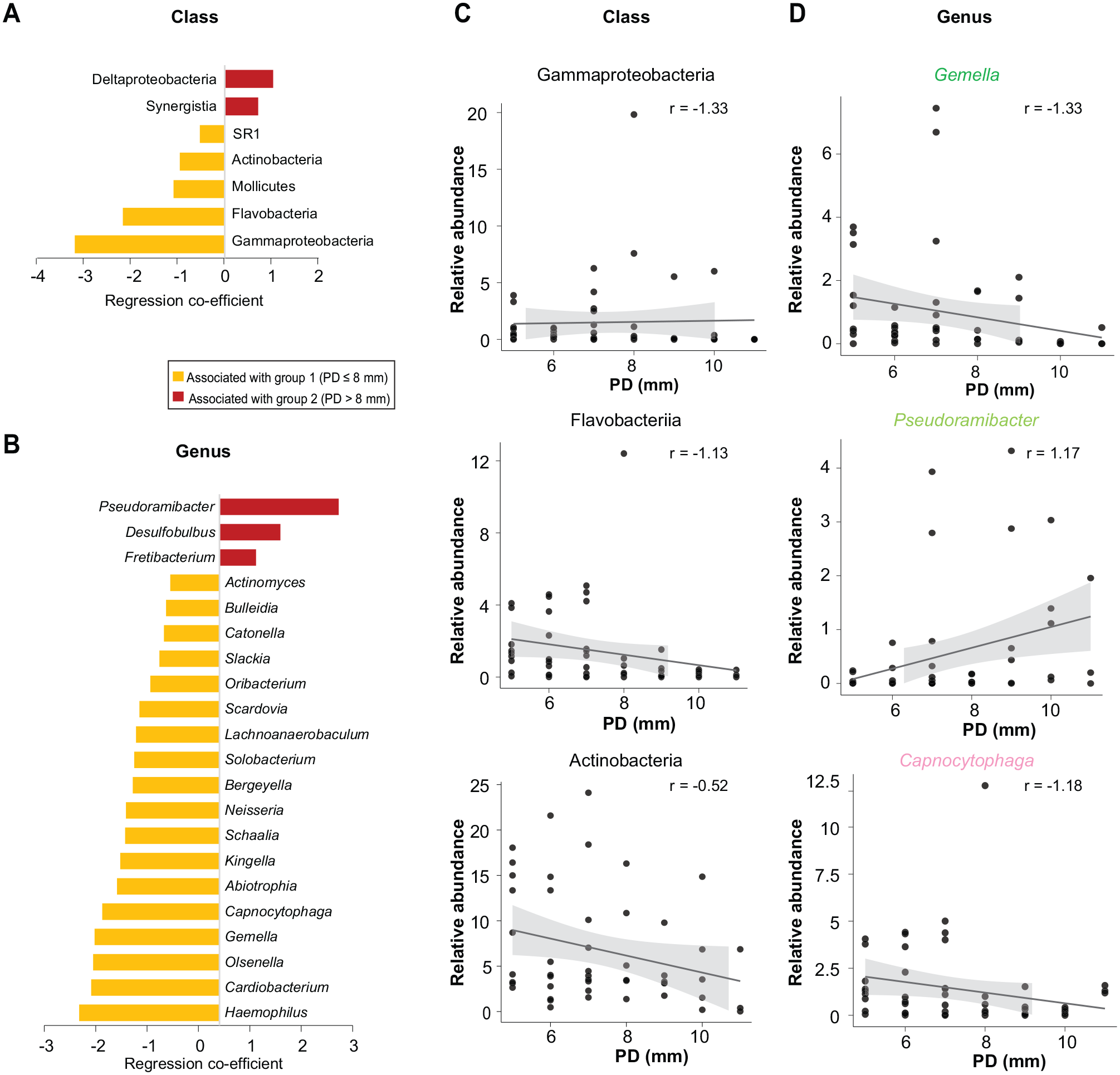
Correlation of microbial taxa with PD groups and distinct PD levels. (
Next, we analyzed PD as a continuous variable, as displayed in Figure 2C and D, and identified microbial taxa significantly associated (adjusted
The comparison of different approaches (PD continuous vs PD groups) and methods (DESeq2 and MaAsLin2) showed the highest correlation (
Prediction of Functional Potential of Biofilms in Peri-implantitis
Prediction of functional potential within biofilms based on the 16S rRNA marker gene revealed 261 MetaCyc pathways with relative abundances >0.1% in at least 1 peri-implantitis sample. The nMDS ordination plot illustrates a substantial overlap of PDs across MDS1, with a slight tendency of clustering among higher PD samples (Fig. 3A). A heat map displays the relative abundances of significant predicted MetaCyc functional pathways (MaAsLin2 default adjusted
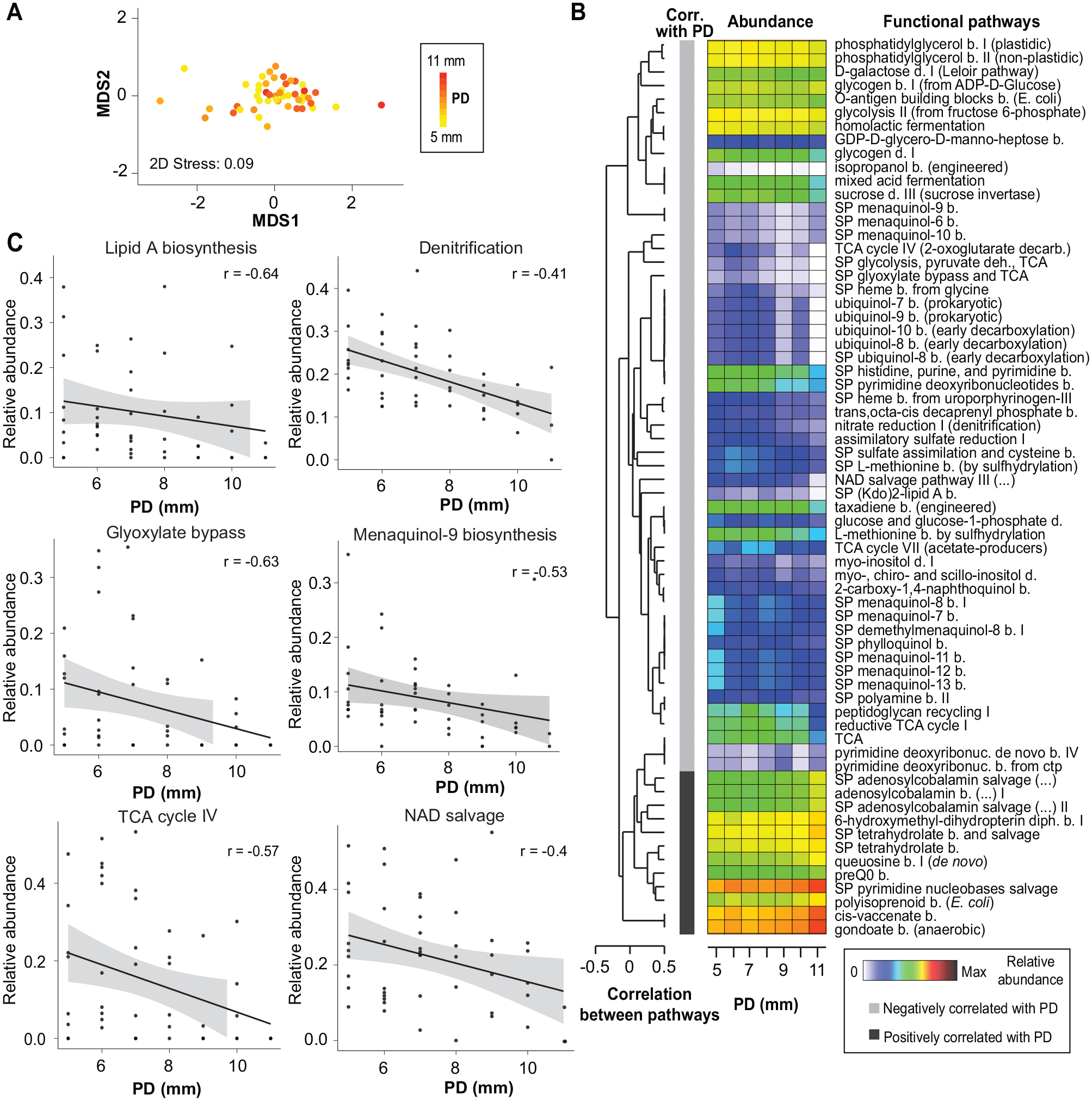
Correlation of predicted MetaCyc functional pathways at different levels of PD. (
Metatranscriptomic Functional Activities at Different Levels of Peri-implantitis Severity
In contrast to PICRUSt2-based predicted enzyme functions, the RNAseq revealed a high number of true enzymatic activities that were significantly associated with PD (adjusted
Metatranscriptomic ECs Significantly Correlated with PD (Adjusted
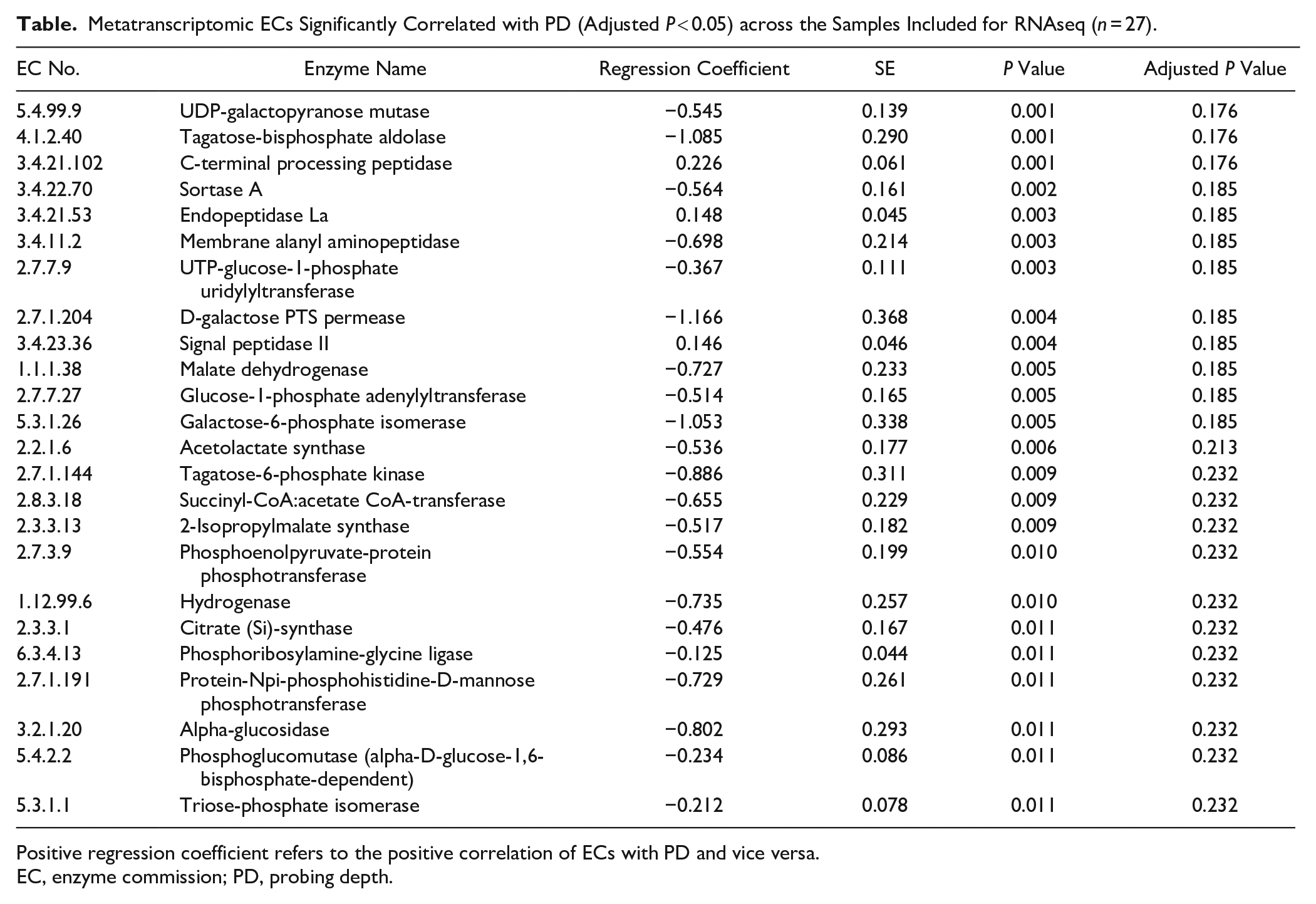
Positive regression coefficient refers to the positive correlation of ECs with PD and vice versa.
EC, enzyme commission; PD, probing depth.
Extended Microbial Dysbiosis Index for Peri-implantitis Severity
To quantify the microbial dysbiosis related to severity and to develop a clinically relevant index with a reduced number of features, we first calculated the microbial dysbiosis index (MDI) using 3 significant PD-associated genus-level taxa:
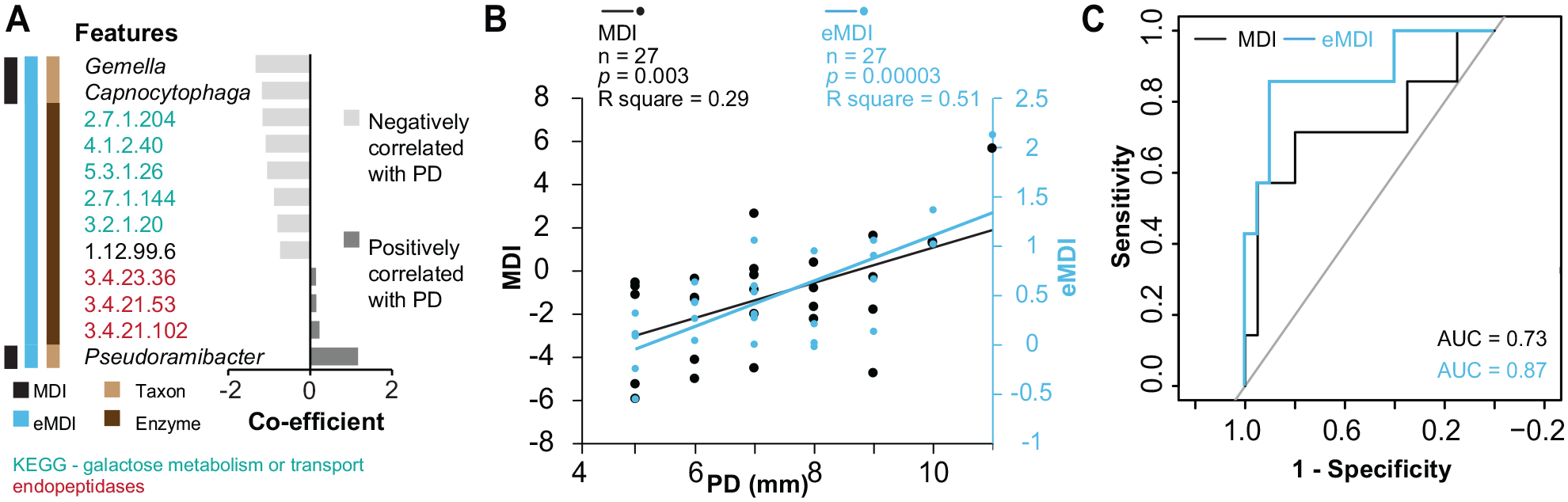
Correlation of MDI and eMDI with different PD levels. (
Discussion
The present study comprised 49 submucosal taxonomic and functional profiles of implants diagnosed with peri-implantitis. We observed clear differences in the microbiome with changes in PD and identified significant associations of microbial taxa, potential functional pathways, and transcriptome-level specific enzymatic activities with peri-implantitis severity. A microbiome-based index (eMDI) was introduced for the first time, which is specific to peri-implantitis severity and integrates taxa and their enzymatic activities.
Our study integrated 2 high-throughput approaches, full-16S and metatranscriptomics, to characterize the severity of peri-implantitis. Sequencing the full-length 16S rRNA gene (~1,500 base pairs) increased reliability and precision of taxonomic assignments (e.g., identification of species for a greater proportion of sequences) as compared with short-read sequencing platforms (Johnson et al. 2019). Furthermore, it is recognized that community composition alone does not adequately reflect the intricate biological processes occurring within peri-implant biofilms representing a range of molecular elements (Solbiati and Frias-Lopez 2018; Belibasakis and Manoil 2021). We employed metatranscriptomics for 27 peri-implantitis samples to identify RNA-level traits revealing the true microbial activities associated with severity of peri-implantitis, thus addressing the need for functional biomarker candidates (Belibasakis and Manoil 2021) while contributing to a deeper understanding of oral dysbiosis (Solbiati and Frias-Lopez 2018). To date, only 17 peri-implantitis metatranscriptomes have been published, originating from 2 studies (Shiba et al. 2016; Ganesan et al. 2022), both of which compared peri-implantitis with either periodontitis or health, without specifically addressing disease severity. Our data provide a foundation for scaling up these comparisons, which is critically needed (Belibasakis and Manoil 2021). Metatranscriptomics has uncovered host-microbiome interactions (Ganesan et al. 2022), presenting an intriguing aspect for further studies across varying PDs.
Our full-16S approach revealed that although peri-implantitis biofilms represent highly complex and diverse microbial communities, core and severity-associated taxa can be identified.
Another highlight of our study was the functional analyses employing 16S-based predictions. Abundances of certain metabolic pathways and their contributing taxa were associated with PD. For instance, the nitrate reduction pathway and
Our study is the first to utilize metatranscriptomics to assess the expressed enzymatic activities (ECs) associated with peri-implantitis severity. Although full-16S–based functional prediction revealed broad, pathway-level significant associations with PD, it lacked the resolution of metatranscriptomics and did not capture a high number of specific enzymatic functions that were uniquely detected through metatranscriptomics. The strongest negative correlations were observed for ECs that belong to galactose metabolism, while 2 serine endopeptidases and an aspartic endopeptidase showed positive correlation with deeper peri-implant pockets. Previous periodontal and peri-implant metatranscriptomic studies have positively linked the expression of various peptidases to disease, whereas multiple carbohydrate-related pathways, including galactose metabolism and phosphotransferase systems, are shown to have an inverse association (Duran-Pinedo and Frias-Lopez 2015; Szafranski et al. 2015; Yost 2015; Nowicki et al. 2018). Utilization of galactose (Takahashi 2015) present on the surfaces of host proteins appeared to be an important adaptation in periodontopathogens to the early peri-implantitis environment, while the production of proteolytic activities potentially capable of degrading host-derived glycoproteins and tissue components could serve as a reliable marker for disease severity (Eley and Cox 2003). Further analysis of peptidases showed that
In this context, the eMDI for peri-implantitis severity proposed by us showed high accuracy for severity-based peri-implantitis stratification and may be relevant for clinical evaluation of peri-implantitis progression. Previous efforts (Kröger et al. 2018; Shi et al. 2022; Feng et al. 2024) have successfully quantified the microbial dysbiosis in peri-implant diseases and underscored the association between the level of dysbiosis and PD. However, there still remains a need for an objective microbiome-based index that uses a smaller number of high-sensitivity and high-accuracy biomarkers, making it suitable for chair-side diagnostics. This gap could be addressed with the index presented in this study. The key advantage of eMDI lies in integrating taxonomic and taxon-independent enzymatic features as biomarkers. The panel of limited candidate markers—including 3 genera (
This study reveals important microbiome differences linked to peri-implantitis severity. However, limitations should be considered. The cross-sectional design limits causal inference and prognostic implications, underscoring the need for longitudinal validation. Moreover, a relatively small number of severe peri-implantitis samples (PD >8 mm,
In conclusion, this study is the first to integrate DNA-based taxonomic and RNA-based functional analyses of peri-implantitis microbiomes in relation to disease severity. Our findings demonstrated significant associations of microbiome composition and functional activities with peri-implantitis disease severity. Additionally, we proposed a microbiome-based index (eMDI) that strongly correlates with different disease severities. Our high-resolution characterization of the peri-implantitis microbiome is an important step toward disease stratification and the development of targeted therapeutic strategies based on the level of peri-implantitis severity.
Author Contributions
A.A. Joshi, contributed to data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; S.P. Szafrański, contributed to data acquisition, analysis, and interpretation, critically revised the manuscript; M. Steglich, I. Yang, W. Behrens, contributed to data analysis, critically revised the manuscript; P. Schaefer-Dreyer, contributed to data acquisition and analysis, critically revised the manuscript; J. Grischke, contributed to data acquisition, critically revised the manuscript; S. Häussler, M. Stiesch, contributed to data conception and design, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345251352809 – Supplemental material for The Submucosal Microbiome Correlates with Peri-implantitis Severity
Supplemental material, sj-docx-1-jdr-10.1177_00220345251352809 for The Submucosal Microbiome Correlates with Peri-implantitis Severity by A.A. Joshi, S.P. Szafrański, M. Steglich, I. Yang, W. Behrens, P. Schaefer-Dreyer, J. Grischke, S. Häussler and M. Stiesch in Journal of Dental Research
Supplemental Material
sj-docx-2-jdr-10.1177_00220345251352809 – Supplemental material for The Submucosal Microbiome Correlates with Peri-implantitis Severity
Supplemental material, sj-docx-2-jdr-10.1177_00220345251352809 for The Submucosal Microbiome Correlates with Peri-implantitis Severity by A.A. Joshi, S.P. Szafrański, M. Steglich, I. Yang, W. Behrens, P. Schaefer-Dreyer, J. Grischke, S. Häussler and M. Stiesch in Journal of Dental Research
Footnotes
Acknowledgements
We thank Dr. Andreas Winkel for the administrative support and Marly Dalton and Rainer Schreeb for technical assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Deutsche Forschungsgemeinschaft (German Research Foundation; SFB/TRR-298-SIIRI, project 426335750) and under Germany’s Excellence Strategy (EXC 2155, project 390874280).
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
