Abstract
Orofacial cleft (OFC) is a common congenital anomaly in humans with variable birth prevalence in different ethnic groups. Although genome-wide association studies (GWAS) have identified single nucleotide polymorphisms (SNPs) associated with nonsyndromic OFC (nsOFC), understanding of the underlying biological mechanisms of causative SNPs and genes in nsOFC remains limited. Here, we report that the noncoding SNP, rs3741442, has an expression quantitative trait locus (eQTL) effect on epithelial genes associated with periderm differentiation. Using a combination of epigenetic markers and in silico analysis, we prioritized the intergenic SNP rs3741442 as a potential causal factor in nsOFC. The risk allele of rs3741442 is prevalent in East Asian populations, and its presence in CRISPR-edited cells leads to reduced expression of neighboring KRT18 and EIF4B. The transcription factor SP1 differentially binds the risk versus nonrisk alleles of rs3741442. Alongside this cis-eQTL impact, rs3741442 has a trans-eQTL effect on the epithelial gene TP63 that is associated with syndromic forms of OFC and psoriasis. These findings provide insights into the mechanism by which an intergenic SNP can affect palatogenesis through the modulation of gene expression.
Introduction
Orofacial cleft (OFC) is one of the most common congenital anomalies. Genome-wide association studies (GWAS) have uncovered nonsyndromic OFC (nsOFC)–related noncoding single nucleotide polymorphisms (SNPs), improving our understanding of genetic factors in nsOFC (Leslie et al. 2015; Sun et al. 2015; Thieme and Ludwig 2017; Yu et al. 2017; Howe et al. 2018; Butali et al. 2019). SNPs linked to gene transcript levels are denoted as expression quantitative trait loci (eQTLs), which can play a crucial role in unravelling essential details regarding gene expression and signaling pathways that are likely to contribute to disease (Võsa et al. 2021). Cis-eQTLs are characterized by gene expression levels influenced by a noncoding SNP near the gene. In contrast, trans-eQTLs involve SNPs located distal to the target gene or on another chromosome, providing insights into more distant regulatory interactions (Võsa et al. 2021). GWAS susceptibility SNPs in nsOFC are potential candidates that contribute to the incidence of nsOFC; however, the interpretation of the regulatory role of such variants in nsOFC incidence and the identification of their target genes remain compelling challenges for future research owing to the limited availability of functional datasets from human tissues that accurately represent relevant embryonic sites and developmental stages (Thieme and Ludwig 2017).
Lead SNPs associated with nsOFC near the KRT8 and KRT18 locus have been reported (Liu et al. 2020). These genes are highly expressed in the periderm, a simple squamous epithelium that forms the outermost layer of the developing oral epithelium (Vaziri Sani et al. 2010). A SNP in this region (rs3741442) is a candidate causal SNP associated with nsOFC, with P values reaching formal genome-wide significance (Yu et al. 2017) and is one of several SNPs near KRT18 previously reviewed (Liu et al. 2020), but it has not been followed up functionally. The chromatin status of embryonic relevant data shows that rs3741442 is potentially in an open chromatin region (Wilderman et al. 2018; Liu et al. 2020). To fully understand the disease mechanism, it is necessary to identify as many causal SNPs as possible. As a potential causal element for the incidence of nsOFC, here we investigated the cis-eQTL and trans-eQTL effects of the rs3741442 risk allele on gene expression. Our findings provide new insights into the prevalence of nsOFC and mechanisms that potentially underlie increased risk.
Materials and Methods
Bioinformatic and Chromatin Analyses
Histone marks in the 12q13.13 locus were analyzed in all available tissue types in ENCODE Phase 3. Plots were generated using the UCSC Genome Browser. Summary statistics of the UK Biobank cohort were obtained from the GeneATLAS database. The confidence interval for the odds ratio was calculated as previously described (Altman and Bland 2011). To identify genes potentially regulated by SNPs in nsOFC, we searched all available tissue data for eQTL analysis using the GTEx project. mRNA expression data were obtained from GEPIA using the GTEx and the Cancer Genome Atlas databases. WEB resources are listed in the Appendix Methods. The comparative interaction heatmap was provided by the 3D-Genome Interaction Viewer.
The differential effect of the rs3741442 allele on transcription factor (TF) binding was predicted using JASPAR for all human TF motif sets, with a relative score of >0.80 as the threshold for significance. Twenty base pairs surrounding rs3741442 were evaluated, and the putative TF-binding motifs of rs3741442-C and -T were compared.
Plasmids
pSpCas9(BB)-2A-GFP (#48138; Addgene) and pSpCas9(BB)-2A-Puro V2.0 (#62988; Addgene) were gifted by Feng Zhang (Broad Institute of MIT and Harvard). Plasmid construction for genome editing was performed as previously described (Funato et al. 2024). DNA fragments of the KRT18 promoter (–297/+1) and EIF4B promoter (–348/+31) were amplified from the human BAC RP11 clone (ID 1104D21; ThermoFisher) and cloned into the pGL2-Basic vector (Promega) using standard restriction enzyme cloning. We prepared 25-bp complementary nucleotides for the nonrisk C and risk T alleles of rs3741442, and they were cloned downstream of the target gene promoter-driven luciferase using the SalI sites of pGL2-Basic vector. The full-length cDNAs of SP1 and YY1 were synthesized using GenScript and subcloned into the pcDNA3.1 (+)/C-(K)-DYK vector. The sequences used for plasmid construction are listed in Appendix Methods.
Cell Culture
HeLa cells (Tohoku University Cell Bank), MCF7 cells (a kind gift of Kinji Ito), and A549 cells (RIKEN Cell Bank) were cultured in DMEM supplemented with 10% fetal bovine serum and 1% antibiotics.
Genome Editing in Human Cells
Using a dual-guide RNA strategy with 2 Cas9-guide RNA constructs, we generated 3 independent HeLa clones with biallelic deletion of 62-bp noncoding sequence around rs3741442. In addition, we used CRISPR/Cas9-mediated targeting with oligonucleotide donors in HeLa cells to generate 3 independent clones each, carrying either homozygous C/C or T/T alleles. A detailed description of the genome-editing methods are included in Appendix Methods.
Quantitative PCR Analysis
Total RNA extraction and quantitative polymerase chain reaction (qPCR) analysis were performed as previously described (Funato et al. 2024). mRNA expression was normalized to that of GAPDH and TBP. For TP63, a TaqMan probe (ThermoFisher; Hs00978340_m1) was used. The primer sequences used for qPCR are listed in Appendix Methods.
Luciferase Reporter Assays
Luciferase reporter assays were performed as previously described (Funato et al. 2024). All assays were performed in technical duplicates in 3 or 4 independent experiments.
Electrophoretic Mobility Shift Assays (EMSAs)
The EMSA was performed as previously described (Funato et al. 2024). We prepared probes for the nonrisk C and risk T alleles of rs3741442 by annealing 25-bp complementary nucleotides. Nuclear proteins were prepared from SP1-overexpressing HeLa cells. For competition assays, nuclear proteins were preincubated with excess unlabeled probes or 2 µg of DYK antibody (Wako Chemicals; 1E6) before adding biotin-labeled probes in band shift buffer.
Statistical Analysis
Experiments were repeated at least 3 times independently, and data are presented as the mean ± SEM for n experiments. Data were analyzed using PRISM software (GraphPad). Unpaired 2-tailed Student’s t tests were used to compare 2 groups of independent samples. One-way analysis of variance (ANOVA) with Dunnett’s post hoc test was used to compare 3 or more groups. Two-way ANOVA with Sidak’s multiple comparison test was used to determine the transcriptional activity between genotypes and TF overexpression. Correlations between the mRNA–mRNA pairs of the gene set were analyzed by calculating Spearman’s correlation coefficient.
Results
rs3741442 Is an Intergenic SNP within a Putative Regulatory Element
We investigated the functional role of the rs3741442 risk allele (Appendix Fig. 1A). The SNP rs3741442 was identified in East Asians and Brazilian populations by GWAS of nsOFC (Yu et al. 2017; Machado et al. 2022). The nsOFC-associated C>T substitution at rs3741442 was more prevalent and homozygous for the minor allele in East Asians, whereas the rs3741442-T allele was virtually absent in Europeans (Fig. 1A). rs3741442 falls within a block of mammalian sequence conservation at the 12q13.13 locus (Fig. 1B). In addition, rs3741442 overlaps a predicted TF binding site cluster and enrichment for activating markers (H3K4me1 and H3K4me3) (Fig. 1B).
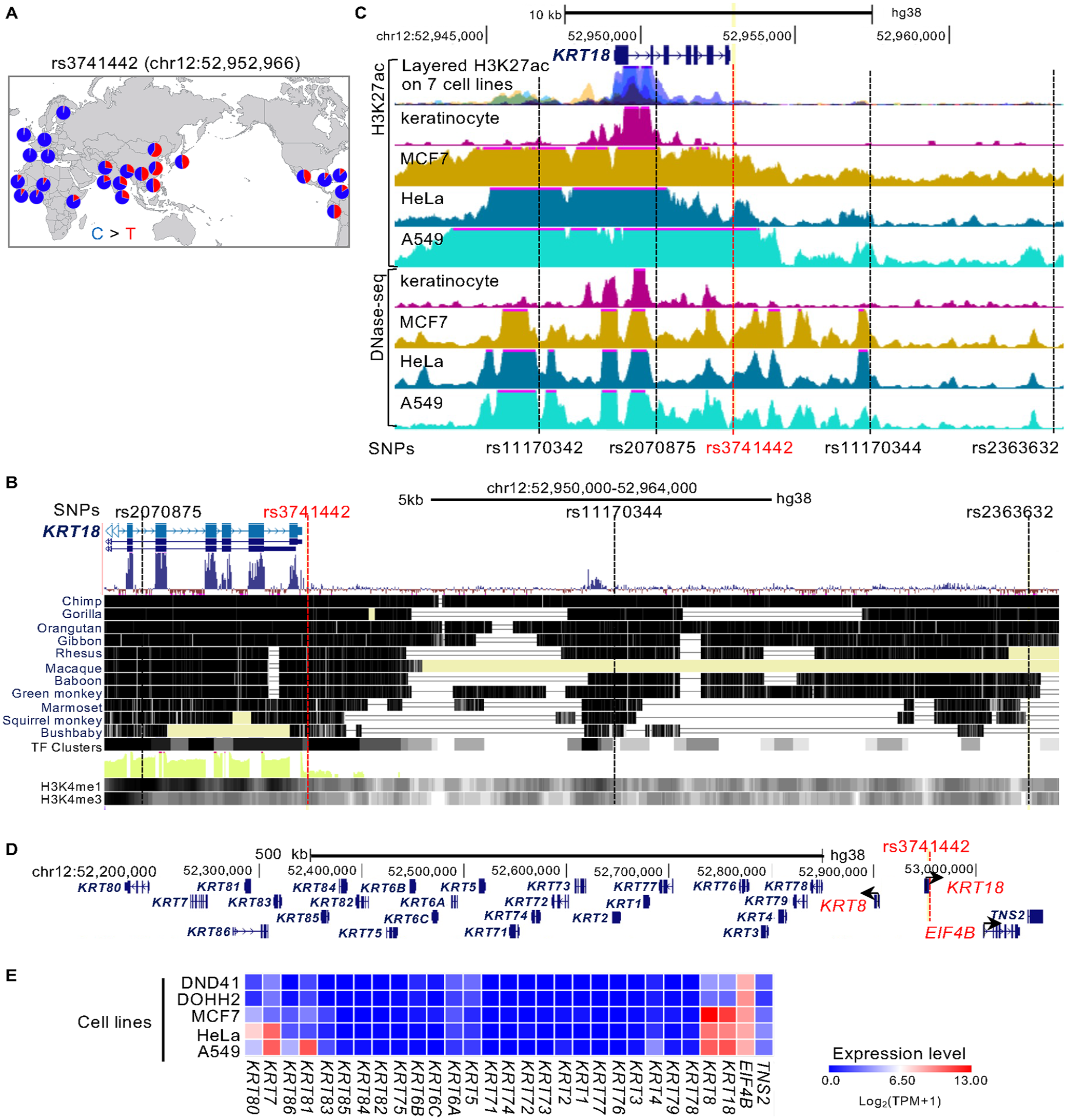
rs3741442 is located in a regulatory region downstream of KRT18. (
To find cell types in which rs3741442 has a relevant regulatory function, we examined the occupancy of histone marks at rs3741442 in epithelial cell lines annotated in the ENCODE project. Active histone modification peaks (H3K27ac) and DNase I–sensitive regions were enriched at rs3741442 in human epithelial–derived tumor cell lines (Fig. 1C; Appendix Fig. 1B, C). In particular, HaploReg analysis indicated enrichment of histone marks and DNase I signals at rs3741442 in HeLa cells (Appendix Table 1). In contrast, normal keratinocytes and 7 other cell lines did not show H3K27 acetylation at rs3741442, suggesting that this putative regulatory region exhibits cell type–specific activity and that the active chromatin state may be tightly regulated. MCF7, HeLa, and A549 cells showed higher expression of KRT18, KRT8, and EIF4B, whereas most other genes in the chr12q13.13 region showed lower expression (Fig. 1D, E). These data suggested that the genomic region encompassing rs3741442 could be involved in cell-specific regulation of these genes.
Targeted Deletion at rs3741442 Decreases KRT18 and EIF4B Expression
To determine whether rs3741442 influences phenotypic traits in humans, we searched for associations between the rs3741442 polymorphism and phenotypes. The T allele of rs3741442 is more frequent in nsOFC than in control cases (Machado et al. 2022). In the UK Biobank records, the effect allele of rs3741442-T was associated with an increased risk of skin disorders such as atrophic disorders and psoriasis (Fig. 2A). These findings suggest that this SNP may have gene-regulatory functions in epithelial tissues. The predicted rs3741442–gene interactions using 3DSNP suggested that rs3741442 is located in the same 3D chromatin loops as KRT8, KRT18, and EIF4B in the epithelium (Appendix Fig. 2). The keratin genes KRT8 and KRT18 are expressed in epithelial-derived tissues, whereas EIF4B, which encodes eukaryotic translation initiation factor 4B, is expressed ubiquitously (Fig. 2B). To assess whether the region including rs3741442 affects the expression of the neighboring genes, we performed genome-editing experiments. First, we used a dual-guide RNA strategy with 2 single-guide RNAs to generate a deletion of the noncoding sequence around rs3741442 (Fig. 2C; restricted to 62 bp due to gRNA design constraints). Three HeLa cell clones with a biallelic 62-bp deletion were generated (Fig. 2D). In the 62-bp noncoding sequence around rs3741442 (Appendix Fig. 3A), JASPAR predicted 264 putative sites (Appendix Table 2). Deletion of the 62-bp flanking rs3741442 resulted in significantly lower expression of KRT18 and EIF4B compared with wild-type cell lines (Fig. 2E), suggesting that this region could be involved in regulating the expression of these genes. We also edited HeLa cells to compare the effect of homozygous rs3741442 C or T alleles on the expression of these genes (Fig. 2F). Risk T/T HeLa cells demonstrated a significantly lower expression of KRT18 and EIF4B than nonrisk C/C cells (Fig. 2G). The difference in protein expression changes between T/T and C/C cell lines was not significant (Appendix Fig. 3B), although the reason remains unclear. These results indicate that the rs3741442 risk allele could lead to reduced expression of EIF4B and KRT18.
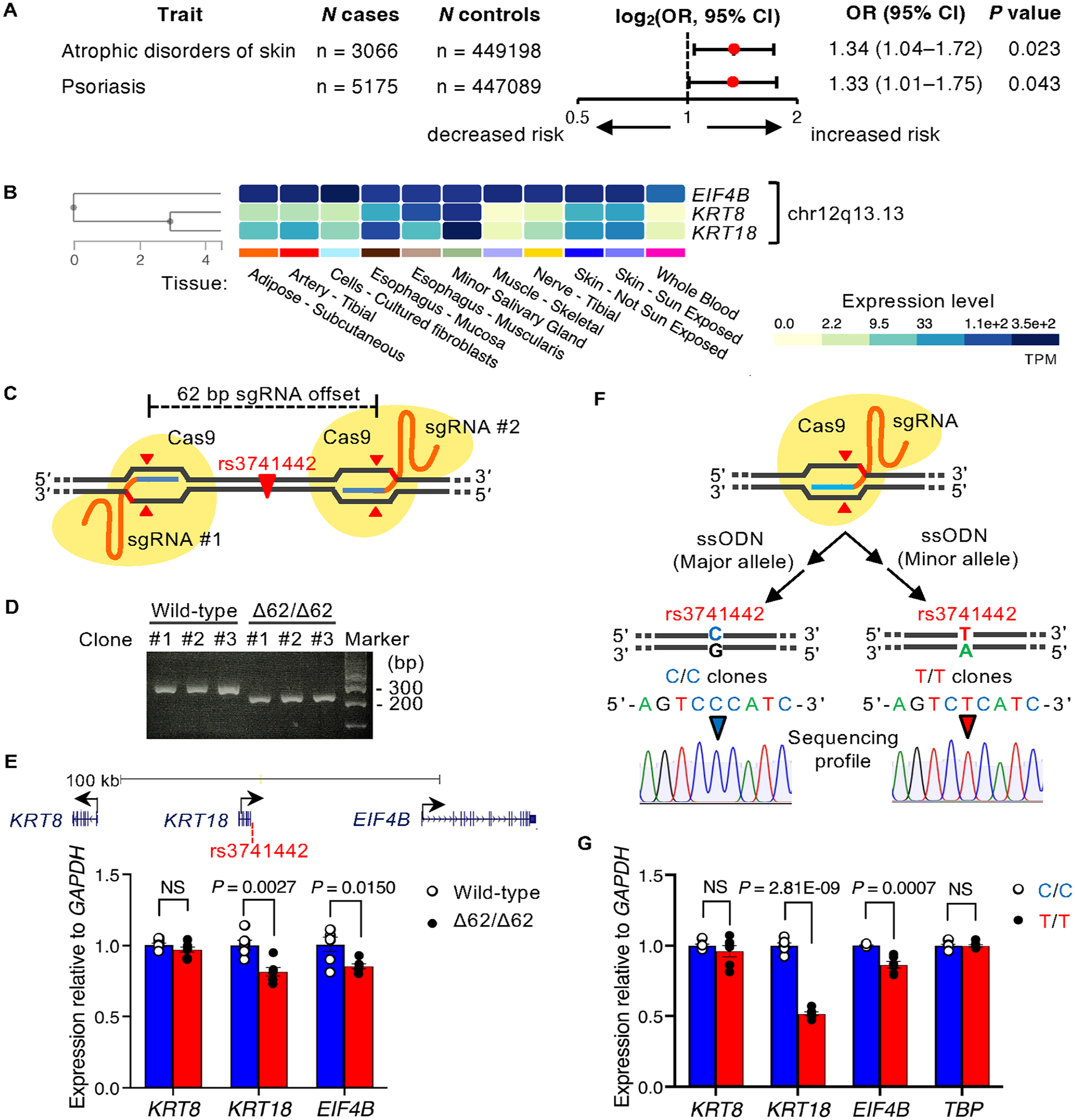
The intergenic single nucleotide polymorphism (SNP) rs3741442 affects the expression of KRT18 and EIF4B. (
SP1 Differentially Binds the Risk versus Nonrisk Alleles of rs3741442
To characterize cis-regulation by rs3741442, we constructed luciferase reporter vectors by cloning the KRT18 promoter and inserting 25 nucleotides containing either the nonrisk allele C or the risk allele T of rs3741442 (Fig. 3A). We examined the effect of rs3741442 on KRT18 promoter activity (Fig. 3B). We found that the risk allele T significantly reduced KRT18 promoter activity compared with the nonrisk allele C in all cell lines (Fig. 3B). The allele-specific reporter activity of rs3741442 may be due to the different binding affinities of TFs. JASPAR identified the motif of SP1 and YY1 as candidate TFs that could bind to rs3741442-C (Fig. 3C). Chromatin immunoprecipitation and sequencing data from the ENCODE database showed similar enrichment of SP1 within the region surrounding rs3741442, and 11 potential SP1 binding sites are located within the KRT18 promoter (Appendix Fig. 4A, B). When co-transfected with SP1, the effect of rs3741442 on the KRT18 promoter activity showed a significant allelic difference (Fig. 3D). The nonrisk rs3741442-C mediated SP1 responsiveness, whereas the nsOFC-associated rs3741442-T reduced responsiveness (Fig. 3D), indicating that rs3741442 has allele-specific enhancer activity for the KRT18 promoter by SP1. In contrast to SP1, overexpression of YY1 reduced luciferase activity in Hela and MCF7 cells, with an allele-specific difference apparent in the MCF7 assay (Fig. 4). Increasing concentrations of YY1 led to a graded reduction in SP1-driven luciferase activity in MCF7 cells, indicating that YY1 acts as a transcriptional repressor in this context. We also examined SP1 and YY1 in EIF4B promoter construct assays containing either the nonrisk or risk allele. SP1 activated the EIF4B promoter constructs, which includes 10 potential SP1 binding sites (Appendix Fig. 4C), but the effect of rs3741442 on promoter activity showed no significant allelic differences (Appendix Fig. 5), indicating that the enhancer activity of rs3741442 was sensitive to the context of the promoter. These data demonstrated that rs3741442 modulates SP1-dependent activity to regulate the transcription of KRT18.
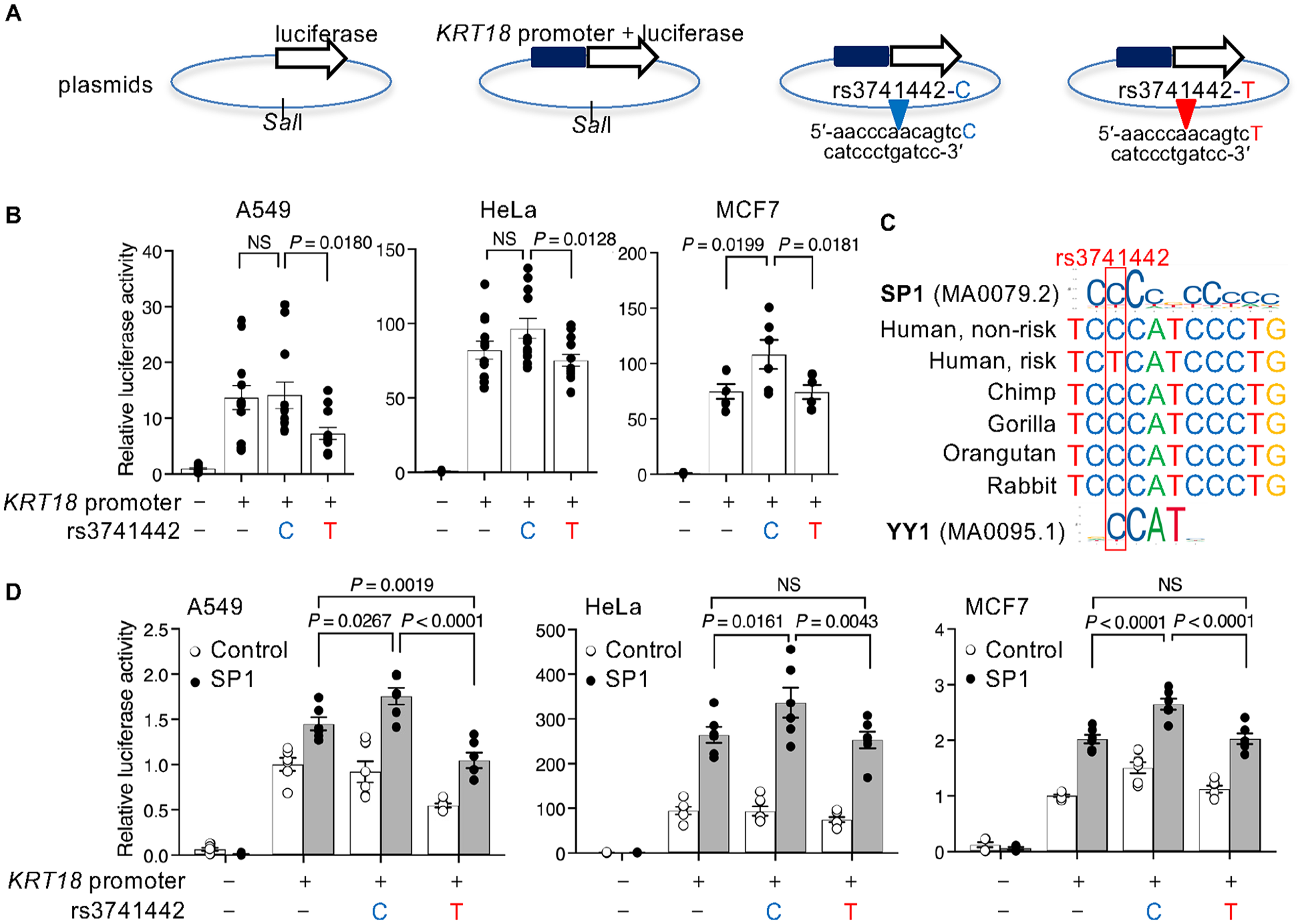
The rs3741442 risk allele alters the SP1 binding site and reduces KRT18 promoter activity. (
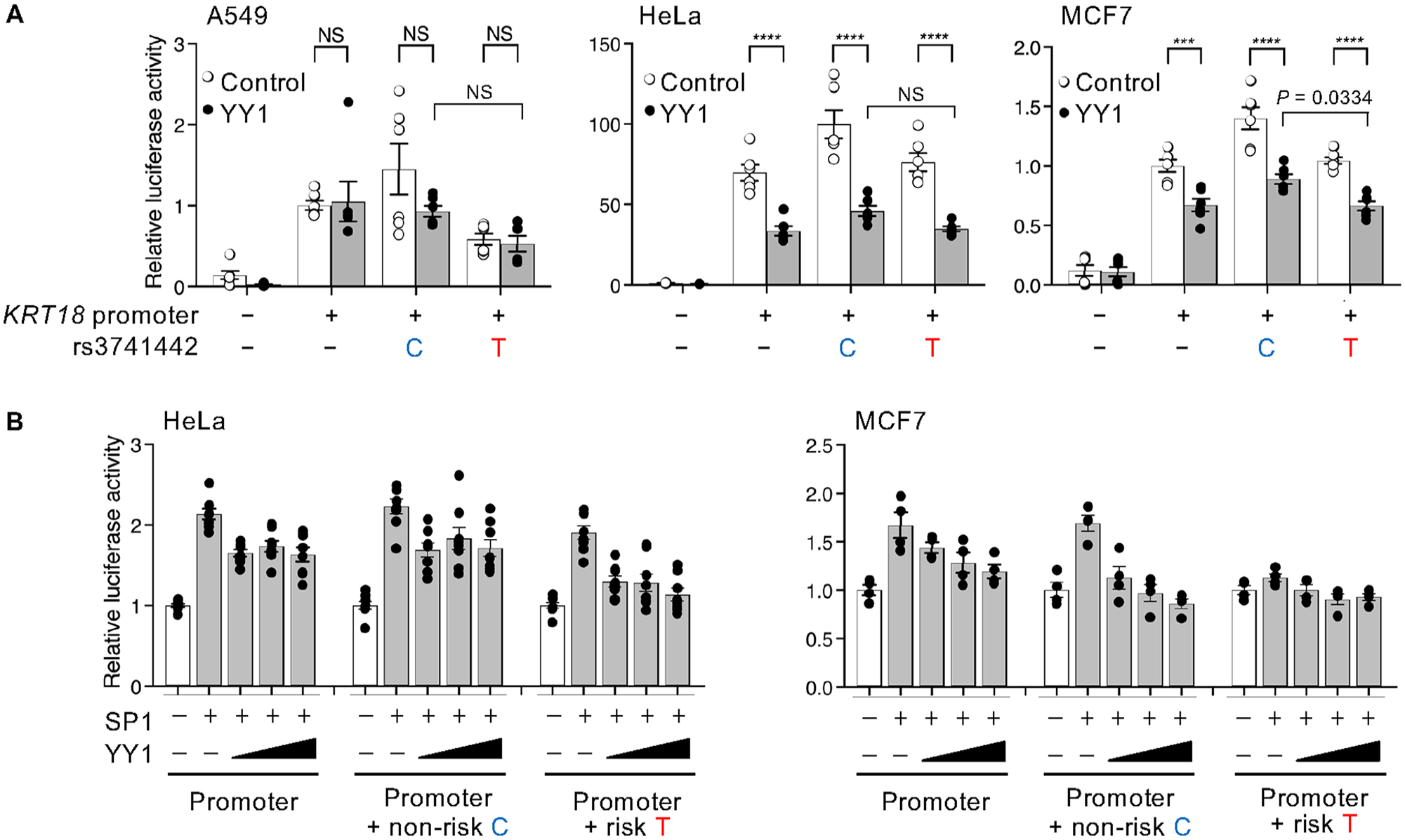
Overexpression of YY1 reduced luciferase activity in Hela and MCF7 cells. (
The Risk Allele of rs3741442 Modulates SP1 Binding
The allele-specific enhancer activity of rs3741442 may be attributed to the different binding affinities of SP1. EMSAs were performed to examine the binding of SP1 to the rs3741442 allele. Consistent with the differences in promoter activity, as compared with the rs3741442-T allele, the rs3741442-C allele preferentially bound to nuclear proteins (Fig. 5A), indicating that SP1 binds to rs3741442 in cells in an allele-specific manner. We experimentally validated the binding of SP1 to this region. EMSA showed that the bands were decreased by antibodies for both the nonrisk and risk alleles of rs3741442, indicating that the band was formed by the binding of SP1 to the probe (Fig. 5B). Differentiated epithelial keratinocyte 3C assays support the interaction between elements at the rs3741442 region and the KRT18 and EIF4B promoters (Appendix Fig. 6). Interestingly, the 3D interaction frequency was significantly elevated during the differentiation of primary epidermal keratinocytes. Most of these interactions were detected with the KRT18 promoter; however, weak EIF4B interactions were observed in differentiated epidermal keratinocytes.
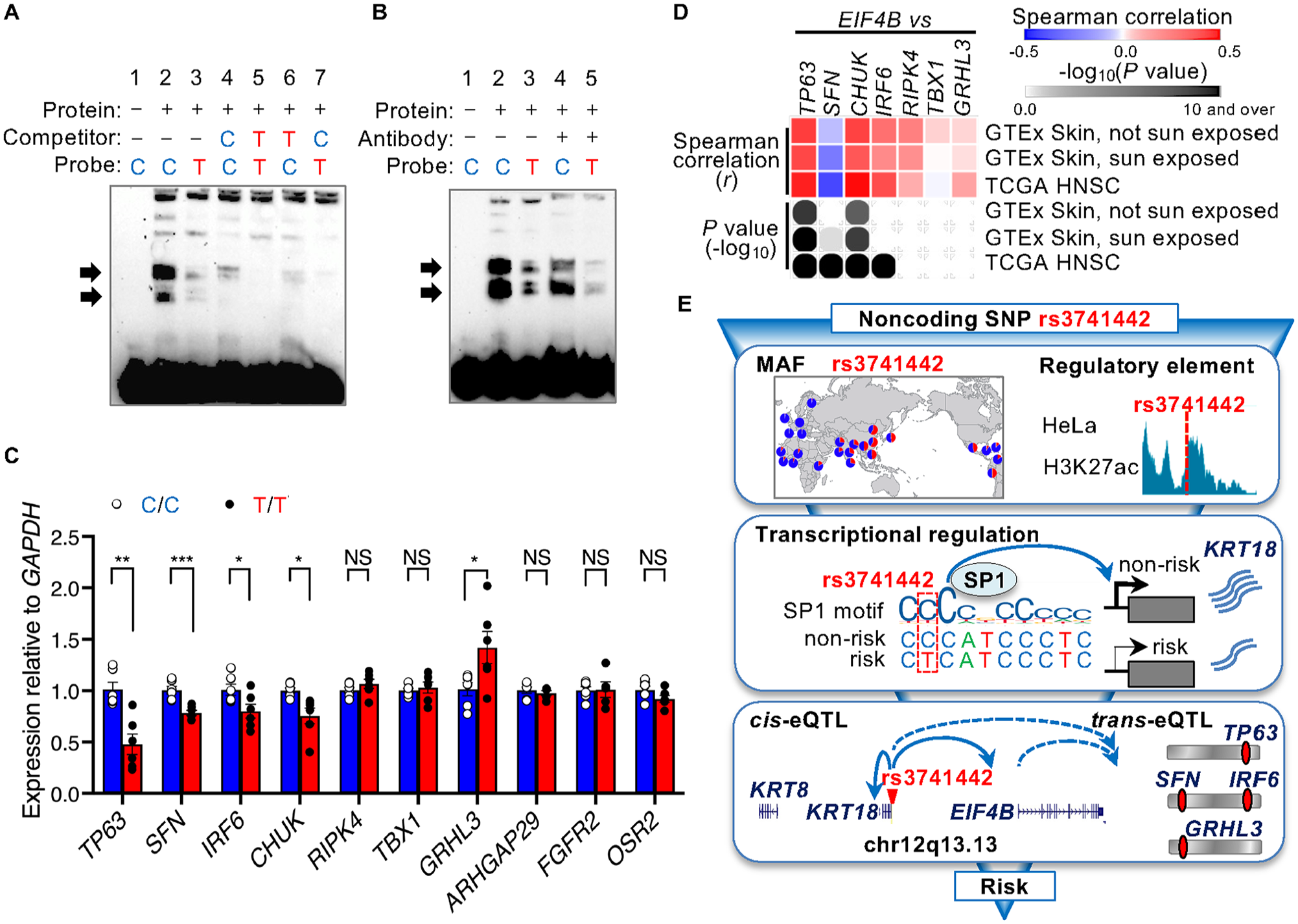
The rs3741442 single nucleotide polymorphism (SNP) affects SP1 binding as well as the expression levels of genes associated with syndromic forms of orofacial cleft (OFC). (
rs3741442 Has trans-eQTL Effects on Epithelial Genes Associated with Syndromic Forms of OFC
As cis-eQTLs generally explain only a modest proportion of disease heritability, some of the genetics of complex traits may be mediated by gene expression–level effects in trans (Yao et al. 2020). Some of the genes associated with syndromic forms of OFC in humans (Schuler et al. 2022) are expressed in epithelial tissues and craniofacial development tissues (Appendix Fig. 7; Appendix Table 3). Therefore, we determined if the enhancer activity associated with rs3741442 has a trans-eQTL effect on these epithelial genes using the genome-edited HeLa cells. Interestingly, the expression of TP63 was significantly lower, and the expression of GRHL3 was significantly higher, in rs3741442-T/T cells than in control C/C cells. The expression of SFN and IRF6 in T/T HeLa cells was also reduced (Fig. 5C). These results indicate that rs3741442 has a trans-eQTL effect on genes associated with periderm development, alongside the cis-eQTL impact on KRT18 and EIF4B. The expression of EIF4B, a cis-eQTL gene, positively correlates with the expression of TP63, CHUK, and IRF6 in skin tissues (Fig. 5D; Appendix Fig. 8). In summary, our results demonstrate that the SNP rs3741442 T allele, prevalent in East Asians, modulates the expression of cis-eQTL genes KRT18 and EIF4B. In addition to the cis-eQTL impact, rs3741442 had trans-eQTL effects on epithelial genes associated with the syndromic forms of OFC (Fig. 5E). This insight into the function of a causative SNP associated with nsOFC contributes to the understanding of the etiology of this condition.
Discussion
The identification of target genes and potential networks of risk variants associated with nsOFC is critical for understanding the etiology of nsOFC. Here, we demonstrated that increased risk of nsOFC can be genetically explained by the functional role of rs3741442 on neighboring gene expression. In silico analyses can prioritize disease-associated SNPs; however, functional tests are essential. The rs3741442 risk allele had both a cis-regulatory effect, resulting in decreased expression of KRT18 and EIF4B, as well as a trans effect on expression of the OFC-related genes TP63 and GRHL3. The function of the risk SNP rs3741442 may be temporal and tissue specific, as no significant eQTL genes were found for this SNP in any of the eQTL tissues in the GTEx database. Interestingly, analysis of SNPs near KRT18 has shed light on OFC risk in this region (Liu et al. 2020). Functional analysis was focused on a KRT18 intron 2 SNP (rs2070875) located in an active enhancer; however, whether this SNP is singularly significant remains uncertain, and possibly an additive effect of a combination of SNPs affecting gene expression is important for disease development.
Enhancer–promoter interactions bring distal regulatory elements and promoters into proximity to control gene expression mediated by TFs that help fold the looped genome. Certain SNPs located in intergenic regions exert control over the expression of distant genes through genomic interactions (Long et al. 2016; Gupta et al. 2017). Our results suggest that SP1 function is affected by binding at rs3741442 modulating the promoter activity of KRT18. SP1, a ubiquitously expressed TF, can interact with several proteins to enable the regulation of its target genes (O’Connor et al. 2016). SP1 can bind to both enhancers and promoters, bringing them into close proximity due to its ability to oligomerize (Mastrangelo et al. 1991). We demonstrated that SP1-dependent transcriptional activity is transcriptionally repressed by YY1 at the KRT18 promoter. YY1 also binds to active enhancers and promoters, facilitating physical and functional communication between them (Beagan et al. 2017; Weintraub et al. 2017). The degradation of YY1 or deletion of YY1-binding sites leads to a loss of enhancer–promoter interactions, resulting in impaired gene expression (Weintraub et al. 2017). Therefore, the risk allele of rs3741442 may alter interactions involved in the normal expression of KRT18. The risk allele also affected the expression of EIF4B, an RNA binding factor necessary for cell survival and proliferation that regulates the translation of prosurvival and proliferative mRNAs (Shahbazian et al. 2010). However, the significance of this finding in relation to OFC is unclear, although EIF4B is required for embryonic development (Chen et al. 2021) and physically interacts with the OFC-related molecules SFN (Wilker et al. 2007) and RAD21 (Panigrahi et al. 2012).
Trans-eQTLs with even modest effects exhibit the capacity to precisely identify genes that are relevant to specific traits (Võsa et al. 2021). Our study shows that rs3741442 is potentially a trans-eQTL that affects the expression of OFC-related genes. The risk allele of rs3741442 was associated with a significant decrease in TP63 expression in keeping with TP63 haploinsufficiency underlying diseases associated with OFC (Lansdon et al. 2023). In addition, the finding that the risk allele was associated with an increased risk of psoriasis in the UK Biobank data is consistent with TP63 downregulation in psoriatic lesions (Gu et al. 2006). The trans-eQTL genes Trp63, Sfn, and Irf6, together with Krt18, play a key role in periderm development (Ingraham et al. 2006; Richardson et al. 2006; Richardson et al. 2017; Liu et al. 2020). Loss of the periderm induces abnormal intraoral adhesions and fusions, allowing interactions between adjacent oral epithelial layers or underlying mesenchymal cells (Ingraham et al. 2006; Richardson et al. 2006; Peyrard-Janvid et al. 2014; Richardson et al. 2014; Richardson et al. 2017). Because the periderm layer serves to prevent premature adhesion between intraoral epithelia, perturbation of these genes may potentially lead to the onset of the OFC phenotype (Twigg and Wilkie 2015) or skin disorders by synergistically affecting epithelial morphogenesis. In mice, Trp63 regulates the expression of Irf6, Sfn, and Chuk (Candi et al. 2006; Richardson et al. 2017), and Irf6−/− and Chuk−/− mouse embryos display epithelial fusions resulting from the failure of periderm development (Li et al. 1999; Ingraham et al. 2006; Richardson et al. 2006). Sfn and Irf6 interact epistatically to control periderm development in mice (Richardson et al. 2014). The relationship between rs3741442 and nsOFC risk could therefore be mediated in part by the role of TP63 in periderm development. Interestingly, we found that the expression of GRHL3 was higher in cells homozygous for the rs3741442 risk allele than in control cells. GRHL3 encodes a TF required for formation and maintenance of the periderm, and variants in this gene are a cause of OFC (Peyrard-Janvid et al. 2014). These variants are thought to act through a dominant negative mechanism, but we note that the open-eye phenotype observed in Grhl3-null fetuses with spina bifida is also evident in Grhl3 transgenics, suggesting that overexpression of Grhl3 is also detrimental (De Castro et al. 2018). Further studies are warranted to elucidate the precise functions of the eQTL target genes and to understand the wider consequences of their dysregulation.
In conclusion, we present evidence that allelic variation in the intergenic SNP rs3741442 leads to dysregulation of cis-eQTL and trans-eQTL genes. An additive effect arising from the combination of cis-eQTL and trans-eQTL gene expression may be involved in disease development. Future investigations are needed to validate other GWAS-implicated variants and unravel the network of SNPs and their associated target genes in the context of nsOFC etiology.
Author Contributions
N. Funato, contributed to conception, design, data acquisition, analysis, or interpretation, drafted and critically revised the manuscript; S.R.F. Twigg, contributed to data acquisition and interpretation, drafted and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345251334385 – Supplemental material for The Functional Impact of the Noncoding SNP rs3741442 on Orofacial Clefting
Supplemental material, sj-docx-1-jdr-10.1177_00220345251334385 for The Functional Impact of the Noncoding SNP rs3741442 on Orofacial Clefting by N. Funato and S.R.F. Twigg in Journal of Dental Research
Footnotes
Acknowledgements
We thank Masataka Nakamura for providing the reagents and Eriko Matsumoto for data collection. The A549 cell line (RCB0098) was provided by RIKEN BRC through the National BioResource Project of the MEXT, Japan.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Japan Society for the Promotion of Science (JSPS; KAKENHI grant number JP23K09149; N.F.), the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (S.R.F.T), and the MRC National Mouse Genetics Network (MC_PC_21044; S.R.F.T.).
Data Availability
The published article includes all datasets generated or analyzed during this study.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
