Abstract
Tissue-specific immune responses are critical determinants of health-maintaining homeostasis and disease-related dysbiosis. In the context of COVID-19, oral immune responses reflect local host-pathogen dynamics near the site of infection and serve as important “windows to the body,” reflecting systemic responses to the invading SARS-CoV-2 virus. This study leveraged multiplex technology to characterize the salivary SARS-CoV-2–specific immunological landscape (37 cytokines/chemokines and 11 antibodies) during early infection. Cytokine/immune profiling was performed on unstimulated cleared whole saliva collected from 227 adult SARS-CoV-2+ participants and 37 controls. Statistical analysis and modeling revealed significant differential abundance of 25 cytokines (16 downregulated, 9 upregulated). Pathway analysis demonstrated early SARS-CoV-2 infection is associated with local suppression of oral type I/III interferon and blunted natural killer–/T-cell responses, reflecting a potential novel immune-evasion strategy enabling infection. This virus-associated immune suppression occurred concomitantly with significant upregulation of proinflammatory pathways including marked increases in the acute phase proteins pentraxin-3 and chitinase-3-like-1. Irrespective of SARS-CoV-2 infection, prior vaccination was associated with increased total α-SARS-CoV-2-spike (trimer), -S1 protein, -RBD, and -nucleocapsid salivary antibodies, highlighting the importance of COVID-19 vaccination in eliciting mucosal responses. Altogether, our findings highlight saliva as a stable and accessible biofluid for monitoring host responses to SARS-CoV-2 over time and suggest that oral-mucosal immune dysregulation is a hallmark of early SARS-CoV-2 infection, with possible implications for viral evasion mechanisms.
Introduction
The oral cavity has emerged as an important site for SARS-CoV-2 infection, and oronasal immunity is a critical first line of defense against COVID-19 (Azkur et al. 2020; Isho et al. 2020; Silva et al. 2021). Disease severity is related to the nature and magnitude of the immunological response to SARS-CoV-2 and is modified by several risk factors, including individual susceptibility, preexisting comorbidities, oral microbiome, as well as viral load and infectivity (Lucas et al. 2020; Williamson et al. 2020). Most studies have sought to define disease mechanisms and host defenses based on systemic responses to infection (Cohen et al. 2023); however, understanding local, tissue-specific immune responses is critically important to uncovering foundational principles of host-pathogen interactions in health and disease (Isho et al. 2020; Alqedari et al. 2023). Further, leveraging saliva as a readily available biofluid holds great public health promise for rapidly diagnosing infection, monitoring disease prevalence, predicting severity, and ascertaining previous exposure; this is particularly relevant as vaccine-resistant variants emerge (Fernandes et al. 2020).
In this study, we aimed to characterize the immunological landscape of the oral cavity during early SARS-CoV-2 infection by comparing the salivary cytokine and antibody profile of COVID-19+ subjects to a control population to understand oral conditions supportive of SARS-CoV-2 infection. Our findings suggest that oral-mucosal immune dysregulation is a previously undescribed hallmark of early SARS-CoV-2 infection, with possible implications for viral evasion mechanisms. Understanding innate and adaptive defense mechanisms in the oral cavity during the early course of SARS-CoV-2 infection holds potential to identify novel diagnostic, prognostic, and therapeutic biomarkers.
Methods
Study Design and Human Subjects
A cross-sectional study was conducted among 314 adults 18 to 65 y, who were enrolled in 2 outpatient randomized controlled trials evaluating oral rinses for impacts on the SARS-CoV-2 virus (identifiers: NCT04584684 [posted October 2020, enrolling December 2020 to October 2021]; NCT05178173 [posted January 2022, enrolling January 2022 to July 2023]) (Appendix Table 1). Participants had physician- or laboratory-confirmed SARS-CoV-2+ via polymerase chain reaction (PCR) or antigen nasopharyngeal swab testing within 10 d of participation (n = 277). Controls (n = 37) were age and gender matched to the SARS-CoV-2+ cohort and tested negative by nasopharyngeal antigen and salivary quantitative PCR assays. Controls were vaccinated against COVID-19, and most had been infected previously but reported having no known infection within 2 mo of participation. Additional details are provided in the appendix.
Sample Collection and Processing
These studies were approved by the UNC Institutional Review Board (#20-2040, #21-3063, #21-2019). Subject data were collected using questionnaires and medical charts including demographics, medical history/comorbidities, oral hygiene practices, COVID-19–related symptomatology, and treatment course (e.g., diagnosis, medications) (Table 1). Participants provided unstimulated saliva samples collected via the passive drool self-collection method; 1 mL aliquots were prepared on the collection day and stored at −80 °C until synchronized batch processing for downstream assays (Fig. 1A). Additional details are provided in the appendix.
Participant Demographics.

Data are presented as number of participants (% of all participants) unless otherwise noted.
Statistical analysis performed using the chi-square test.
Statistical analysis performed using the Wilcoxon test.
Data are presented as mean ± standard deviation.
Potential variants were assessed based on a literature review, looking at the date ranges when a certain variant was more prevalent among the United States population.
Participant received either both doses of a 2-dose series (e.g., Pfizer-BioNTech, Moderna SpikeVax, Novavax) or the single dose of a 1-dose series (e.g., Johnson & Johnson) and at least a single booster dose prior to sample collection.
Participant received both doses of a 2-dose series (e.g., as above) or the single dose of a 1-dose series (e.g., Johnson & Johnson) but did not receive a booster vaccination.
Participant received only a single dose in a 2-dose series (e.g., Pfizer-BioNTech, Moderna SpikeVax, Novavax).
COVID-19 prescription medication information as reported in medical charts, where available.
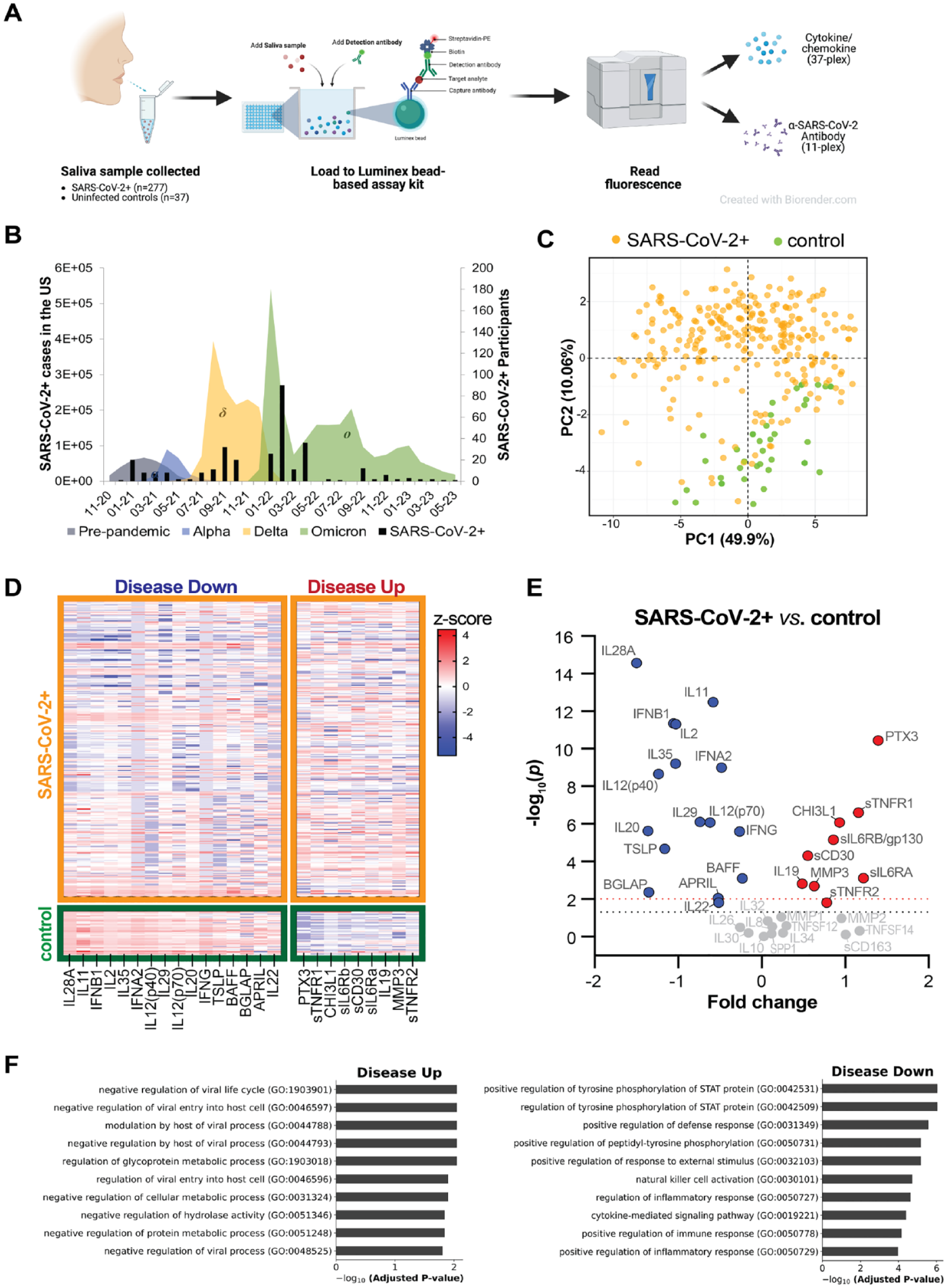
Distinct salivary cytokine and antibody profiles associated with early SARS-CoV-2+ infection. (
Soluble Mediator Analysis
Salivary cytokine abundance was determined using the Bio-Plex Pro Human Inflammation Panel 1, 37-Plex kit (Bio-Rad) without additional sample dilution; salivary SARS-CoV-2+–specific antibodies (Ab) were measured using the Coronavirus Ig Total Human 11-Plex ProcartaPlex Panel (Invitrogen, ThermoFisher Scientific) after 1:1 sample dilution in assay buffer (Appendix Table 2). Assays were performed according to the manufacturer’s instructions and saliva-specific processing recommendations. A 5-parameter logistic regression function (5PL) and 9- (cytokine) or 7-point (Ab) standard curve were used to determine soluble mediator (pg/mL) or Ab (U/mL) concentrations, respectively. Assays were performed on a Bio-Plex-200 platform; Bio-Plex Manager (V6.1) was used for data acquisition and analysis (BioRad). Additional details on pathway analysis are provided in the appendix.
Statistical Analysis
Statistical analyses and graphical visualization were performed using R4.2.2 and GraphPad Prism (V9.5.1). Principal component analysis (PCA) was performed on all log10[pg/mL] values derived from the cytokine 37-plex for all participants comparing SARS-CoV-2+ to control. Patients were grouped by days post–positive test (PT_days), days post–symptom onset (ST_days), and vaccination type (details in the appendix). As we did not assume a distribution, nonparametric 2-sample Wilcoxon rank-sum tests were used to perform comparisons between patient groups or for each pair of the 4 time or vaccine groups. Kruskal-Wallis tests were used for comparisons among multiple time groups and, separately, among vaccine groups. Bonferroni multiple testing adjustment was used to adjust the P values from tests. P ≤ 0.05 was considered significant.
Multiple linear regression
To rigorously compare the level of cytokine and/or Ab concentration (as the dependent variable) between SARS-CoV-2 and control, multiple linear regression was used to adjust the confounding effects from other clinical/demographic variables (with age, sex, days post–positive test, days post–symptom onset, and vaccination status as covariates). Confounders were adjusted by including them in the regression model. Interaction terms involving SARS-CoV-2 status and 2 main covariates (age, gender) were incorporated to assess potential interaction. Resulting coefficient estimates of the multiple linear regression indicate the average expression change comparing SARS-CoV-2+ with controls, with adjustment of all other covariates. This regression model was used to assess the SARS-CoV-2 effect and time effects separately. Additional details are provided in the appendix; statistical tests corresponding to each figure and table are included in legends.
Results
Study Participants
Participant demographics were representative of the local population, including a plurality of European descent, female, and non-Hispanic participants (Table 1). Ages ranged from 18 to 65 y (COVID+: mean 35.2 y, median 32 y; control: mean 34.7 y, median 28 y). On average, SARS-CoV-2+ participants developed symptoms 7 d and tested positive 5 d prior to saliva collection, with a vast majority as outpatients (96%) and not receiving treatment (87%). Of the 13% SARS-CoV-2+ participants who received treatment, most received antiviral therapy. Most participants were vaccinated (73%) and experienced rhinorrhea (74%), cough (72%), and/or fatigue (57%). Several COVID-19 waves and variants are putatively represented, with majority representation likely from the Omicron variant (59%) (Fig. 1B).
SARS-CoV-2+ Is Associated with Altered Abundance of Salivary Immune Mediators
PCA of normalized 37-plex cytokine values revealed a robust separation between SARS-CoV-2+ and control groups, in which the first principal component (PC1) represents the largest variability in the data (49.9% of total variance) and PC2 accounts for 10.06% of total variance (Fig. 1C). Significant differences in the salivary abundance of 25 soluble mediators were observed comparing SARS-CoV-2+ and control (Figs. 1D–E, 2A–Y; Table 2; Appendix Fig. 1A–L). Of these, 64% (16 soluble mediators) were found to be less abundant in the saliva of SARS-CoV-2+ subjects, including type I, II, and III interferons (IFNs; IFN-α2*, IFN-β; IFN-γ*; IFN-λ1/IL-29, IFN-λ2/IL-28A*), multiple interleukin (IL)–20 subfamily cytokines (IL-20*, IL-22*), IL-12 family cytokines (IL-12(p40)*, IL-12(p70)*, IL-35*), and tumor necrosis factor (TNF) ligand superfamily members (APRIL*, BAFF*). Reduced salivary levels of IL-2, IL-11*, thymic stromal lymphopoietin (TSLP*), and osteocalcin/BGLAP were also observed (Fig. 1E, Table 2, Appendix Fig. 2). In contrast, SARS-CoV-2+ was associated with increased salivary abundance of 9 soluble mediators, including proinflammatory soluble receptors (sTNF-R1*, sTNF-R2*, sIL-6Rα* and sIL-6Rβ/gp130*), chitinase 3-like-1 (CHI3L1), soluble CD30 (sCD30)*, matrix metalloproteinase-3 (MMP-3)*, and IL-19*. Notably, we also observed a striking and significant upregulation of pentraxin-3 (PTX3)* that was specific to SARS-CoV-2+ subjects and undetectable in most controls (Fig. 1E, Table 2
Cytokine and Antibody Levels Comparing SARS-CoV-2+ and Control Subjects.

Cytokines (37-plex) and antibodies (11-plex) are shown in the left-most column. Marginal effect: Wilcoxon P values were derived from Wilcoxon rank-sum tests comparing SARS-CoV-2+ versus control. Nominal P values were derived from adjusted multiple linear regression models. Estimate subcolumn (left): directional estimates shown indicate the directionality of computed differences comparing SARS-CoV-2+ versus control; a positive estimate indicates values are increased in SARS-CoV-2+ participants relative to controls, whereas a negative estimate indicates values are decreased in SARS-CoV-2+ participants relative to controls. Interaction effect: For the “Interaction Variables” subcolumn, each potential interaction variable (age, gender, days post–symptom onset [ST_days], and days post–positive test [PT_days]) was tested to determine whether the variable significantly influenced the SARS-CoV-2+ effect on cytokines and antibodies observed. Only significant interaction variables are listed in the “Interaction Variables” subcolumn, and corresponding P values are shown in the “Interaction P Value” subcolumn, where P ≤ 0.05; insignificant variables are designated as NA. Estimate subcolumn (right): directional estimates shown indicate the directionality of interaction effects. For example, salivary IL-2 is significantly reduced in SARS-CoV-2+ participants but how much it decreases varies by the age of the participant (Appendix Fig. 2). Additional details on variables and the statistical approach are included in the appendix.
P ≤ 0.05. Exact P values are shown.
Levels of 28 cytokines positively correlated with SARS-CoV-2 viral load, including 12 downregulated and 8 upregulated cytokines when comparing SARS-CoV-2+ to control, indicated by * above. In addition, positive correlations were found for MMP-1, MMP-2, sCD163, IL-32, IL-34, osteopontin/SPP1, LIGHT/TNFSF14, and IL-10 (Appendix Fig. 3). No cytokines were negatively associated with viral load.
Vaccination Imparts Significant Elevation in α-SARS-CoV-2 Salivary Antibodies and Differentially Affects Soluble Mediators
A multiplex strategy was also used to determine α-SARS-CoV-2–specific salivary Ab abundance comparing SARS-CoV-2+ to controls and separately comparing controls, unvaccinated and vaccinated SARS-CoV-2+ patients. Wilcoxon rank-sum tests revealed that SARS-CoV-2+ participants exhibited reduced salivary abundance of total α-SARS-CoV-2-spike (trimer), -S1 protein, and -nucleocapsid Ig (Table 2, Fig. 2AA–AC, Appendix Fig. 1M–R). Vaccinated participants (SARS-CoV-2+ and controls) exhibited higher total α-SARS-CoV-2-spike (trimer), -S1 protein, -RBD, and -nucleocapsid Ig than unvaccinated SARS-CoV-2+ patients (Appendix Table 3, Appendix Fig. 4).
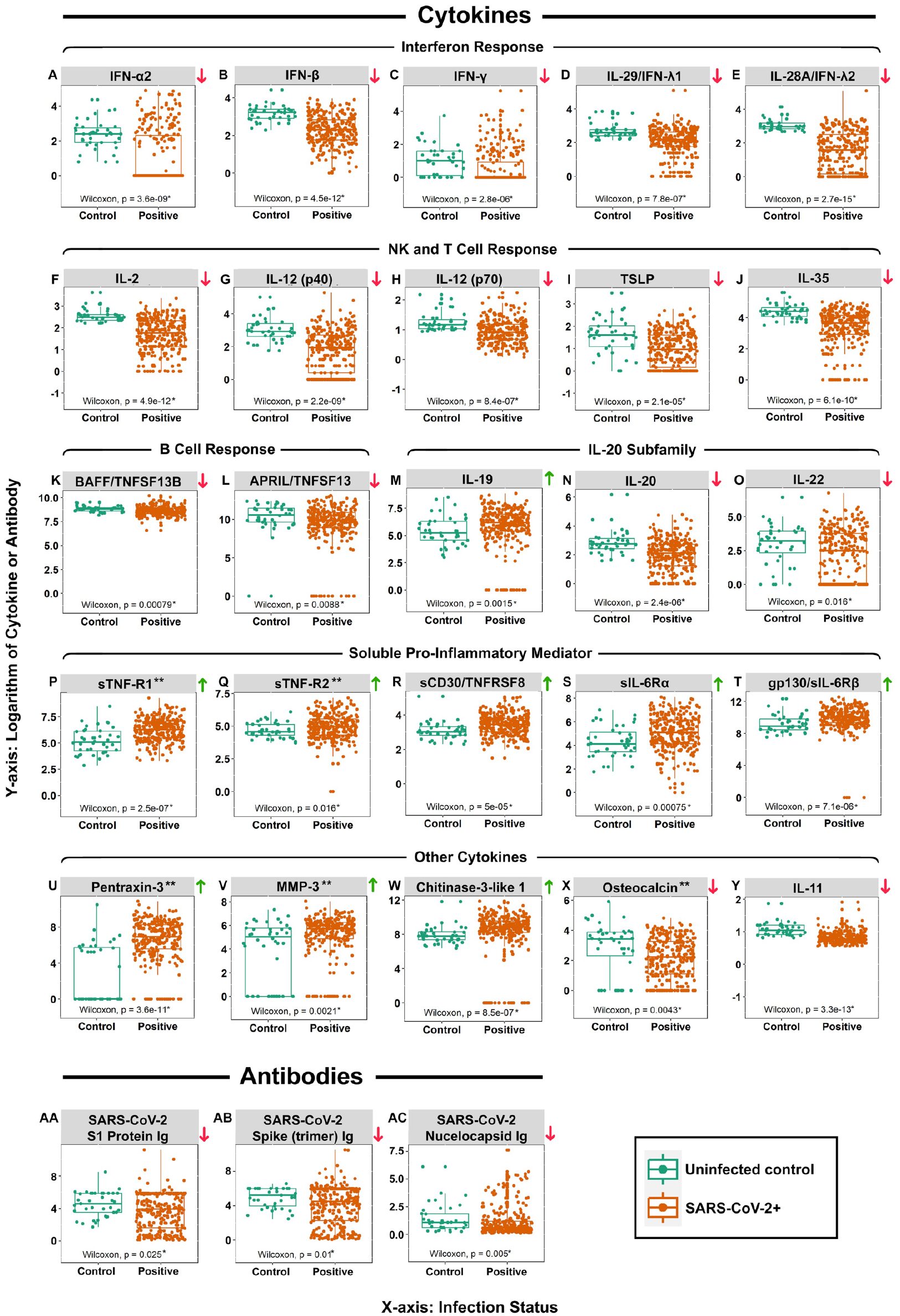
Significant cytokines and antibodies boxplots. Box-and-whisker plots of normalized log10[U/mL Ab] and log10[pg/mL cytokine] values determined to be significantly different (P ≤ 0.05; Wilcoxon rank-sum test) comparing control (green) and SARS-CoV-2+ (orange) subjects. Exact P values for each comparison are shown at the bottom of each plot. Arrows to the immediate right of each plot indicate the mediator is increased (green arrows) or decreased (red arrows) in the SARS-CoV-2 group. (
When comparing whether salivary α-SARS-CoV-2 Abs or cytokines varied as a function of vaccination status/type among SARS-CoV-2+ participants, vaccination was associated with increased total α-SARS-CoV-2-spike (trimer), -S1 protein, and -RBD Ig, irrespective of vaccine type/supplier (Fig. 3A–D). Although vaccine type did not correlate with most investigated cytokines, we observed subtle yet significant differences in levels of IL-12(p40), IL-27(p28), and IFN-λ1/IL-29 between vaccine groups (Fig. 3E–G, Appendix Figs. 5 and 6).
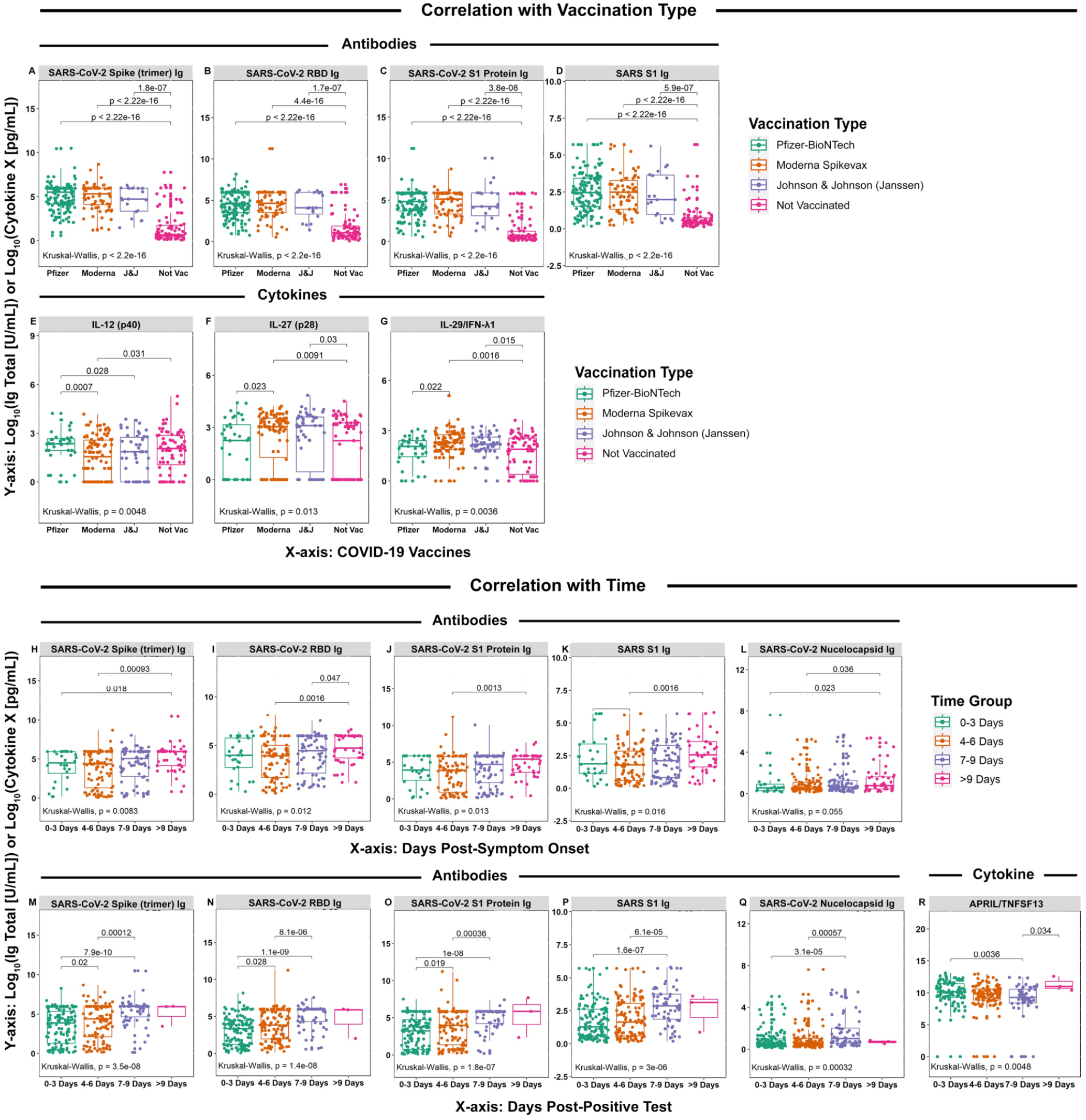
Antibody and cytokine levels relative to vaccination status, days post–symptom onset, and days post–positive test with significant trends, for SARS-CoV-2 subjects. Box-and-whisker plots of normalized log10[U/mL Ab] and log10[pg/mL cytokine] values as indicated above each group of plots among SARS-CoV-2+ participants relative to COVID-19 vaccination status and type (
When evaluating whether SARS-CoV-2 viral load affected total salivary α-SARS-CoV-2 Ig abundance in SARS-CoV-2+ participants, increased viral load was negatively correlated with total α-SARS-CoV-2-spike (trimer), -RBD, -S1 protein, and -nucleocapsid Ig (Appendix Fig. 7).
Adjusted Modeling as a Maximally Conservative Approach
Following the primary analyses (Figs. 1 and 2), random effect multiple regression analysis that had participants as the random term was used to account for possible confounding variables arising within large datasets (including age, sex, days post–positive test, days post–symptom onset, and vaccination status) (Nørgaard et al. 2017). In these analyses, significant nominal P values were obtained for 12 cytokines, including 9 reduced in SARS-CoV-2+ (IFN-β, IL-11, IL-20, IL-2, IL-35, IL-12[p40], IL-12[p70], TSLP, osteocalcin/BGLAP) and 3 increased in SARS-CoV-2+ (MMP-3, sCD30/TNFRSF8, sTNF-R2) relative to controls. There were no differences between controls and SARS-CoV-2+ participants’ salivary Ab (Table 2). These data are consistent with the initial analyses and are maximally conservative.
Dynamic Oral Inflammatory Responses during Early SARS-CoV-2+ Infection
Next, we used a linear regression model to determine whether salivary immune mediators or α-SARS-CoV-2 Abs varied as a function of time (e.g., days post–symptom onset or days post–positive test) during the early course of infection. Salivary cytokine concentration remained similar for both time variables except for APRIL/TNFSF13, which changed as a function of days post–positive test. This occurred concomitant with an increased salivary α-SARS-CoV-2-spike (trimer), -RBD, -S1 protein, and -nucleocapsid Ig, which positively correlated with days post–symptom onset and days post–positive test (Fig. 3H–R, Appendix Tables 4 to 7).
Discussion
Dysregulated systemic immune responses to SARS-CoV-2 are appreciated to underlie COVID-19 severity, mortality, and development of long COVID, where the degree of immune dysfunction appears to correlate with disease severity (Silva et al. 2021). Although hyperinflammatory systemic responses have been described in both mild and severe COVID-19, SARS-CoV-2 also appears to induce immune suppression as a novel evasion strategy; indeed, recent studies indicate that both innate and adaptive immune responses are dysregulated in COVID-19 (Azkur et al. 2020; Qin et al. 2020).
As a biofluid, saliva is a promising, noninvasive diagnostic and prognostic tool for a variety of conditions and diseases, including COVID-19 (Song et al. 2023), where higher viral load correlates with greater disease severity and mortality (Fajnzylber et al. 2020). In addition, salivary mediator analysis can yield important insights into fundamental principles of mucosal homeostasis and host-pathogen interactions (Alqedari et al. 2023). In this study, we leveraged multiplex technology to characterize the impact of SARS-CoV-2 on the oral immune landscape and observed that infection was associated with significant alterations to multiple innate and adaptive immune pathways, relevant to mucosal immune responses.
Multiple IFNs exhibited reduced salivary abundance in early SARS-CoV-2 infection (Fig. 1). Type I (IFN-α2, IFN-β), type II (IFN-γ), and type III (IFN-λ1/IL-29, IFN-λ2/IL-28A) IFNs are potent initiators of antiviral immune responses and provide robust protection at mucosal interfaces (Lazear et al. 2019). Dysregulated type I IFN responses are a hallmark of COVID-19, and type I IFN evasion is a critical virulence property of SARS-CoV-2 (Xia et al. 2020); reduced type I and II IFN in COVID-19 nasopharynx samples have also been reported (Lazear et al. 2019). Intriguingly, among our vaccinated participants, those receiving Moderna had higher levels of IFN-λ1/IL-29 relative to those receiving Pfizer. Moderna recipients are at elevated myocarditis risk compared with Pfizer recipients, although natural infection poses the highest risk of myocarditis (Gargano et al. 2021; Marschner et al. 2023). Given that IFN-λ is the predominant inflammatory cytokine involved in the pathogenesis of myocarditis (Cosper et al. 2012; Tran et al. 2024), future investigations should explore potential interactions among SARS-CoV-2 infection, host responses, and vaccination. Altogether, these data demonstrate that SARS-CoV-2 impairs early host antiviral immune responses through IFN pathway suppression and that modulation of IFN responses is a critical component of SARS-CoV-2 immunity, both natural and induced.
Pathway analysis of reduced abundance mediators, including IL-2 and IL-12 (which promote NK cell survival, activation, and cytotoxicity) revealed a downregulation of pathways associated with NK cell activation (Ross and Cantrell 2018; Wang et al. 2000). These findings are consistent with prior findings that SARS-CoV-2 is associated with NK cell cytopenia and impairs NK cell function, which correlates with COVID-19 disease status and severity (Giamarellos-Bourboulis et al. 2020; van Eeden et al. 2020). In addition, IFN-γ, which we observed to be downregulated, plays a key role in downstream IL-12 production. These data demonstrate that NK cell dysfunction is detectable in saliva during early infection, even among ambulatory patients with mild disease, and provide support for NK cells as a potential therapeutic target (Zafarani et al. 2023). In the future, saliva could identify patients susceptible to severe disease or monitor response to NK-targeting therapies.
Our findings suggest that early SARS-CoV-2 infection may suppress mucosal-associated T-cell responses. In addition to its effects on NK cells, IL-2 is important for T-cell survival and maintenance, and IL-12 plays critical roles in Th1 polarization and IFN-γ production as well as enhancing cytotoxic T-cell responses. TSLP—an IL-2 family epithelial-derived cytokine important for the recruitment, maturation, and maintenance of innate and adaptive immune cells, including T cells—was also significantly reduced in the saliva of SARS-CoV-2+ participants, consistent with previous findings (Fig. 1) (Roan et al. 2019). We also observed a reduced abundance of IL-35, a lymphocyte-secreted IL-12 family cytokine that exerts anti-inflammatory properties through T-cell regulation (Sun et al. 2015; Jiang et al. 2019). Supporting our findings are studies showing that SARS-CoV-2 infection not only damages CD8+ T-cell responses (Gao et al. 2023) but is also associated with salivary immune mediator suppression resulting in oral microbiome perturbation (Alqedari et al. 2023). Altogether, these data suggest that SARS-CoV-2 may suppress and/or modulate mucosal T-cell responses through multiple pathways, potentially leading to compromised immune surveillance and increased oral inflammation. Whether this mucosal T-cell suppression is an etiologic factor in oral manifestations associated with COVID-19 including salivary gland inflammation, oral ulceration, gingivitis, periodontitis, and increased susceptibility to oral candidiasis is an exciting future direction (Gutierrez-Camacho et al. 2022; Carmona Loayza and Lafebre 2023; Nasiri et al. 2023; Pisano et al. 2023).
Our findings also suggest that SARS-CoV-2 may suppress B-cell–mediated immunity (Vincent et al. 2014). Supporting this, our data show that early SARS-CoV-2 is associated with reduced salivary abundance of B-cell survival and maturation mediators including APRIL and BAFF (Nakayamada and Tanaka 2016). In addition, we observed that although α-SARS-CoV-2–specific Ab and BAFF increased among SARS-CoV-2+ participants as a function of time, there was an overall reduction in salivary SARS-CoV-2–specific Ab compared with controls, a difference primarily driven by vaccination status, demonstrating that participants without prior vaccination or infection mount the least robust antibody response (Appendix Fig. 4).
COVID-19–related lymphopenia is a common finding early during infection and correlates with disease severity (Palladino 2021). Altogether, our findings agree with studies indicating that suppression of NK-, T-, and B-cell responses occurs early in infection and demonstrate for the first time that this multipronged SARS-CoV-2–mediated suppression is detectable in saliva and suggest that local mucosal immune suppression may be a major pathogenic feature of SARS-CoV-2+ infection. Extending these findings, antigen-specific salivary Ab profiling during early infection could help reveal oral immune response dynamics with implications for long-term immunity and reinfection susceptibility.
In this study, we observed altered salivary abundance of IL-20 subfamily cytokines in SARS-CoV-2+ participants including IL-19 (increased), IL-20 (decreased), and IL-22 (decreased). Recent studies report elevated salivary IL-19 in SARS-CoV-2 (Saheb Sharif-Askari et al. 2022) and a positive correlation between IL-20 subfamily cytokine expression and severe COVID-19 (Rajamanickam et al. 2023), in line with our data showing viral load is positively correlated with IL-20/-22 abundance. Further, IL-22 was proposed to play a critical role in COVID-19 pathogenesis (Zhang et al. 2023). Mechanistic investigations into SARS-CoV-2-associated perturbation of IL-20 subfamily cytokines, which play a critical role in regulating epithelial inflammation, is a promising future direction.
Our data show that SARS-CoV-2 is also associated with a profound increase in salivary abundance of proinflammatory mediators including soluble TNF and IL-6 receptors (sTNF-R1, -R2; sCD30; sIL-6Rα,β), CHI3L1, PTX3, and MMP-3, many of which were positively correlated with viral load (Appendix Fig. 3). sTNF-R, CHI3L1, and MMP-3 play roles in pathogenic inflammation including COVID-19, where elevated levels in circulation increase with severity and predict poorer outcome (De Lorenzo et al. 2022; Rocha Aciole et al. 2023). Further, CHI3L1 has been proposed as both a promising biomarker and universal therapeutic target in COVID-19 (Kamle et al. 2022; Parlak and Laloğlu 2022). Similarly, the importance of IL-6/IL-6R axis dysfunction in SARS-CoV-2 is now well-known (Copaescu et al. 2020).
Of note, elevated salivary PTX3 was observed in SARS-CoV-2+ subjects but remained undetectable in most controls. PTX3 is an early innate immune cytokine released by several cell types (including endothelial cells) in response to proinflammatory stimuli (Bottazzi et al. 2010). Studies show that serum PTX3 concentration may be an early, strong indicator of COVID-19–related mortality (Capra et al. 2023). Our findings agree with studies showing that PTX3 can be a reliable and sensitive biomarker in SARS-CoV-2 infection with the novel finding that noninvasive salivary monitoring could serve as a surrogate for serum PTX3 assessment. Altogether, our data support development of these analytes as COVID-19 biomarkers and highlight the prognostic potential of saliva in assessing proinflammatory cascades during early disease course.
Inflammatory bone loss has been reported in SARS-CoV-2 animal infection models (Haudenschild et al. 2023). Intriguingly, several of the above-described proinflammatory mediators including sTNF-R1/2, PTX3, and MMP-3 are involved in bone homeostasis and, perhaps of interest to oral health care providers, have established roles in bone turnover for orthodontic tooth movement (OTM) (Awosanya et al. 2022; Haudenschild et al. 2023). Specifically, sTNF-Rs are potent osteoclastogenic factors needed for OTM bone resorption (Abu-Amer et al. 2000), and both PTX3 and MMP-3 participate in extracellular matrix remodeling during periodontal disease and OTM (Letra et al. 2012; Parente et al. 2019). Exploring the role of these mediators in COVID-19–related oral sequalae is a promising area of future investigation.
There were several limitations to this study, which future and ongoing work will aim to address. Control subjects were enrolled “postpandemic” (September 2022 to November 2022), were largely vaccinated with prior SARS-CoV-2 exposure, and had narrower age and demographic ranges. The control sample size was small for some analyses; however, we were adequately powered for 15 of 37 tested cytokines, and we identified significant differences between control and SARS-CoV-2+ participants in 25 cytokines. Moreover, the single-visit study design limited interindividual variability assessment and longitudinal analyses. Changes in cytokine profiles of patients’ saliva suffering from stroke, cancer, COVID-19, and other disease states have been predictive of clinical outcomes and provided valuable diagnostic information (Diesch et al. 2021; Maciejczyk et al. 2021; Alqedari et al. 2023). Future work investigating longitudinal changes in the salivary immunological profile of COVID-19 patients with clinical outcomes could shed light on the oral immune response dynamics with implications for predicting disease course and informing care.
Studies indicate that assessing concomitant changes in salivary immune markers and oral microbiota may predict COVID-19 severity (Alqedari et al. 2023). While microbial factors were not assessed here, future work exploring the relationship between salivary immune markers and the oral microbiome may yield novel insights into host oral immune-microbial interactions during early SARS-CoV-2 infection. This would build upon investigations linking the oral microbiome with inflammatory processes in disease states, including COVID-19 (Kleinstein et al. 2020; Alqedari et al. 2023).
The multiplex Ig assay used did not distinguish between Ig subtypes (IgA, IgM, IgG), although studies have shown that antigen-specific salivary IgA responses correlate with results obtained from total Ig multiplex assays (Li et al. 2022).
It is well-appreciated that significant crosstalk exists between the host immune and stress response systems (Slavish and Szabo 2019). Acute stress increases susceptibility to viral and respiratory infections including that of SARS-CoV-2 (Graham et al. 1986). In addition, salivary stress biomarkers are elevated in hospitalized adults with moderate COVID-19 (Deneva et al. 2022). Whether activation of stress response pathways contributed to the elevated proinflammatory mediators observed is unclear but represents an intriguing avenue for investigation.
In conclusion, we demonstrate that the salivary immune landscape is dramatically altered in early SARS-CoV-2+ infection and marked by NK and adaptive (T- and B-cell) immune suppression; future work will explore whether this reflects a novel viral evasion strategy. This suppression was seen concomitant with increased abundance of innate proinflammatory mediators, several of which have been previously characterized; our work extends these findings to the salivary compartment and highlights the public health potential of salivary biomarker monitoring for COVID-19 diagnosis, prognosis, and vaccine or treatment response (Diesch et al. 2021; Alqedari et al. 2023). Identifying specific cytokines, such as PTX3, or immune pathways, such as IFN, that are dysregulated during early infection may lead to improved diagnostics and targeted interventions to modulate oral immune responses and improve clinical outcomes. Finally, our findings highlight the importance of vaccination in eliciting adaptive SARS-CoV-2–specific mucosal responses.
Author Contributions
C. Graves, contributed to conception or design, data acquisition, analysis, or interpretation, drafted and critically revised the manuscript; E. Babikow, N. Ghaltakhchyan, contributed to data acquisition, analysis, or interpretation, drafted and critically revised the manuscript; T.Q. Ngo, C. Liu, S. Wang, A. Shoji, C. Bocklage, S.T. Phillips, M. Markovetz, S.A. Frazier-Bowers, K. Divaris, M. Freire, D. Wu, contributed to data acquisition, analysis, or interpretation, critically revised the manuscript; S. Wallet, contributed to data conception or design, critically revised the manuscript; L.A. Jacox, contributed to conception or design, data acquisition, analysis, or interpretation, drafted and critically revised the manuscript. All authors gave their final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345241271943 – Supplemental material for Immune Dysregulation in the Oral Cavity during Early SARS-CoV-2 Infection
Supplemental material, sj-docx-1-jdr-10.1177_00220345241271943 for Immune Dysregulation in the Oral Cavity during Early SARS-CoV-2 Infection by C. Graves, E. Babikow, N. Ghaltakhchyan, T.Q. Ngo, C. Li, S. Wang, A. Shoji, C. Bocklage, S.T. Phillips, M. Markovetz, S.A. Frazier-Bowers, K. Divaris, M. Freire, S. Wallet, D. Wu and L.A. Jacox in Journal of Dental Research
Supplemental Material
sj-jpeg-2-jdr-10.1177_00220345241271943 – Supplemental material for Immune Dysregulation in the Oral Cavity during Early SARS-CoV-2 Infection
Supplemental material, sj-jpeg-2-jdr-10.1177_00220345241271943 for Immune Dysregulation in the Oral Cavity during Early SARS-CoV-2 Infection by C. Graves, E. Babikow, N. Ghaltakhchyan, T.Q. Ngo, C. Li, S. Wang, A. Shoji, C. Bocklage, S.T. Phillips, M. Markovetz, S.A. Frazier-Bowers, K. Divaris, M. Freire, S. Wallet, D. Wu and L.A. Jacox in Journal of Dental Research
Footnotes
Acknowledgements
We thank the participants for their time and engagement with this study. We thank all members of the Jacox, Wallet, and Graves laboratories for thoughtful discussion, as well as the UNC Respiratory TRACTS Core for technical support. We are grateful for the support and guidance of the UNC School of Dentistry GoHealth Clinical Research Unit (CRU), specifically Tammy McGonagle, Carol Culver, and Wendy Lamm.
Correction (November 2024):
Article updated to correct the spelling of the fifth author’s name.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Southern Association of Orthodontists Research Award (to E.B.), the American Association of Orthodontists Foundation (AAOF) Resident Research Aid Award (to E.B.), and the Robert Boyd Biomedical Research Award (to L.J. and S.F.B). The project was funded by the National Institutes of Health (NIH), through the National Institutes of Dental and Craniofacial Research (NIDCR) via a K08 award (to L.J.), with grant award No. 1K08DE030235-01A1 and R03 award (to L.J. and C.G.) with grant award No. 1R03DE031301. This work was supported by research grants from the NIH: NIDCR R03DE028983 (to D.W.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Data Availability Statement
The data supporting the findings of this study are available within the article and its supplementary materials.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
