Abstract
The odontoclast is a rarely studied cell type that is overly active in many dental pathologies, leading to tooth loss. It is difficult to find diphyodont mammals in which either physiological or pathological root resorption can be studied. Here we use the adult leopard gecko, which has repeated cycles of physiological tooth resorption and shedding. RNA-seq was carried out to compare gene expression profiles of functional teeth to developing teeth. Genes more highly expressed in bell-stage developing teeth were related to morphogenesis (PTHLH, SFRP2, SHH, EDAR). Some genes expressed in osteoclasts (ACP5, CTSK, CSF1R) were relatively more abundant in functional teeth compared with developing teeth. There was, however, no differential expression of RANKL (TNFSF11) in the 2 tooth types. In addition, functional teeth expressed proteolysis genes not found in osteoclasts (ADAMTS2, 3, 4, 14; CTSA, CTSH, CTSS). We used tartrate acid resistant phosphatase and cathepsin K (CTSK) staining to identify odontoclasts in and around the gecko dentition. There were 3 populations of CTSK cells: (1) large, functional multinucleated odontoclasts in the crown of the tooth with a ruffled border inside resorption pits; (2) smaller, precursor cells in the pulp with fewer nuclei; and (3) flattened external precursor cells next to the root and bone of attachment. We found a positive relationship between developing teeth and the population of CTSK+ cells on the root surface. We tested a candidate signal that may be involved in CTSK+ cell presence. An antagonist of CSF1R was delivered to developing teeth in vivo, which resulted in a significant decrease in CTSK and CSF1R compared with DMSO controls. Thus, the CSF1 signaling pathway is upstream of CTSK in teeth. This is the first work to detail the molecular characteristics of odontoclasts during physiological tooth shedding and to demonstrate that in vivo, local drug delivery is possible in the gecko model.
Introduction
Tooth replacement occurs once in humans and carnivores (diphyodont), but most vertebrates have tooth replacement throughout life (polyphyodonty) (Henriquez and Richman 2024). Instead of examining tooth initiation and development, here we focus on a less studied part of the tooth life cycle: the resorption of roots followed by shedding. Humans shed their teeth on a schedule, and odontoclasts are responsible for physiological root resorption (Domon et al. 1994; Sahara et al. 1996; Takada et al. 2004). Unwanted odontoclast activity occurs after orthodontic treatment, trauma, and pulpal inflammation, which increases the risk of root resorption and ultimately tooth loss (Iglesias-Linares and Hartsfield 2017). The prevailing assumption is that odontoclasts are just a different version of osteoclasts but with a different target tissue (Xiao et al. 2022). However, others have shown that some aspects of odontoclast biology are not the same as osteoclasts (Takada et al. 2004; Ogawa et al. 2019). To determine whether odontoclasts are indeed similar to osteoclasts, it is necessary to study physiological root resorption.
Here we developed an unconventional reptilian model that has polyphyodonty in order to study odontoclasts. As described for more than 100 y (Woerdeman 1921), reptilian teeth are resorbed and replaced in a periodic pattern independent of wear or injury. Four reptiles have been used to study molecular aspects of tooth development but not tooth resorption in adults: leopard geckos (Eublepharis macularius; Handrigan et al. 2010; Brink et al. 2021), bearded dragons (Pogona vitticeps; Salomies et al. 2019), iguanas (Iguana iguana; Brink et al. 2020), and alligators (Alligator mississippiensis; Wu et al. 2013). These studies confirm that signaling molecules in teeth are highly conserved in comparison with the mouse. Furthermore, when agonists or antagonists are applied to the dentition of reptiles, the expected response genes are activated or inhibited (Handrigan and Richman 2010; Wu et al. 2013). In contrast to these other reptiles, the leopard gecko has many teeth per quadrant that are exchanged rapidly (teeth per quadrant: 40 in geckos, 21 in alligators, 19 to 20 in iguanas; replacement interval: 4 to 5 wk in geckos, 4 to 6 mo in iguanas, and 10 to 20 mo in alligators). The functional teeth in geckos have a pleurodont attachment where the labial and lingual side of the root dentin merges with bone (bone of attachment). The lingual side of the root is much longer than the labial side is. In addition, the lingual side of the root is connected to the dental lamina, an epithelial layer connecting all teeth throughout the jaw. In contrast to mammals and alligators (Bertin et al. 2018), the developing teeth of geckos are not embedded in bony sockets but instead are attached to soft tissues and can be easily removed or treated (Brink et al. 2021). Since geckos shed their teeth in regular waves, there are odontoclasts present throughout the dentition similar to lizards and snakes (Fuenzalida et al. 1999; LeBlanc et al. 2023).
Here we carry out a transcriptome analysis of functional and developing teeth extracted from the jaw. Previous reptilian transcriptomes profiled only developing teeth and did not include odontoclasts (Tsai et al. 2016; Salomies et al. 2019). We identified many proteolytic enzymes and other markers that were unique to gecko functional teeth, possibly representing the odontoclasts. Using cathepsin K (CTSK) antibodies, we identified 3 stages in the odontoclast life cycle from precursors in the connective tissue to entering the pulp of the functional tooth to resorption of dentin in the crown and root. Finally, we performed in vivo experiments that showed signals from the developing tooth regulated CTSK+ cells near the functional tooth.
Methods
Leopard gecko eggs (Just Geckos) were hatched and housed at the University of British Columbia as described (Brink et al. 2021) (ethics protocol No. A19-0308; A23-0292). This study complied with the ARRIVE guidelines 2.0. Teeth were removed under sedation as described (Appendix methods; Fig. 1A) and used for RNA-seq analysis (accession: GSE220653; aligned to the genome for Eublepharis macularius; GCF_028583425.1). The role of colony stimulating factor 1 (CSF1) in gene expression was studied by exposing the developing teeth to the CSF1R antagonist BLZ945 (20 mg/mL) or DMSO in vivo. The developing teeth were removed 24 or 72 h later for quantitative reverse transcription polymerase chain reaction (qRT-PCR) or for histological analysis. See the appendix for full experimental details.
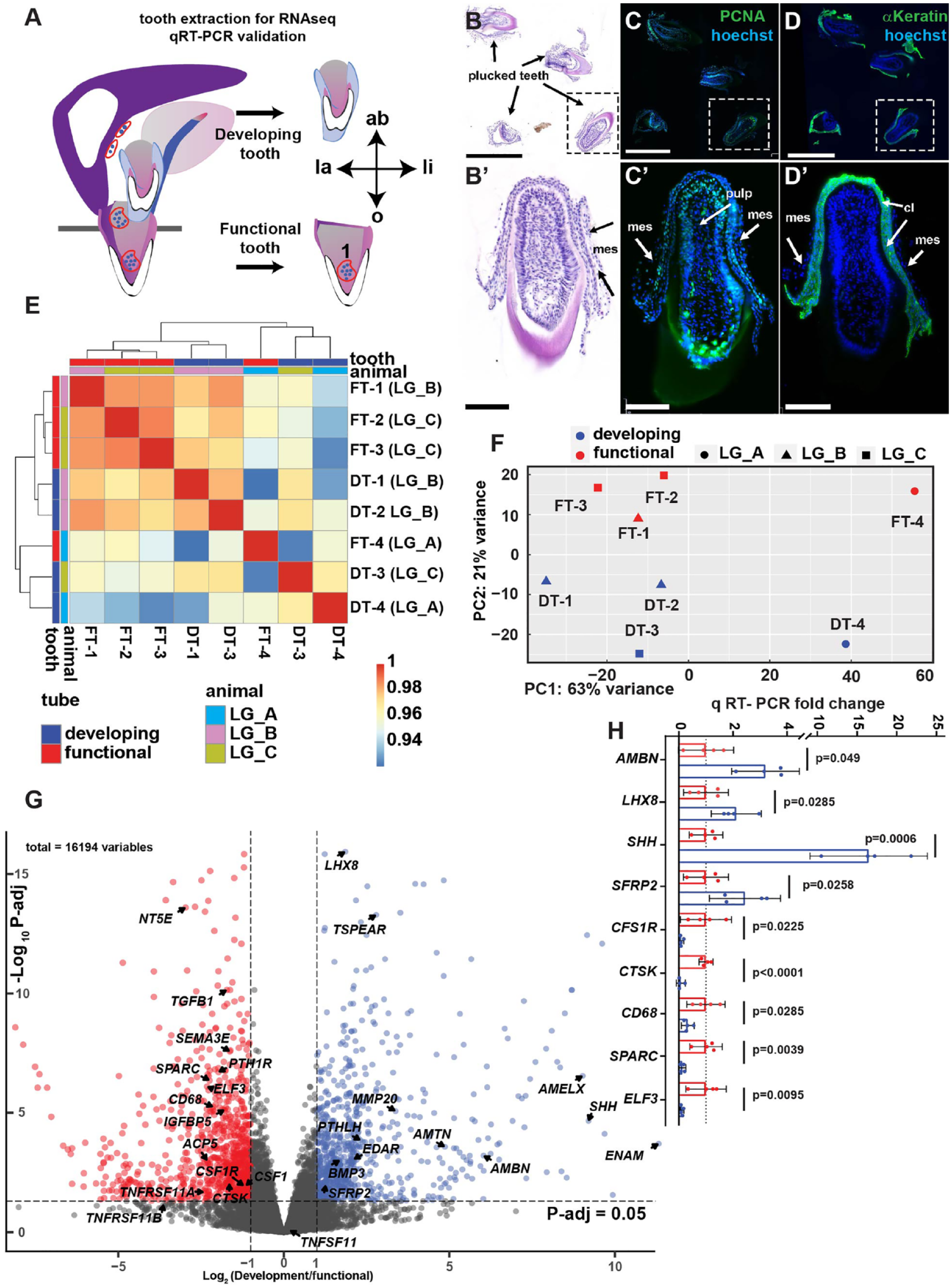
Differential expression analysis of gecko functional and developing teeth. (
Results
Differential Gene Expression in Developing versus Functional Gecko Teeth
We carried out bulk RNA-seq to compare the expression of genes in developing versus functional teeth. Functional teeth were separated from bone; therefore, no osteoclasts or osteoblasts were included in the dissection (Fig. 1A). The developing teeth were removed along with attached dental epithelium and mesenchyme (Fig. 1B, B′–D, D′), while excluding the successional and dental lamina. Quality control analysis of the expression data confirmed tooth types clustered together (Fig. 1E, F). In the PCA analysis, 2 samples (FT-4 and DT-4) clustered separately from the other samples along PC1 due to a batch effect but grouped with similar tooth types in PC2 (Fig. 1F). The analysis of 4 replicates had sufficient power to identify more than 3,000 genes. There were 1,745 genes more highly expressed in functional teeth and 1,355 genes more highly expressed in developing teeth (adjusted P < 0.05; Fig. 1G; Appendix Tables 5, 6).
Developing teeth expressed genes associated with tooth morphogenesis, differentiation, and proliferation (Appendix Fig. 1A; Appendix Table 5). Bell stage enamel organ genes included EDAR (Ectodysplasin A receptor), SHH (Sonic Hedgehog), PTHLH (Parathyroid Hormone Like Hormone; Appendix Fig. 2), and dental mesenchyme genes such as LHX8 (LIM homeobox 8) (Fig. 1G, H). Amelogenesis genes included AMBN (Ameloblastin), ENAM (Enamelin), AMELX (Amelogenin), AMTN (Amelotin), and MMP20 (codes for Enamelysin) (Fig. 1H). The developing teeth had higher expression of chemotaxis genes SEMA3F, 3G, 6A, and SLIT1 (Research data 1).
The most highly represented Gene Ontology (GO) function in developing teeth was cell division (Appendix Fig. 1A). The noncanonical, planar cell polarity WNT pathway was enriched in developing teeth including 2 WNT antagonists SFRP1 and SFRP2 (Fig. 1G, H). Several FGF ligands were also in the top 50 most highly expressed in developing teeth (FGF3, 10, 18), suggesting activity of this signaling pathway (Appendix Table 5).
Spatially Restricted Localization of SFRP2, PTHLH, and SEMA3E Transcripts in Developing Teeth
We mapped the expression of several genes with unknown expression patterns in gecko teeth using in situ hybridization. SFRP2 expression was abundant in the mesenchyme surrounding the cap-stage enamel organs and successional laminae (Fig. 2A–D). PTHLH was mapped to the stellate reticulum of developing tooth enamel organs (Fig. 2E, E′), which was in agreement with mammalian data (Philbrick et al. 1998). We found high levels of SEMA3E in the odontoblasts of functional and developing teeth (Fig. 2F, G) which would explain why we did not find differential expression using qRT-PCR (Appendix Fig. 2). In addition, there was strong expression of SEMA3E outside the teeth next to the root dentin and bone of attachment (Fig. 2F, G).
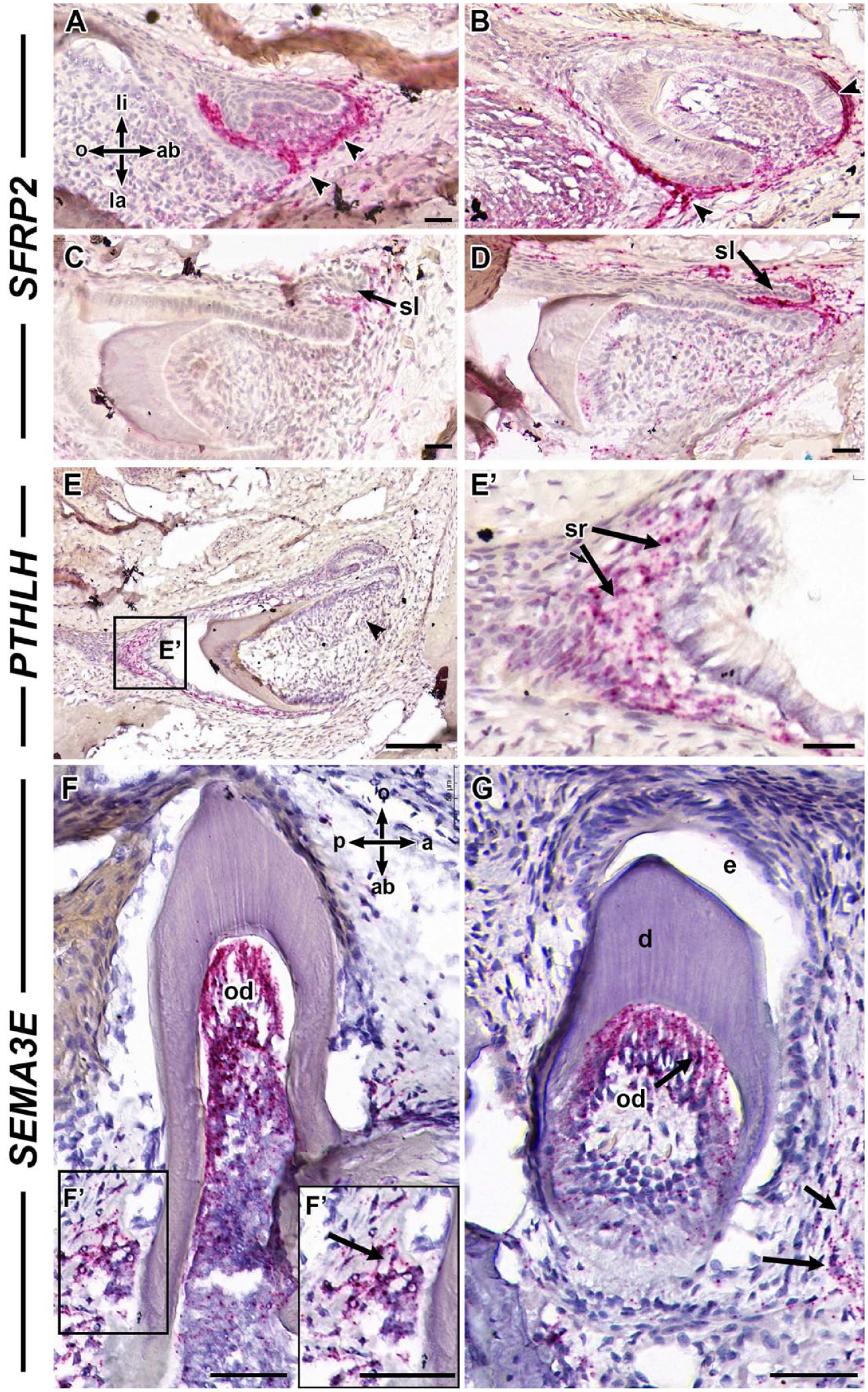
SFRP2, PTHLH, and SEMA3E expression in gecko teeth. Frontal sections of gecko jaws hybridized to RNAscope probes where the signal is chromogenically detected (pink) and sections are counterstained with hematoxylin. (
Comparison of Gene Expression in Odontoclasts in the Gecko Dentition to Osteoclasts
The GO Biological process enrichment in functional teeth included dentinogenesis genes SPARC (Secreted acidic cysteine rich glycoprotein or Osteonectin) and DMP1 (Dentin matrix acidic phosphoprotein 1) (Fig. 1G, H; Research data 1). Osteoclast induction genes CSF1 and CSF1R receptor and TNFRSF11A (codes for RANK receptor) were more highly expressed in functional teeth. In contrast, the soluble receptor TNFRSF11B (codes for Osteoprotegerin [OPG]) and the ligand TNFSF11 (RANKL) were not differentially expressed. We found expression of TNFSF11 in odontoblasts in functional and developing teeth as well as, dental mesenchyme, and enamel organs of developing teeth (Appendix Fig. 3A–E). Interestingly, there was no expression of RANKL in odontoclasts (Appendix Fig. 3C, C′, D). Functional teeth differentially expressed higher levels of genes involved in macrophage chemotaxis, replacement ossification (occurs after dental trauma), regulation of ruffled border, and osteoclast differentiation (Appendix Fig. 1B). A major gene required for osteoclast fusion, DCSTAMP, was expressed in functional teeth. The main transcription factor involved in differentiation of osteoclasts was also detected (NFATC1). Genes required for dissolution of bone matrix (CTSK, ACP5, MMP2, MMP9) were highly expressed in functional teeth (Table 1; Fig. 1G, H; Research data 1). ACP5 codes for tartrate acid resistant phosphatase (TRAP), which resorbs mineral. CTSK resorbs collagen after dissolution of mineral (Drake et al. 1996). MMP9 and CTSK are expressed in odontoclasts inside dentin resorption pits (Tsuchiya et al. 2008). The data support the idea that extracted functional teeth contain odontoclasts in addition to normal pulpal cells.
Genes More Highly Expressed in Functional Teeth from RNA-seq Analysis.
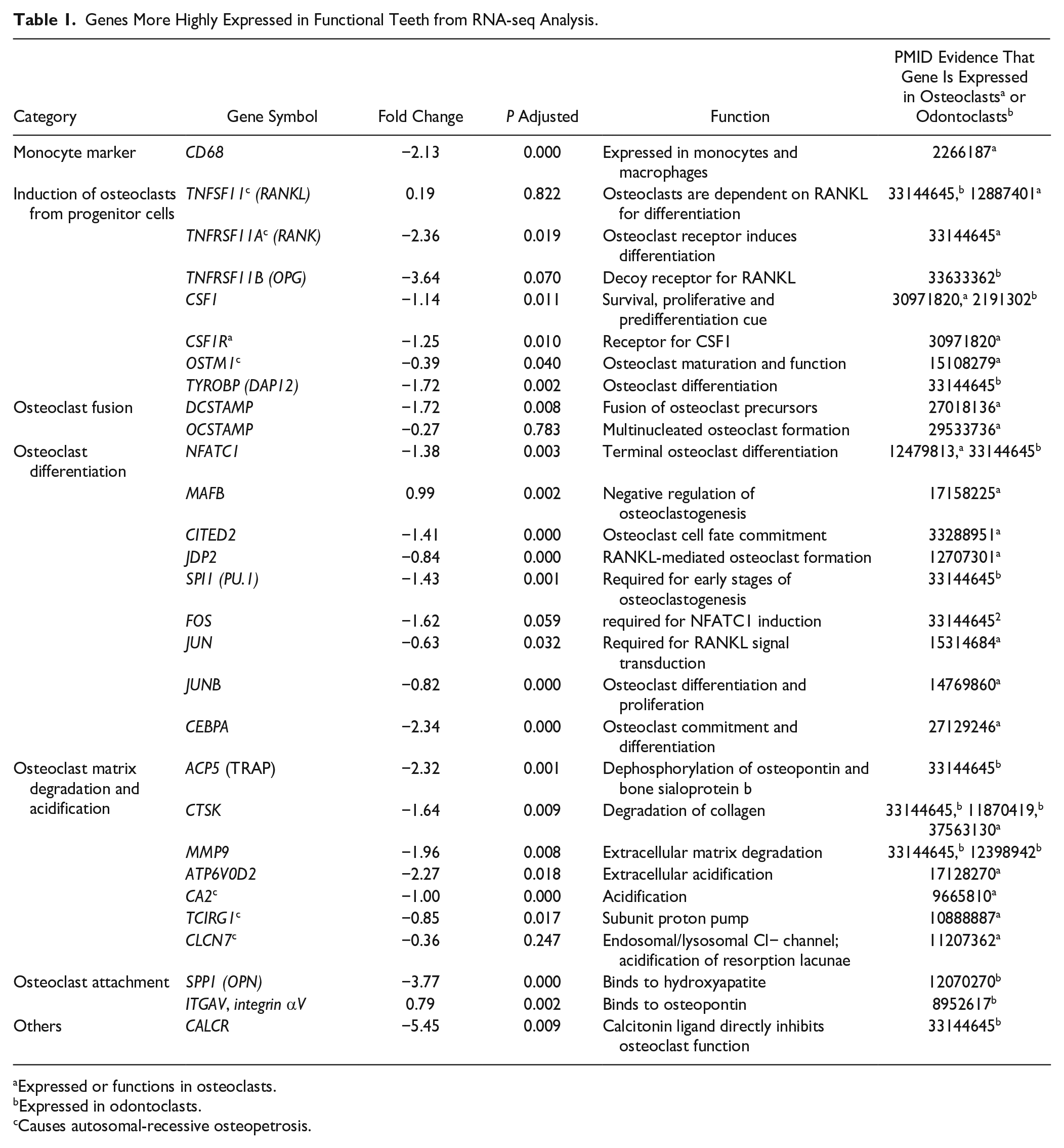
Expressed or functions in osteoclasts.
Expressed in odontoclasts.
Causes autosomal-recessive osteopetrosis.
We encountered other genes responsible for proteolysis in the pulp of functional teeth (fibrinolysis, collagen catabolic process) (Appendix Fig. 1B; Appendix Table 7; research data file 1). Enzymes included MMP23B (FC = −2.47), ADAMTS2 (FC = −1.85), ADAMTS3 (FC = −1.76), ADAMTS4 (FC = −2.03), ADAMTS14 (FC = −2.37), CTSA (FC = −1.2), CTSH (FC = −1.2) (Wang et al. 2023), and CTSS (FC = −2.26) (Smyth et al. 2022). Expression of these genes or proteins has not been reported in osteoclasts (Drake et al. 1996; Tsukasaki et al. 2020), and their spatial localization in the gecko dentition needs to be verified.
We next used TRAP and CTSK staining to identify putative odontoclasts. The CTSK antibody recognizes osteoclasts in regenerating tails of anoles (Vonk et al. 2023), and we confirmed this antibody identified gecko odontoclasts (Fig. 3A–F). Prior to resorption, functional teeth have an intact odontoblast layer and bone of attachment merging root dentin with the bone (Appendix Fig. 4A, A″). Odontoclasts are positioned on the root dentin and bone of attachment of the apical third and dentin in the middle third (Appendix Fig. 5A, C,C″; Appendix Fig. 6C; Appendix Table 8). Then odontoclasts entered functional teeth through the apical end (Appendix Fig. 4A′), while no CTSK+ cells reached the coronal pulp (Appendix Fig. 4A″, B″′). Inside the pulp, there were 2 populations of CTSK+ and TRAP+ cells, smaller cells with fewer nuclei, and larger, multinucleated cells inside resorption pits (Fig. 3B, B′, C, C′, G–I; Appendix Fig. 4B, B″; App. Fig. 5A–C; Appendix Fig. 6A–C; Appendix Table 8). The smaller pulpal cells were the likely precursors of active odontoclasts (Appendix Fig. 5B′). Eventually, the full odontoblast layer next to the dentin was replaced by multinucleated, CTSK+, TRAP+ cells. The cells in resorption pits had ruffled borders, and the Golgi apparatus was located between the nucleus and ruffled edge (Appendix Fig. 5B′). The ruffled border indicated that the collagenous matrix was actively being resorbed.
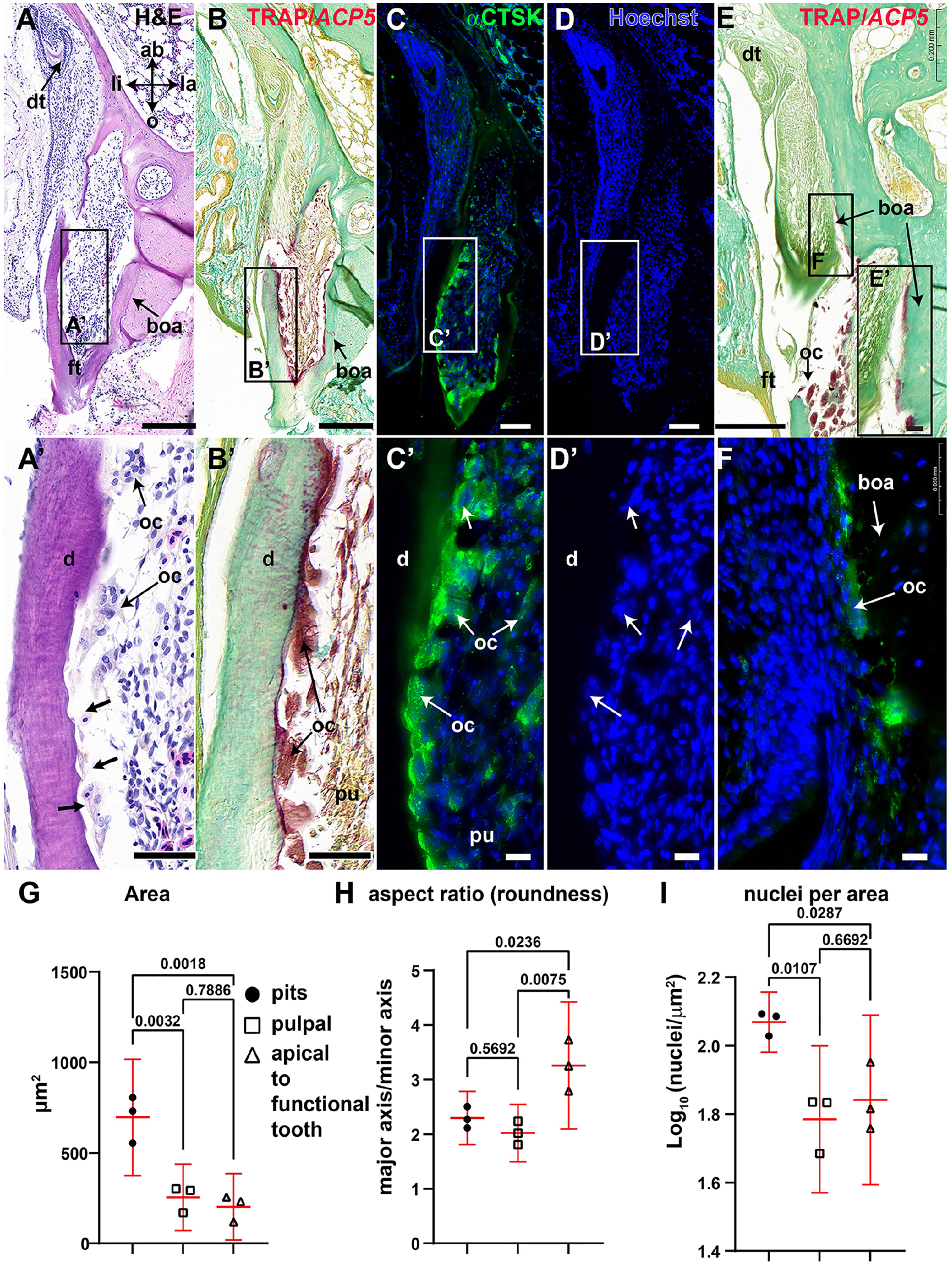
Histology and morphometry of clastic cells in the adult gecko dentition. Frontal sections of maxillary gecko teeth. (
The TRAP and CTSK+ cells attached to root dentin, bone of attachment, or bone had no visible ruffled border and had Golgi apparatus superimposed on the nuclei, which correlated with a lack of resorptive activity (Appendix Fig. 5C′, D). Transient CTSK expression was also found in a restricted group of secretory ameloblasts in the cusp tip (Appendix Fig. 7A–C′) and in the odontoblasts just at the onset of dentin secretion (Appendix Fig. 7D–F).
Selective Removal of Developing Teeth Reduced the Surface Covered with CTSK+ Cells
Next, we tested whether developing teeth provide signals that support the presence of CSTK+ cells. Our previous analysis showed that selective tooth removal delays de novo tooth formation until such time as the dental lamina heals (between 3 and 4 wk) (Brink et al. 2021). We divided the lingual root surface of functional teeth into thirds and measured the percentage of the surface that was covered CTSK+ cells (Fig. 4A). The oral third encompasses dentin close to the crown of the tooth, the middle third includes the root dentin, and the aboral third consists of apical root dentin and bone of attachment. The distance covered by CTSK+ cells was not significantly different between conditions when considered whole (Fig. 4B). However, the middle third was covered with significantly more CTSK+ surface than the oral third was (Fig. 4C; Appendix Fig. 8A, B; Appendix Table 9). Selective removal of developing teeth resulted in significantly less of the middle and aboral third surfaces covered by CTSK+ cells (Fig. 4C, P = 0.0359; Appendix Fig. 8C, D; Appendix Table 9). Therefore, removal of developing teeth either prevented survival or reduced migration or the generation of new odontoclasts attached to the root dentin.
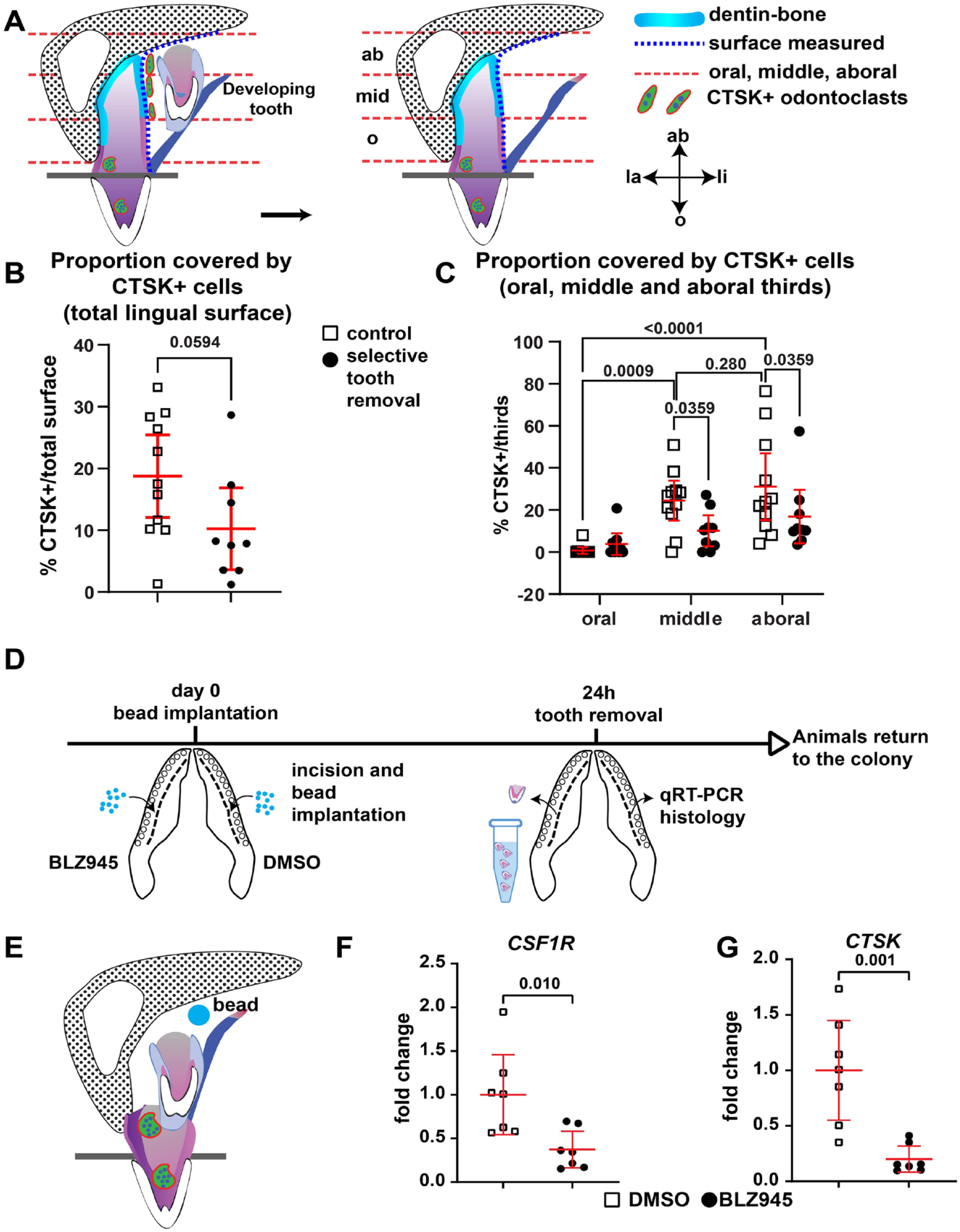
Effect of tooth removal and in vivo drug treatment on CTSK+ cells. (
Local Delivery of BLZ945, a CSF1R Antagonist, Decreases in the Expression of CSF1R and CTSK in Developing Teeth
To test an early signal that may be involved in the induction of odontoclasts, we blocked CSF1, a selective CSF1R antagonist (BLZ945) using bead implants (Fig. 4D; Appendix Fig. 9A–C′). Control beads soaked in a fluorescent dye—CM-DiI—confirmed that dye is released locally in the tissues (Appendix Fig. 9A–C′). Once attached to cell membranes, the dye does not spread unless via cell migration or cell division. BLZ945 exposure for 24 h caused a significant decrease in CSF1R and CTSK expression in developing tooth samples (Fig. 4E–G; Appendix Fig. 9E, F) while other potential target genes were not affected (CSF1, DCSTAMP, ACP5; Appendix Fig. 10A, C). Gene expression returned to control levels by 72 h (Appendix Figs. 9E, F; 10A, B). The decrease in the expression of CSF1R may be due to decreased survival of CSF1R+ cells, as shown in mice treated with other CSF1R antagonists (Rosin et al. 2018).
These results show that CSF signaling is upstream of CTSK and that local release of drugs around developing teeth is sufficient to measure gene expression changes. These methods can be used to build a hierarchy of signaling responses.
Discussion
Here, we characterize the differences between functional and developing teeth using histological and molecular analyses. The novel findings include a full transcriptome of gecko teeth in 2 stages of the life cycle and includes the first sequencing data on teeth undergoing physiological tooth resorption. We defined 3 populations of odontoclasts by their morphology, size, and position. These populations express TRAP and CTSK and may represent 2 stages of odontoclast precursors and the functional cell type. We also tested the relationship between developing and functional teeth and showed that the developing teeth maintain the presence of odontoclast precursors near or attached to functional teeth. We also showed that one of the signals that positively regulates CTSK expression in gecko teeth in vivo is CSF1.
The Developing Teeth and the Relationship to Resorption of Functional Teeth
The mechanisms to shed teeth evolved in conjunction with polyphyodonty. In animals with one set of teeth such as the mouse or rat, odontoclasts are not normally present except in a limited time window (Tsuchiya et al. 2008). In diphyodont mammals, there is a transient appearance of odontoclasts that attach to the roots of primary teeth. The absence of the permanent tooth bud delays the process of tooth shedding (Obersztyn 1963; Lin et al. 2012). In a classic study carried out on dogs, it is clear that the primary tooth roots remain fully intact for 5 mo after the coronal or full follicle is removed (Marks and Cahill 1987). On the other hand, if the coronal follicle is left in place, the primary tooth is never present in the image, implying that the tooth has been resorbed and shed. Similarly, in humans where there is agenesis of premolars resulting in an absent follicle, there is often prolonged retention of the primary molars into adulthood (Bjerklin et al. 2008).
The dental follicle in diphyodont mammals is critical for root resorption. There are data on the expression of genes in mouse dental follicle (Sfrp2, Postn, Aldh1a2, Acta2, Tagln, Gdf10, Cldn2, and Igfbp5) (Krivanek et al. 2020). Thus far, we have identified SFRP2 expression in the mesenchyme surrounding cap-stage teeth and the successional lamina. We removed SFRP2-expressing cells along with the extracted developing teeth, which correlated with a reduction in CTSK+ cells in the middle and aboral thirds. The removal of developing gecko teeth plus attached cells likely delays resorption of functional teeth (Brink et al. 2021).
Commonalities and Differences between Odontoclasts and Osteoclasts
The 2 strengths of the gecko model are that gene expression is conserved with mammals, but unlike mammals such as the mouse and rat, geckos have a steady supply of odontoclasts to study. It makes sense that some features of odontoclasts are the same as osteoclasts based on their likely common origin in fish (Chen et al. 2016). However, in a landmark study that clearly separates observations on odontoclasts from osteoclasts, orthodontic tooth movement induced odontoclasts and root resorption (Ogawa et al. 2019). These authors used a germ-line knockout of Tnfα and treated these animals with orthodontic appliances. They showed that Tnfα is required for odontoclast and osteoclast induction (Ogawa et al. 2019). We extend these findings and show that CSF1 is upstream of CTSK in the odontoclast differentiation pathway similar to osteoclasts (McDonald et al. 2021).
Other studies imply that odontoclasts have separate properties from osteoclasts. Strong evidence from a bone marrow transplant experiment in mice showed that odontoclasts have different origins than osteoclasts. Bone marrow contains osteoclast progenitor cells (McDonald et al. 2021). Mice injected with bone marrow from a donor knockout animal lacking Tnfα showed that odontoclasts were still present in the chimeric animal even though osteoclasts were lacking (Ogawa et al. 2019). These data strongly suggest that the origins of odontoclasts are from other tissues in the host, such as the connective tissue stroma (Ogawa et al. 2019). These data fit our observations that CTSK+ cells seem to be resident apical functional teeth.
Another line of evidence for osteoclasts and odontoclasts not being the same cell type comes from attempts to block root resorption using drugs that are used to treat osteoporosis in animal models. In a model in which root resorption is induced by extracting teeth and replanting them, topical treatment of the roots with bisphosphonates did not prevent resorption (Najeeb et al. 2017). Different results were obtained in a model of orthodontic-induced root resorption in which root resorption was inhibited at the same time as the inhibition of bone remodeling (Liu et al. 2004). Studies using denosumab, a monoclonal antibody that blocks RANK, have reported that there is increased risk of root resorption in patients. A retrospective study consisting of patients taking denosumab without taking bisphosphonates (Alyahya and Myers 2021) showed an association with increased root resorption; however, the authors admitted that the exact timing of drug therapy was unknown, so they could not rule out a rebound effect (Deeb et al. 2019). In a case report, a patient actively receiving denosumab therapy that successfully treated the osteoporosis developed rampant, external cervical resorption with no other risk factors (Mikuskova et al. 2022). Therefore, there is a need to test denosumab further in a model such as the gecko to determine whether there is a separate, RANK-independent pathway that regulates odontoclast function. Other proteolytic enzymes identified in this study should also be examined to determine whether they are expressed in odontoclasts and whether they mediate odontoclast root resorption.
Correlation with Human Genes Associated with Overretention of Deciduous Teeth
We found clinical correlates of the genes differentially expressed in functional teeth with genes that cause osteopetrosis, a bone disease characterized by increased bone density due to impaired osteoclast differentiation or function. We were intrigued to find that RNA-seq profiling of functional teeth included 6 genes out of the 9 that cause autosomal recessive osteopetrosis (OSTM1, CA2, TC1RG1, TNFSF11, CLCN7, TNFRSF11A; Table 1). We hypothesized that if the genes were expressed in odontoclasts, then patients with osteopetrosis would have overretained deciduous teeth. Indeed, osteopetrosis type 2 (OMIM 259710) caused by variants of TNFSF11 have overretained primary teeth (Kahler et al. 1984). All forms of osteopetrosis have dental eruption problems, but not all were diagnosed as a specific failure to resorb the primary teeth. More careful dental phenotyping in children is needed. We have used CTSK+ as a marker of odontoclasts in this study. Human variants in CTSK cause pycnodysostosis (OMIM #265800), which often includes a double row of teeth due to persistence of primary teeth (Edelson et al. 1992). Thus, we have 2 promising approaches for future work: targeting the initial induction of odontoclasts and inhibiting the function of the differentiated odontoclasts. Taken together, the human correlates and conserved gene expression profiles support the continued use of the gecko for preclinical odontoclast research.
Here, we identified a panel of proteolysis enzymes that correlate with higher numbers of odontoclasts in functional teeth. Most of these enzymes are either not found in osteoclasts or are expressed at low levels, some of which may be more adapted to degrading dentin matrix. Dentin matrix has proportionately higher levels of dentin matrix protein (DMP1) and dentin sialoprotein (DSP) compared with bone (Butler et al. 2003). Osteopontin is another noncollagenous glycoprotein in dentin that is required for odontoclast but not osteoclast formation in response to orthodontic tooth movement (Chung et al. 2008).
Author Contributions
J.I. Henriquez, J.M. Richman, contributed to conception, design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; S. Flibotte, contributed to data analysis, critically revised the manuscript; K. Fu, contributed to data acquisition and analysis, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-pdf-1-jdr-10.1177_00220345241304756 – Supplemental material for Molecular Profiling of Odontoclasts during Physiological Tooth Replacement
Supplemental material, sj-pdf-1-jdr-10.1177_00220345241304756 for Molecular Profiling of Odontoclasts during Physiological Tooth Replacement by J.I. Henriquez, S. Flibotte, K. Fu and J.M. Richman in Journal of Dental Research
Research Data 1
sj-xlsx-2-jdr-10.1177_00220345241304756 – Supplemental material for Molecular Profiling of Odontoclasts during Physiological Tooth Replacement
sj-xlsx-2-jdr-10.1177_00220345241304756 for Molecular Profiling of Odontoclasts during Physiological Tooth Replacement by J.I. Henriquez, S. Flibotte, K. Fu and J.M. Richman in Journal of Dental Research
Research Data 2
sj-xlsx-3-jdr-10.1177_00220345241304756 – Supplemental material for Molecular Profiling of Odontoclasts during Physiological Tooth Replacement
sj-xlsx-3-jdr-10.1177_00220345241304756 for Molecular Profiling of Odontoclasts during Physiological Tooth Replacement by J.I. Henriquez, S. Flibotte, K. Fu and J.M. Richman in Journal of Dental Research
Footnotes
Acknowledgements
We thank Dr. P. Panwar for technical advice on TRAP staining and quantification of odontoclasts.
Correction (February 2025):
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Natural Sciences and Engineering Research Council of Canada grants RGPIN-2016-05477 and RGPIN-2023-05145 and National Institutes of Health grant 5R21DE026839-02 to JMR.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
