Abstract
The molecular mechanisms controlling the differentiation of stem cells into specialized cells and tissues are enormously complex. Deciphering the mechanisms behind this precisely regulated process is essential not only for understanding ontogenesis but also for interpretations of evolutionary dynamics. A deep understanding of differentiation processes in various cell types would open the way for safe and targeted tissue engineering. This was previously limited by technical constraints, but the recent development of multiomic approaches has overcome these barriers, providing unprecedented insight into the intricate workings of this molecular machinery. Based on single-cell RNA-seq analyses of continuously growing mouse teeth, 4 transcription factors (Etv4, Gsc, Sall1, and Nupr1) were selected. Their role during tooth development has not been previously evaluated. Here, we provide evidence about their function in the differentiation of odontoblasts—dental cells responsible for dentin production. Our results revealed that controlled overexpression of specific transcription factors directly in mouse-induced pluripotent stem cells is sufficient to guide their differentiation into cells with an odontoblast-like phenotype, both in vitro and in vivo. The differentiated cells exhibited upregulated expression of odontoblast-specific molecular markers (DMP1 and DSP) as well as production of collagenous and mineralized tissue. We demonstrate that a deep insight into fundamental developmental events can provide a powerful basis for innovative cell differentiation approaches. Taken together, this research might serve as a proof of concept for using large-omics data in the generation of specific, differentiated cell types by controlled expression of a particular transcription factor code. This may represent a turning point in both developmental biology and regenerative medicine fields.
Introduction
Odontoblasts are highly polarized cells that produce dentin. They originate from neural crest, which migrates during early embryonic development into the first pharyngeal arch. Here, at the prospective site of dental germ formation, a complex reciprocal interaction of the migrated ectomesenchyme with the odontogenic epithelium subsequently occurs. These interactions involve the activation of a number of transcription factors (TFs) that control and regulate this process (Thesleff 2003; Kim et al. 2017; Balic 2018). Odontoblasts not only produce the main hard dental matrix during tooth development but are also able to respond to tooth damage and contribute to the reparative processes that are essential for the longevity of dentition (Neves and Sharpe 2018). Thus, they are an important target for biologically based regenerative dental medicine (Balic 2018).
To investigate the process of gradual odontoblast differentiation from their mesenchymal progenitors, continuously growing teeth of rodents are traditionally used as a model system (Krivanek et al. 2020; Lavicky et al. 2022). Their continuous growth is secured by permanently active mesenchymal and epithelial stem cell niches in their apical region. The incisor constantly recapitulates all stages of odontoblast (and ameloblast) differentiation along its apical–distal axis. This enables investigation of the continuous differentiation of all dental cell lineages including transient cell states in a single organ at any time during the lifetime (Krivanek et al. 2017). Furthermore, it was shown that molecular mechanisms driving the formation of the tooth are highly conserved across species, including humans and mice (Tucker and Fraser 2014; Hulsey et al. 2020; Hermans et al. 2021).
Several recent omics-based technological advances accelerated research in developmental biology and opened an unexpected range of possibilities. In continuously growing teeth, comprehensive works including expression databases have been published. These have analyzed the sequential differentiation steps of tooth-forming cells at an unprecedently detailed molecular level (Sharir et al. 2019; Chiba et al. 2020; Krivanek et al. 2020). Importantly, recent works suggest how to apply these fundamental dental development–related large data to obtain tooth-forming cells. This is the basic prerequisite for bridging the fields of developmental biology and regenerative dentistry (Alghadeer et al. 2023; Hanson-Drury et al. 2023). Based on single-cell transcriptomics data, these studies proposed a strategy for manipulating different signaling pathways in multi-/pluripotent cells using specific proteins and small molecules to differentiate them into odontoblasts or ameloblasts in vitro. However, the potential of TFs to mediate this process remains largely unrecognized. Therefore, the aim of this study was to elucidate whether information about TFs’ activity during odontoblast development can be used to obtain a code of TFs capable of controlling the differentiation of odontoblasts directly from pluripotent cells.
Previous work done by Krivanek et al. (2020) had mapped the developmental trajectory of odontoblasts at a detailed transcriptional level. Analyzing these data, 4 transcription factors/transcriptional regulators (TFs) were selected: ETV4, GSC, SALL1, and NUPR1 (detailed information concerning functions of selected genes can be found in the Appendix). These were predicted to have a high potential for controlling the differentiation of dentin-forming cells—odontoblasts. While these TFs play important roles in a wide variety of developmental processes, only one of them (SALL1) was recently found to be involved in tooth development and odontoblast differentiation (Lin et al. 2021). The role of the remaining TFs during odontoblast differentiation, however, remains unclear.
This work demonstrates that the controlled overexpression of the selected development-related TFs guides the differentiation of mouse induced pluripotent stem cells into lineages capable of expressing late odontoblast markers and possessing the capacity of generating mineralized matrix.
Materials and Methods
Details concerning the methodology can be found in the Appendix Materials and Methods. The article complies with the ARRIVE 2.0 checklist.
Results
Selection and Validation of Odontoblast-Specific TF Code
The selected odontoblast-specific TFs were selected based on a secondary analysis of previously published scRNA-seq data (Krivanek et al. 2020). First, the (pre)odontoblast cluster containing odontoblasts and their progenitors was defined within the mouse incisor dataset using the expression of known marker genes and their consensual expression patterns (Fig. 1A). Subsequently, based on the consensual expression pattern similarity search, an unbiased analysis of potential target genes was performed (Appendix Figs. 3, 4). Subsequently, based on their expression specificity (Fig. 1B) and regulon activity analysis (Fig. 1C) genes Etv4, Gsc, Sall1, and Nupr1 have been selected to be used. Moreover, we have also used the current knowledge concerning their function as well as the bioinformatic analysis of their regulon members to support this selection (Appendix Figs. 5–9).
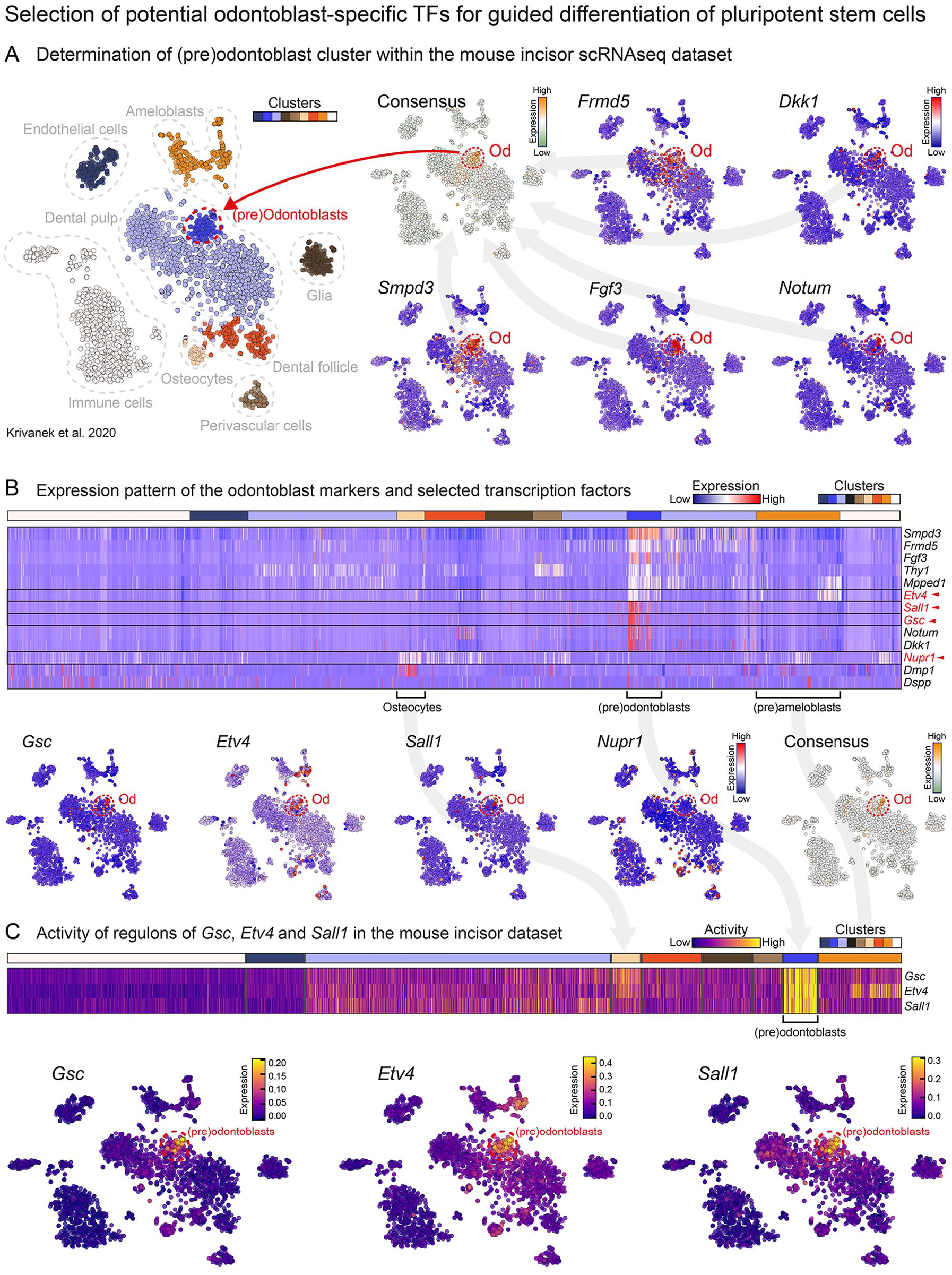
Analysis of the odontoblast differentiation trajectory—background for the selection of the utilized transcription factors. t-SNE visualization of the whole mouse incisor scRNA dataset with simplified cluster labels. Using the consensus expression pattern of previously established (Krivanek et al. 2020) odontoblast markers (Frmd5, Dkk1, Smpd3, Fgf3, and Notum) the (pre)odontoblast subpopulation was defined within the dataset (
The identity of four subgroups of cells within the odontoblast differentiation trajectory was defined (Fig. 2A). The expression pattern of the four selected TFs was incorporated into an expression heatmap of the used marker genes to elucidate during which phases of odontoblast differentiation they are being expressed (Fig. 2B). To validate our scRNA-seq–based gene expression pattern predictions, the expression of the selected TFs was mapped onto the odontoblast trajectory, and in situ gene expression analysis of the TFs in continuously growing incisor (Fig. 2C) and during molar development (Appendix Fig. 10) was performed. The developmentally earliest expressed TF was Etv4, located in the near-cervical loop area of the incisor. This area has been previously identified as a progenitor region (Krivanek et al. 2020, 2023). In addition, the expression of Etv4 was also observed in pre-odontoblasts and adjacent ameloblastic epithelium (Fig. 2C). The second TF, Gsc, was highly expressed in the preodontoblastic region on the labial aspect of the incisor and continued up to mature odontoblasts (Fig. 2C). The expression of the third TF, Sall1, was initiated in the preodontoblastic region in both the labial and lingual sides of the incisor. However, contrary to the Etv4 and Gsc, the Sall1 expression in mature odontoblasts continued further in the odontoblast layer on the labial side (Fig. 2C). The last transcriptional regulator Nupr1 was expressed exclusively in mature odontoblasts on both the labial and lingual sides (Fig. 2C). While selected genes start at different stages of odontoblast differentiation, a natural expression overlap of these genes exists in mature odontoblasts expressing Dspp and Dmp1 (Fig. 2B, C). Moreover, in parallel to the incisor, we mapped the expression pattern of the selected TFs also during molar development. It reflected the expression pattern observed in continuously growing incisor (Appendix Fig. 10).
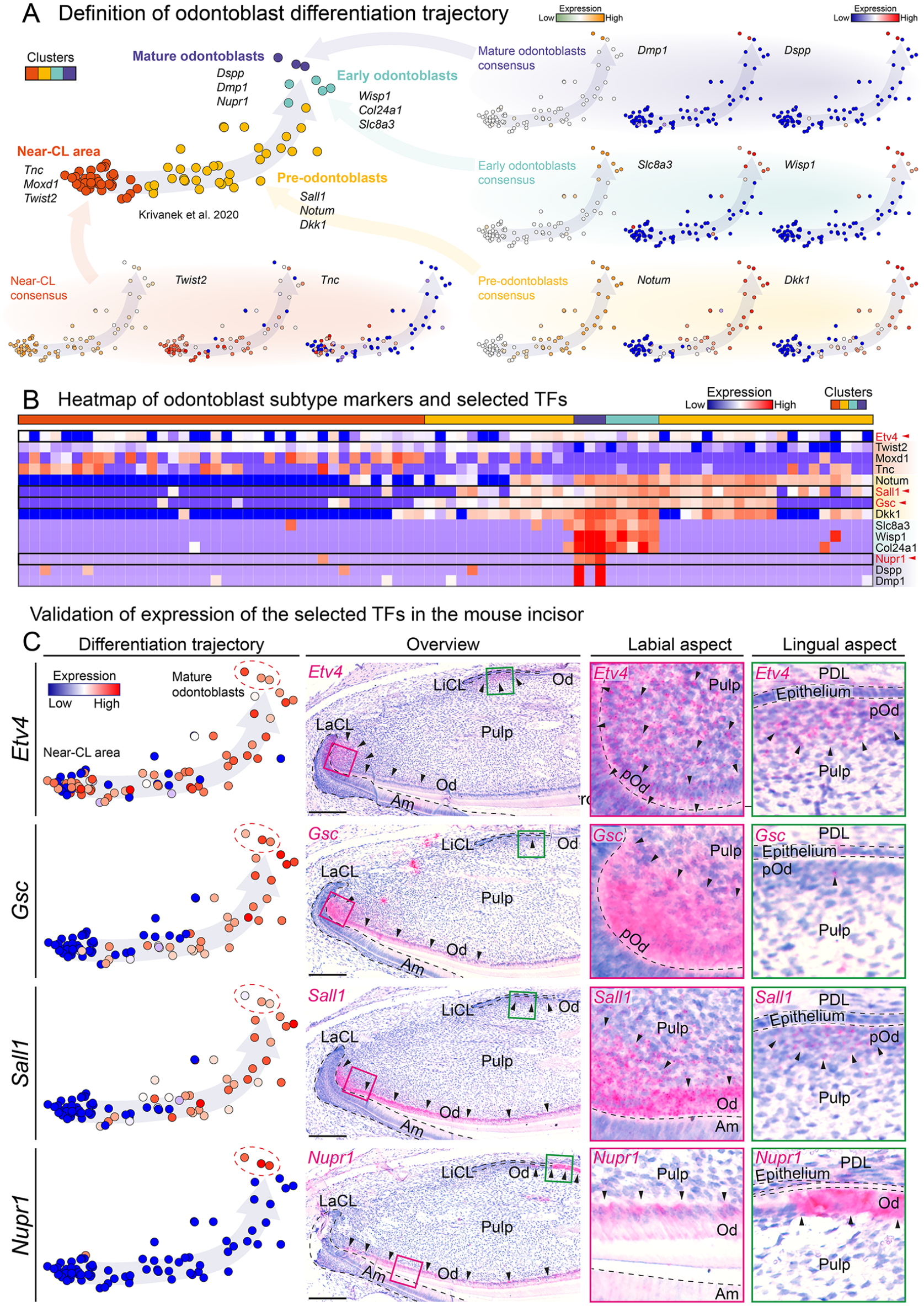
Visualization of the expression of the four selected genes in the odontoblast differentiation trajectory and its in situ validation in the continuously growing mouse incisor. The definition of the odontoblast differentiation trajectory including its subgroups using the consensus patterns of known marker genes (
Overexpression of Etv4, Gsc, Sall1, and Nupr1 in Induced Pluripotent Stem Cells Results in the Upregulation of Odontoblast Molecular Markers and Induction of Mineralization In Vitro
To understand the role of selected TFs during odontoblast differentiation, we performed an odontogenic differentiation assay on freshly isolated dental pulp stem cells (DPSCs) (Fig. 3A). The expression of Gsc was stable during the differentiation. Etv4’s expression displayed a trend with the highest mean at D7 and significantly upregulated expression at D14. The expression of Sall1 mirrored that of Runx2 with low expression at D3 and D7 and with stable significant upregulation at D14 and D21. Finally, Nupr1 displayed a steadily growing expression over the whole culture period.
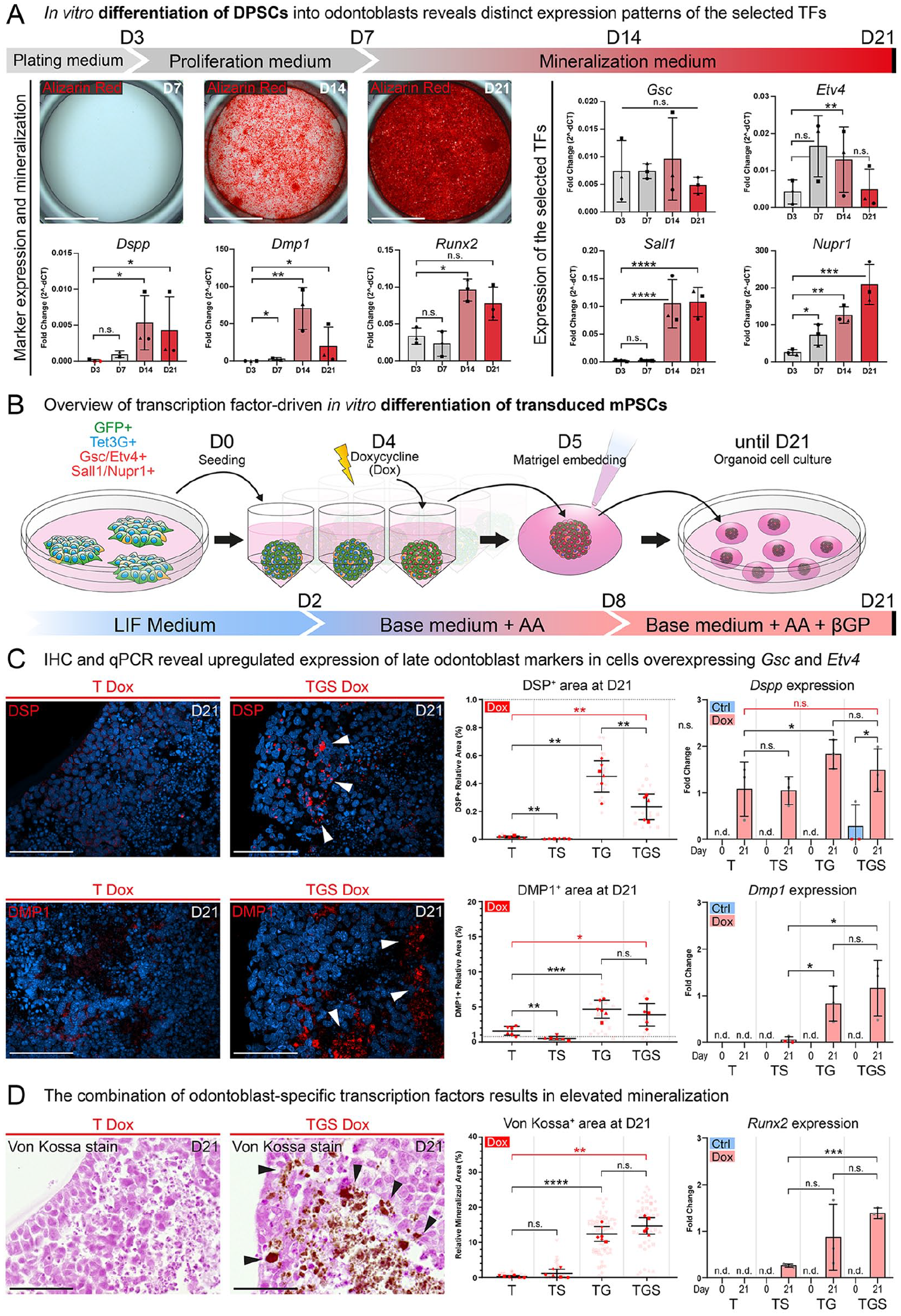
The expression of Etv4, Gsc, Sall1, and Nupr1 displays distinct expression patterns during in vitro differentiation of DPSCs and their overexpression in miPSCs leads to the upregulation of odontoblast molecular markers and initiation of mineralization in vitro. Results of in vitro odontogenic differentiation of DPSCs (
The mouse induced pluripotent stem cell line derived from the DSPP-Cerulean/DMP1-Cherry reporter strain was transduced with lentiviral vectors to enable the inducible overexpression of the selected TFs (Appendix Fig. 11). To test the effects of the overexpression of the selected TFs, in vitro differentiation experiments were performed (Fig. 3B). The growth dynamics, proliferation, and apoptosis rates were analyzed in cultured spheroids during the differentiation. The analysis showed their growth and formation of necrotic centers (Appendix Fig. 13). The expression of DSP and DMP1 was significantly upregulated in spheroids generated from cell lines containing the Gsc/Etv4 vector (Fig. 3C). Both DSP and DMP1 formed granular aggregates inside the spheroids (Fig. 3C). Co-staining illustrating their overlap is in Appendix Fig. 15. Observed trends were quantified as a positive relative area for the given protein. Interestingly, due to the background activity of the TET3G transactivator (Appendix Fig. 11), spheroids generated from TG and TGS cell lines displayed a significant upregulation of marker proteins even without added doxycycline (this known phenomenon is further discussed in Appendix Figs. 11,12). For this reason, the T-Dox condition was used as a negative control instead of conditions without added doxycycline.
To further investigate the expression of Dspp and Dmp1, reverse transcriptase quantitative polymerase chain reaction (RT-qPCR) was performed (Fig. 3C). The analysis of Dmp1 expression mirrored the results of the immunohistochemistry (IHC) analysis (Fig. 3C). While a significant upregulation was observed in the TG and TGS conditions, low or no expression was detected in the remaining conditions. The expression of Dspp was less robust, mirroring the protein expression trends in only the TG-Dox condition.
To demonstrate the mineralization capability of the engineered cells, von Kossa staining was performed (Fig. 3D). This approach highlighted the presence of mineralized areas in TS, TG, and TGS conditions. Interestingly, mineralized particles exhibited similar size and distribution to the DSP and DMP1-positive particles identified by IHC (Fig. 3C, D). Quantification of the von Kossa stain uncovered the same trends as seen in the IHC analysis of DSP and DMP1. Both TG-Dox and TGS-Dox conditions were significantly more mineralized than the T-Dox condition. To complement the mineralization analysis, an RT-qPCR analysis of Runx2 was performed (Fig. 3C). Its expression mirrored the mineralization pattern, in which Runx2 was upregulated in TS, TG, and TGS conditions, while the control condition displayed no expression of Runx2 (further discussed in Appendix Fig. 14). Taken together, the features of the odontoblast-like phenotype observed after the TFs’ overexpression-driven differentiation of miPSCs mirrored those observed in the odontogenic differentiation of DPSCs.
Overexpression of Selected TFs In Vivo Results in an Odontoblast-like Phenotype
To achieve more natural growth of engineered cells and to show their full differentiation potential, doxycycline-primed miPSC spheroids were implanted under the kidney capsule of immunodeficient mice 5 d after seeding (Fig. 4A). Based on the previous in vitro experiments and to minimize the number of experimental mice (3R), 2 borderline experimental groups were used: the T-Dox as a control group and the TGS-Dox as an experimental condition. IHC analysis of the TGS-Dox condition uncovered the presence of distinct structures expressing both DSP and DMP1 (Fig. 4B, C) Their co-staining showed their stratification and overlap in the structures (Appendix Fig. 15). In contrast, the T-Dox condition displayed no such organized, epithelium-like features. Quantifications of the DSP+ and DMP1+ area have shown significantly upregulated DSP and DMP1 expression in the TGS-Dox condition. The quantification of the IHC has revealed a certain level of cell population heterogeneity within implants (Fig. 4B; Appendix Fig. 19). To validate the observed trends on the transcriptional level, the expression of the fluorescent reporters DSPP-Cerulean/DMP1-Cherry was investigated. The expression pattern of the Dspp and Dmp1 reporters was generally comparable with the protein expression observed via IHC detection (Fig. 4C). Similar to the in vitro experiments, the expression of Dspp quantified by RT-qPCR was not significantly upregulated in the TGS-Dox condition. The expression of Dmp1 was more robust, displaying significant upregulation in the TGS-Dox condition (Fig. 4C).
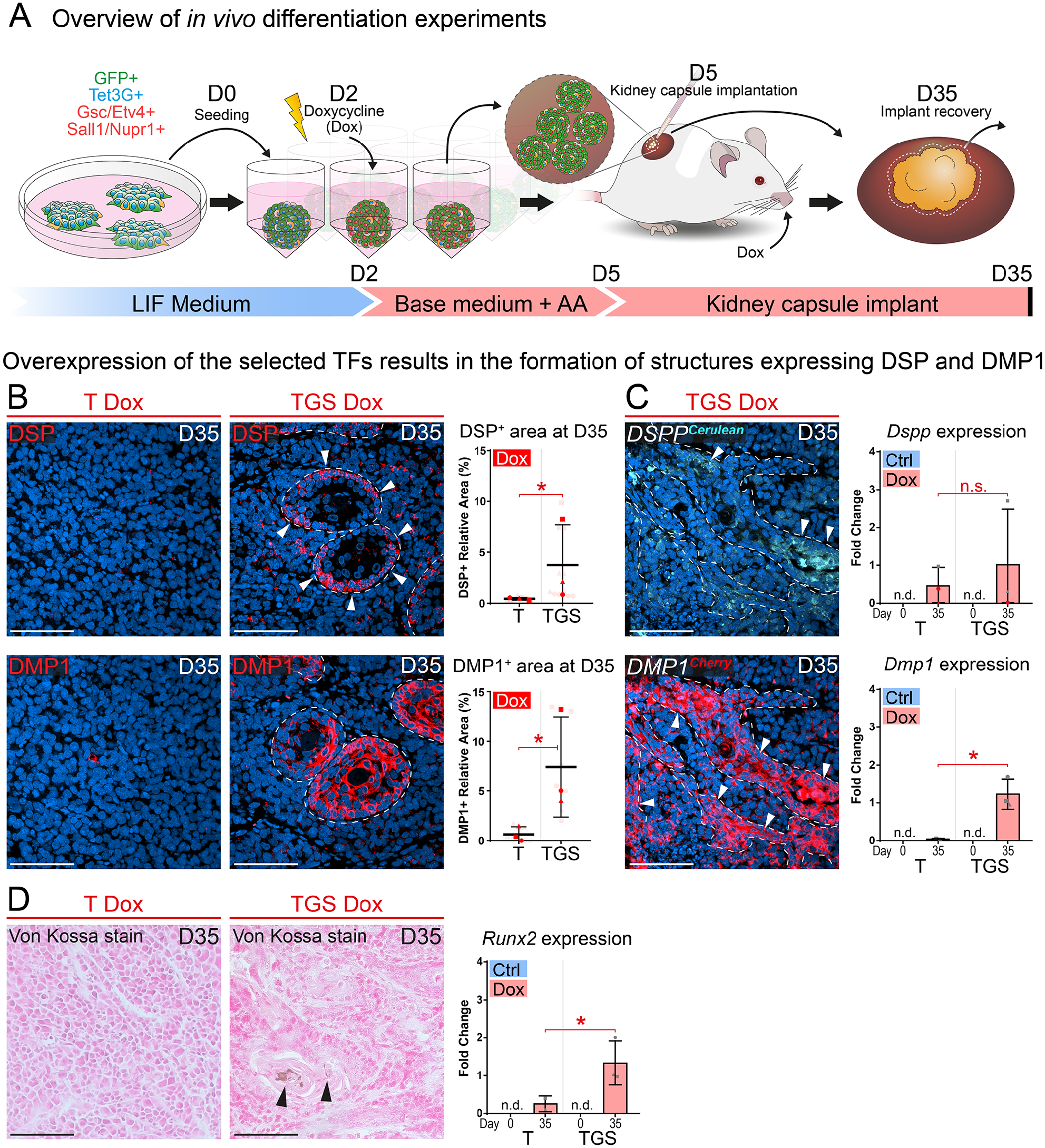
The overexpression of Etv4, Gsc, Sall1, and Nupr1 leads to acquisition of odontoblast characteristics in vivo. The process of the in vivo experiment is summarized in (
To uncover whether any mineralization occurs inside the DSP/DMP1-positive structures, von Kossa staining was performed (Fig. 4D). The TGS-Dox condition displayed mineralized nodules inside the DSP and DMP1-positive structures in contrast to the control T-Dox condition. Moreover, RT-qPCR analysis of Runx2 expression was performed, and it revealed significant upregulation of Runx2 in the TGS-Dox condition (Fig. 4D).
Dual Nature of Odontoblast-like Cells
To further investigate whether collagenous matrix, characteristic of (pre)dentin, is produced, Masson’s trichrome staining was performed on the kidney implant sections (Fig. 5A). The staining revealed that the DSP and DMP1-positive structures in the TGS-Dox condition display the presence of collagenous matrix in their centers. However, it also revealed a red-stain–positive matrix being present in the TGS-Dox implants, suggesting the presence of either enamel (Cho et al. 2013), keratin (Yang et al. 2016), or muscle tissue (Chang and Kessler 2008). Since some DSP+/DMP1+ regions displayed an epithelium-like arrangement, IHC for epithelial (CDH1) and keratinocyte (KRT10) markers was performed to further characterize the nature of the engineered tissue. The staining revealed CDH1-positive epithelial structures in the TGS-Dox condition (Fig. 5B). The centers of these regions were positive for the Masson’s trichrome red stain and generally expressed KRT10 (Fig. 5A). Co-staining with odontoblast markers (DMP1, DSP, Nestin) further revealed that these structures display both epithelial and odontoblast markers (Fig. 5B, Appendix Figs. 16, 17). Quantification showed that while the expression of CDH1 was not significantly upregulated in the TGS-Dox condition, 2 of the 3 replicates displayed approximately 10 times higher relative area (mean = 36.4% ± 1.9%) than those in the T-Dox condition (mean = 3.5% ± 2.1%) (Fig. 5C). To confirm the expression of keratinocyte markers, RT-qPCR was performed and showed upregulated expression of Krt10 and Ivl (Involucrin).
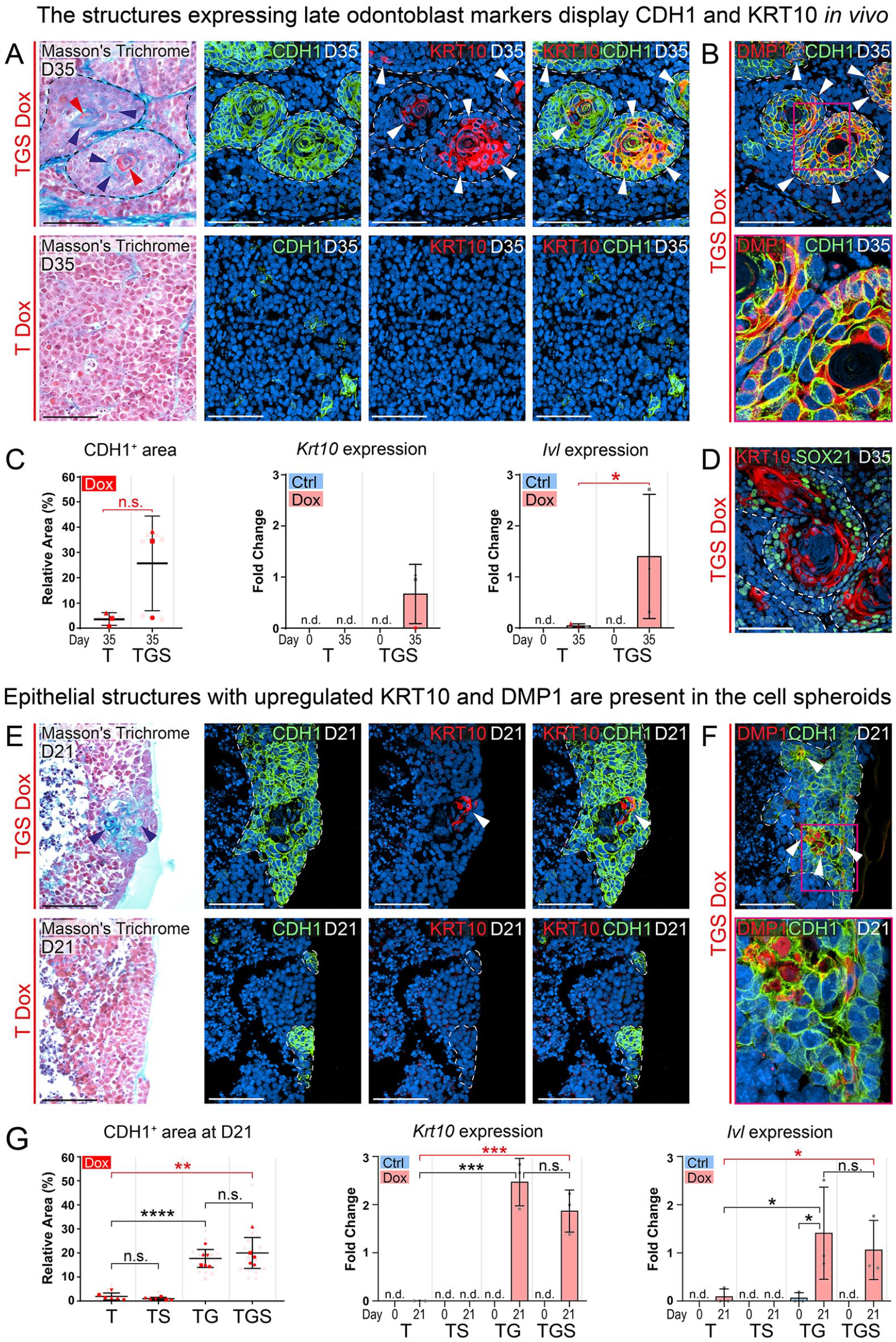
The structures expressing odontoblast markers also demonstrate epithelial characteristics and express keratinocyte markers. Masson’s trichrome staining of the kidney capsule implants (
To investigate a possible mechanism behind the observed keratinocyte-like phenotype, SOX21 expression was analyzed (Fig. 5D, Appendix Fig. 18). It revealed that SOX21 is expressed predominantly on the periphery of the epithelial structures. KRT10 co-staining showed that centrally located KRT10+ cells display generally lower or no SOX21 expression. Taken together, this suggests that the unknown matrix in the TGS-Dox condition is keratinous.
Similarly, in the in vitro–generated cell spheroids, the Masson’s trichrome staining uncovered the presence of a collagenous matrix in cell lines overexpressing Gsc and Etv4 (TG and TGS), albeit without any signs of keratinous matrix (Fig. 5E). These collagenous regions were surrounded by cells expressing CDH1 and contained KRT10-positive cells similarly to the kidney implants. Subsequently, IHC staining revealed that some of the epithelial areas displayed DMP1 and DSP expression (Fig. 5F, Appendix Fig. 15). Quantification of the CDH1-positive area revealed significant upregulation of CDH1 in the Gsc and Etv4 overexpressing conditions (TG and TGS) (Fig. 5G). Furthermore, the RT-qPCR results for Krt10 and Ivl mirrored the CDH1 expression as well as the previous DSP, DMP1, and Runx2 quantifications (Fig. 3C, D). Both Krt10 and Ivl were significantly upregulated in conditions overexpressing Gsc and Etv4 when compared with the T-Dox condition.
Taken together, the expression trends and localization of the odontoblast markers DSP and DMP1 (Fig. 5B, E; Appendix Fig. 14) mirrored those of CDH1, Krt10, and Ivl (Fig. 5). This suggests a potential developmental connection or a partially shared transcriptional program between odontoblasts and keratinocytes. This is especially apparent in the kidney implants, as both the keratinocyte and odontoblast markers were generally more overlapping in the same structures.
Discussion
The boom of modern single-cell–based transcriptomics approaches has provided unprecedented insight into the complex transcriptomic landscape that regulates cell behavior during development (Sharir et al. 2019; Krivanek et al. 2020; Pagella et al. 2021). This in-depth knowledge serves as an important resource for elucidating how organs and tissues are formed, cells interact, differentiate, and fulfill their function. Previously, Krivanek et al. (2020) generated a dental cell–type atlas using scRNA-seq of the continuously growing mouse incisor. The analysis of the ever-growing incisor enabled the in silico gene expression–based reconstruction of a differentiation trajectory of one of the main tooth hard-tissue-forming cells: odontoblasts. This trajectory, mapping their continuous differentiation from mesenchymal progenitors, was used as a basis for the selection of potential regulatory TFs driving odontoblast differentiation.
Suitable TFs have been selected by a combination of bioinformatic approaches as well as previously obtained knowledge on the function of the potential candidate genes. The expression patterns suggested by the differentiation trajectory closely resembled those found in the labial side of the incisor. Based on in vivo localization and regulon analyses, newly uncovered TFs in teeth Etv4 and Gsc are likely essential for early stages of odontoblast differentiation. In addition, the localization of Etv4 expression suggested its importance for the differentiation of (pre)ameloblastic epithelium. The observed expression pattern of Sall1 aligned with recent studies (Lin et al. 2021; Lavicky et al. 2022). This, along with bioinformatic analysis, suggests its significance for both odontoblast differentiation and function. Finally, transcriptional regulator Nupr1 displayed striking specificity to cells expressing odontoblast markers Dspp and Dmp1. Its strong expression, specific for mature odontoblasts, suggests its important role in dentin production.
The in vitro differentiation of transduced miPSCs uncovered the formation of mineralized DSP and DMP1-positive particles in the conditions overexpressing Gsc and Etv4. The formation of these mineralized particles, along with the formation of structures producing collagen, suggested the differentiation of the engineered cells into those with an odontoblast-like phenotype. Conversely, the overexpression of Sall1 and Nupr1 caused a downregulation of odontoblast markers. Intriguingly, it also slightly increased the formation of mineralized particles and the upregulation of Runx2. These findings mirrored odontogenic differentiation of DPSCs and are consistent with previous studies (Lin et al. 2021). Taken together, our data suggest that Sall1 and Nupr1 are important for the mineralization capability of odontoblasts but are not sufficient to trigger direct odontogenic differentiation from miPSCs alone without Gsc and Etv4.
The in vivo differentiation experiments highlighted the same trends observed in the in vitro experiments. The expression of DSP, DMP1 was significantly upregulated in the TGS-Dox conditions. Moreover, the expression of both DSP and DMP1 was localized into distinct structures that produced a collagenous matrix, strongly suggesting their odontoblast-like phenotype. This further supported our hypothesis that overexpression of selected TFs leads to the acquisition of an odontoblast-like phenotype in a portion of the cell population.
The dual nature of a subpopulation of transduced cells was found in the conditions overexpressing Gsc and Etv4 both in vivo and in vitro. The epithelial structures that formed in the kidney implants displayed an inward stratification, similar to ameloblast epithelium organoids generated by Hemeryck et al. (2022). The outer cells displayed DSP, DMP1, CDH1 as well as rare Nestin expression. This, apart from the CDH1 expression, suggested an odontoblast-like phenotype. The inner cells displayed a keratinocyte-like phenotype suggested by the KRT10 and CDH1 expression (Wang et al. 2020) and formation of keratinous matrix. Interestingly, the inner, keratinocyte-like cells also displayed DMP1 but no DSP expression. This downregulation of DSP might be explained by a change in the transcriptional landscape as the cells move toward the keratinocyte-like phenotype while Dmp1 remains transcriptionally active. Nonetheless, the incomplete overlap of Dspp and Dmp1 expression is present during regular tooth growth as well (Appendix Fig.4) (Lavicky et al. 2022).
To our knowledge, no naturally occurring cell types exhibit this combination of odontoblast and keratinocyte molecular hallmarks in vivo. We hypothesize that via an artificial overexpression of Gsc and Etv4, we have generated cells with a dual phenotype that partially display the characteristics of both odontoblasts and a keratinocyte-like epithelium. The presence of keratinocyte markers can be explained by the function of Etv4, which is also involved in keratinocyte differentiation during hair formation (Finnegan et al. 2019). Therefore, Our results may highlight the developmental similarities between different ectodermal appendages: hair follicles and teeth. Their embryonic development shares many molecular and morphological similarities, including an Etv4 expression in the epithelium (Biggs and Mikkola 2014). The co-expression of odontoblast and keratinocyte markers may indicate the initiation of an odontogenic transcriptional program in conjunction with a hair-forming program. A similar remarkable level of cellular plasticity has been suggested before in studies highlighting the change of stem cell fate from one ectodermal appendage to another (Yoshizaki et al. 2014; Jimenez-Rojo et al. 2019). This notion is further supported by the presence of SOX21 on the KRT10-negative periphery of the epithelial structures. This suggests that the conversion between the DSP-expressing outer cells and KRT10-expressing inner cells occurred via the mechanism described previously by Saito et al. (2020). Presented data, together with the research of other teams (Kurpios et al. 2009), highlight the importance of Etv4 and Gsc in the development of ectodermal appendages. This is supported by the performed bioinformatic analysis of their regulons, which suggests their involvement in appendage and epithelial morphogenesis. In addition, we show that even relatively short overexpression of the selected genes is enough to determine the fate of stem cells toward specific cell types, which is in line with the currently used approaches of generating neuronal lineages in vitro (Ng et al. 2021).
Taken together, our results demonstrate that controlled overexpression of a few select TFs is sufficient to guide the differentiation of miPSCs into matrix-producing cells with an odontoblast-like phenotype. However, because TFs usually serve multiple roles during differentiation, which is dynamically and precisely regulated, further research is needed to reliably generate single, specific cell types using this approach. This work shows how to utilize new knowledge arising from the rapidly advancing field of -omics research, bringing it closer to precise and reliable applications in tissue engineering and regenerative medicine.
Author Contributions
J. Lavický, J. Krivanek, contributed to conception, design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; M.G. Lopez, V. Rakultsev, V. Bryja, D. Bohačiaková, T. Bárta, contributed to data acquisition and analysis, critically revised the manuscript; V. Chochola, J. Raška, H. Tuaima, J. Cotney, contributed to data analysis, critically revised the manuscript; P. Kompaníková, E.W. Winchester, contributed to data analysis and interpretation, critically revised the manuscript; L. Englmaier, J. Verner, E. Švandová, contributed to data acquisition, critically revised the manuscript; M. Buchtová, contributed to data acquisition and interpretation, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345251348148 – Supplemental material for Unraveling the Transcription Factor Code of Odontoblast Differentiation
Supplemental material, sj-docx-1-jdr-10.1177_00220345251348148 for Unraveling the Transcription Factor Code of Odontoblast Differentiation by J. Lavický, M. Gonzalez Lopez, V. Chochola, P. Kompaníková, V. Rakultsev, J. Raška, E. Wentworth Winchester, H. Tuaima, L. Englmaier, J. Verner, E. Švandová, M. Buchtová, J. Cotney, V. Bryja, D. Bohačiaková, T. Bárta and J. Krivanek in Journal of Dental Research
Footnotes
Acknowledgements
We would like to sincerely thank Katarína Marečková and Lucie Vrlíková for their help with histological sample preparation. Furthermore, we would like to sincerely thank Dr. Květoslava Matulová from the Department Pathology at the University Hospital Brno for her assistance with sample imaging. We would like to sincerely thank Zdeněk Andrysík for his kind help with exploring bioinformatic analyses. We would like to thank Kateřina Syrová for her kind help with proofreading the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research of JK was supported by Czech Science Foundation (23-06160S) and MB by the Ministry of Education, Youth and Sports of the Czech Republic from the Operational Programme Research, Development and Education (CZ.02.1.01/0.0/0.0/15_003/0000460). We acknowledge funds from the Faculty of Medicine of Masaryk University (MUNI/A/1738/2024). PK and VB have been supported by Czech Science Foundation (grant No. GA25-17360S). We acknowledge the core facility CELLIM supported by the Czech Bioimaging large RI project (LM2023050 funded by MEYS CR) for their support with obtaining scientific data presented in this paper. JL is a Brno PhD Talent Scholarship Holder, funded by the Brno City Municipality.
Data Availability Statement
All raw data are available upon request from the corresponding author.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
