Abstract
Periodontitis, a prevalent and costly oral disease, remains incompletely understood in its etiopathogenesis. The conventional model attributes it to pathogenic bacteria, but emerging evidence suggests dysbiosis involving bacteria, herpesviruses, and an exaggerated host immune response. Among herpesviruses, Epstein-Barr virus (EBV) closely links to severe periodontitis, yet the mechanisms underlying EBV-related pathogenesis remain elusive. This study examined the presence, methylation patterns, and infection states of EBV in gingival tissues from healthy patients and those with periodontitis. It also assessed gene expression differences associated with EBV through whole-genome transcriptomic profiling in healthy and periodontitis-affected tissues. EBV DNA was found at similar frequencies in healthy and periodontitis tissues, suggesting common EBV infection even before disease manifestation. In healthy tissues, mostly unmethylated EBV genomes indicated lytic infection in gums, consistent with the literature on lytic EBV spread in epithelia and continual significant virus release in the saliva of healthy carriers. Conversely, EBV DNA in periodontitis tissues showed both methylated and unmethylated patterns, suggesting a mix of latent and lytic genomes. This indicates the coexistence of latent EBV in B-cells and lytic EBV in plasma cells (PCs), linking EBV presence with both cell types in periodontitis. Whole-genome transcriptomic analysis revealed distinct expression profiles in EBV-positive periodontitis tissues, with upregulated genes associated with inflammatory/immune responses and B-cell and PC markers, while downregulated genes were related to epithelial structure and organization. The EBV-positive periodontitis signature differed distinctly from that of EBV-positive healthy gums, eliciting only a typical viral-induced immune response. These findings provide new insights into EBV physiopathology in the gum, notably assigning a direct etiopathogenetic contribution to EBV in periodontitis. The results suggest a model where EBV can commonly, and apparently asymptomatically, spread in healthy gingiva but may also aggravate inflammation in the context of gum dysbiosis, involving infiltration of B-cells and PCs and loss of epithelial integrity.
Keywords
Introduction
Periodontitis, an infection-driven inflammatory disease, damages tooth-supporting tissues, causing tooth loss, masticatory dysfunction, nutritional deficiencies, and reduced quality of life (Könönen et al. 2019). Linked to >50 systemic diseases (Beck et al. 2019) and various cancers (Michaud et al. 2018), periodontitis is a significant global health burden with >1 billion severe periodontitis cases worldwide (Jain et al. 2024). Current treatments offer only stabilization, highlighting the need for better solutions.
Despite its impact, the etiopathogenesis of periodontitis remains unclear. The conventional model blames bacteria, but an emerging hypothesis suggests complex interactions among bacteria, host immune responses, and human herpesviruses (HHVs; Slots 2015). Notably, Epstein-Barr virus (EBV) has been detected in periodontal lesions at varying levels, depending on the severity of periodontitis and detection procedures (Slots 2015; Tonoyan et al. 2021). It is plausible that bacteria and HHVs collaborate in periodontal lesions to exacerbate immune responses and damage the periodontium (Slots 2007; Chen et al. 2020), gaining credibility with increasing evidence of viral-bacterial synergy in various diseases (Zhang et al. 2024). These polymicrobial synergisms offer new insights into pathogenesis and therapies.
EBV, a prevalent γ-HHV, infects approximately 95% of people asymptomatically, primarily targeting B-cells and oral epithelial cells (ECs) via saliva transmission (Thorley-Lawson 2001). It establishes latent infection in B-cells, with sporadic lytic reactivation associated with terminal B-cell differentiation into plasma cells (PCs; Thorley-Lawson 2015). Long-term latent EBV infection is linked to lymphomas and carcinomas (Ayee et al. 2020). EBV is also implicated in several autoimmune diseases (Houen and Trier 2021), including as a direct culprit in multiple sclerosis (Bjornevik et al. 2022).
Our prior research uncovered significant insights into EBV’s role in periodontitis. We found that EBV commonly infected periodontal junctional ECs (Vincent-Bugnas et al. 2013; Tonoyan et al. 2024). Moreover, while it was established that periodontitis lesions contain numerous PCs (Li et al. 2020), we showed that these PCs were mostly EBV positive (Olivieri et al. 2020). These studies marked the first evidence of active EBV presence in periodontal lesions, suggesting that it worsens local inflammation and contributes to periodontitis pathogenesis. However, is EBV the primary suspect?
This study aims to detail differential gene expression signatures and related pathways linked to EBV presence in both periodontitis-affected and healthy gingiva, providing foundational insight into EBV’s role in periodontitis.
Materials and Methods
Donors, Gingival Tissue Sampling, and Nucleic Acid Extraction
Biomedical sample collection was conducted under DC-2022-5040 as approved by the Comité de Protection des Personnes Sud Méditerannée, and informed consent was obtained from all participants.
A total of 54 participants were recruited: 27 patients with periodontitis and 27 healthy controls (Appendix Table 1). Periodontitis diagnosis was based on established criteria (Caton et al. 2018). Patients with periodontitis displayed tooth sites with probing depth ≥6 mm, clinical attachment level ≥5 mm, and bleeding on probing. Healthy controls exhibited no signs of periodontal disease or gum inflammation.
Periodontitis-affected tissues were collected as surgical waste during periodontal therapy. Healthy gingival tissues were collected from the area just posterior to the second molar at the maxillary tuberosity region by carefully elevating the sulcus epithelium and periodontal attachment with a periosteal elevator in healthy donors. These tissues were considered surgical waste during gingival graft procedures for aesthetic purposes.
Tissues were homogenized and total RNA/DNA extracted with Qiagen kits. The concentration and purity of DNA/RNA were assessed by a SimpliNano spectrophotometer (Biochrom), and RNA integrity was confirmed with an Agilent Bioanalyzer. RNA samples with an RNA integrity number (RIN) ≥6 were selected for transcriptomic analysis.
EBV Detection
Extracted DNA was tested for the presence and quantity of EBV load via the Argene EBV quantification kit (Biomérieux), according to the manufacturer’s instructions. Following the EBV detection, all patient samples were categorized as EBV-negative and EBV-positive.
Methylation-Specific Quantitative Polymerase Chain Reaction for Latent vs Lytic EBV
EBV-positive samples (Appendix Tables 1 and 2) underwent methylation-specific quantitative polymerase chain reaction (methyl-qPCR) to assess methylation levels, distinguishing latent and lytic states (Borde et al. 2022).
Transcriptome Microarrays
Sixteen tissue samples underwent transcriptomic analysis: 4 healthyEBV-negative (H_NEG), 4 healthy EBV-positive (H_EBV), 4 periodontitis EBV-negative (P_NEG), and 4 periodontitis EBV-positive (P_EBV). Briefly, RNA was transcribed to cDNA and labeled by the single-color Low Input Quick Amp Labeling Kit (Agilent). Labeled cRNA was hybridized onto SurePrint G3 Human Gene Expression (8 × 60 K) microarrays, with fluorescent signals detected by Agilent’s scanner. Data were analyzed with limma and normalized by the quantile method. Pathway analysis was conducted by Ingenuity Pathway Analysis (IPA; Qiagen; Fall Release 2022) software. The microarray data are deposited in the GEO database (GSE223328).
High-Throughput Microfluidics-Based Reverse Transcription–qPCR for Target Gene Validation
Sixteen samples from the microarray assay and 16 new independent samples (Appendix Table 1) were used to validate the transcriptomic microarray results. Selected genes were further analyzed by microfluidics-based reverse transcription–qPCR (RT-qPCR) technology on the Biomark system (Standard BioTools), following the manufacturer’s instructions and previously described methods (Fassy et al. 2021). Thirty-one genes related to tissue markers, 30 immune and inflammatory genes, 5 B-cell and 6 PC markers, and 2 reference genes (
Statistics
EBV prevalence percentage and 95% confidence intervals (CIs) were calculated via the Clopper-Pearson method. A chi-square (χ2) test assessed EBV’s association with periodontitis. Differences in viral loads were evaluated with a Mann-Whitney
See Appendix methods for more details.
Results
Prevalence of EBV in Health and Periodontitis
EBV was detected in 75.9% (95% CI, 62.4%–86.5%) of gingival tissue samples among total participants (Fig. 1A). EBV prevalence was slightly higher in periodontitis tissues than in healthy controls (81.5% vs 70.4%) but was not statistically different (
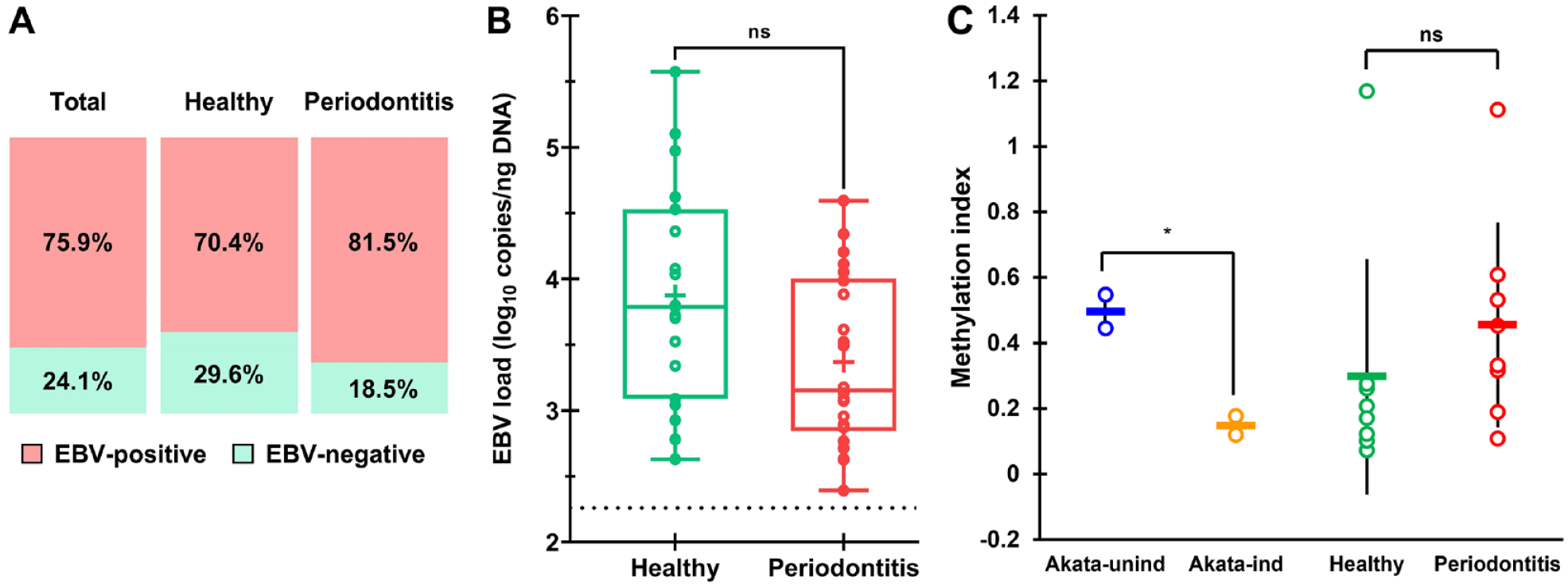
Presence of EBV in periodontitis and healthy gingiva. (
EBV genome methylation levels were examined to discern their latent or lytic states. For validation, methylation levels were assessed in EBV-infected Akata cells during latency and lytic reactivation induced by antisurface IgG. The mean methylation index dropped from 0.5 in uninduced Akata cells to 0.15 in induced ones (Fig. 1C), consistent with the findings of Borde et al. (2022). These results confirm that methyl-qPCR distinguishes latent from lytic states. In EBV-positive healthy patients, a low average methylation index indicated predominantly lytic EBV, with only 1 patient showing a high index, suggesting latent EBV (Fig. 1C, Appendix Table 2). In contrast, patients with periodontitis had both methylated and unmethylated EBV genomes, indicating a mix of latent and lytic EBV.
Identification of Up- and Downregulated Genes
Microarray-based transcriptomic analysis of healthy and periodontitis-affected gingival tissues with and without EBV presence was conducted to understand EBV’s role in periodontitis. The microarray results were subjected to IPA analysis, and differentially expressed genes (DEGs) were identified. The highest number of DEGs was in EBV-infected periodontitis tissues (P_EBV vs P_NEG), followed by EBV-negative periodontitis tissues (P_NEG vs H_NEG) and EBV-infected healthy tissues (H_EBV vs H_NEG; Fig. 2A). These results imply that EBV could have caused an extra burden on the host in addition to the periodontal disease state itself.
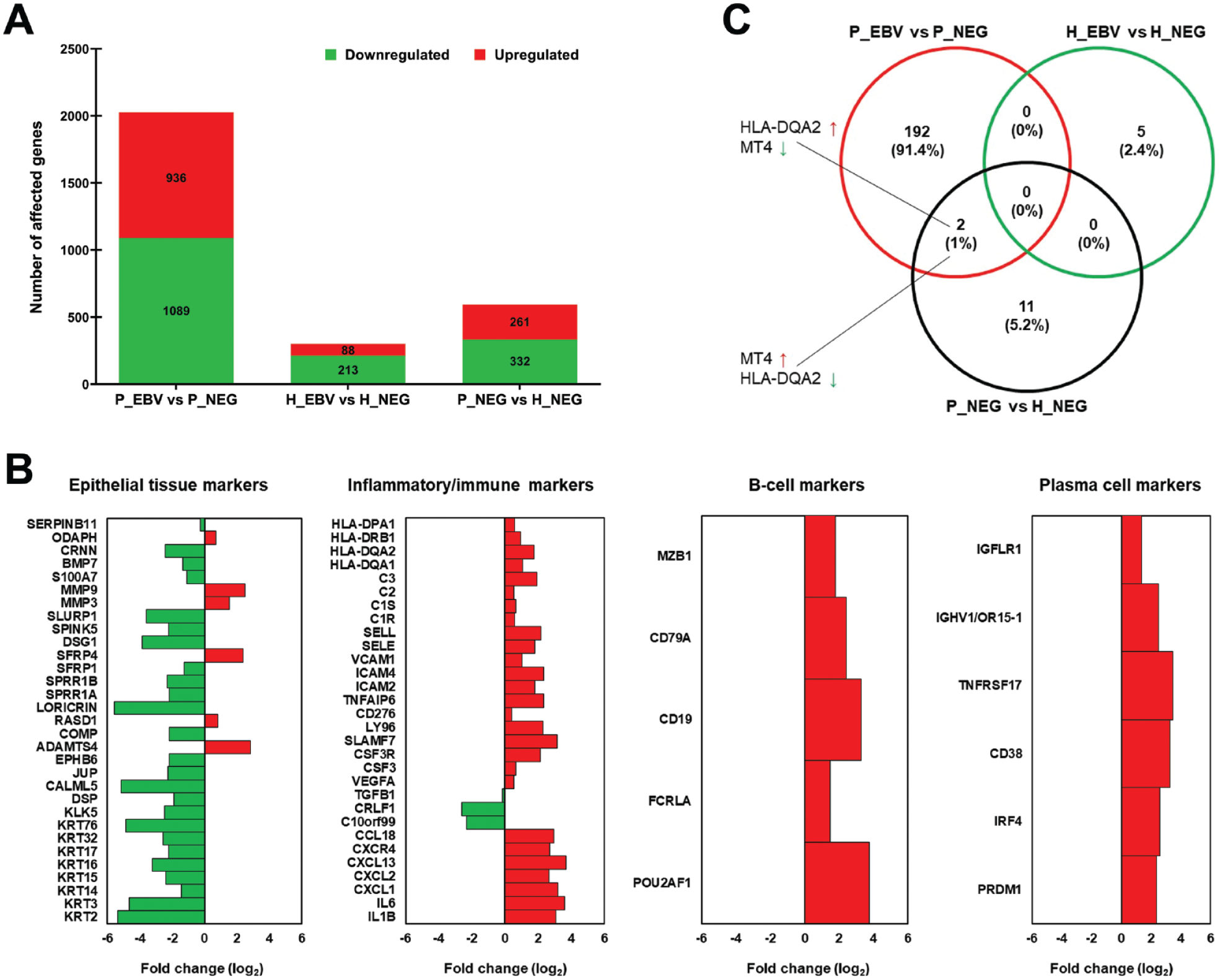
Overview of gene expression profiles. (
When we examined EBV’s impact in periodontitis (P_EBV vs P_NEG; Fig. 2B; Appendix Data Set 1), significant upregulation occurred in inflammatory molecules (IL1B, IL6, CXCL1, CXCL2, CXCL13, CSF3, ICAM4), B-cell markers (MZB1, POU2AF1, CD19, FCRLA, FCRL5, CD79A, TCL1A), and PC markers (TNFRSF17, IRF4, PRDM1, CD38, and numerous immunoglobulin transcripts), indicating an intensified inflammatory/immune response. Conversely, genes involved in epithelial cell development, keratinization, and cell adhesion were downregulated (
Second, when we investigated EBV’s impact in healthy gingiva (H_EBV vs H_NEG; Appendix Data Set 1), significantly upregulated DEGs featured
Third, as a control, we compared the states of health and periodontal disease in EBV-negative samples (P_NEG vs H_NEG; Appendix Data Set 1) and found that significantly upregulated DEGs included
Overall, DEG analysis showed poor overlap across the 3 scenarios, indicating distinct patterns linked to EBV presence and/or absence in healthy and/or periodontitis-affected gum. Specifically, only 2 genes overlapped between P_EBV vs P_NEG and P_NEG vs H_NEG, being expressed in opposing directions (Fig. 2C).
Validation of Microarray Gene Expression by Microfluidics-Based RT-qPCR
The expression trends of selected genes identified as DEGs by microarray were validated by microfluidics-based RT-qPCR. Results confirmed downregulation of epithelial tissue markers and upregulation in inflammatory/immune response and B-cell and PC markers for P_EBV vs P_NEG (Fig. 3A). Pearson correlation analysis revealed a significant positive relationship between the expression levels obtained by both techniques (
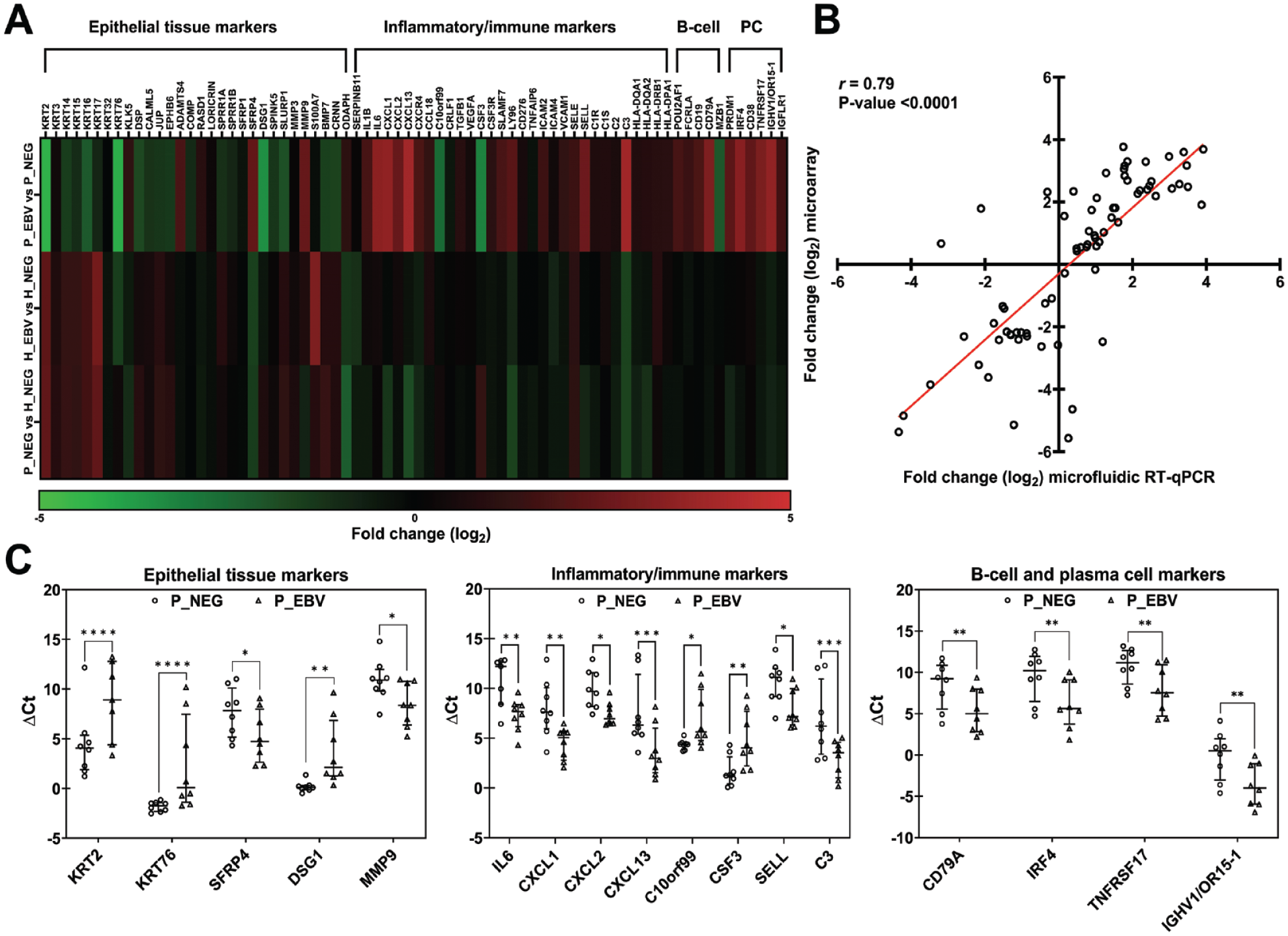
Validation of transcriptomic microarray results with microfluidics-based RT-qPCR. (
Identification of Canonical Pathways
IPA analysis was conducted to assess canonical pathway (CP) enrichments in each scenario (Appendix Data Set 2). In P_EBV vs P_NEG, 169 significant pathways were identified. The most pronounced activations (determined by z-score metric) were observed in pathways related to cellular immune response and cytokine signaling (Fig. 4A), which could lead to chronic inflammation, gingival tissue degradation, and necrosis. In H_EBV vs H_NEG, 54 significant CPs were identified. Only 2 CPs, both categorized under cellular immune response, were notably activated (Fig. 4B). In P_NEG vs H_NEG, among the 40 significantly involved CPs, the activated pathways included those related to neuroprotection and regulation of lipid metabolism and inflammation, while the inhibited pathway was associated with immune/inflammatory response (Fig. 4C). These findings suggest distinct pathway dynamics between periodontitis-affected and healthy gingival tissues in the absence or presence of EBV.
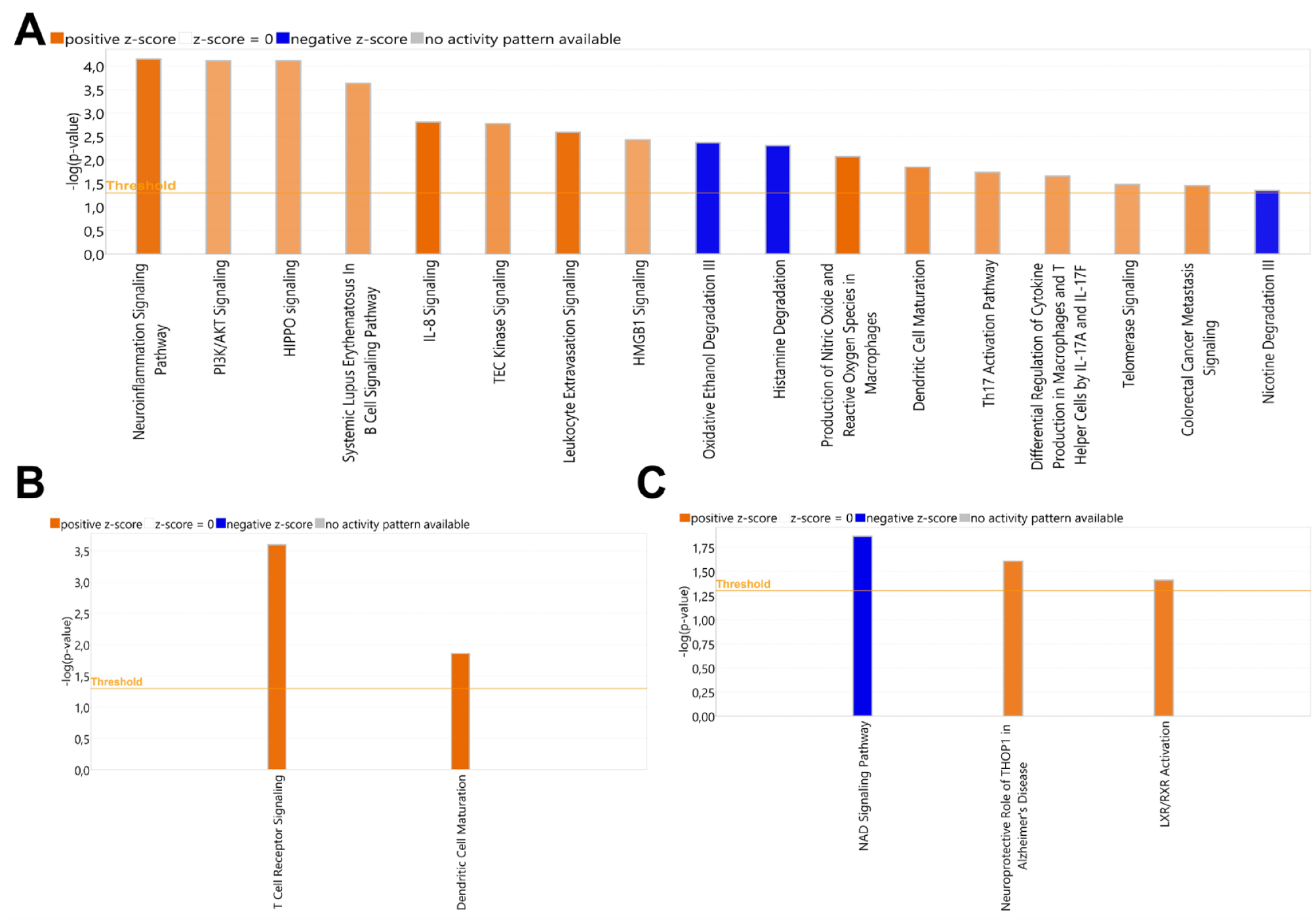
Enriched canonical pathways. (
Discussion
Identifying true pathogens and the underlying biological processes during the transition from periodontal health to disease remains elusive. Transcriptomic analysis has emerged as a leading method for studying diseases at the molecular level and uncovering mechanisms. Several studies have performed transcriptomic analyses on gingival tissues from patients with periodontitis and reported gene expression variations in periodontitis-affected vs healthy tissues related to inflammation, antimicrobial humoral response, antigen presentation, leukocyte transendothelial migration, cell communication, apoptosis, tissue degradation, response to wound healing, proliferation, regulation of metabolic processes, signal transduction, angiogenesis, etc. (Demmer et al. 2008; Abe et al. 2011; Davanian et al. 2012; Becker et al. 2014; Lundmark et al. 2015; Kim et al. 2016). Papapanou et al. (2009) even coupled 2 key drivers of periodontitis: host and microbial factors. They investigated if bacterial content in the periodontal pocket affected gene expression in nearby gingival tissues, finding a strong correlation. Nevertheless, no studies to date have investigated gingival tissue gene expression specifically in response to EBV infection in both healthy and periodontitis tissues.
This study reveals the widespread presence of EBV in both periodontitis-affected and healthy gingival tissues. EBV was detected in 82% of periodontitis cases and, unexpectedly, 70% of healthy tissues as well, highlighting its ubiquitous occurrence in gingival tissues regardless of inflammation. While EBV presence in healthy gingiva was previously suggested (Vincent-Bugnas et al. 2013), most studies reported higher prevalence in periodontitis (Slots and Slots 2019). This discrepancy between studies may stem from several factors, such as variations in detection methods, a lack of standardized procedures, and differences in sample types and sources. A consensus on EBV detection and quantification in the periodontal samples is lacking due to the diverse amplification methods and EBV genes employed across studies (Tonoyan et al. 2021), complicating comparisons. Here, we employed a standard qPCR assay, commonly used for detecting EBV in biological samples (Engelmann et al. 2018), to directly identify EBV in tissues. Another key aspect of this study was the use of tissue lysates, which offer better sensitivity for the unbiased detection of viral infections as compared with sulcular fluid or gingival smears, as they provide a concentrated source of viral genetic material. As an illustration of how sample type and detection method influence EBV prevalence, our earlier study (Vincent-Bugnas et al. 2013) detected EBV in ECs collected via cytospin from periodontal pockets and healthy sulci using Epstein-Barr virus-encoded RNA
Epigenetic modifications, such as DNA methylation of EBV gene promoters, significantly influence the entire process of EBV infection, specifically governing the transition between latent and lytic states (Zhang et al. 2022). To investigate EBV’s latent/lytic state in gingival tissues, this study employed methyl-qPCR, a method not previously documented for detecting latent/lytic EBV in gingival tissues. It is worth mentioning that EBV plays a crucial role in B-cell biology: it infects naïve B-cells and persists latently in memory B-cells. EBV replication occurs in terminally differentiated B-cells, namely PCs, where EBV transitions from latency to lytic cycle, releasing virions for dissemination. The presence of EBV in periodontitis is thus closely associated with the existence of both EBV-positive B-cells and PCs. Interestingly, we observed a similar proportion of latent and lytic EBV genomes in patients with periodontitis, suggesting the coexistence of latent EBV in B-cells and lytic EBV in PCs within the gingival tissues of these patients. Conversely, in healthy gingival tissues, a predominant lytic form of EBV was observed. Using the same method, Borde et al. (2022) similarly detected lytic EBV DNA in the saliva of EBV-positive healthy donors. Lytic EBV in healthy gingiva implies EBV spreading in epithelia, as EBV infection is primarily lytic in ECs (Eichelberg et al. 2019). This supports prior findings, showing that healthy EBV carriers release significant virus levels in saliva, highlighting ECs’ crucial role (Hadinoto et al. 2009). Although the sample size was relatively small and larger cohort analyses are necessary to validate these findings, our results highlight the distinction between the predominantly lytic state in healthy gingival tissues as compared with the balanced lytic/latent state in periodontitis. These observations imply divergent pathogenesis profiles in EBV infection between healthy and periodontitis tissues.
In a comparison of EBV-positive and EBV-negative healthy tissues, activated pathways mainly involved the cellular immune response, with DEGs mostly related to the major histocompatibility complex (Appendix Data Set 2). This suggests that immune response activation in EBV-positive healthy tissues may function as a defense mechanism against viral challenge rather than direct pathology. Conversely, altered immune-related pathways in EBV presence in healthy tissues may signal disease onset. Our results indicate that EBV can be silently present in apparently healthy tissues, likely maintained by a local equilibrium or immune tolerance between the virus and host. Disruptions to this balance, potentially from bacterial dysbiosis and/or environmental factors, could shift EBV from a silent presence to a more detrimental role by altering local immunity. However, these speculations require further investigation.
When periodontal disease was compared with health, several pathways associated with tissue damage and immune/inflammatory responses were observed, aligning with existing literature and expectations, as they reflect the pathologic processes characteristic of periodontitis.
The scenario of EBV in periodontitis was much more complex. Although CP categories in P_EBV vs P_NEG resembled those of P_NEG vs H_NEG, the sets of DEGs comprising each CP were extensive yet somewhat grouped into specific cohorts. Generally, inflammatory molecules involve cytokines, growth factors, enzymes, and complement-related molecules (Bagyinszky et al. 2017). And so, in the data set of P_EBV vs P_NEG, a wide range of mentioned inflammatory molecules was highly upregulated, such as interleukins, chemokines, colony-stimulating factors, adhesion molecules, complement-related molecules, etc. Further assessment of the transcriptomic profile of P_EBV vs P_NEG revealed panels of upregulated B-cell and PC signature genes, as well as extensive immunoglobulin transcripts that should correspond to an abundant PC infiltration. These data largely overlapped with gene expression patterns of known B-cell and PC profiles (Streicher et al. 2014), confirming the infiltration of B-cells and PCs into EBV-harboring periodontitis tissues. Interestingly, these data also confirmed our previous results that periodontal lesions were abundant with EBV-infected PCs (Olivieri et al. 2020). The presence of PCs in periodontitis has long been considered a sign of disease progression (Mahanonda et al. 2016). As we formerly proposed (Tonoyan et al. 2019), EBV-positive PCs in periodontitis may contribute to the disease through various mechanisms. These include local differentiation of EBV-infected B-cells, leading to autoantibody production. EBV-infected PCs, alongside immunoglobulin production, secrete proinflammatory molecules (e.g., IL1B, IL8, CXCL1, CXCL2), triggering significant inflammation. This heightened inflammatory response may disrupt host immune defenses, potentially leading to bacterial superinfection. The idea that EBV escalates PC infiltration in periodontitis lesions is groundbreaking, prompting further exploration of the pathogenic roles of PCs and EBV in periodontitis.
Gingival tissue destruction, another feature of periodontitis, was also evident in P_EBV vs P_NEG. Interestingly, we found DEGs known to be involved in epithelial cell development, keratinization, and cell adhesion, such as downregulated kallikreins, numerous keratins, small proline-rich proteins, desmogleins, desmoplakins, junction plakoglobins, cartilage oligomeric matrix proteins, and upregulated matrix metalloproteinases. These data suggest that the presence of EBV in periodontitis lesions may increase epithelial tissue destruction and injury. These transcriptomic results also agree with our previous study (Vincent-Bugnas et al. 2013), indicating that periodontal ECs were commonly infected with EBV and prone to apoptosis, potentially compromising epithelial integrity and contributing to periodontal destruction. Obviously, in periodontitis, the abundant clusters of EBV-infected PCs in close vicinity with ECs should greatly favor infection of periodontal epithelia and EBV oral spreading.
In conclusion, this study explored EBV’s impact on both healthy and periodontitis-affected tissues through gene expression profiles, revealing potential molecular mechanisms underlying EBV’s involvement in periodontitis pathogenesis. In particular, the continuous presence of lytic EBV in healthy gingiva may suggest the presence of yet unknown tolerogenic mechanisms that facilitate EBV persistence without eliciting inflammation. Our transcriptomic results support this assumption, revealing that EBV provoked only typical host immune surveillance against the viral challenge and no inflammatory cascade in periodontally healthy tissues. Consistent with the literature, in periodontitis-affected tissues, the host bioprocesses linked to immune/inflammatory response and tissue degradation were documented. While in periodontitis-affected tissues, the presence of EBV significantly intensified these processes: EBV exacerbated the deleterious state of the disease by recruiting an army of inflammatory molecules, summoning assistance from B-cells and PCs, and precipitating tissue destruction.
One limitation of this study is that, although tissues were classified as healthy or periodontitis based on clinical signs, their cellular composition remained heterogeneous and uncertain. Future studies should use cell type–specific analyses to better understand EBV infection in the diverse cellular environment of periodontitis-affected tissues. Additionally, the small sample size in this study highlights the need for future research to include larger cohorts. Another limitation is the lack of data on bacterial infection status: while we focused on EBV, understanding its interaction with bacterial pathogens would provide a more comprehensive view of disease mechanisms. Future research should further incorporate pathogenic bacteria into the periodontitis jigsaw puzzle, establishing a model that encompasses the interplay among bacteria, EBV, and host immune response. Hence, we propose a tentative model of periodontitis: EBV-positive PCs infiltrate the periodontitis lesions, leading to EBV replication, massive inflammation, suppression of immune surveillance, and superinfection by pathogenic bacteria, thus managing periodontitis pathogenesis (Fig. 5). These findings shed new light on periodontitis pathogenesis, marking the first direct involvement of EBV in periodontitis pathogenesis. The results could revolutionize disease management by suggesting unconventional treatments, such as antivirals for EBV-positive patients.
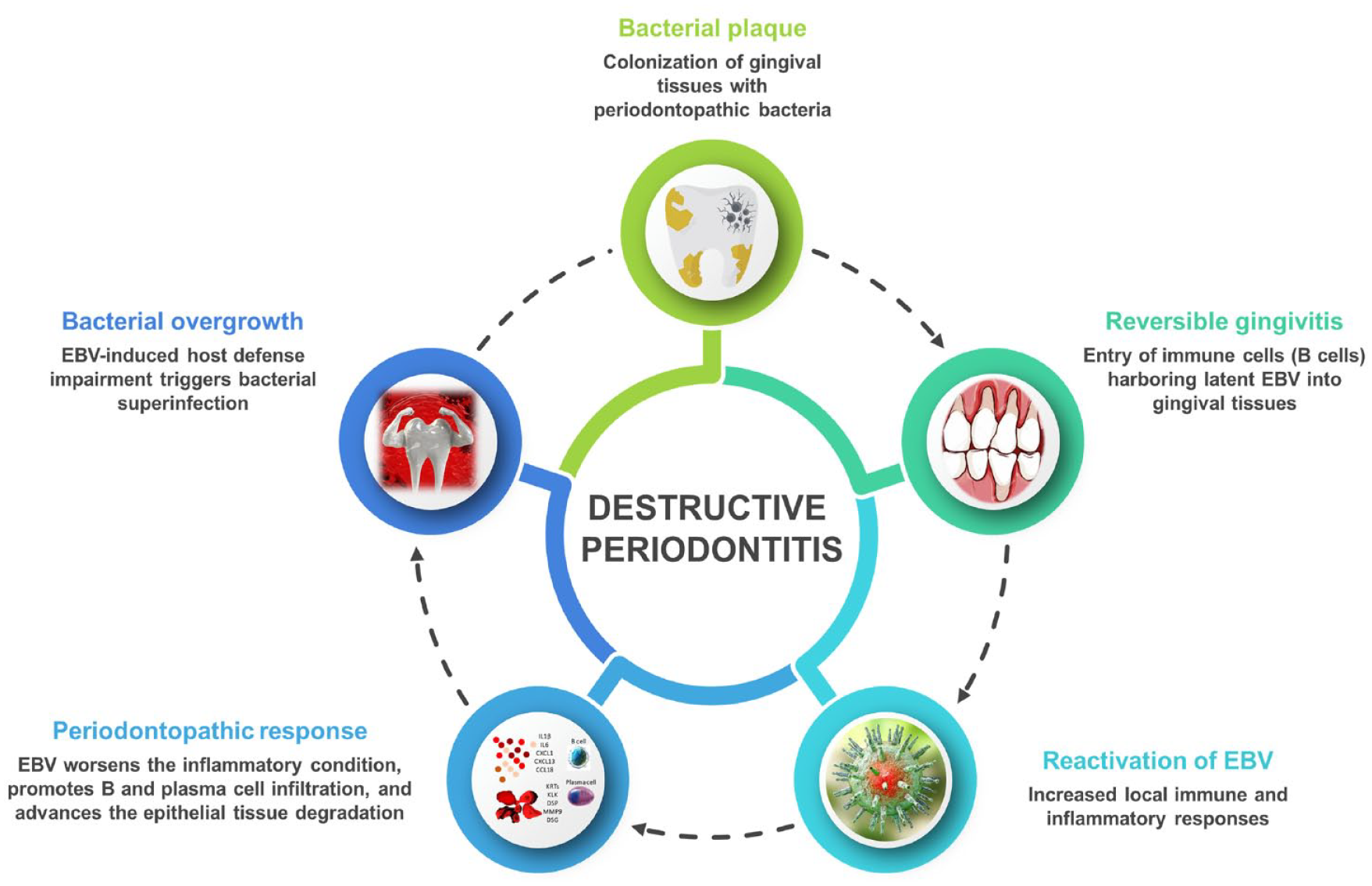
Proposed model of EBV-associated pathogenesis in periodontitis in the context of bacterial-viral-host synergistic interactions. This model suggests that episodic overgrowth of periodontopathic bacteria in the gingiva may trigger transient inflammation (gingivitis). If left untreated, this inflammation disrupts the balance between the host immune system and endogenous EBV, which persists within the tissues. Persistent EBV facilitates viral evasion and exacerbates the infection. Increased viral activity leads to heightened local immune and inflammatory responses, worsening the inflammatory condition, promoting B-cell and plasma cell infiltration, and advancing epithelial tissue degradation. EBV-induced impairment of host defenses results in bacterial superinfection, culminating in destructive periodontitis. This model underscores the intricate interplay among host immune responses, EBV, and bacterial dysbiosis, challenging traditional paradigms that attribute periodontal disease solely to bacterial infection. EBV, Epstein-Barr virus.
Author Contributions
L. Tonoyan, A. Doglio, contributed to conception, design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; C. Mounier, J. Fassy, contributed to design, data acquisition, analysis, and interpretation, critically revised the manuscript; S. Leymarie, S. Mouraret, P. Monneyron, S. Vincent-Bugnas, contributed to data acquisition and interpretation, critically revised the manuscript; B. Mari, contributed to conception, design, data acquisition, analysis, and interpretation, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345241303138 – Supplemental material for Unveiling the Etiopathogenic Role of Epstein-Barr Virus in Periodontitis
Supplemental material, sj-docx-1-jdr-10.1177_00220345241303138 for Unveiling the Etiopathogenic Role of Epstein-Barr Virus in Periodontitis by L. Tonoyan, C. Mounier, J. Fassy, S. Leymarie, S. Mouraret, P. Monneyron, S. Vincent-Bugnas, B. Mari and A. Doglio in Journal of Dental Research
Supplemental Material
sj-xlsx-2-jdr-10.1177_00220345241303138 – Supplemental material for Unveiling the Etiopathogenic Role of Epstein-Barr Virus in Periodontitis
Supplemental material, sj-xlsx-2-jdr-10.1177_00220345241303138 for Unveiling the Etiopathogenic Role of Epstein-Barr Virus in Periodontitis by L. Tonoyan, C. Mounier, J. Fassy, S. Leymarie, S. Mouraret, P. Monneyron, S. Vincent-Bugnas, B. Mari and A. Doglio in Journal of Dental Research
Supplemental Material
sj-xlsx-3-jdr-10.1177_00220345241303138 – Supplemental material for Unveiling the Etiopathogenic Role of Epstein-Barr Virus in Periodontitis
Supplemental material, sj-xlsx-3-jdr-10.1177_00220345241303138 for Unveiling the Etiopathogenic Role of Epstein-Barr Virus in Periodontitis by L. Tonoyan, C. Mounier, J. Fassy, S. Leymarie, S. Mouraret, P. Monneyron, S. Vincent-Bugnas, B. Mari and A. Doglio in Journal of Dental Research
Footnotes
Acknowledgements
The authors thank Prof. Vincent Marechal for helpful advice regarding methylation-specific quantitative polymerase chain reaction assays.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement 896201 and from GIRCI-Méditerrannée (PHRC 2020, HERPARO project).
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
