Abstract
Early childhood caries (ECC) is the most common noncommunicable childhood disease—an important health problem with known environmental and social/behavioral influences lacking consensus genetic risk loci. To address this knowledge gap, we conducted a genome-wide association study of ECC in a multiancestry population of U.S. preschool-age children (N = 6,103) ages 3 to 5 y participating in a community-based epidemiologic study of early childhood oral health. Calibrated examiners used International Caries Detection and Assessment System criteria to measure ECC; the primary trait was the number of primary tooth surfaces with caries experience (i.e., dmfs index). We estimated heritability and concordance rates and conducted genome-wide association analyses to estimate overall genetic effects as well as stratified by sex, household water fluoride, and dietary sugar and leveraged combined gene/gene-environment effects using 2-degree-of-freedom joint tests. Common genetic variants explained 24% of ECC phenotypic variance among unrelated individuals, while concordance rates were 0.64 (95% confidence interval [CI] = 0.42–0.79) among monozygotic twins and 0.44 (95% CI = 0.34–0.53) among first-degree relatives. Across all analyses, we identified 21 novel nonoverlapping genome-wide significant loci (P < 5 × 10−8) and 1 genome-wide significant gene (TAAR6) associated with ECC. The taste receptor activity gene set, with known roles in chemosensing, bacterial recognition, and innate immunity in the oral cavity, was strongly associated with ECC. While no locus remained significant after studywise multiple-testing correction, 3 loci were nominally significant (P < 0.05) and directionally consistent in external cohorts of 285,248 adults (rs1442369, DLGAP1 and rs74606067, RP11-856F16.2) and 18,994 children (rs71327750, SLC41A3). Meanwhile, the strongest marker known to be associated with adult caries (rs1122171, tagging the long noncoding RNA PITX1-AS1) was nominally significant (P = 0.01) and directionally consistent with ECC in our study. Taken together, the results of this study add to the genomics knowledge base for early childhood caries, offer several plausible candidates for future mechanistic studies, and underscore the importance of accounting for sex and pertinent environmental exposures in genetic investigations.
Keywords
Introduction
Early childhood caries (ECC) is the most common noncommunicable disease of childhood, with a reported global prevalence of 46% (Kazeminia et al. 2020). It is an early-onset form of dental disease defined by the presence of 1 or more primary tooth surfaces with caries experience in a child younger than 6 y. Efforts to better understand, treat, and prevent this persistent disease must include disentangling its social/behavioral and biological determinants and represent populations that experience high burdens of disease but may be underrepresented in research.
Dental caries is now understood as a complex dysbiotic disease resulting from the interplay between environmental and genetic etiologic factors (Divaris 2016). About a dozen genome-wide association studies (GWAS) of dental caries in children and adults have been reported; however, only 2 studies have interrogated the early-onset, severe form of disease that is captured in ECC (Borgio et al. 2022; Orlova et al. 2022). To date, 7 loci for caries in children (including those older than 6 y) have been reported, yet studies have been limited by small sample sizes, a focus on European populations (~90%), heterogeneous phenotypic characterization, and wide age intervals.
As with most complex diseases, genetic effects on ECC may differ according to environmental exposures. Nongenetic factors that play an important role in the etiology of dental caries include sugar consumption, fluoride exposure, oral hygiene, the oral microbiome, and biological sex, among others. Indeed, genetic studies have demonstrated that accounting for environmental heterogeneity can aid the detection of genetic associations that may be under the radar in main-effects analyses alone (Kraft et al. 2007). Yet, there is a paucity of genetic studies on ECC, both overall and with consideration of environmental heterogeneity. To add to the evidence base of ECC genomics, we carried out a GWAS leveraging potential gene-environment (GxE) interactions to identify genetic risk loci associated with ECC in a multiancestry population of preschool-age children.
Methods
Study Population
The analytical sample comprised a multiethnic cohort of 6,103 preschool-age children ages 3 to 5 y participating in the ZOE 2.0 study in North Carolina, United States (Appendix Fig. 1) (Divaris et al. 2020). Approximately 48% were non-Hispanic African Americans, 20% Hispanic Americans, 18% non-Hispanic Whites, among others, and 50% were females (Appendix Table 1). More information on study methodology is presented in the supplemental material (Appendix).
Phenotypes
The primary quantitative phenotype (“cavitated decay” or d3-6mfs) was defined as the number of caries-affected primary tooth surfaces (with caries lesions considered at the International Caries Detection and Assessment System [ICDAS] ≥ 3 threshold) (Pitts and Ekstrand 2013), missing or filled due to dental caries (i.e., the decayed-missing-filled surfaces [dmfs] index). Three secondary ECC traits considered were a quantitative ECC dmfs index (“clinical decay” or d1-6mfs) that included early-stage caries lesions (i.e., both cavitated and noncavitated lesions, ICDAS ≥ 1) (Ginnis et al. 2019) and 2 dichotomous ECC traits (i.e., dmfs > 0) corresponding to the 2 quantitative traits defined above (Appendix Table 2) and consistent with the international consensus definition of ECC defined as 1 or more caries-affected primary tooth surfaces in a child younger than 6 y (Pitts et al. 2019).
Genotyping and Imputation
High-density genotyping of purified DNA that was obtained from saliva was performed at the Center for Inherited Disorders Research (CIDR) at Johns Hopkins University, using the Infinium™ Global Diversity Array-8 v1.0 (Illumina). Imputation was carried out at CIDR for 6,103 unique, genotyped study participants using the Trans-Omics for Precision Medicine (TOPMed) imputation server.
Heritability and Concordance Estimates
Heritable variance (h2) of ECC attributable to all GWAS single nucleotide polymorphisms (SNPs) was estimated among 5,580 unrelated participants using genome-wide complex trait analysis using genotyped and high-quality imputed SNPs (R2 > 0.7), excluding SNPs with minor allele frequency (MAF) < 5%, and adjusting for age, sex, 8 ancestry principal components, and self-reported race/ethnicity (Yang et al. 2013). The concordance of ECC among 682 related individuals was estimated using Cohen’s kappa for the 2 dichotomous traits and intraclass correlation coefficients (ICCs) for the 2 quantitative traits.
Statistical Analyses
Considerations regarding phenotype modeling as well as accounting for population stratification and aspects of study design are elaborated in the appendix (Appendix notes). Briefly, we used 3 approaches for genome-wide association testing (Fig. 1). Approach 1 (main discovery) was a GWAS in the entire study sample, using linear and logistic regression mixed-effects models, for quantitative and dichotomous traits, respectively, adjusting for age, sex, race/ethnicity, first 8 ancestry principal components, sugary snacks/beverages, and fluoride content of household water (i.e., fixed effects) (Appendix Table 3). In the second approach (SNPjoint), we investigated single variant associations with ECC, testing the hypothesis that a variant has a main and/or interaction effect on ECC using a joint 2-degree-of-freedom (2df) test, in the full study sample and in separate models for each environmental exposure (Kraft et al. 2007). This approach leveraged potential gene-environment interaction effects in the development of ECC, accounting for the heterogeneity of genetic effects across different strata of (1) sex, (2) daily between-meal consumption frequency of high (2 or more) versus low (0–1) sugary snacks and beverages, and (3) exposure to optimal (≥0.6 ppm) versus suboptimal levels (<0.6 ppm) of domestic water-source fluoride. The third approach entailed stratified analyses, wherein the sample was split in two, for each of the 3 environmental exposures, followed by stratum-specific GWAS. For loci demonstrating genome-wide significant evidence of association in any of the previous analyses, we calculated the P value for difference (P difference) between the stratum-specific beta-coefficients of lead SNPs to screen for evidence of GxE interaction effects.
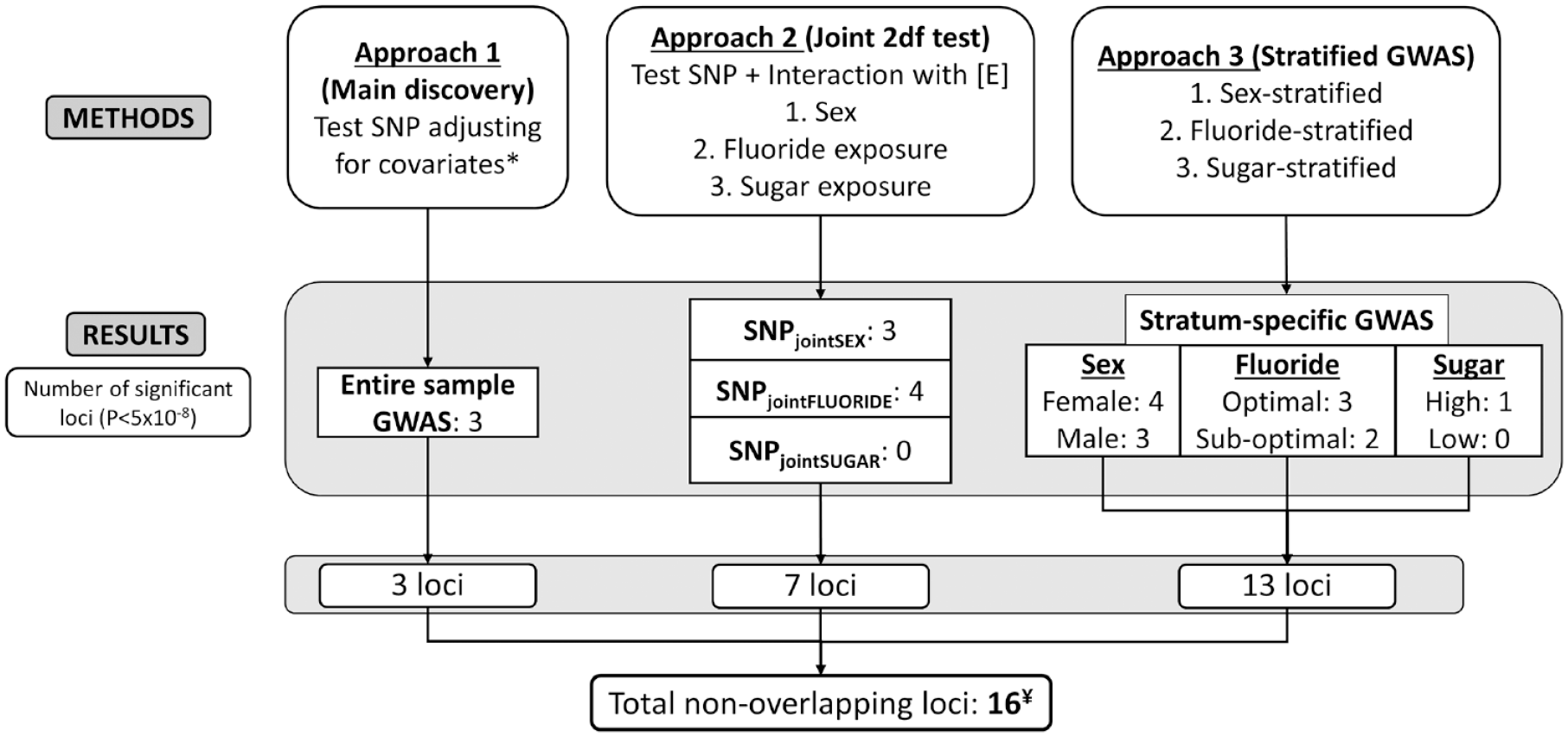
Summary of the study’s design and results. In approach 1, we conducted a GWAS in the entire sample. In approach 2, we conducted a joint 2-degree-of-freedom test to test single markers’ main and interaction effects jointly accounting for interactions with 3 environmental exposures: biological sex (male or female), fluoride exposure (optimal or suboptimal fluoride content of household water), and sugar exposure (frequency of daily consumption of sugary snacks and beverages in between meals). In approach 3, we conducted stratified GWAS for the 3 dichotomous environmental exposures. *Covariates in all association tests included age, sex, race/ethnicity, first 8 ancestry principal components, fluoride exposure, and sugar exposure. ¥Further, 5 loci were independently genome-wide significant for the secondary ECC traits (Appendix Table 7). 2df, 2-degree-of-freedom; [E], environmental factor; GWAS, genome-wide association study; SNP, single nucleotide polymorphism.
The R package EasyStrata was used to perform quality control, generate Manhattan and quantile-quantile (Q-Q) plots, and conduct 1df, 2df, and P-difference tests (Winkler et al. 2015). We used SAIGE for all genetic association analyses and accounted for relatedness using a genetic relationship matrix (Zhou et al. 2018). We excluded variants with MAF < 1%, R2 < 0.3, and those with small effective sample sizes (effN < 20 for combined and effN < 40 for stratified analyses), resulting in the test of ~14 million autosomal SNPs (Appendix Table 4). We used a conventional genome-wide statistical significance criterion of P < 5 × 10−8 to identify novel loci (Uffelmann et al. 2021) and a stringent studywide Bonferroni-corrected threshold of P < 9.6 × 10−10 to account for all analyses carried out in this study. To reduce the likelihood of reporting false positives, we report significant loci only if the lead SNP had effN ≥ 100. In addition, we explored the stability of identified signals’ effect sizes across groups of self-reported race/ethnicity, via population-specific GWAS.
Generalization
We examined the summary estimates of the genome-wide statistically significant SNPs in our study for directional consistency and Bonferroni-corrected statistical significance (P < 2.4 × 10−3) in 2 genome-wide meta-analyses of dental caries conducted among children (Haworth et al. 2018) and adults (Shungin et al. 2019) in addition to a multiethnic GWAS conducted among participants of an orofacial cleft study (Alotaibi et al. 2021). We considered the variants fulfilling both these criteria in any of these study results as generalized.
Functional Annotation
We used functional mapping and annotation of genome-wide association studies (FUMA GWAS) for functional annotation of single variant testing results (Watanabe et al. 2017). In addition, we used FATHMM-XF, HaploReg, GTeX, GeneCards, and GWAS-catalog for further biological annotations.
Gene-Based Test, Gene-Set Analysis, and Pathway Enrichment Test
MAGMA v1.6 was used to perform gene-centric analyses within the SNP2GENE process of FUMA. Genes prioritized from SNP2GENE were tested in the GENE2FUNC process using hypergeometric tests to evaluate pathway enrichment in predefined gene sets from MsigDB, WikiPathways, and the GWAS catalog.
Results
Heritability and Trait Concordance
We estimated that a quarter of the variance in the quantitative cavitated decay trait was explained by common GWAS SNPs (Table 1, h2 = 0.24, SE = 0.07, P = 9.8 × 10−5). A genome-wide interaction term for optimal fluoride exposure was statistically significant (P = 2.9 × 10−3) and resulted in the variance explained increasing to 28% (h2 = 0.28, SE = 0.07, P = 3.7 × 10−2), a 17% relative increase. A weaker genome-wide interaction was found for frequency of sugary snacks consumption (P = 7.1 × 10−2). As expected, we found higher concordance among participants with a higher level of relatedness; for example, the concordance for dmfs was 0.64 (95% confidence interval [CI] = 0.42–0.79) for monozygotic twins, 0.44 (95% CI = 0.34–0.53) for first-degree relatives, and 0.13 (95% CI = 0.03–0.23) for second- and third-degree relatives.
SNP-Based Heritability Estimates with and without the Inclusion of Gene × Environment Interaction Terms with Sugar Exposure and Fluoride Level in Household Water among Unrelated Individuals. a

ECC, early childhood caries; GxE, GxE interaction term; h2, heritability estimate; P, P value; P(G), P value for the variance of main genetic effect; P(GxE), P value for the variance of the gene-environment interaction effect; SE, standard error; SNP, single nucleotide polymorphism.
Concordance (kappa or intraclass correlation coefficients and corresponding 95% confidence intervals) of quantitative (i.e., dmfs) and dichotomous (i.e., caries affection, dmfs > 0) ECC traits among pairs of related individuals at different levels of relatedness (assessed using the kinship coefficient).
The analysis used age, sex, and race/ethnicity model-adjusted Pearson residuals.
Natural log-transformed count.
All estimates are adjusted for age, sex, race/ethnicity, and the first 8 ancestry principal components.
Genome-wide Association Analysis
Population stratification was well controlled, as evidenced by lambdas ranging between 0.98 and 1.02 (Appendix Table 5) and Q-Q plots (Appendix Fig. 2). We identified 16 genome-wide significant loci for the primary quantitative ECC trait. We identified 3 loci in the main discovery GWAS (Fig. 2). 6 additional ones in the joint main effect/GxE approach (i.e., SNPjoint [2 for SNPjointSEX, 4 for SNPjointFLUORIDE]), 5 additional ones in the sex-stratified, 1 in the fluoride-stratified, and 1 in the sugar-stratified analysis (Fig. 1). We summarize the findings from approaches 2 and 3 in Table 2 (details in Appendix Tables 6 and 7; Manhattan plots in Appendix Figs. 3 and 4).
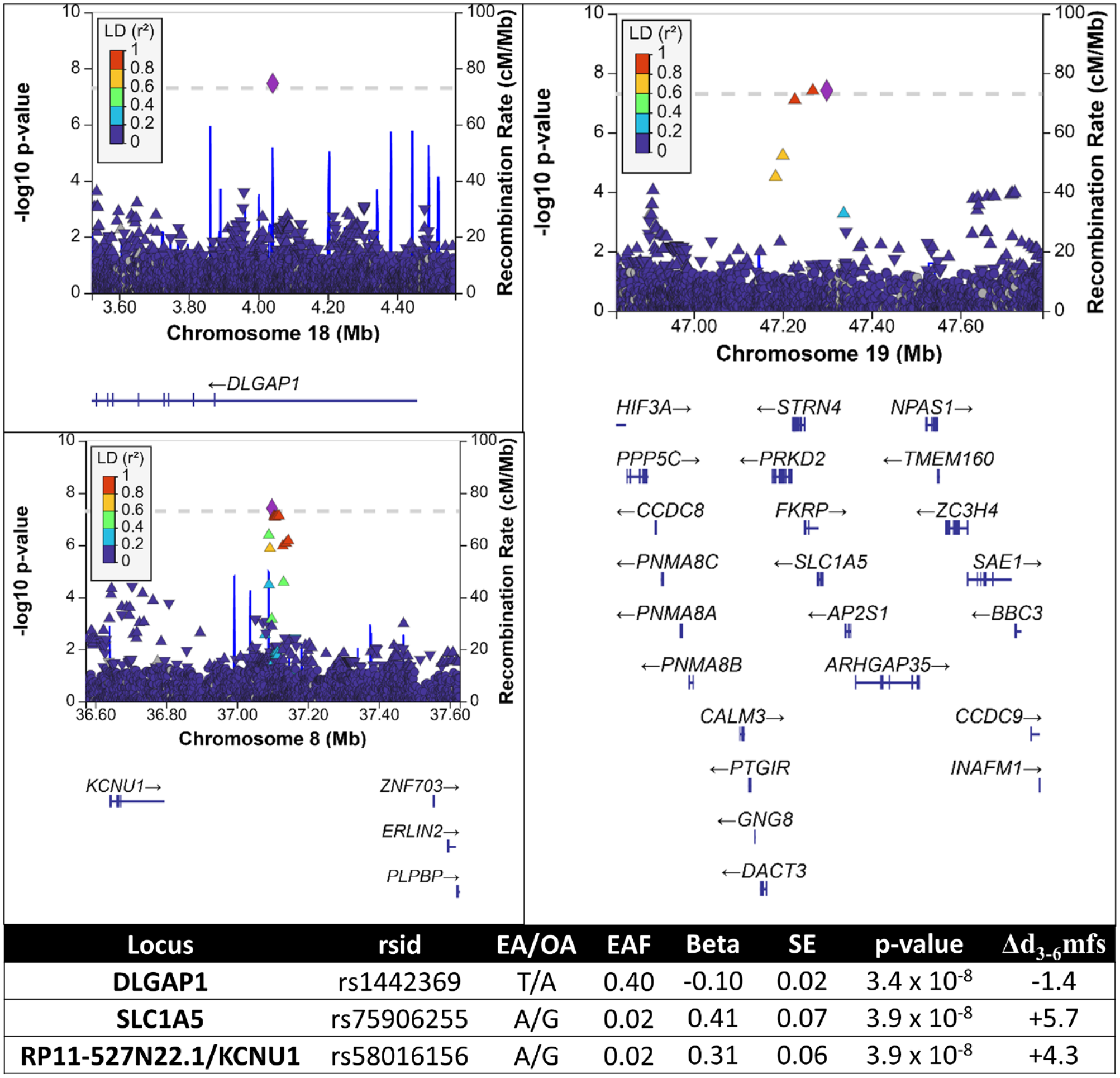
Regional association plots and summary of association results of the 3 genetic risk loci for ECC from the main discovery analysis (approach 1) in a multiancestry population of preschool-age children. (
Summary of Association Results for the 14 Loci That Met Genome-wide Significance Criteria (P < 5 × 10−8) in Analyses That Accounted for Heterogeneity by Sex, Fluoride, and Sugar Exposure, Using 2-Degree-of-Freedom Tests and Stratified Analyses.

2df, 2-degree-of-freedom; b, beta coefficient; chr, chromosome; EA, effect allele; EAF, effect allele frequency; ECC, early childhood caries; OA, other allele; P, P value; P-diff, P-difference (derived from contrasting 2 stratum-specific estimates). Boldface signifies P values less than 5 × 10–8.
Discovery GWAS
In the main discovery analysis (Appendix Table 8), we identified 3 genome-wide significant loci on chromosomes 8 (lead SNP: rs58016156), 18 (rs1442369), and 19 (rs75906255). Rs1442369 (effect allele frequency [EAF] [T]: 0.40, P = 3.4 × 10−8, beta = −0.10) is a variant intronic to DLGAP1. Rs75906255 (EAF[A]: 0.02, P = 3.9 × 10−8, beta = 0.41) is adjacent (~7 Kb) to SLC1A5 and in LD with potentially functional variants rs77394147 (R2 = 0.93, RegulomeDB score [RDB]: 2b) and rs76308698 (R2 = 0.80, CADD score [CADD]: 10.4). Rs58016156 (EAF[A]: 0.02, P = 3.9 × 10−8, beta = 0.31) is an intergenic variant adjacent to (~89 Kb) RP11-527N22.1 and 305 Kb upstream of the KCNU1 gene.
SNPjoint tests
Accounting for sex led to the identification of 2 additional loci on chromosomes 14 (rs2255032, EAF[G]: 0.03; NPAS3) and 5 (rs192232327, EAF[T]: 0.03; AC005740.4) (Table 2). Both loci were also genome-wide significant in male- and female-stratified analyses, respectively. Rs2255032 (intronic to NPAS3) has a CADD score of 11.7 and is in LD with the potentially functional variant rs74775070 (R2 = 0.67, CADD score: 17.4). Accounting for fluoride led to the identification of 4 additional loci, all of which were also genome-wide significant in fluoride-stratified analyses: chromosomes 3 (rs71327750, EAF[T]: 0.14; SLC41A3), 17 (rs650314, EAF[T]: 0.86; RP1-62O9.3), 16 (rs76985043, EAF[C]: 0.02, RP11-525K10.3), and 6 (rs3861977, EAF[T]: 0.05; ZDHHC14). Rs71327750 is intronic to SLC41A3 and in LD with multiple potential functional variants, for example, rs1077620 (R2 = 0.74, RDB: 2b), rs6796610 (R2 = 0.65, RDB: 1f), rs13100420 (R2 = 0.64, CADD: 12.6), rs4314124 (R2 = 0.65, RDB: 1f), and rs35839813 (R2 = 0.61, RDB: 2b). The less common variant, rs76985043 in RP11-525K10.3, is also in LD with multiple potentially functional variants (rs77285614 [R2 = 0.81, CADD: 10.1], rs76805928 [R2 = 0.73, CADD: 12.0], and rs79906923 [R2 = 0.75, RDB: 2b]). Accounting for sugar exposure highlighted a chromosome 2 locus (rs12052352, EAF[T]: 0.43; AC092635.1), which emerged as genome-wide significant in sugar-stratified analyses.
Stratified GWAS
The sex-stratified analysis revealed 5 additional genome-wide significant loci. Three emerged among only females (AC016970.1, AC027119.1, UBBP5) and 2 among males (BNC2, RP11-856F16.2). Fluoride-stratified analysis revealed 1 additional locus that emerged in the suboptimal fluoride stratum. The lead SNP is rs13256016 intronic to NCOA2 and in LD with potentially functional variants: rs13269274 (R2 = 0.72, CADD: 13.7) and rs11784848 (R2 = 0.71, RDB: 2b). Finally, the sugar-stratified GWAS identified a significant locus (AC092635.1) in the high-sugar stratum. Upon comparison of stratum-specific estimates, we found that most of the identified signals remained significantly different after a Bonferroni correction (Fig. 3).
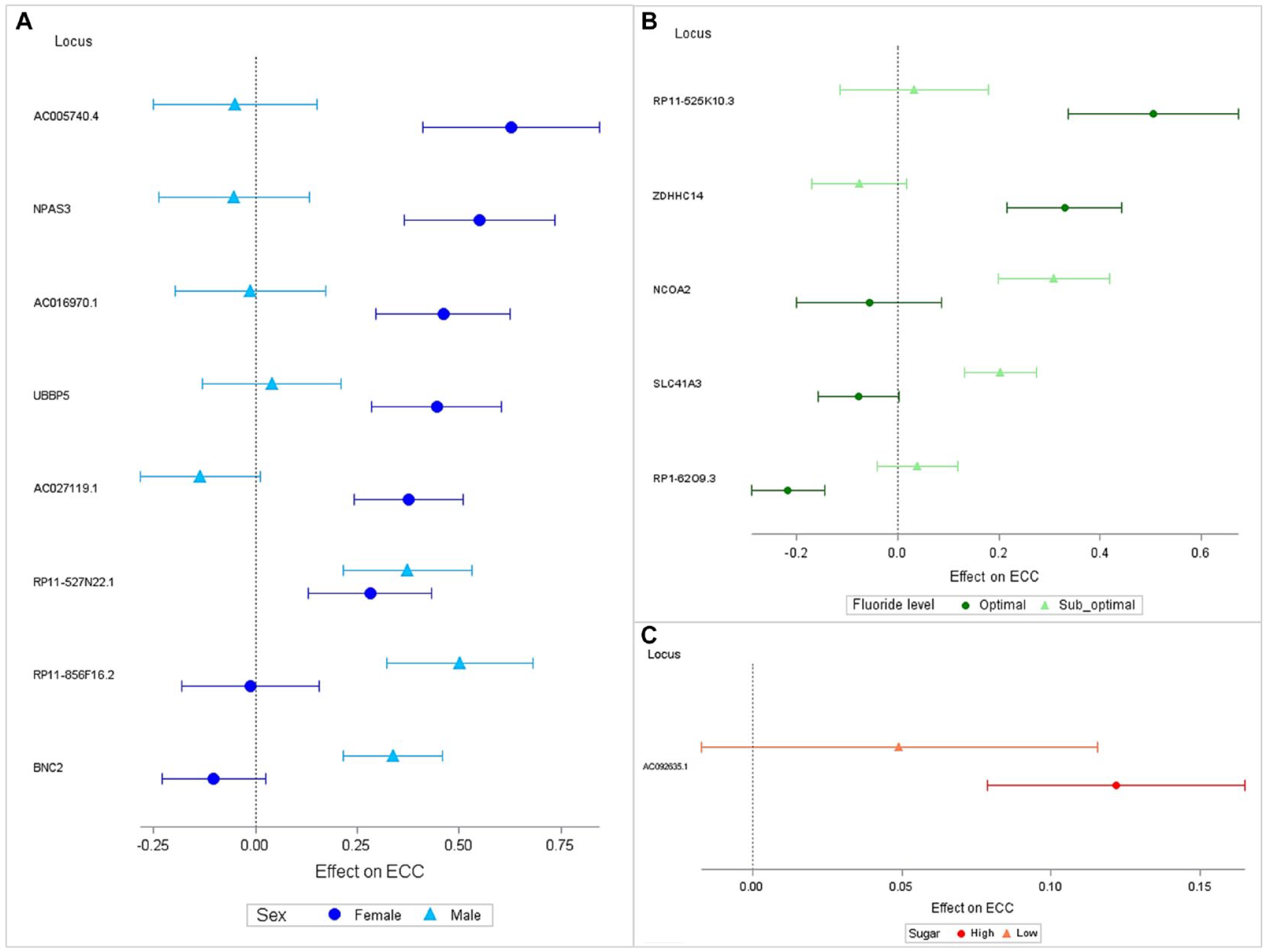
Forest plots demonstrating heterogeneity of genetic effect due to (
Secondary ECC Traits
We discovered 5 genome-wide significant loci for the 3 secondary ECC traits (Appendix Table 7). In the female-specific analysis for the dmfs trait including early-stage lesions, we identified a genome-wide significant signal led by rs4899701 (EAF[T]: 0.31, P = 3.6 × 10−8, beta = −0.17) located 10 Kb downstream of the NRXN3 (Neurexin 3) locus. This gene has been associated with obesity, autism spectrum disorder, schizophrenia, and alcohol dependence (Heard-Costa et al. 2009). In the optimal fluoride-specific analysis, the MACROD1 locus was genome-wide significant. A gene-sex interaction effect has been reported for MACROD1 in early-onset periodontitis (Freitag-Wolf et al. 2021). Furthermore, rs11231965, an intergenic variant near (~147 Kb) CTD-2555I5.1, was genome-wide significant in the joint 2df test for the sugar-stratified analysis and in the high-sugar stratum–specific analysis (EAF[G]: 0.11, P = 7.8 × 10−9, beta = −0.21, P-joint = 3.0 × 10−8). For the dichotomous cavitated decay trait (d3-6mfs > 0), we identified 1 genome-wide significant locus in females (RP11-215I16.1) and 1 in males (RP11-933H2.4/NUDT16P1; rs35487488).
Cross-Trait and Cross-Test Relevance of Identified Loci
We inspected all 21 loci’s estimates of association across all analyses and traits interrogated in this study (Appendix Table 9). While no signal remained significant after a studywise multiple-testing correction (i.e., a stringent P < 9.6 × 10−10), 7 loci were genome-wide significant in 2 analyses, for example, rs58016156 in the RP11-527N22.1/KCNU1 locus in the main GWAS and the joint test for the sex-stratified analysis. Of note, SLC1A5 and RP11-527N22.1/KCNU1 were associated with ECC at a suggestive significance level (P < 5 × 10−6) in 7 and 6 different analyses, respectively.
Population-Specific Subanalyses
We examined the stability of the 21 identified associations across strata of self-reported race/ethnicity and found an overarching consistency, with 16 loci demonstrating directionally consistent associations in these subanalyses (Appendix Table 10).
Generalization in External Cohorts of Children and Adults and Previously Reported Loci
While no locus generalized after a multiple-testing correction, 3 signals were nominally significant (P < 0.05) and directionally consistent in the external cohorts. These were rs1442369 (DLGAP1) and rs74606067 (RP11-856F16.2) in the GLIDE-adults cohort (EAF[A]: 0.53, P = 0.03, N = 285,246 and EAF[T]: 0.06, P = 0.01, N = 285,248), rs71327750 (SLC41A3) in the GLIDE-children cohort examining caries in primary teeth (EAF[T]: 0.20, P = 0.02, N = 18,994) (Appendix Tables 11–13), and DLGAP1 in the permanent dentition caries analysis in the multiethnic orofacial cleft GWAS (Appendix Table 14).
Among previously reported risk loci (P < 5 × 10−8) for caries (Appendix Table 15), rs1122171 (PITX1-AS1), a variant showing the strongest known GWAS association with caries in adults (Shungin et al. 2019), was nominally significant and directionally consistent with ECC (dmfs) in our study. We did not find any associations listed for the statistically significant variants from our study or their proxies (R2 ≥ 0.8) in the GWAS-catalog.
Functional annotation results are presented in Appendix Table 16. We considered and summarized all protein-coding genes within ~250 Kb of the independently significant variants (research data). We identified several genes near genome-wide significant loci with potential roles in the development of dental tissues. For example, SPRY4 near rs192232327 antagonizes fibroblast growth factor (Klein et al. 2006), which is important at different stages of tooth development (Thesleff 2006), and PHOSPHO1 near rs650314 is involved in mineralization (Pandya et al. 2017).
Gene-Based, Gene-Set, and Pathway Enrichment Analyses
We identified 1 genome-wide significant gene for the primary ECC trait, TAAR6 (Appendix Table 17 and Appendix Fig. 5) and 11 genome-wide significant gene sets for primary and secondary ECC traits (Appendix Table 18) including “taste receptor activity.” We also identified several enriched curated gene sets, including the positional gene set chr20q12 (Appendix Table 19); PLCG1, ZHX3, LPIN3, EMILIN3, and CHD6 were the prioritized genes overlapping with this gene set. Imhof and colleagues (2020) demonstrated an upregulation of EMILIN-3 in dentin caries lesions.
Discussion
In this GWAS among a well-characterized, community-based, multiethnic cohort of preschool-age children, we demonstrate that a quarter of the variance in ECC can be explained by common genetic variation and that the joint consideration of established environmental factors such as sugar consumption and fluoride exposure increases the phenotypic variance explained. We leveraged approaches accounting for gene-environment interactions to detect 21 novel loci for ECC, 3 of which were nominally significant and directionally consistent in external, independent populations of adults and children. While no locus met studywide multiple-testing corrected discovery and generalization statistical significance criteria, the study offers several novel candidates with plausible biological roles in the pathogenesis of ECC that are promising targets for future investigations.
The most prominent novel identified loci included DLGAP1, SLC1A5, and KCNU1. Rs1442369 (DLGAP1) and rs75906255 (SLC1A5) showed relatively consistent evidence of association in all stratified analyses and DLGAP1 nominally generalized in the external cohorts of adults. Consistent with the majority (~93%) of disease- and trait-associated variants emerging from GWAS being in noncoding sequences and likely involved transcriptional regulatory mechanisms (Maurano et al. 2012), these lead SNPs were in noncoding regions—their functional implications or regulatory roles will need to be further studied. The KCNU1 locus (rs58016156) is related to the sweet taste signaling pathway (Safran et al. 2021). The finding of genome-wide evidence of the association between ECC, this gene, and the “taste receptor activity” gene set in general is crucial. Taste receptors have broad chemosensory properties beyond taste throughout the body, possess an ability to detect bacterial communications (Kouakou and Lee 2023), and have key roles with relevance to innate immunity and oral health (Xi et al. 2022). For example, the bitter taste receptor T2R14 is expressed in gingival epithelial cells (GEC) and was recently found to sense Streptococcus mutans’ competence stimulating peptides, thereby mediating GEC’s innate immune responses (Medapati et al. 2021).
It is noteworthy that the strongest GWAS marker known to be associated with adult caries (rs1122171, tagging the long noncoding RNA PITX1-AS1) was nominally significant and directionally consistent with ECC in our study—adding to the evidence base supporting its role in dental caries across the life course. None of the previously reported GWAS loci for childhood caries were generalized in our study (Appendix Table 20). The sample size of ~6,000, the participants’ narrow age range (3–5 y), and the detailed phenotypic characterization enabling inquiry of quantitative traits at different detection thresholds are relative improvements. The inclusion of traditionally underrepresented racial/ethnic backgrounds in this study addresses some equity issues in oral and genetic research and has been shown to confer analytical advantages (Agler and Divaris 2020; Lin et al. 2021). Meanwhile, several clinical subtypes of ECC have been proposed based on expert opinion (e.g., severe ECC; Drury et al. 1999) and data-driven analyses of large population data (Simancas-Pallares et al. 2023). Because the etiology of ECC subtypes may vary, future GWAS should seek to formally investigate differences in their genetic underpinning.
The study was limited by its small sample sizes in stratified analyses, multiple populations introducing ancestry latent structures (Haworth et al. 2019), and the utilization of multiple correlated phenotypes that increase the risk of false-positive findings. We must also acknowledge that the correlation between genetic variants and environmental exposures such as fluoride in drinking water can also induce false positives (Lindström et al. 2009).
Among biological factors, sex-related differences may partly explain the difference in caries experience (Klein and Palmer 1938; Lukacs and Largaespada 2006). Only 1 study to date has interrogated gene-sex interactions in dental caries comparing sex-specific heritability estimates and genetic correlations (Shaffer et al. 2015). In our study, we leveraged sex heterogeneity via joint tests and discovered 2 additional independent signals including SPRY4 (suggestively associated with erosive tooth wear; Alaraudanjoki et al. 2019) and tyrosine kinase signaling, expressed in the mesenchyme of the tooth germs (Klein et al. 2006; Thesleff 2006). MTRR, near the LOC729506 locus, was identified in the female-specific GWAS and has been associated with ECC and being underweight in a candidate gene study (Antunes et al. 2017). Finally, PHOSPHO1, a gene that emerged in fluoride-stratified analyses, is involved in bone mineralization and maturation, plays a role in mineralization of enamel (Pandya et al. 2017), and is associated with childhood hypophosphatasia (Reibel et al. 2009).
Taken together, the results of this study add to the genomics knowledge base for ECC, offer several plausible candidates for future mechanistic studies, and underscore the importance of accounting for sex and pertinent environmental exposures in genetic investigations. Validation of these findings in future investigations, in different populations, and using mechanistic studies is warranted to establish the identified loci as replicable and causal.
Author Contributions
P. Shrestha, contributed to analysis and interpretation, drafted and critically revised the manuscript; M. Graff, contributed to analysis and interpretation, critically revised the manuscript; Y. Gu, Y. Wang, contributed to data analysis, critically revised the manuscript; C.L. Avery, D.Y. Lin, M.L. Marazita, contributed to data interpretation, critically revised the manuscript; J. Ginnis, M.A. Simancas-Pallares, contributed to data acquisition, critically revised the manuscript; A.G. Ferreira Zandoná, G.D. Slade, contributed to data design, critically revised the manuscript; R.N. Alotaibi, E. Orlova, H.S. Ahn, K.N. Nguyen, H.M. Highland, contributed to data analysis, critically revised the manuscript; J.S. Preisser, contributed to design, data interpretation, critically revised the manuscript; K.E. North, contributed to conception and design, data interpretation, critically revised the manuscript; K. Divaris, contributed to conception and design, data acquisition and interpretation, critically revised the manuscript. All authors gave their final approval and agreed to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345241291528 – Supplemental material for Multiancestry Genome-Wide Association Study of Early Childhood Caries
Supplemental material, sj-docx-1-jdr-10.1177_00220345241291528 for Multiancestry Genome-Wide Association Study of Early Childhood Caries by P. Shrestha, M. Graff, Y. Gu, Y. Wang, C.L. Avery, J. Ginnis, M.A. Simancas-Pallares, A.G. Ferreira Zandoná, R.N. Alotaibi, E. Orlova, H.S. Ahn, K.N. Nguyen, H.M. Highland, D.Y. Lin, J.S. Preisser, G.D. Slade, M.L. Marazita, K.E. North and K. Divaris in Journal of Dental Research
Supplemental Material
sj-xlsx-1-jdr-10.1177_00220345241291528 – Supplemental material for Multiancestry Genome-Wide Association Study of Early Childhood Caries
Supplemental material, sj-xlsx-1-jdr-10.1177_00220345241291528 for Multiancestry Genome-Wide Association Study of Early Childhood Caries by P. Shrestha, M. Graff, Y. Gu, Y. Wang, C.L. Avery, J. Ginnis, M.A. Simancas-Pallares, A.G. Ferreira Zandoná, R.N. Alotaibi, E. Orlova, H.S. Ahn, K.N. Nguyen, H.M. Highland, D.Y. Lin, J.S. Preisser, G.D. Slade, M.L. Marazita, K.E. North and K. Divaris in Journal of Dental Research
Footnotes
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: During the preparation of this manuscript, J.S. Preisser served on a data safety and monitoring board of a study funded by the National Institute of Dental and Craniofacial Research (NIDCR). The remaining authors declare no competing interests.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by research grants from the National Institutes of Health, National Institute for Dental and Craniofacial Research U01DE025046 (K.D., A.G.F.Z, D.Y.L, J.S.P, G.D.S., K.E.N.) and the National Human Genome Research Institute X01HG010871 (K.D.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the funders.
The authors thank CIDR investigators and staff at Johns Hopkins University for carrying out genotyping and imputation for the project with support from a resource-allocation grant (NIH/NIHGR X01-HG010871); Dr. Patricia V. Basta and her team at the UNC-Chapel Hill Biospecimen Processing facility for the accessioning, storage, and disbursement of the saliva and extracted nucleic acid samples in the ZOE studies; and all study participants and their families for their contributions.
A supplemental appendix to this article is available online.
Data Availability
Genotype and phenotype data for ZOE 2.0 are publicly available via phs002232.v1.p1 “TOPDECC-Trans-omics for Precision Dentistry and Early Childhood Caries: Genome-Wide Genotyping (CIDR) and Microbiome in the ZOE 2.0 Study.” Genomic summary results from the GLIDE adults study are available at https://data.bris.ac.uk/data/dataset/2j2rqgzedxlq02oqbb4vmycnc2 and GLIDE-children at ![]() .
.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
